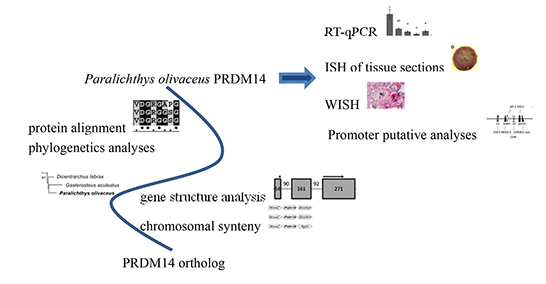
Identification and Characterization of a PRDM14 Homolog in Japanese Flounder (Paralichthys olivaceus)
Abstract
:1. Introduction
2. Results
2.1. Molecular Characterization of Po-prdm14
2.1.1. Cloning and Analysis of Po-prdm14


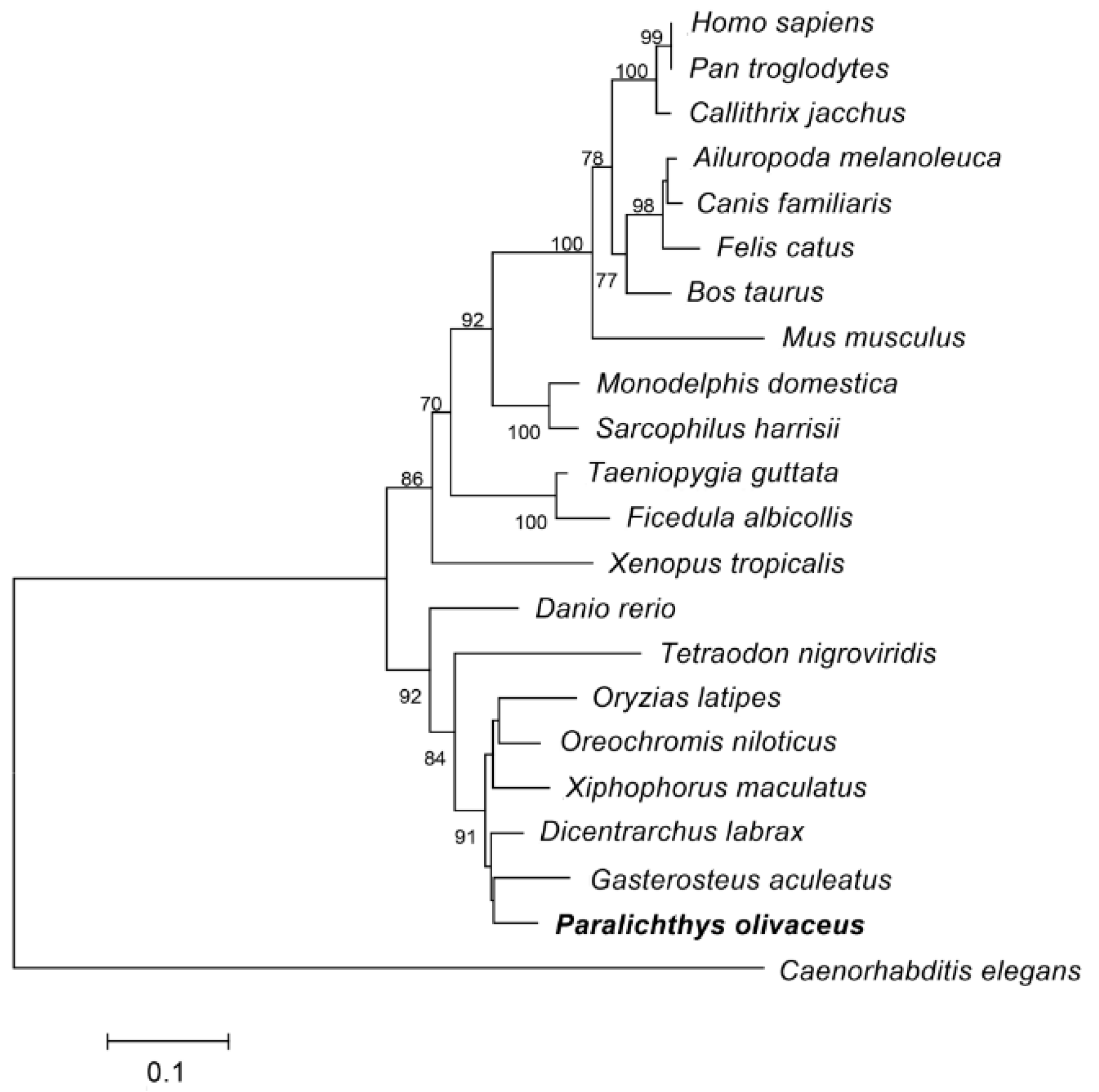
2.1.2. Homology Analysis of Po-PRDM14
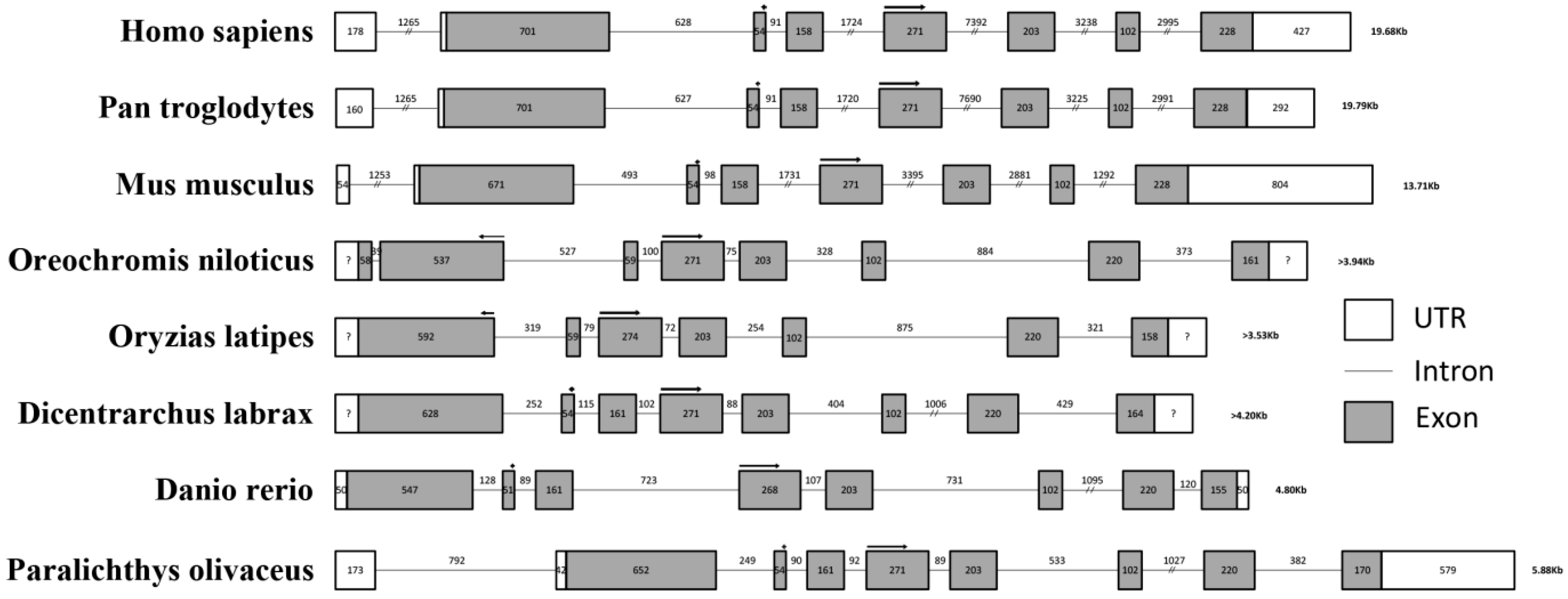
2.2. Genomic Analysis of Po-prdm14

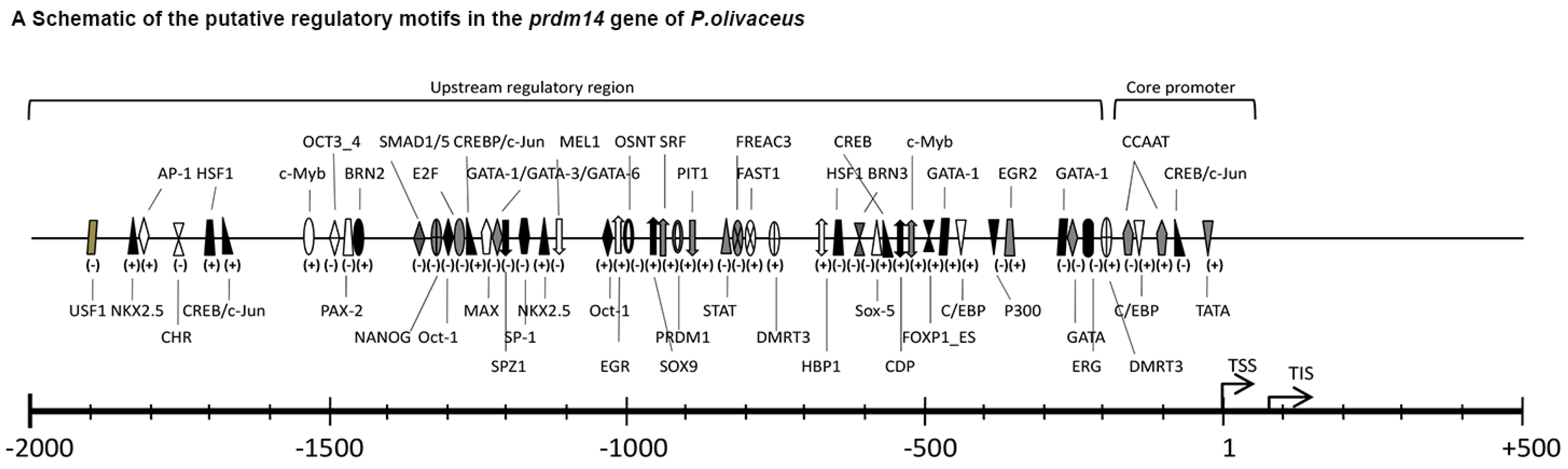

2.3. Expression and ISH of Po-prdm14 in Different Developmental Stages
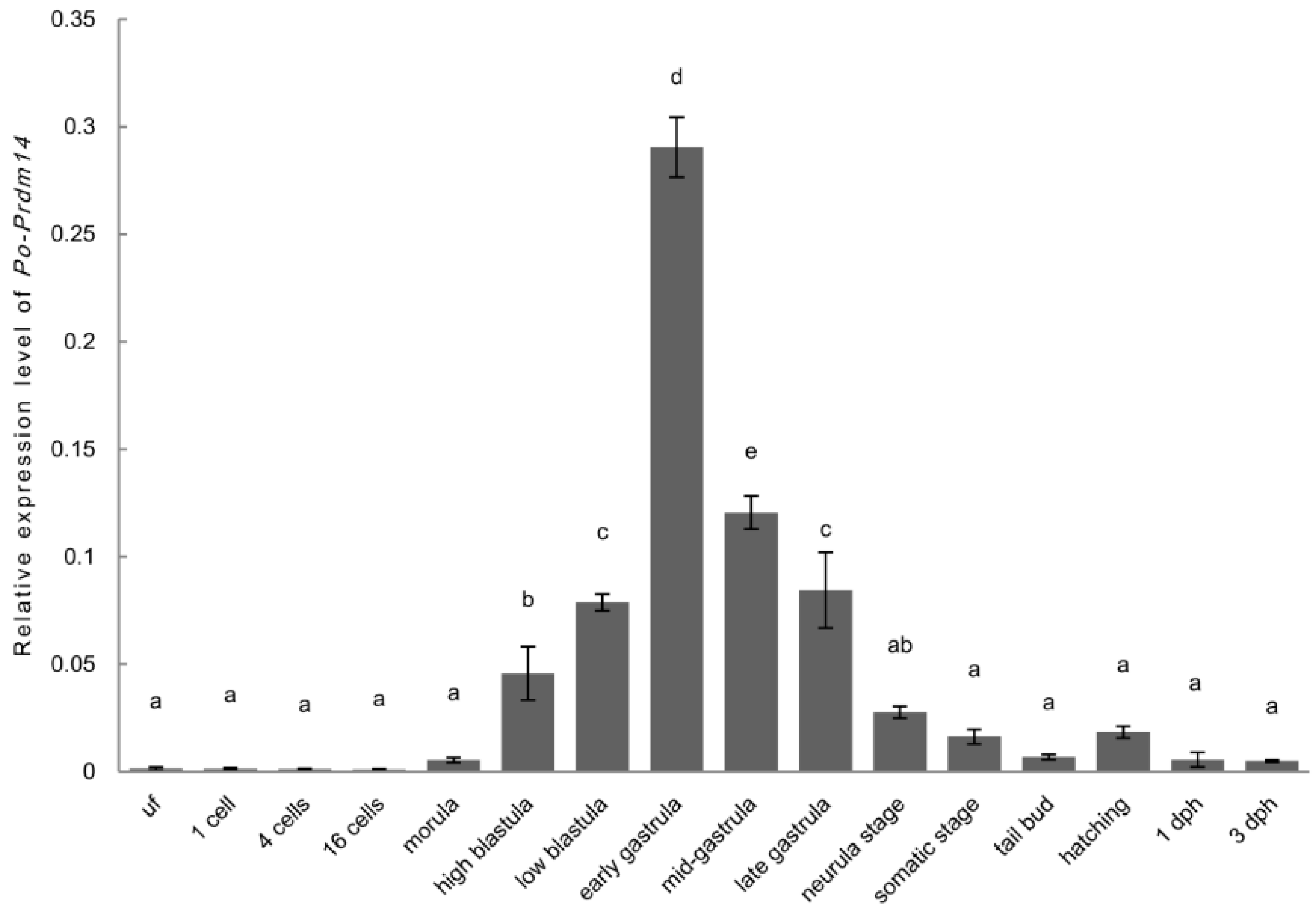
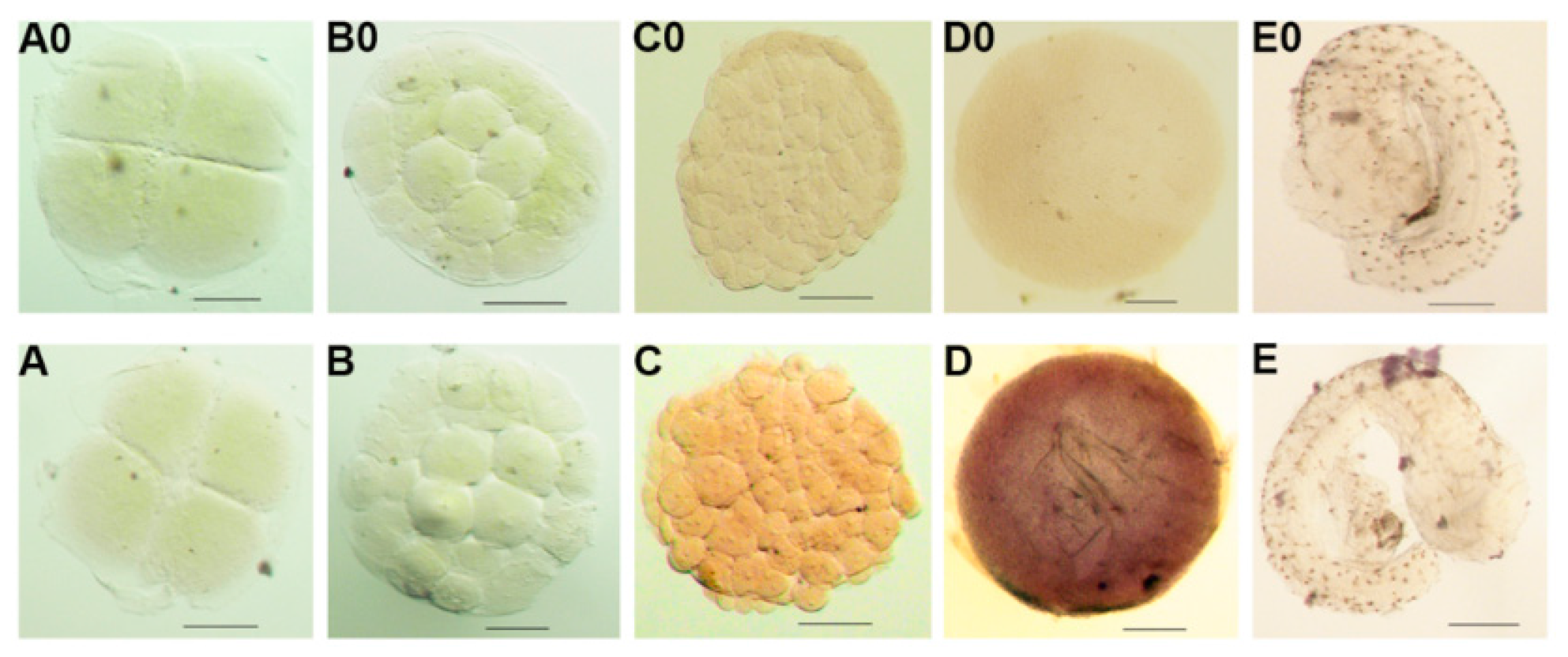
2.4. ISH of Po-prdm14 in the Gonad Sections
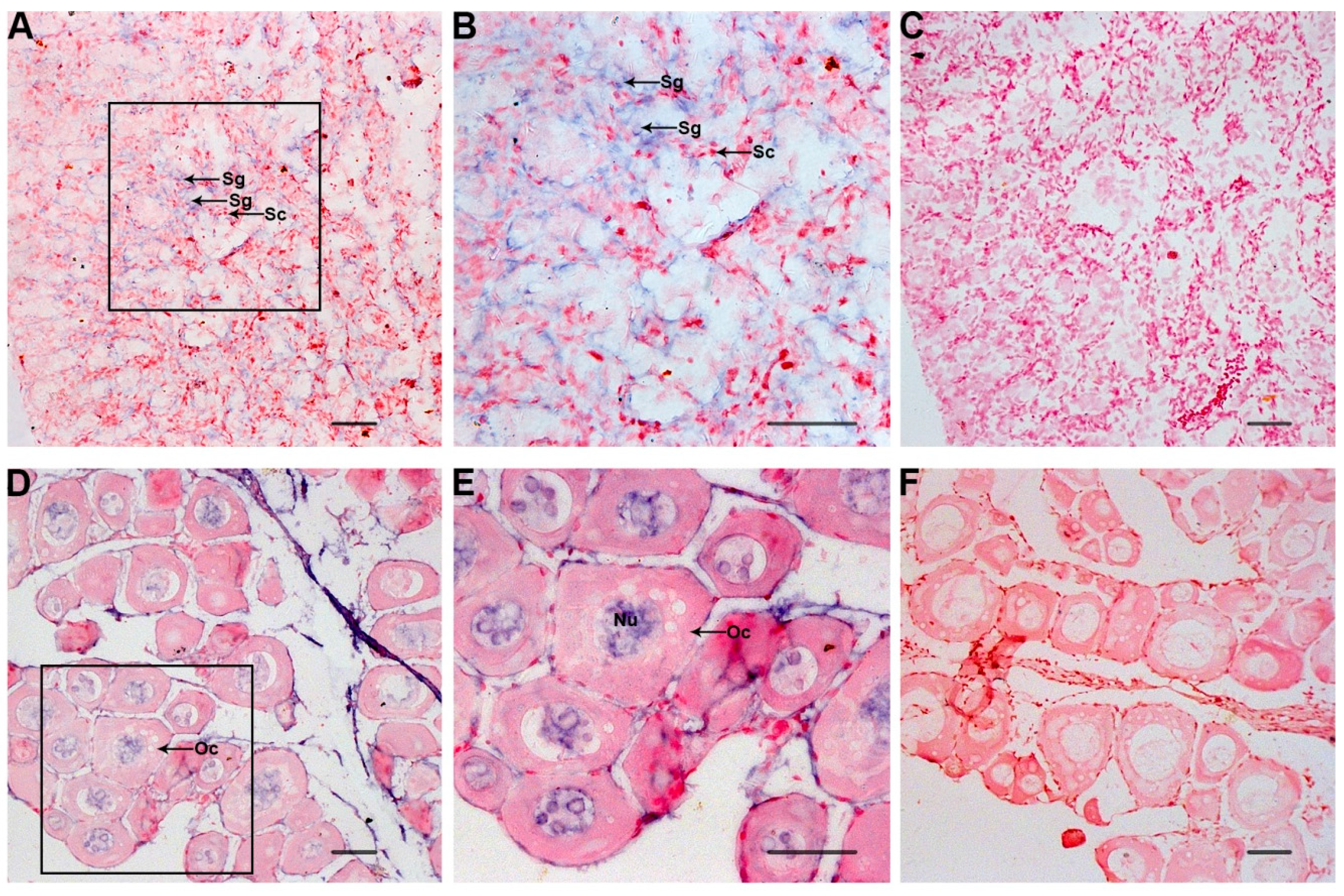
3. Discussion
4. Experimental Section
4.1. Embryo and Fish Collection
4.2. RNA and Genomic DNA Extraction
4.3. Cloning and Sequencing of the Po-prdm14 Gene
| Primers | Sequence (5'–3') | Usage |
|---|---|---|
| prdm14-core1-Fw | ATGGGTCACGCTCTATCTGG | Core fragment PCR |
| prdm14-core1-Rv | ATCCTCAGTCGGTCCCTCTT | Core fragment PCR |
| prdm14-core2-Fw | AGAAGAGGGACCGACTGAGG | Core fragment PCR |
| prdm14-core2-Rv | AGGACGGAAGGACTGTGCTG | Core fragment PCR |
| prdm14-ORF-Fw | CTCTGAAGTTTGCAGAAGTGG | Verify ORF |
| prdm14-ORF-Rv | AACGTTCACAAAAGCATCGC | Verify ORF |
| prdm14-DNA1-Fw | TTCCTCTATTTACTGGCTGATTGGT | Full-length PCR |
| prdm14-DNA1-Rv | CCGTTATCTGGAGAAGAGGACACAC | Full-length PCR |
| prdm14-DNA2-Fw | TTGAGAACGGCAGGTTGAGTC | Full-length PCR |
| prdm14-DNA2-Rv | GAAAGTCAAGGCGAATGATGCT | Full-length PCR |
| prdm14-DNA3-Fw | AGTCGTCCAGTCTCAACAAACACAT | Full-length PCR |
| prdm14-DNA3-Rv | GAGCGGAGGTCGTTGTGAGTAT | Full-length PCR |
| prdm14-promoter-Fw | ACACCCGCATCCACCTTC | Verify promoter |
| prdm14-promoter-Rv | AGTGGAGCGTGGTAGAAGC | Verify promoter |
| prdm14-3'RACE-1 | CGAGAAGAGGGACCGACTGAG | 3'RACE PCR |
| prdm14-3'RACE-2 | ACACATCCGCCAGCATTCC | 3'RACE PCR |
| prdm14-5'RACE-1 | TGAGAGGGATGCCCAGGAAC | 5'RACE PCR |
| prdm14-5'RACE-2 | GAGATGCTGCTCTTGTCCGTG | 5'RACE PCR |
| prdm14-5'RACE-3 | ATGAAGTGAGGCGGGTGGC | 5'RACE PCR |
| prdm14-RT-Fw | TCTGAGTCTCACATCCTTCCA | RT-qPCR |
| prdm14-RT-Rv | CTCAAACACCTCCCACATCAG | RT-qPCR |
| β-actin-Fw | GAGATGAAGCCCAGAGCAAGAG | Semi-RT-PCR |
| β-actin-Rv | CAGCTGTGGTGGTGAAGGAGTAG | Semi-RT-PCR |
| 18S-RT-Fw | GGTAACGGGGAATCAGGGT | RT-qPCR |
| 18S-RT-Rv | TGCCTTCCTTGGATGTGGT | RT-qPCR |
| UbcE-RT-Fw | TTACTGTCCATTTCCCCACTGAC | RT-qPCR |
| UbcE-RT-Rv | GACCACTGCGACCTCAAGATG | RT-qPCR |
| prdm14-ISH-Fw | ATTTAGGTGACACTATAGAAACAGCACAGTCCTTCCGTCCT | ISH-probe |
| prdm14-ISH-Rv | TAATACGACTCACTATAGGGTTTGCAGCCTTTCGTTACAGT | ISH-probe |
4.4. RT-qPCR Amplification
4.5. ISH Analysis
4.6. Sequence Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Huang, S.; Shao, G.; Liu, L. The PR domain of the Rb-binding zinc finger protein RIZ1 is a protein binding interface and is related to the SET domain functioning in chromatin-mediated gene expression. J. Biol. Chem. 1998, 273, 15933–15939. [Google Scholar] [CrossRef] [PubMed]
- Fog, C.K.; Galli, G.G.; Lund, A.H. PRDM proteins: Important players in differentiation and disease. BioEssays 2012, 34, 50–60. [Google Scholar] [CrossRef] [PubMed]
- Hohenauer, T.; Moore, A.W. The PRDM family: Expanding roles in stem cells and development. Development 2012, 139, 2267–2282. [Google Scholar] [CrossRef] [PubMed]
- Buyse, I.M.; Shao, G.; Huang, S. The retinoblastoma protein binds to RIZ, a zinc-finger protein that shares an epitope with the adenovirus E1A protein. Proc. Natl. Acad. Sci. USA 1995, 92, 4467–4471. [Google Scholar] [CrossRef] [PubMed]
- Huang, S. Histone methyltransferases, diet nutrients and tumour suppressors. Nat. Rev. Cancer 2002, 2, 469–476. [Google Scholar] [CrossRef] [PubMed]
- Schneider, R.; Bannister, A.J.; Kouzarides, T. Unsafe SETs: Histone lysine methyltransferases and cancer. Trends Biochem. Sci. 2002, 27, 396–402. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.J.; Xu, P.F.; Zhou, T.; Hu, M.; Fu, C.T.; Zhang, Y.; Jin, Y.; Chen, Y.; Chen, S.J.; Huang, Q.H. Genome-wide survey and developmental expression mapping of zebrafish SET domain-containing genes. PLoS ONE 2008, 3, e1499. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Min, J.H.; Lunin, V.V.; Antoshenko, T.; Dombrovski, L.; Zeng, H.; Allali-Hassani, A.; Campagna-Slater, V.; Vedadi, M.; Arrowsmith, C.H. Structural biology of human H3K9 methyltransferases. PLoS ONE 2010, 5, e8570. [Google Scholar] [CrossRef] [PubMed]
- Derunes, C.; Briknarova, K.; Geng, L.; Li, S.; Gessner, C.R.; Hewitt, K.; Wu, S.; Huang, S.; Woods, V.I., Jr.; Ely, K.R. Characterization of the PR domain of RIZ1 histone methyltransferase. Biochem. Biophys. Res. Commun. 2005, 333, 925–934. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Ferguson, J.E.; Wang, H.; Kelley, R.; Ren, R.; McDonough, H.; Meeker, J.; Charles, P.C.; Wang, H.; Patterson, C. PRDM6 is enriched in vascular precursors during development and inhibits endothelial cell proliferation, survival, and differentiation. J. Mol. Cell. Cardiol. 2008, 44, 47–58. [Google Scholar] [CrossRef] [PubMed]
- Eom, G.H.; Kim, K.; Kim, S.M.; Kee, H.J.; Kim, J.Y.; Jin, H.M.; Kim, J.R.; Kim, J.H.; Choe, N.; Kim, K.B. Histone methyltransferase PRDM8 regulates mouse testis steroidogenesis. Biochem. Biophys. Res. Commun. 2009, 388, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, I.; Margueron, R.; Shukeir, N.; Eisold, M.; Fritzsch, C.; Richter, F.M.; Mittler, G.; Genoud, C.; Goyama, S.; Kurokawa, M. PRDM3 and PRDM16 are H3K9me1 methyltransferases required for mammalian heterochromatin integrity. Cell 2012, 150, 948–960. [Google Scholar] [CrossRef] [PubMed]
- Gewies, A.; Castineiras-Vilarino, M.; Ferch, U.; Jahrling, N.; Heinrich, K.; Hoeckendorf, U.; Przemeck, G.K.; Munding, M.; Gross, O.; Schroeder, T. Prdm6 is essential for cardiovascular development in vivo. PLoS ONE 2013, 8, e81833. [Google Scholar] [CrossRef] [PubMed]
- Koh-Stenta, X.; Joy, J.; Poulsen, A.; Li, R.; Tan, Y.; Shim, Y.; Min, J.H.; Wu, L.; Ngo, A.; Peng, J. Characterisation of the histone methyltransferase PRDM9 utilising biochemical, biophysical and chemical biology techniques. Biochem. J. 2014, 461, 323–334. [Google Scholar] [CrossRef] [PubMed]
- Chan, Y.S.; Goke, J.; Lu, X.; Venkatesan, N.; Feng, B.; Su, I.H.; Ng, H.H. A PRC2-dependent repressive role of PRDM14 in human embryonic stem cells and induced pluripotent stem cell reprogramming. Stem Cells 2012, 31, 682–692. [Google Scholar] [CrossRef]
- Fumasoni, I.; Meani, N.; Rambaldi, D.; Scafetta, G.; Alcalay, M.; Ciccarelli, F.D. Family expansion and gene rearrangements contributed to the functional specialization of PRDM genes in vertebrates. BMC Evol. Biol. 2007, 7, 187. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Swigut, T.; Valouev, A.; Rada-Iglesias, A.; Wysocka, J. Sequence-specific regulator PRDM14 safeguards mouse ESCs from entering extraembryonic endoderm fates. Nat. Struct. Mol. Biol. 2011, 18, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Yamaji, M.; Ueda, J.; Hayashi, K.; Ohta, H.; Yabuta, Y.; Kurimoto, K.; Nakato, R.; Yamada, Y.; Shirahige, K.; Saitou, M. PRDM14 ensures naive pluripotency through dual regulation of signaling and epigenetic pathways in mouse embryonic stem cells. Cell Stem Cell 2013, 12, 368–382. [Google Scholar] [CrossRef] [PubMed]
- Keller, A.D.; Maniatis, T. Only two of the five zinc fingers of the eukaryotic transcriptional repressor PRDI-BF1 are required for sequence-specific DNA binding. Mol. Cell. Biol. 1992, 12, 1940–1949. [Google Scholar] [PubMed]
- Myers, S.; Bowden, R.; Tumian, A.; Bontrop, R.E.; Freeman, C.; MacFie, T.S.; McVean, G.; Donnelly, P. Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science 2010, 327, 876–879. [Google Scholar] [CrossRef] [PubMed]
- Chia, N.Y.; Chan, Y.S.; Feng, B.; Lu, X.; Orlov, Y.L.; Moreau, D.; Kumar, P.; Yang, L.; Jiang, J.; Lau, M.S. A genome-wide RNAi screen reveals determinants of human embryonic stem cell identity. Nature 2010, 468, 316–320. [Google Scholar] [CrossRef] [PubMed]
- Payer, B.; Rosenberg, M.; Yamaji, M.; Yabuta, Y.; Koyanagi-Aoi, M.; Hayashi, K.; Yamanaka, S.; Saitou, M.; Lee, J.T. Tsix RNA and the germline factor, PRDM14, link X reactivation and stem cell reprogramming. Mol. Cell 2013, 52, 805–818. [Google Scholar] [CrossRef] [PubMed]
- Gillich, A.; Bao, S.; Grabole, N.; Hayashi, K.; Trotter, M.W.; Pasque, V.; Magnusdottir, E.; Surani, M.A. Epiblast stem cell-based system reveals reprogramming synergy of germline factors. Cell Stem Cell 2012, 10, 425–439. [Google Scholar] [CrossRef] [PubMed]
- Yamaji, M.; Seki, Y.; Kurimoto, K.; Yabuta, Y.; Yuasa, M.; Shigeta, M.; Yamanaka, K.; Ohinata, Y.; Saitou, M. Critical function of Prdm14 for the establishment of the germ cell lineage in mice. Nat. Genet. 2008, 40, 1016–1022. [Google Scholar] [CrossRef] [PubMed]
- Margueron, R.; Reinberg, D. The polycomb complex PRC2 and its mark in life. Nature 2011, 469, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Burton, A.; Muller, J.; Tu, S.; Padilla-Longoria, P.; Guccione, E.; Torres-Padilla, M.E. Single-cell profiling of epigenetic modifiers identifies PRDM14 as an inducer of cell fate in the mammalian embryo. Cell Rep. 2013, 5, 687–701. [Google Scholar] [CrossRef] [PubMed]
- Okashita, N.; Kumaki, Y.; Ebi, K.; Nishi, M.; Okamoto, Y.; Nakayama, M.; Hashimoto, S.; Nakamura, T.; Sugasawa, K.; Kojima, N. PRDM14 promotes active DNA demethylation through the Ten-eleven translocation (TET)-mediated base excision repair pathway in embryonic stem cells. Development 2014, 141, 269–280. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Ma, W.; Su, W.; Zhang, J. PRDM14 acts upstream of islet2 transcription to regulate axon growth of primary motoneurons in zebrafish. Development 2012, 139, 4591–4600. [Google Scholar] [CrossRef] [PubMed]
- Bʼejar, J.; Hong, Y.H.; Alvarez, M.C. An ES-like cell line from the marine fish Sparus aurata: Characterization and chimaera production. Transgenic Res. 2002, 11, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Yi, M.; Hong, N.; Hong, Y. Generation of medaka fish haploid embryonic stem cells. Science 2009, 326, 430–433. [Google Scholar] [CrossRef] [PubMed]
- Parameswaran, V.; Shukla, R.; Bhonde, R.; Hameed, A.S. Development of a pluripotent ES-like cell line from Asian sea bass (Lates calcarifer)—An oviparous stem cell line mimicking viviparous ES cells. Mar. Biotechnol. 2007, 9, 766–775. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.H.; Schartl, M. Isolation and differentiation of medaka embryonic stem cells. Methods Mol. Biol. 2006, 329, 3–16. [Google Scholar] [PubMed]
- Chen, S.L.; Sha, Z.X.; Ye, H.Q. Establishment of a pluripotent embryonic cell line from sea perch (Lateolabrax japonicus) embryos. Aquaculture 2003, 218, 141–151. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alvarez, M.C.; Bejar, J.; Chen, S.; Hong, Y. Fish ES cells and applications to biotechnology. Mar. Biotechnol. 2007, 9, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Morita, T.; Kumakura, N.; Morishima, K.; Mitsuboshi, T.; Ishida, M.; Hara, T.; Kudo, S.; Miwa, M.; Ihara, S.; Higuchi, K. Production of donor-derived offspring by allogeneic transplantation of spermatogonia in the yellowtail (Seriola quinqueradiata). Biol. Reprod. 2012, 86, 176. [Google Scholar] [CrossRef] [PubMed]
- Matsui, Y.; Zsebo, K.; Hogan, B.L.M. Derivation of pluripotential embryonic stem cells from murine primordial germ cells in culture. Cell 1992, 70, 841–847. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Wang, J.; Jiang, J.; Fan, L.; Wang, W.; Liu, J.; Zhang, Q.; Wang, X. Identification and characterization of a nanog homolog in Japanese flounder (Paralichthys olivaceus). Gene 2013, 531, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Nakaki, F.; Saitou, M. PRDM14: A unique regulator for pluripotency and epigenetic reprogramming. Trends Biochem. Sci. 2014, 39, 289–298. [Google Scholar] [CrossRef] [PubMed]
- Ng, H.H.; Chia, N.Y.; Feng, B. Method for inducing pluripotency in human somatic cells with PRDM14 or NFRKB. U. S. Patent US20130078720 A1, 28 March 2013. [Google Scholar]
- Ravi, V.; Venkatesh, B. Rapidly evolving fish genomes and teleost diversity. Curr. Opin. Genet. Dev. 2008, 18, 544–550. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.S.; Peer, Y.V.; Braasch, I.; Meyer, M. comparative genomics provides evidence for an ancient genome duplication event in fish. Philos. Trans. R. Soc. Lond. B 2001, 356, 1661–1679. [Google Scholar] [CrossRef]
- Semon, M.; Wolfe, K.H. Rearrangement rate following the whole-genome duplication in teleosts. Mol. Biol. Evol. 2007, 24, 860–867. [Google Scholar] [CrossRef] [PubMed]
- Boyer, L.A.; Lee, T.I.; Cole, M.F.; Johnstone, S.E.; Levine, S.S.; Zucker, J.P.; Guenther, M.G.; Kumar, R.M.; Murray, H.L.; Jenner, R.G. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005, 122, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Ohinata, Y.; Ohta, H.; Shigeta, M.; Yamanaka, K.; Wakayama, T.; Saitou, M. A signaling principle for the specification of the germ cell lineage in mice. Cell 2009, 137, 571–584. [Google Scholar] [CrossRef] [PubMed]
- Kurimoto, K.; Yamaji, M.; Seki, Y.; Saitou, M. Specification of the germ cell lineage in mice. Cell Cycle 2008, 7, 3514–3518. [Google Scholar] [CrossRef] [PubMed]
- Saitou, M.; Barton, S.C.; Surani, M.A. A molecular programme for the specification of germ cell fate in mice. Nature 2002, 418, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Saitou, M. Germ cell specification in mice. Curr. Opin. Genet. Dev. 2009, 19, 386–395. [Google Scholar] [CrossRef] [PubMed]
- Yabuta, Y.; Kurimoto, K.; Ohinata, Y.; Seki, Y.; Saitou, M. Gene expression dynamics during germline specification in mice identified by quantitative single-cell gene expression profiling. Biol. Reprod. 2006, 75, 705–716. [Google Scholar] [CrossRef] [PubMed]
- Seki, Y.; Yamaji, M.; Yabuta, Y.; Sano, M.; Shigeta, M.; Matsui, Y.; Saga, Y.; Tachibana, M.; Shinkai, Y.; Saitou, M. Cellular dynamics associated with the genome-wide epigenetic reprogramming in migrating primordial germ cells in mice. Development 2007, 134, 2627–2638. [Google Scholar] [CrossRef] [PubMed]
- Seki, Y.; Hayashi, K.; Itoh, K.; Mizugaki, M.; Saitou, M.; Matsui, Y. Extensive and orderly reprogramming of genome-wide chromatin modifications associated with specification and early development of germ cells in mice. Dev. Biol. 2005, 278, 440–458. [Google Scholar] [CrossRef] [PubMed]
- Grabole, N.; Tischler, J.; Hackett, J.A.; Kim, S.; Tang, F.; Leitch, H.G.; Magnusdottir, E.; Surani, M.A. Prdm14 promotes germline fate and naive pluripotency by repressing FGF signalling and DNA methylation. EMBO Rep. 2013, 14, 629–637. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Yang, M.; Guo, H.; Yang, L.; Wu, J.; Li, R.; Liu, P.; Lian, Y.; Zheng, X.; Yan, J. Single-cell RNA-Seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 2013, 20, 1131–1139. [Google Scholar] [CrossRef] [PubMed]
- Xue, Z.; Huang, K.; Cai, C.; Cai, L.; Jiang, C.Y.; Feng, Y.; Liu, Z.; Zeng, Q.; Cheng, L.; Sun, Y.E. Genetic programs in human and mouse early embryos revealed by single-cell RNA sequencing. Nature 2013, 500, 593–597. [Google Scholar] [CrossRef] [PubMed]
- Tsuneyoshi, N.; Sumi, T.; Onda, H.; Nojima, H.; Nakatsuji, N.; Suemori, H. PRDM14 suppresses expression of differentiation marker genes in human embryonic stem cells. Biochem. Biophys. Res. Commun. 2008, 367, 899–905. [Google Scholar] [CrossRef] [PubMed]
- Assou, S.; le Carrour, T.; Tondeur, S.; Strom, S.; Gabelle, A.; Marty, S.; Nadal, L.; Pantesco, V.; Reme, T.; Hugnot, J.P. A meta-analysis of human embryonic stem cells transcriptome integrated into a web-based expression atlas. Stem Cells 2007, 25, 961–973. [Google Scholar] [CrossRef] [PubMed]
- Nishikawa, N.; Toyota, M.; Suzuki, H.; Honma, T.; Fujikane, T.; Ohmura, T.; Nishidate, T.; Ohe-Toyota, M.; Maruyama, R.; Sonoda, T. Gene amplification and overexpression of PRDM14 in breast cancers. Cancer Res. 2007, 67, 9649–9657. [Google Scholar] [CrossRef] [PubMed]
- Detman, E.J.; Justice, M.J. The zinc finger SET domain gene Prdm14 is overexpressed in lymphoblastic lymphomas with retroviral insertions at Evi32. PLoS ONE 2008, 3, e3823. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989. [Google Scholar]
- Zhong, Q.; Zhang, Q.; Wang, Z.; Qi, J.; Chen, Y.; Li, S.; Sun, Y.; Li, C.; Lan, X. Expression profiling and validation of potential reference genes during Paralichthys olivaceus embryogenesis. Mar. Biotechnol. 2008, 10, 310–318. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.; Milpetz, F.; Bork, P.; Ponting, C.P. SMART, a simple modular architecture research tool: Identification of signaling domains. Proc. Natl. Acad. Sci. USA 1998, 95, 5857–5864. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res. 2012, 40, D302–D305. [Google Scholar] [CrossRef]
- Cartharius, K.; Frech, K.; Grote, K.; Klocke, B.; Haltmeier, M.; Klingenhoff, A.; Frisch, M.; Bayerlein, M.; Werner, T. MatInspector and beyond: Promoter analysis based on transcription factor binding sites. Bioinformatics 2005, 21, 2933–2942. [Google Scholar] [CrossRef] [PubMed]
- Tsunoda, T.; Takagi, T. Estimating transcription factor bindability on DNA. Bioinformatics 1999, 15, 622–630. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fan, L.; Jiang, J.; Gao, J.; Song, H.; Liu, J.; Yang, L.; Li, Z.; Chen, Y.; Zhang, Q.; Wang, X. Identification and Characterization of a PRDM14 Homolog in Japanese Flounder (Paralichthys olivaceus). Int. J. Mol. Sci. 2015, 16, 9097-9118. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms16059097
Fan L, Jiang J, Gao J, Song H, Liu J, Yang L, Li Z, Chen Y, Zhang Q, Wang X. Identification and Characterization of a PRDM14 Homolog in Japanese Flounder (Paralichthys olivaceus). International Journal of Molecular Sciences. 2015; 16(5):9097-9118. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms16059097
Chicago/Turabian StyleFan, Lin, Jiajun Jiang, Jinning Gao, Huayu Song, Jinxiang Liu, Likun Yang, Zan Li, Yan Chen, Quanqi Zhang, and Xubo Wang. 2015. "Identification and Characterization of a PRDM14 Homolog in Japanese Flounder (Paralichthys olivaceus)" International Journal of Molecular Sciences 16, no. 5: 9097-9118. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms16059097