EGFR Mutations in Surgically Resected Fresh Specimens from 697 Consecutive Chinese Patients with Non-Small Cell Lung Cancer and Their Relationships with Clinical Features
Abstract
:1. Introduction
2. Results
2.1. Patient Characteristics
2.2. EGFR Mutation Status
3. Discussion
4. Experimental Section
4.1. Patients and Specimen Sampling
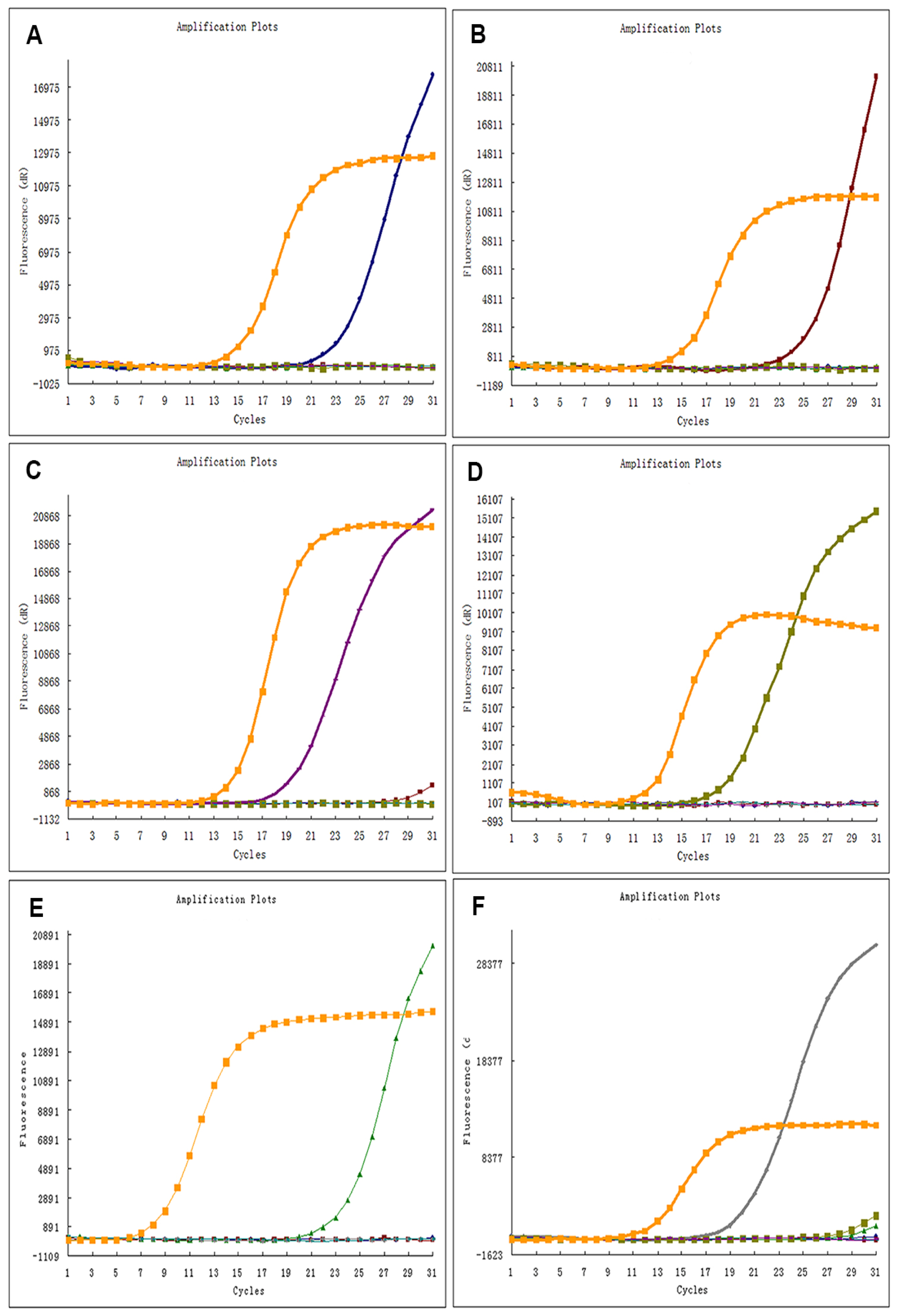
4.2. EGFR Mutation Analysis Using ARMS
4.3. Statistical Analysis
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin 2011, 61, 69–90. [Google Scholar]
- Kosaka, T.; Yatabe, Y.; Endoh, H.; Kuwano, H.; Takahashi, T.; Mitsudomi, T. Mutations of the epidermal growth factor receptor gene in lung cancer: Biological and clinical implications. Cancer Res 2004, 64, 8919–8923. [Google Scholar]
- Lynch, T.J.; Bell, D.W.; Sordella, R.; Gurubhagavatula, S.; Okimoto, R.A.; Brannigan, B.W.; Harris, P.L.; Haserlat, S.M.; Supko, J.G.; Haluska, F.G.; et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med 2004, 350, 2129–2139. [Google Scholar]
- Paez, J.G.; Janne, P.A.; Lee, J.C.; Tracy, S.; Greulich, H.; Gabriel, S.; Herman, P.; Kaye, F.J.; Lindeman, N.; Boggon, T.J.; et al. EGFR mutations in lung cancer: Correlation with clinical response to gefitinib therapy. Science 2004, 304, 1497–1500. [Google Scholar]
- Bell, D.W.; Lynch, T.J.; Haserlat, S.M.; Harris, P.L.; Okimoto, R.A.; Brannigan, B.W.; Sgroi, D.C.; Muir, B.; Riemenschneider, M.J.; Iacona, R.B.; et al. Epidermal growth factor receptor mutations and gene amplification in non-small-cell lung cancer: Molecular analysis of the IDEAL/INTACT gefitinib trials. J. Clin. Oncol 2005, 23, 8081–8092. [Google Scholar]
- Soung, Y.H.; Lee, J.W.; Kim, S.Y.; Seo, S.H.; Park, W.S.; Nam, S.W.; Song, S.Y.; Han, J.H.; Park, C.K.; Lee, J.Y.; et al. Mutational analysis of EGFR and K-RAS genes in lung adenocarcinomas. Virchows Arch 2005, 446, 483–488. [Google Scholar]
- Tsao, A.S.; Tang, X.M.; Sabloff, B.; Xiao, L.; Shigematsu, H.; Roth, J.; Spitz, M.; Hong, W.K.; Gazdar, A.; Wistuba, I. Clinicopathologic characteristics of the EGFR gene mutation in non-small cell lung cancer. J. Thorac. Oncol 2006, 1, 231–239. [Google Scholar]
- Kang, S.M.; Kang, H.J.; Shin, J.H.; Kim, H.; Shin, D.H.; Kim, S.K.; Kim, J.H.; Chung, K.Y.; Chang, J. Identical epidermal growth factor receptor mutations in adenocarcinomatous and squamous cell carcinomatous components of adenosquamous carcinoma of the lung. Cancer 2007, 109, 581–587. [Google Scholar]
- Sasaki, H.; Endo, K.; Yukiue, H.; Kobayashi, Y.; Yano, M.; Fujii, Y. Mutation of epidermal growth factor receptor gene in adenosquamous carcinoma of the lung. Lung Cancer 2007, 55, 129–130. [Google Scholar]
- Kim, Y.T.; Kim, T.Y.; Lee, D.S.; Park, S.J.; Park, J.Y.; Seo, S.J.; Choi, H.S.; Kang, H.J.; Hahn, S.; Kang, C.H.; et al. Molecular changes of epidermal growth factor receptor (EGFR) and KRAS and their impact on the clinical outcomes in surgically resected adenocarcinoma of the lung. Lung Cancer 2008, 59, 111–118. [Google Scholar]
- Jang, T.W.; Oak, C.H.; Chang, H.K.; Suo, S.J.; Jung, M.H. EGFR and KRAS mutations in patients with adenocarcinoma of the lung. Korean J. Intern. Med 2009, 24, 48–54. [Google Scholar]
- Rosell, R.; Moran, T.; Queralt, C.; Porta, R.; Cardenal, F.; Camps, C.; Majem, M.; Lopez-Vivanco, G.; Isla, D.; Provencio, M.; et al. Screening for epidermal growth factor receptor mutations in lung cancer. N. Engl. J. Med 2009, 361, 958–967. [Google Scholar]
- Tanaka, T.; Matsuoka, M.; Sutani, A.; Gemma, A.; Maemondo, M.; Inoue, A.; Okinaga, S.; Nagashima, M.; Oizumi, S.; Uematsu, K.; et al. Frequency of and variables associated with the EGFR mutation and its subtypes. Int. J. Cancer 2010, 126, 651–655. [Google Scholar]
- Mitsudomi, T.; Yatabe, Y. Mutations of the epidermal growth factor receptor gene and related genes as determinants of epidermal growth factor receptor tyrosine kinase inhibitors sensitivity in lung cancer. Cancer Sci 2007, 98, 1817–1824. [Google Scholar]
- Mok, T.S.; Wu, Y.L.; Thongprasert, S.; Yang, C.H.; Chu, D.T.; Saijo, N.; Sunpaweravong, P.; Han, B.; Margono, B.; Ichinose, Y.; et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N. Engl. J. Med 2009, 361, 947–957. [Google Scholar]
- Mitsudomi, T.; Morita, S.; Yatabe, Y.; Negoro, S.; Okamoto, I.; Tsurutani, J.; Seto, T.; Satouchi, M.; Tada, H.; Hirashima, T.; et al. Gefitinib versus cisplatin plus docetaxel in patients with non-small-cell lung cancer harbouring mutations of the epidermal growth factor receptor (WJTOG3405): An open label, randomised phase 3 trial. Lancet Oncol 2010, 11, 121–128. [Google Scholar]
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; Tarpey, P.; et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med 2012, 366, 883–892. [Google Scholar]
- Newton, C.R.; Graham, A.; Heptinstall, L.E.; Powell, S.J.; Summers, C.; Kalsheker, N.; Smith, J.C.; Markham, A.F. Analysis of any point mutation in DNA. The amplification refractory mutation system (ARMS). Nucleic Acids Res 1989, 17, 2503–2516. [Google Scholar]
- Ellison, G.; Donald, E.; McWalter, G.; Knight, L.; Fletcher, L.; Sherwood, J.; Cantarini, M.; Orr, M.; Speake, G. A comparison of ARMS and DNA sequencing for mutation analysis in clinical biopsy samples. J. Exp. Clin. Cancer Res 2010, 29, 132. [Google Scholar]
- Mok, T.; Wu, Y.L.; Zhang, L. A small step towards personalized medicine for non-small cell lung cancer. Discov. Med 2009, 8, 227–231. [Google Scholar]
- Kimura, H.; Fujiwara, Y.; Sone, T.; Kunitoh, H.; Tamura, T.; Kasahara, K.; Nishio, K. High sensitivity detection of epidermal growth factor receptor mutations in the pleural effusion of non-small cell lung cancer patients. Cancer Sci 2006, 97, 642–648. [Google Scholar]
- Tokumo, M.; Toyooka, S.; Kiura, K.; Shigematsu, H.; Tomii, K.; Aoe, M.; Ichimura, K.; Tsuda, T.; Yano, M.; Tsukuda, K.; et al. The relationship between epidermal growth factor receptor mutations and clinicopathologic features in non-small cell lung cancers. Clin. Cancer Res 2005, 11, 1167–1173. [Google Scholar]
- Sonobe, M.; Manabe, T.; Wada, H.; Tanaka, F. Mutations in the epidermal growth factor receptor gene are linked to smoking-independent, lung adenocarcinoma. Br. J. Cancer 2005, 93, 355–363. [Google Scholar]
- Pao, W.; Miller, V.A.; Politi, K.A.; Riely, G.J.; Somwar, R.; Zakowski, M.F.; Kris, M.G.; Varmus, H. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med 2005, 2, e73. [Google Scholar]
- Turner, N.C.; Reis-Filho, J.S. Genetic heterogeneity and cancer drug resistance. Lancet Oncol 2012, 13, e178–e185. [Google Scholar]
- Brambilla, E.; Travis, W.D.; Colby, T.V.; Corrin, B.; Shimosato, Y. The new World Health Organization classification of lung tumours. Eur. Respir. J 2001, 18, 1059–1068. [Google Scholar]
| Characteristics | N | Mutation *,% | p-value |
|---|---|---|---|
| Age | 0.066 | ||
| <60 years | 378 | 116, 30.7% | |
| ≥60 years | 319 | 119, 37.3% | |
| Gender | <0.001 | ||
| Male | 476 | 108, 22.7% | |
| Female | 221 | 127, 57.5% | |
| Smoking status | 0.047 | ||
| Never | 331 | 124, 37.5% | |
| Ever | 366 | 111, 30.3% | |
| Histology | <0.001 | ||
| Squamous-cell carcinoma | 282 | 41, 14.5% | |
| Adenocarcinoma ** | 293 | 155, 52.9% | |
| Adenosquamous carcinoma | 86 | 34, 39.5% | |
| Large-cell carcinoma | 11 | 1, 9.1% | |
| Sarcomatoid carcinoma | 18 | 2, 11.1% | |
| Mucoepidermoid carcinoma | 7 | 2, 28.6% | |
| Stage | 0.898 | ||
| Ia | 105 | 34, 32.4% | |
| Ib | 110 | 40, 36.4% | |
| IIa | 153 | 49, 32.0% | |
| IIb | 159 | 56, 35.2% | |
| IIIa | 155 | 53, 34.2% | |
| IIIb | 12 | 2, 16.7% | |
| IV | 3 | 1, 33.3% |
| Mutation spot | Gender | p-value | |
|---|---|---|---|
| Male | Female | ||
| 19-Del | 48 | 54 | 0.864 |
| L858R | 51 | 62 | |
| L861Q | 1 | 3 | |
| G719X | 8 | 8 | |
| Mutation spot | Smoking Status | p-value | |
|---|---|---|---|
| Ever | Never | ||
| 19-Del | 22 | 80 | 0.202 |
| L858R | 34 | 79 | |
| L861Q | 1 | 3 | |
| G719X | 7 | 9 | |
| Mutation spot | Pathology | ||||
|---|---|---|---|---|---|
| Squamous-cell carcinoma | Adeno-carcinoma | Adeno-squamous carcinoma | Others * | p-value | |
| 19-Del | 17 | 66 | 17 | 2 | 0.671 |
| L858R | 19 | 76 | 16 | 2 | |
| L861Q | 0 | 4 | 0 | 0 | |
| G719X | 5 | 9 | 1 | 1 | |
| Subtype | Age (years) | Gender | Smoking status | Tumor Histology | Stage |
|---|---|---|---|---|---|
| T790M | 53 | Male | Ever | Squamous-cell carcinoma | IIIa |
| T790M | 67 | Male | Ever | Adenocarcinoma | IIIb |
| T790M | 65 | Male | Never | Squamous-cell carcinoma | Ib |
| T790M | 50 | Female | Never | Squamous-cell carcinoma | IIa |
| T790M | 68 | Female | Never | Adenocarcinoma | IIb |
| T790M | 71 | Female | Never | Adenocarcinoma | Ib |
| 20-Ins | 73 | Female | Never | Adenocarcinoma | IIIa |
| 20-Ins | 62 | Female | Never | Adenocarcinoma | IIb |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lai, Y.; Zhang, Z.; Li, J.; Sun, D.; Zhou, Y.; Jiang, T.; Han, Y.; Huang, L.; Zhu, Y.; Li, X.; et al. EGFR Mutations in Surgically Resected Fresh Specimens from 697 Consecutive Chinese Patients with Non-Small Cell Lung Cancer and Their Relationships with Clinical Features. Int. J. Mol. Sci. 2013, 14, 24549-24559. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms141224549
Lai Y, Zhang Z, Li J, Sun D, Zhou Y, Jiang T, Han Y, Huang L, Zhu Y, Li X, et al. EGFR Mutations in Surgically Resected Fresh Specimens from 697 Consecutive Chinese Patients with Non-Small Cell Lung Cancer and Their Relationships with Clinical Features. International Journal of Molecular Sciences. 2013; 14(12):24549-24559. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms141224549
Chicago/Turabian StyleLai, Yuanyang, Zhipei Zhang, Jianzhong Li, Dong Sun, Yong'an Zhou, Tao Jiang, Yong Han, Lijun Huang, Yifang Zhu, Xiaofei Li, and et al. 2013. "EGFR Mutations in Surgically Resected Fresh Specimens from 697 Consecutive Chinese Patients with Non-Small Cell Lung Cancer and Their Relationships with Clinical Features" International Journal of Molecular Sciences 14, no. 12: 24549-24559. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms141224549




