Development and Application of Loop-Mediated Isothermal Amplification Assays for Rapid Visual Detection of cry2Ab and cry3A Genes in Genetically-Modified Crops
Abstract
:1. Introduction
2. Results and Discussion
2.1. Extraction of Genomic DNA from Samples
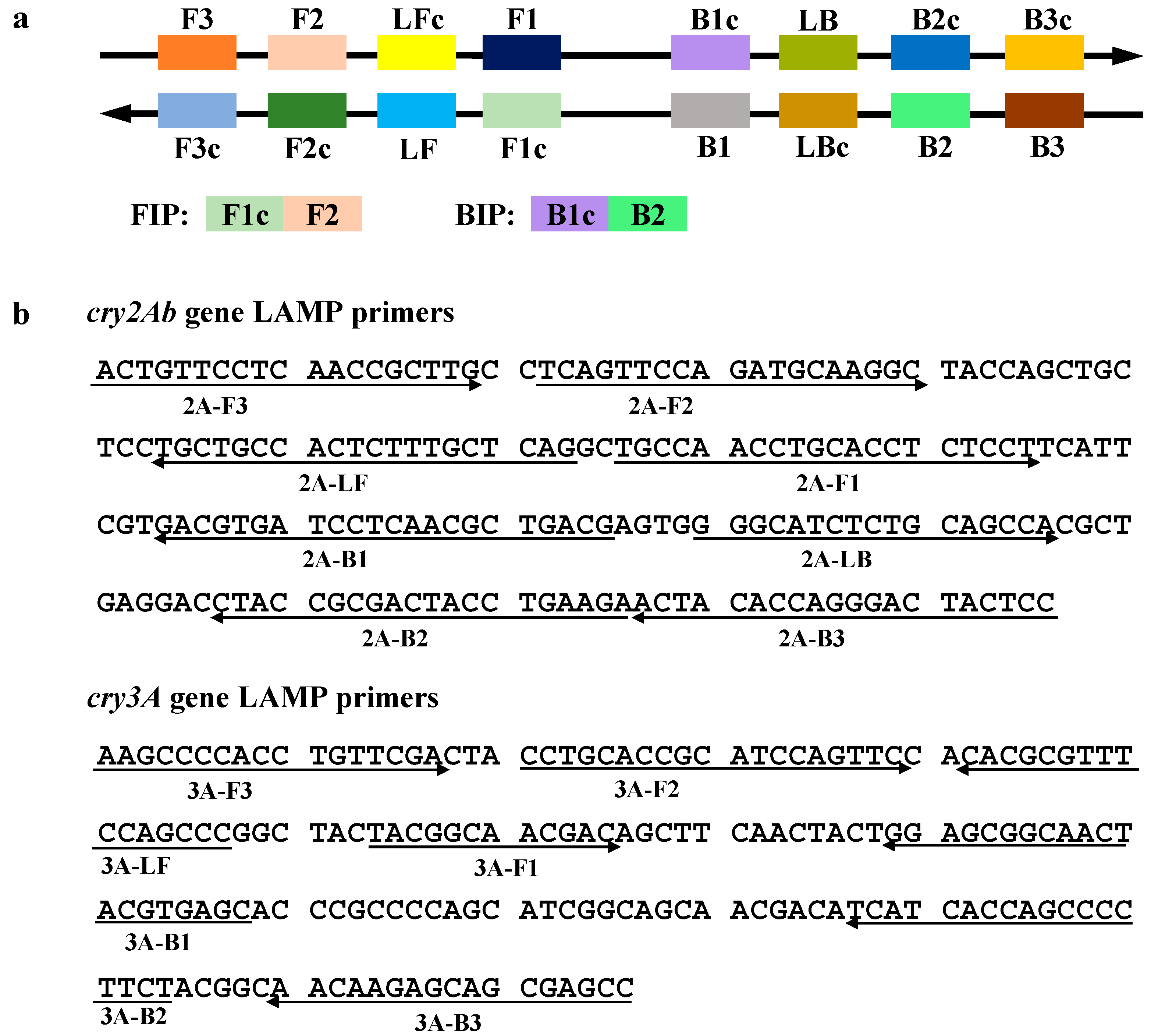
2.2. Design of the Loop-Mediated Isothermal Amplification (LAMP) Primers for cry2Ab and cry3A Genes
| Primers Name | Sequence (5'–3') | Target | Amplicon Size (bp) | Reference |
|---|---|---|---|---|
| 2A-F3 | ACTGTTCCTCAACCGCTTG | cry2Ab gene | 196 | This work |
| 2A-B3 | GGAGTAGTCCCTGGTGTAGT | |||
| 2A-FIP | AGGAGAGGTGCAGGTTGGCACTCAGTTCCAGATGCAAGGC | |||
| 2A-BIP | GACGTGATCCTCAACGCTGACGTCTTCAGGTAGTCGCGGTAG | |||
| 2A-LF | CTGAGCAAAGAGTGGCAGCA | |||
| 2A-LB | GGGCATCTCTGCAGCCA | |||
| 3A-F3 | AAGCCCCACCTGTTCGA | cry3A gene | 176 | This work |
| 3A-B3 | GGCTCGCTGCTCTTGTTG | |||
| 3A-FIP | AGTTGAAGCTGTCGTTGCCGTACCTGCACCGCATCCAGTTC | |||
| 3A-BIP | GGAGCGGCAACTACGTGAGCAGAAGGGGCTGGTGATGA | |||
| 3A-LF | GGGCTGGAAACGCGTGT |
2.3. Specificity of the LAMP Assays
| Species | GM Crops | Contained Bt Genes | cry2Ab | cry3A | ||
|---|---|---|---|---|---|---|
| LAMP | PCR | LAMP | PCR | |||
| Maize | Bt11 | cry1Ab | − | − | − | − |
| Bt176 | cry1Ab | − | − | − | − | |
| MON810 | cry1Ab | − | − | − | − | |
| TC1507 | cry1F | − | − | − | − | |
| MON89034 | cry1A.105, cry2Ab | + | + | − | − | |
| MON863 | cry3Bb | − | − | − | − | |
| MON88017 | cry3Bb | − | − | − | − | |
| MIR604 | cry3A | − | − | + | + | |
| 59122 | cry34Ab1, cry35Ab1 | − | − | − | − | |
| Cotton | MON531 | cry1Ac | − | − | − | − |
| MON15985 | cry1Ac, cry2Ab | + | + | − | − | |
| Rice | TT51-1 | cry1Ab/Ac | − | − | − | − |
| KF-6 | cry1Ab | − | − | − | − | |
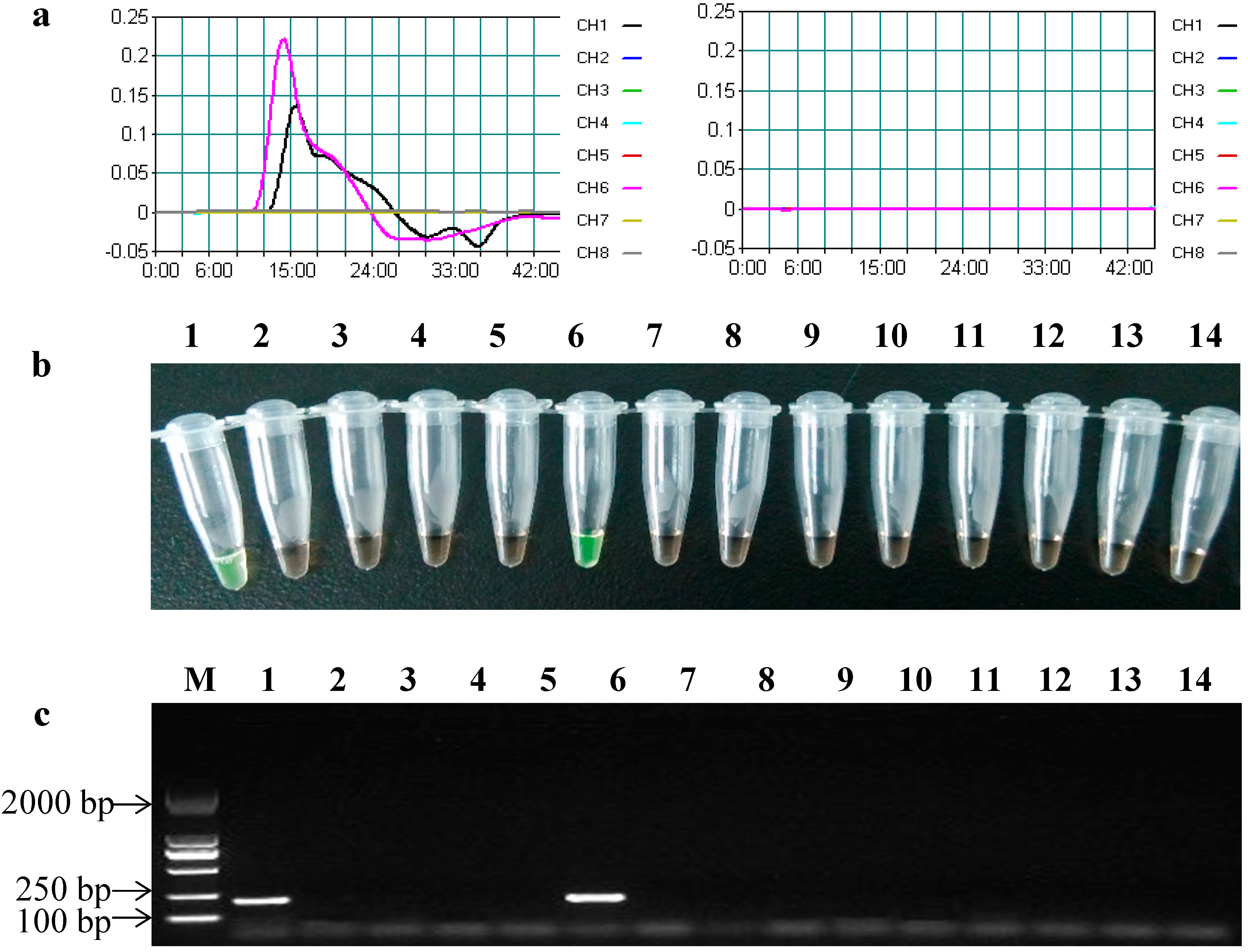
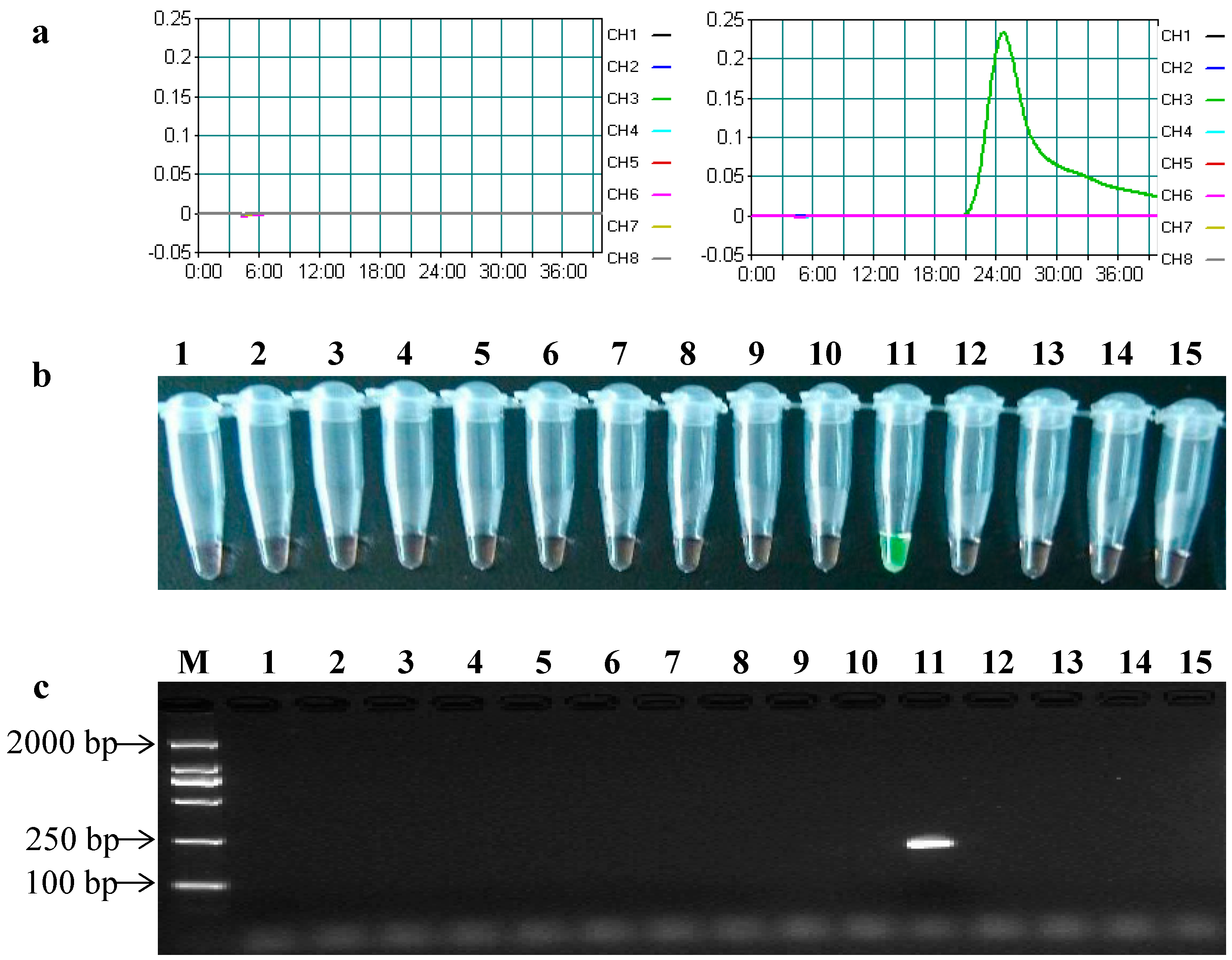
2.4. Sensitivity of the LAMP Assays
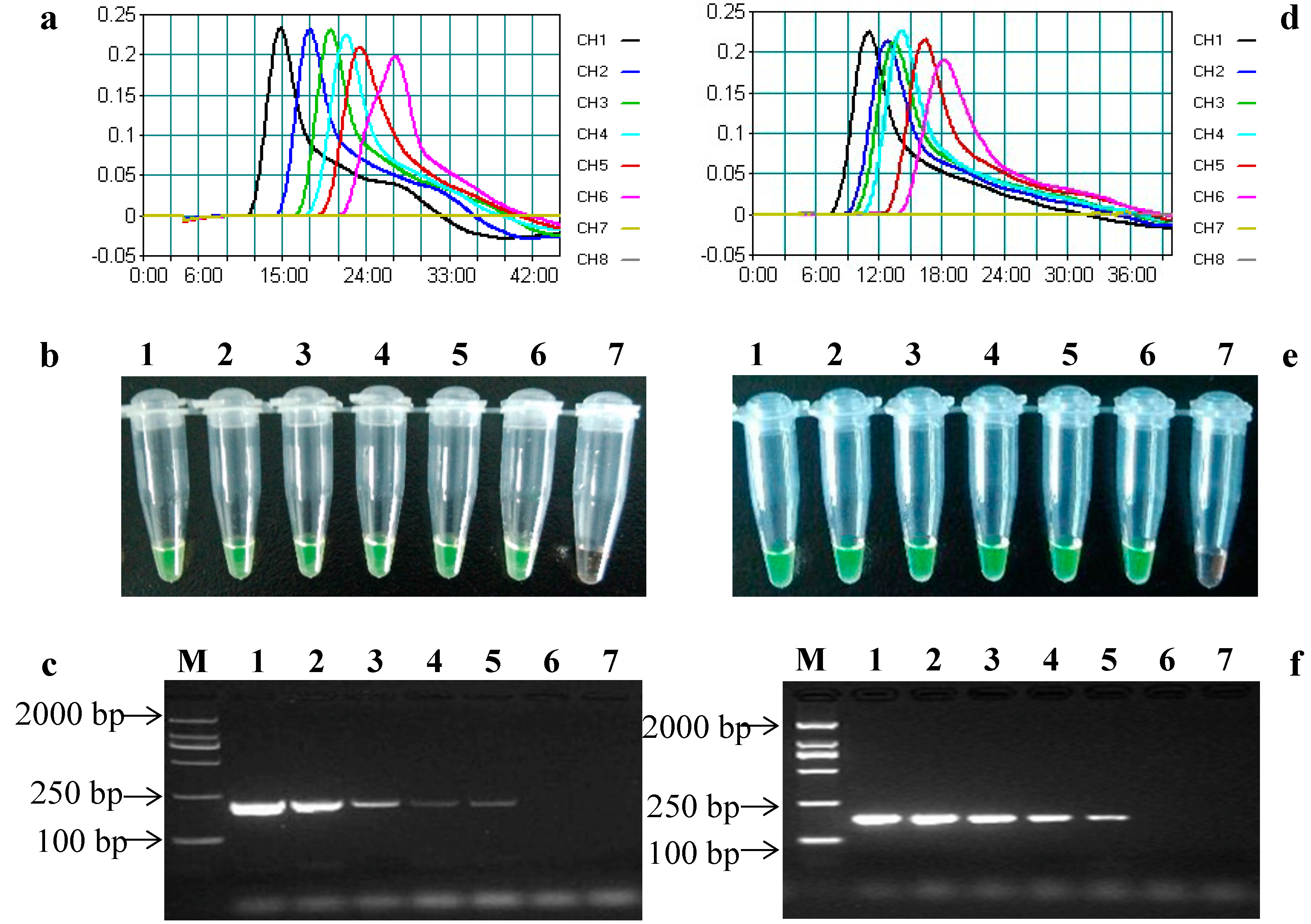
3. Experimental Section
3.1. Plant Material
3.2. DNA Isolation
3.3. Primers Design
3.4. LAMP Assay
3.5. Analysis of LAMP Products
3.6. Conventional PCR Assay
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Clive, J. Global Status of Commercialized Biotech/GM Crops: 2013; ISAAA Brief No. 46; International Service for the Acquisition Agri-biotech Applications (ISAAA): Ithaca, NY, USA, 2013. [Google Scholar]
- Kamle, S.; Ali, S. Genetically modified crops: Detection strategies and biosafety issues. Gene 2013, 522, 123–132. [Google Scholar] [CrossRef]
- Zhang, D.; Guo, J. The development and standardization of testing methods for genetically modified organisms and their derived products. J. Integr. Plant. Biol. 2011, 53, 539–551. [Google Scholar] [CrossRef]
- Debode, F.; Janssen, E.; Berben, G. Development of 10 new screening PCR assays for GMO detection targeting promoters (pFMV, pNOS, pSSuAra, pTA29, pUbi, pRice actin) and terminators (t35S, tE9, tOCs, tg7). Eur. Food Res. Technol. 2013, 236, 659–669. [Google Scholar] [CrossRef]
- Holst-Jensen, A.; Bertheau, Y.; de Loose, M.; Grohmann, L.; Hamels, S.; Hougs, L.; Morisset, D.; Pecoraro, S.; Pla, M.; van den Bulcke, M.; et al. Detecting un-authorized genetically modified organisms (GMOs) and derived materials. Biotechnol. Adv. 2012, 30, 1318–1335. [Google Scholar] [CrossRef]
- Broeders, S.R.; de Keersmaecker, S.C.; Roosens, N.H. How to deal with the upcoming challenges in GMO detection in food and feed. J. Biomed. Biotechnol. 2012, 2012, 402418. [Google Scholar]
- Shao, N.; Jiang, S.M.; Zhang, M.; Wang, J.; Guo, S.J.; Li, Y.; Jiang, H.W.; Liu, C.X.; Zhang, D.B.; Yang, L.T.; et al. MACRO: A combined microchip-PCR and microarray system for high-throughput monitoring of genetically modified organisms. Anal. Chem. 2014, 86, 1269–1276. [Google Scholar] [CrossRef]
- Wu, H.; Zhang, Y.; Zhu, C.; Xiao, X.; Zhou, X.; Xu, S.; Shen, W.; Huang, M. Presence of CP4-EPSPS component in roundup ready soybean-derived food products. Int. J. Mol. Sci. 2012, 13, 1919–1932. [Google Scholar] [CrossRef]
- Kamle, S.; Ojha, A.; Kumar, A. Development of an enzyme linked immunosorbant assay for the detection of Cry2Ab protein in transgenic plants. GM Crops 2011, 2, 118–125. [Google Scholar] [CrossRef]
- Zhang, Y.; Lai, C.; Su, R.; Zhang, M.; Xiong, Y.; Qing, H.; Deng, Y. Quantification of Cry1Ab in genetically modified maize leaves by liquid chromatography multiple reaction monitoring tandem mass spectrometry using 18O stable isotope dilution. Analyst 2012, 137, 2699–2705. [Google Scholar] [CrossRef]
- Xu, W.; Zhai, Z.; Huang, K.; Zhang, N.; Yuan, Y.; Shang, Y.; Luo, Y. A novel universal primer-multiplex-PCR method with sequencing gel electrophoresis analysis. PLoS One 2012, 7, e22900. [Google Scholar]
- Guo, J.; Chen, L.; Liu, X.; Gao, Y.; Zhang, D.; Yang, L. A multiplex degenerate PCR analytical approach targeting to eight genes for screening GMOs. Food Chem. 2012, 132, 1566–1573. [Google Scholar] [CrossRef]
- Holst-Jensen, A. Testing for genetically modified organisms (GMOs): Past, present and future perspectives. Biotechnol. Adv. 2009, 27, 1071–1082. [Google Scholar] [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, e63. [Google Scholar] [CrossRef]
- Tomita, N.; Mori, Y.; Kanda, H.; Notomi, T. Loop-mediated isothermal amplification (LAMP) of gene sequences and simple visual detection of products. Nat. Protoc. 2008, 3, 877–882. [Google Scholar] [CrossRef]
- Xu, J.; Zheng, Q.; Yu, L.; Liu, R.; Zhao, X.; Wang, G.; Wang, Q.; Cao, J. Loop-mediated isothermal amplification (LAMP) method for detection of genetically modified maize T25. Food Sci. Nutr. 2013, 1, 432–438. [Google Scholar] [CrossRef]
- Guan, X.; Guo, J.; Shen, P.; Yang, L.; Zhang, D. Visual and rapid detection of two genetically modified soybean events using loop-mediated isothermal amplification method. Food Anal. Method 2010, 3, 313–320. [Google Scholar] [CrossRef]
- Adams, E.R.; Schoone, G.J.; Ageed, A.F.; Safi, S.E.; Schallig, H.D. Development of a reverse transcriptase loop-mediated isothermal amplification (LAMP) assay for the sensitive detection of Leishmania parasites in clinical samples. Am. J. Trop. Med. Hyg. 2010, 82, 591–596. [Google Scholar] [CrossRef]
- Wang, C.; Shen, X.; Lu, J.; Zhang, L. Development of a reverse transcription-loop-mediated isothermal amplification (RT-LAMP) system for rapid detection of HDV genotype 1. Lett. Appl. Microbiol. 2013, 56, 229–235. [Google Scholar] [CrossRef]
- Mori, Y.; Kanda, H.; Notomi, T. Loop-mediated isothermal amplification (LAMP): Recent progress in research and development. J. Infect. Chemother. 2013, 19, 404–411. [Google Scholar] [CrossRef]
- Parida, M.; Sannarangaiah, S.; Dash, P.K.; Rao, P.V.; Morita, K. Loop mediated isothermal amplification (LAMP): A new generation of innovative gene amplification technique; perspectives in clinical diagnosis of infectious diseases. Rev. Med. Virol. 2008, 18, 407–421. [Google Scholar] [CrossRef]
- Bahkali, A.; Moslem, M.; Amin, O.E.; Niessen, L. An optimized protocol for DNA extraction from wheat seeds and loop-mediated isothermal amplification (LAMP) to detect fusarium graminearum contamination of wheat grain. Int. J. Mol. Sci. 2011, 12, 3459–3472. [Google Scholar] [CrossRef]
- Ueda, S.; Kuwabara, Y. The rapid detection of Salmonella from food samples by loop-mediated isothermal amplification (LAMP). Biocontrol. Sci. 2009, 14, 73–76. [Google Scholar] [CrossRef]
- Vaagt, F.; Haase, I.; Fischer, M. Loop-mediated isothermal amplification (LAMP)-based method for rapid mushroom species identification. J. Agric. Food Chem. 2013, 61, 1833–1840. [Google Scholar] [CrossRef]
- Focke, F.; Haase, I.; Fischer, M. Loop-mediated isothermal amplification (LAMP): Methods for plant species identification in food. J. Agric. Food Chem. 2013, 61, 2943–2949. [Google Scholar] [CrossRef]
- Zhou, D.; Guo, J.; Xu, L.; Gao, S.; Lin, Q.; Wu, Q.; Wu, L.; Que, Y. Establishment and application of a loop-mediated isothermal amplification (LAMP) system for detection of cry1Ac transgenic sugarcane. Sci. Rep. 2014, 4, 4912. [Google Scholar]
- Huang, X.; Chen, L.; Xu, J.; Ji, H.F.; Zhu, S.; Chen, H. Rapid visual detection of phytase gene in genetically modified maize using loop-mediated isothermal amplification method. Food Chem. 2014, 156, 184–189. [Google Scholar] [CrossRef]
- Huang, S.; Xu, Y.; Yan, X.; Shang, Y.; Zhu, P.; Tian, W.; Xu, W. Development and application of a quantitative loop-mediated isothermal amplification method for detecting genetically modified Maize MON863. J. Sci. Food Agric. 2014. [Google Scholar] [CrossRef]
- Randhawa, G.J.; Singh, M.; Morisset, D.; Sood, P.; Zel, J. Loop-mediated isothermal amplification: Rapid visual and real-time methods for detection of genetically modified crops. J. Agric. Food Chem. 2013, 61, 11338–11346. [Google Scholar] [CrossRef]
- Li, Q.; Fang, J.; Liu, X.; Xi, X.; Li, M.; Gong, Y.; Zhang, M. Loop-mediated isothermal amplification (LAMP) method for rapid detection of cry1Ab gene in transgenic rice (Oryza sativa L.). Eur. Food Res. Technol. 2013, 236, 589–598. [Google Scholar] [CrossRef]
- Chen, X.; Wang, X.; Jin, N.; Zhou, Y.; Huang, S.; Miao, Q.; Zhu, Q.; Xu, J. Endpoint visual detection of three genetically modified rice events by loop-mediated isothermal amplification. Int. J. Mol. Sci. 2012, 13, 14421–14433. [Google Scholar] [CrossRef]
- Chen, L.; Guo, J.; Wang, Q.; Kai, G.; Yang, L. Development of the visual loop-mediated isothermal amplification assays for seven genetically modified maize events and their application in practical samples analysis. J. Agric. Food Chem. 2011, 59, 5914–5918. [Google Scholar] [CrossRef]
- Liu, M.; Luo, Y.; Tao, R.; He, R.; Jiang, K.; Wang, B.; Wang, L. Sensitive and rapid detection of genetic modified soybean (Roundup Ready) by loop-mediated isothermal amplification. Biosci. Biotechnol. Biochem. 2009, 73, 2365–2369. [Google Scholar] [CrossRef]
- Lee, D.; la Mura, M.; Allnutt, T.R.; Powell, W. Detection of genetically modified organisms (GMOs) using isothermal amplification of target DNA sequences. BMC Biotechnol. 2009, 9, 7. [Google Scholar] [CrossRef]
- Development Center of Science and Technology, M.o.A. Detection of Genetically Modified Plants and Derived Products—Target-Taxon-Specific Qualitative PCR Method for Maize; Announcement by the Ministry of Agriculture No. 1861-3-2012; China Agriculture Press: Beijing, China, 2012; pp. 1–9. [Google Scholar]
- Development Center of Science and Technology, M.o.A. Detection of Genetically Modified Plants and Derived Products—Target-Taxon-Specific Qualitative PCR Method for Cotton; Announcement by the Ministry of Agriculture No. 1943-1-2013; China Agriculture Press: Beijing, China, 2013; pp. 1–9. [Google Scholar]
- Development Center of Science and Technology, M.o.A. Detection of Genetically Modified Plants and Derived Products—Target-Taxon-Specific Qualitative PCR Method for Rice; Announcement by the Ministry of Agriculture No. 1861-1-2012; China Agriculture Press: Beijing, China, 2012; pp. 1–10. [Google Scholar]
- Nagamine, K.; Hase, T.; Notomi, T. Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol. Cell. Probes 2002, 16, 223–229. [Google Scholar] [CrossRef]
- Arumuganathan, K.; Earle, E.D. Nuclear DNA content of some important plant species. Plant. Mol. Biol. Rep. 1991, 9, 208–218. [Google Scholar] [CrossRef]
- GMO Detection method Database (GMDD). Available online: http://gmdd.shgmo.org/sequence/view/49 (accessed on 10 June 2013).
- GMO Detection method Database (GMDD). Available online: http://gmdd.shgmo.org/sequence/view/38 (accessed on 10 June 2013).
- Primer Explorer V4 Software. Operation Procedure for Designing Regular/Loop Primers. Available online: http://primerexplorer.jp/elamp4.0.0/index.html (accessed on 11 June 2013).
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, F.; Yan, W.; Long, L.; Qi, X.; Li, C.; Zhang, S. Development and Application of Loop-Mediated Isothermal Amplification Assays for Rapid Visual Detection of cry2Ab and cry3A Genes in Genetically-Modified Crops. Int. J. Mol. Sci. 2014, 15, 15109-15121. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms150915109
Li F, Yan W, Long L, Qi X, Li C, Zhang S. Development and Application of Loop-Mediated Isothermal Amplification Assays for Rapid Visual Detection of cry2Ab and cry3A Genes in Genetically-Modified Crops. International Journal of Molecular Sciences. 2014; 15(9):15109-15121. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms150915109
Chicago/Turabian StyleLi, Feiwu, Wei Yan, Likun Long, Xing Qi, Congcong Li, and Shihong Zhang. 2014. "Development and Application of Loop-Mediated Isothermal Amplification Assays for Rapid Visual Detection of cry2Ab and cry3A Genes in Genetically-Modified Crops" International Journal of Molecular Sciences 15, no. 9: 15109-15121. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms150915109




