Characterization of Phase I Hepatic Metabolites of Anti-Premature Ejaculation Drug Dapoxetine by UHPLC-ESI-Q-TOF
Abstract
:1. Introduction
2. Results and Discussion
2.1. Metabolite Identification
2.2. HLM Biotransformation of Dapoxetine
2.3. Hepatic Biotransformation Pathways
2.4. In Silico Assessment of Toxicity
3. Materials and Methods
3.1. Chemicals and Reagents
3.2. In Vitro Simulation of Metabolism by HLM
3.3. Analytical Procedures
3.4. In Silico Assessment of the Toxicity of Dapoxetine and Its Metabolites
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- Almazroo, O.A.; Miah, M.K.; Venkataramanan, R. Drug Metabolism in the Liver. Clin. Liv. Dis. 2017, 21, 1–20. [Google Scholar] [CrossRef]
- Remmer, H. The role of the liver in drug metabolism. Am. J. Med. 1970, 49, 617–629. [Google Scholar] [CrossRef]
- Danielson, P.B. The Cytochrome P450 Superfamily: Biochemistry, Evolution and Drug Metabolism in Humans. Available online: http://www.eurekaselect.com/64118/article (accessed on 19 January 2020).
- McDonnell, A.M.; Dang, C.H. Basic Review of the Cytochrome P450 System. J. Adv. Pract. Oncol. 2013, 4, 263–268. [Google Scholar] [PubMed]
- Meunier, B.; de Visser, S.P.; Shaik, S. Mechanism of Oxidation Reactions Catalyzed by Cytochrome P450 Enzymes. Chem. Rev. 2004, 104, 3947–3980. [Google Scholar] [CrossRef] [PubMed]
- Njoku, D.B. Drug-Induced Hepatotoxicity: Metabolic, Genetic and Immunological Basis. Int. J. Mol. Sci. 2014, 15, 6990–7003. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asha, S.; Vidyavathi, M. Role of human liver microsomes in in vitro metabolism of drugs-a review. Appl. Biochem. Biotechnol. 2010, 160, 1699–1722. [Google Scholar] [CrossRef] [PubMed]
- Knights, K.M.; Stresser, D.M.; Miners, J.O.; Crespi, C.L. In Vitro Drug Metabolism Using Liver Microsomes. Curr. Protoc. Pharmacol. 2016, 74, 7.8.1–7.8.24. [Google Scholar] [CrossRef]
- Cuyckens, F. Mass spectrometry in drug metabolism and pharmacokinetics: Current trends and future perspectives. Rapid Commun. Mass Spectrom. 2019, 33, 90–95. [Google Scholar] [CrossRef]
- Chowdhury, S. Identification and Quantification of Drugs, Metabolites and Metabolizing Enzymes by LC-MS; Elsevier: Amsterdam, The Netherlands, 2005; ISBN 978-0-08-045606-5. [Google Scholar]
- Ho, C.; Lam, C.; Chan, M.; Cheung, R.; Law, L.; Lit, L.; Ng, K.; Suen, M.; Tai, H. Electrospray Ionisation Mass Spectrometry: Principles and Clinical Applications. Clin. Biochem. Rev. 2003, 24, 3–12. [Google Scholar]
- Prakash, C.; Shaffer, C.L.; Nedderman, A. Analytical strategies for identifying drug metabolites. Mass Spectrom. Rev. 2007, 26, 340–369. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.S.; Kerns, E.H. LC/MS applications in drug development. Mass Spectrom. Rev. 1999, 18, 187–279. [Google Scholar] [CrossRef]
- Zhu, M.; Zhang, H.; Humphreys, W.G. Drug Metabolite Profiling and Identification by High-resolution Mass Spectrometry. J. Biol. Chem. 2011, 286, 25419–25425. [Google Scholar] [CrossRef] [Green Version]
- Gawlik, M.; Savic, V.; Jovanovic, M.; Skibiński, R. Mimicking of Phase I Metabolism Reactions of Molindone by HLM and Photocatalytic Methods with the Use of UHPLC-MS/MS. Molecules 2020, 25, 1367. [Google Scholar] [CrossRef] [Green Version]
- Press, D. Dapoxetine, a Novel Selective Serotonin Transport Inhibitor for the Treatment of Premature Ejaculation. Available online: https://www.dovepress.com/dapoxetine-a-novel-selective-serotonin-transport-inhibitor-for-the-tre-peer-reviewed-article-TCRM (accessed on 19 October 2020).
- Hatzimouratidis, K.; Amar, E.; Eardley, I.; Giuliano, F.; Hatzichristou, D.; Montorsi, F.; Vardi, Y.; Wespes, E. Guidelines on Male Sexual Dysfunction: Erectile Dysfunction and Premature Ejaculation. Eur. Urol. 2010, 57, 804–814. [Google Scholar] [CrossRef]
- Ashburn, T.T.; Thor, K.B. Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 2004, 3, 673–683. [Google Scholar] [CrossRef]
- Modi, N.B.; Dresser, M.J.; Simon, M.; Lin, D.; Desai, D.; Gupta, S. Single- and Multiple-Dose Pharmacokinetics of Dapoxetine Hydrochloride, a Novel Agent for the Treatment of Premature Ejaculation. J. Clin. Pharmacol. 2006, 46, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Hoy, S.M.; Scott, L.J. Dapoxetine. Drugs 2010, 70, 1433–1443. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, C.L.; Cornpropst, J.D. Determination of dapoxetine, an investigational agent with the potential for treating depression, and its mono- and di-desmethyl metabolites in human plasma using column-switching high-performance liquid chromatography. J. Chromatogr. B Biomed. Appl. 1993, 612, 253–261. [Google Scholar] [CrossRef]
- Zhang, W.; Qiang, W.; Wang, Y.; Ming, S.; Wang, R. Development and validation of a sensitive UPLC-MS/MS method for the simultaneous determination of dapoxetine and its two metabolites in human plasma. J. Pharm. Biomed. Anal. 2016, 119, 45–49. [Google Scholar] [CrossRef] [PubMed]
- Tambe, V.S.; Deodhar, M.N.; Prakya, V. Stability-indicating UPLC-MS/UV method for simultaneous determination of sildenafil citrate and dapoxetine hydrochloride from bulk abd formulation. Indian J. Pharm. Sci. 2016, 78, 663–672. [Google Scholar] [CrossRef] [Green Version]
- Lantz, R.J.; Gillespie, T.A.; Rash, T.J.; Kuo, F.; Skinner, M.; Kuan, H.-Y.; Knadler, M.P. Metabolism, excretion, and pharmacokinetics of duloxetine in healthy human subjects. Drug Metab. Dispos. 2003, 31, 1142–1150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bakhtyari, N.G.; Raitano, G.; Benfenati, E.; Martin, T.; Young, D. Comparison of In Silico Models for Prediction of Mutagenicity. J. Environ. Sci. Health Part C 2013, 31, 45–66. [Google Scholar] [CrossRef] [PubMed]
- Kang, M.J.; Song, W.H.; Shim, B.H.; Oh, S.Y.; Lee, H.Y.; Chung, E.Y.; Sohn, Y.; Lee, J. Pharmacologically active metabolites of currently marketed drugs: Potential resources for new drug discovery and development. Yakugaku Zasshi 2010, 130, 1325–1337. [Google Scholar] [CrossRef] [Green Version]
- Cassano, A.; Raitano, G.; Mombelli, E.; Fernández, A.; Cester, J.; Roncaglioni, A.; Benfenati, E. Evaluation of QSAR Models for the Prediction of Ames Genotoxicity: A Retrospective Exercise on the Chemical Substances Registered Under the EU REACH Regulation. J. Environ. Sci. Health Part C 2014, 32, 273–298. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, P.; Mumtaz, M.; Gombar, V. Assessing the toxic effects of ethylene glycol ethers using Quantitative Structure Toxicity Relationship models. Toxicol. Appl. Pharmacol. 2011, 254, 198–205. [Google Scholar] [CrossRef]

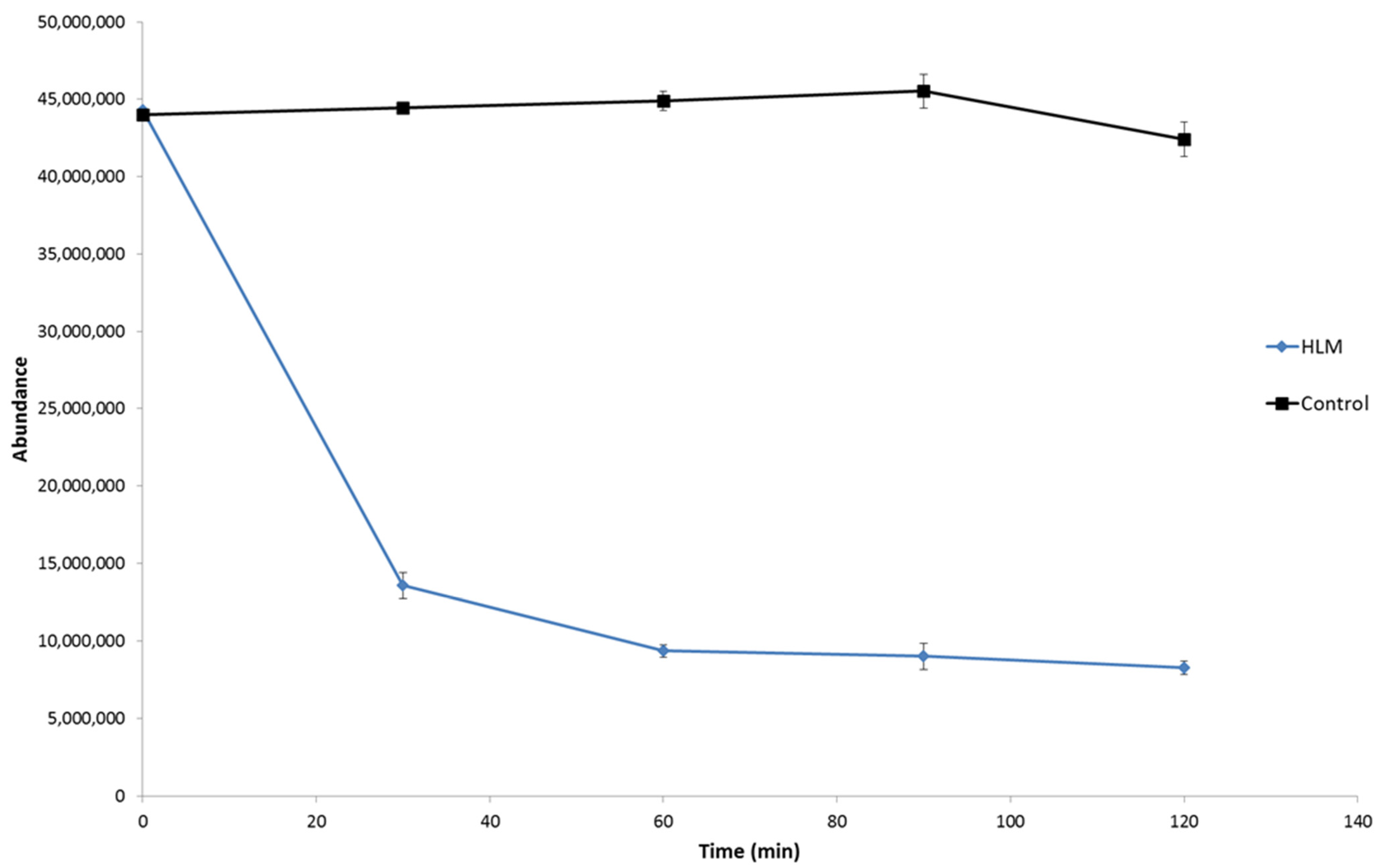
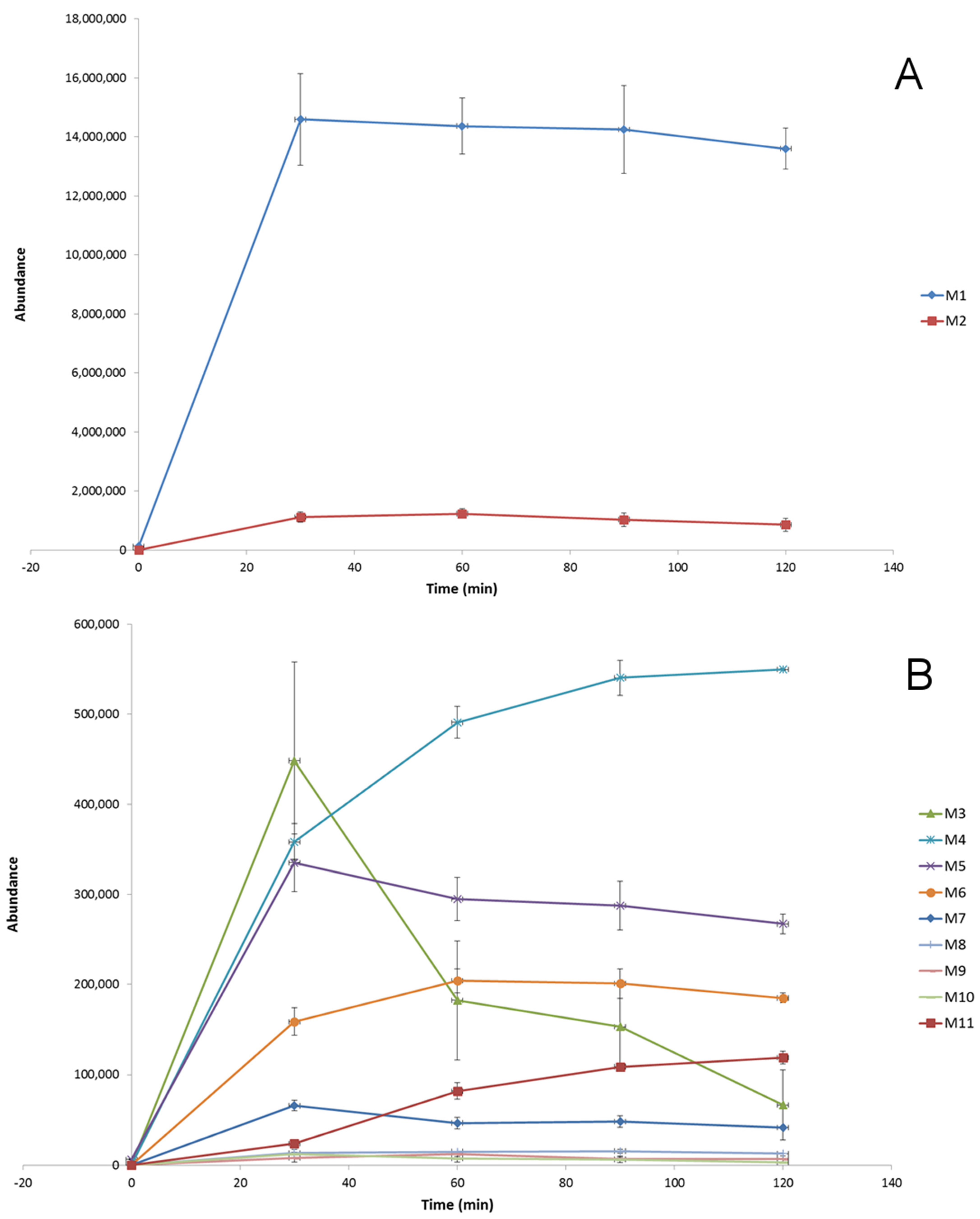
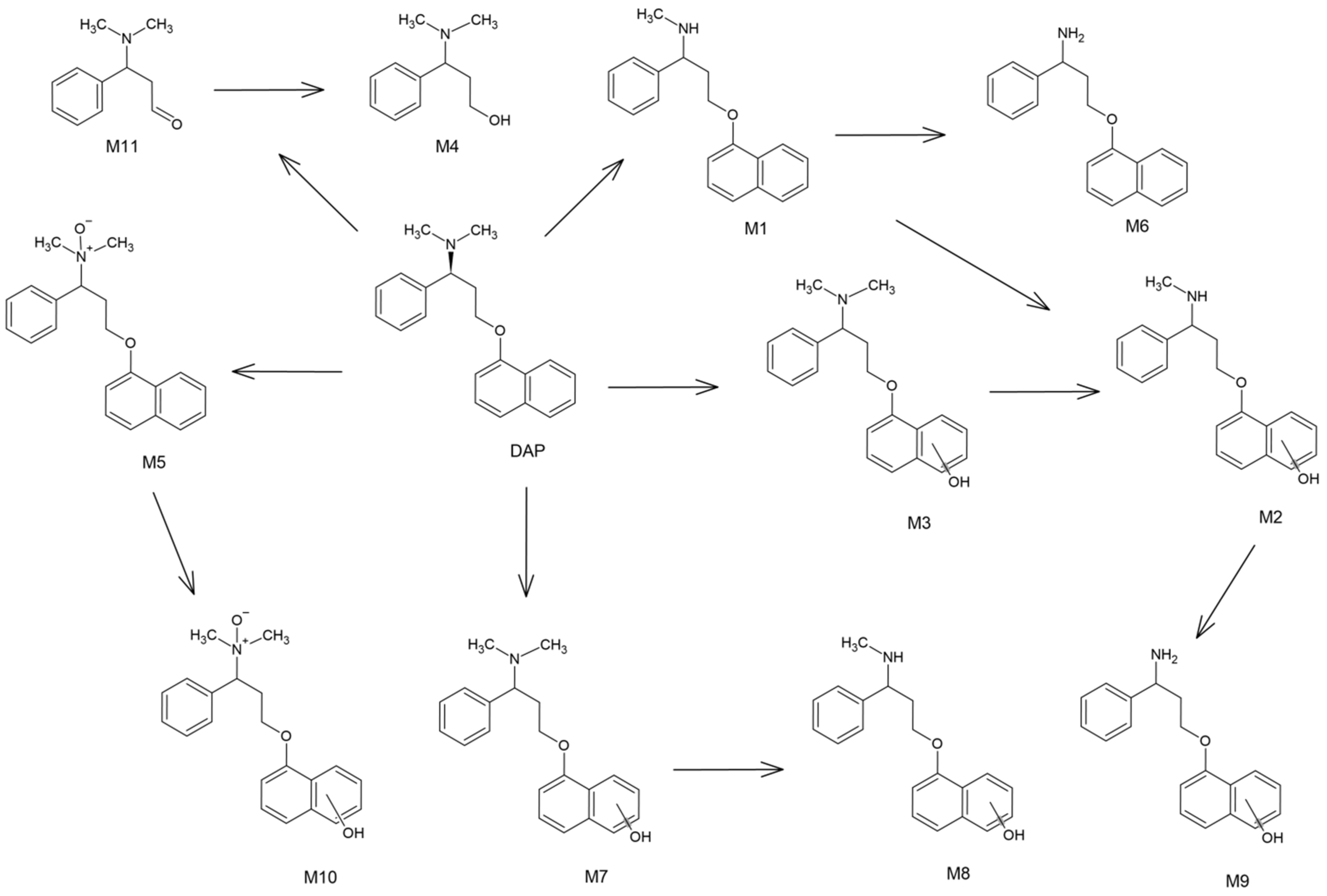
| Name | Reaction Type | Retention TIMe (min) | Measured Mass (m/z) | Theoretical Mass (m/z) | Mass Error (ppm) | Molecular Formula ([M + H]+) | MS/MS Fragmentation (m/z) and Formulas ([M + H]+) |
|---|---|---|---|---|---|---|---|
| DAP | – | 7.55 | 306.1843 | 306.1852 | −2.94 | C21H24NO | 261.1287 (C19H17O); 233.0967 (C17H13O); 215.0877 (C17H11); 183.0809 (C13H11O); 157.0649 (C11H9O); 145.0638 (C10H9O); 129.0697 (C10H7); 117.0698 (C9H9); 91.0544 (C7H7); 77.0396 (C6H5) |
| M1 | Dealk | 7.43 | 292.1687 | 292.1696 | −3.08 | C20H22NO | 261.1272 (C19H17O); 243.1159 (C19H15); 233.0955 (C17H13O); 215.0852 (C17H11); 183.0804 (C13H11O); 157.0645 (C11H9O); 145.0638 (C10H9O); 129.0698 (C10H7); 117.0701 (C9H9); 101.0384 (C8H5); 91.0549 (C7H7); 77.0392 (C6H5) |
| M2 | Dealk Ar-OH | 5.31 | 308.1634 | 308.1645 | −3.57 | C20H22NO2 | 277.1220 (C19H17O2); 249.0909 (C17H13O2); 231.0830 (C17H11O); 199.0748 (C13H11O2); 173.0597 (C11H9O2); 159.0451 (C10H9O2); 145.0655 (C10H9O); 127.0545 (C10H7); 117.0696 (C9H9); 105.0703 (C8H9); 91.0548 (C7H7) |
| M3 | Ar-OH | 5.36 | 322.1770 | 322.1801 | −9.62 | C21H24NO2 | 277.1277 (C19H17O2); 249.0906 (C17H13O2); 199.0749 (C13H11O2); 173.0593 (C11H9O2); 145.0649 (C10H9O); 117.0702 (C9H9) |
| M4 | Dearyl Red | 0.75 | 180.1373 | 180.1383 | −5.55 | C11H18NO | 135.0797 (C9H11O); 117.0689 (C9H9); 105.0698 (C8H9); 91.0546 (C7H7) |
| M5 | N-ox | 7.86 | 322.1770 | 322.1801 | −9.62 | C21H24NO2 | 261.1271 (C19H17O); 233.0940 (C17H13O); 183.0802 (C13H11O); 157.0638 (C11H9O); 129.0694 (C10H7); 117.0698 (C9H9); 74.0979 (C4H12N); 62.0610 (C2H8NO) |
| M6 | Dealk | 7.04 | 278.1528 | 278.1539 | −3.95 | C19H20NO | 261.1263 (C19H17O); 233.0973 (C17H13O); 183.0804 (C13H11O); 157.0646 (C11H9O); 129.0687 (C10H7); 117.0690 (C9H9) |
| M7 | Ar-OH | 6.46 | 322.1770 | 322.1801 | −9.62 | C21H24NO2 | 277.1218 (C19H17O2); 199.0774 (C13H11O2); 145.0640 (C10H9O); 117.0704 (C9H9) |
| M8 | Dealk Ar-OH | 6.39 | 308.1634 | 308.1645 | −3.57 | C20H22NO2 | 277.1231 (C19H17O2); 145.0640 (C10H9O); 117.0703 (C9H9) |
| M9 | Dealk Ar-OH | 4.90 | 294.1479 | 294.1489 | −3.40 | C19H20NO2 | 277.1225 (C19H17O2); 199.0744 (C13H11O2); 173.0583 (C11H9O2); 177.0699 (C9H9) |
| M10 | N-ox Ar-OH | 5.62 | 338.1724 | 338.1751 | −7.98 | C21H24NO3 | 277.1218 (C19H17O2); 199.0737 (C13H11O2); 173.0600 (C11H9O2); 117.0698 (C9H9); 62.0615 (C2H8NO) |
| M11 | Dearyl | 0.88 | 178.1216 | 178.1226 | −5.61 | C11H16NO | 136.9301 (C9H14N); 117.0709 (C9H9); 105.0710 (C8H9); 91.0551 (C7H7) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Skibiński, R.; Trawiński, J.; Gawlik, M. Characterization of Phase I Hepatic Metabolites of Anti-Premature Ejaculation Drug Dapoxetine by UHPLC-ESI-Q-TOF. Molecules 2021, 26, 3794. https://0-doi-org.brum.beds.ac.uk/10.3390/molecules26133794
Skibiński R, Trawiński J, Gawlik M. Characterization of Phase I Hepatic Metabolites of Anti-Premature Ejaculation Drug Dapoxetine by UHPLC-ESI-Q-TOF. Molecules. 2021; 26(13):3794. https://0-doi-org.brum.beds.ac.uk/10.3390/molecules26133794
Chicago/Turabian StyleSkibiński, Robert, Jakub Trawiński, and Maciej Gawlik. 2021. "Characterization of Phase I Hepatic Metabolites of Anti-Premature Ejaculation Drug Dapoxetine by UHPLC-ESI-Q-TOF" Molecules 26, no. 13: 3794. https://0-doi-org.brum.beds.ac.uk/10.3390/molecules26133794






