Identification and Characterization of MicroRNAs in Snakehead Fish Cell Line upon Snakehead Fish Vesiculovirus Infection
Abstract
:1. Introduction
2. Results
2.1. Overview of the Illumine Hiseq2500 Sequencing Data of Small RNAs

2.2. Identification of Conserved miRNAs in SSN-1 Cells
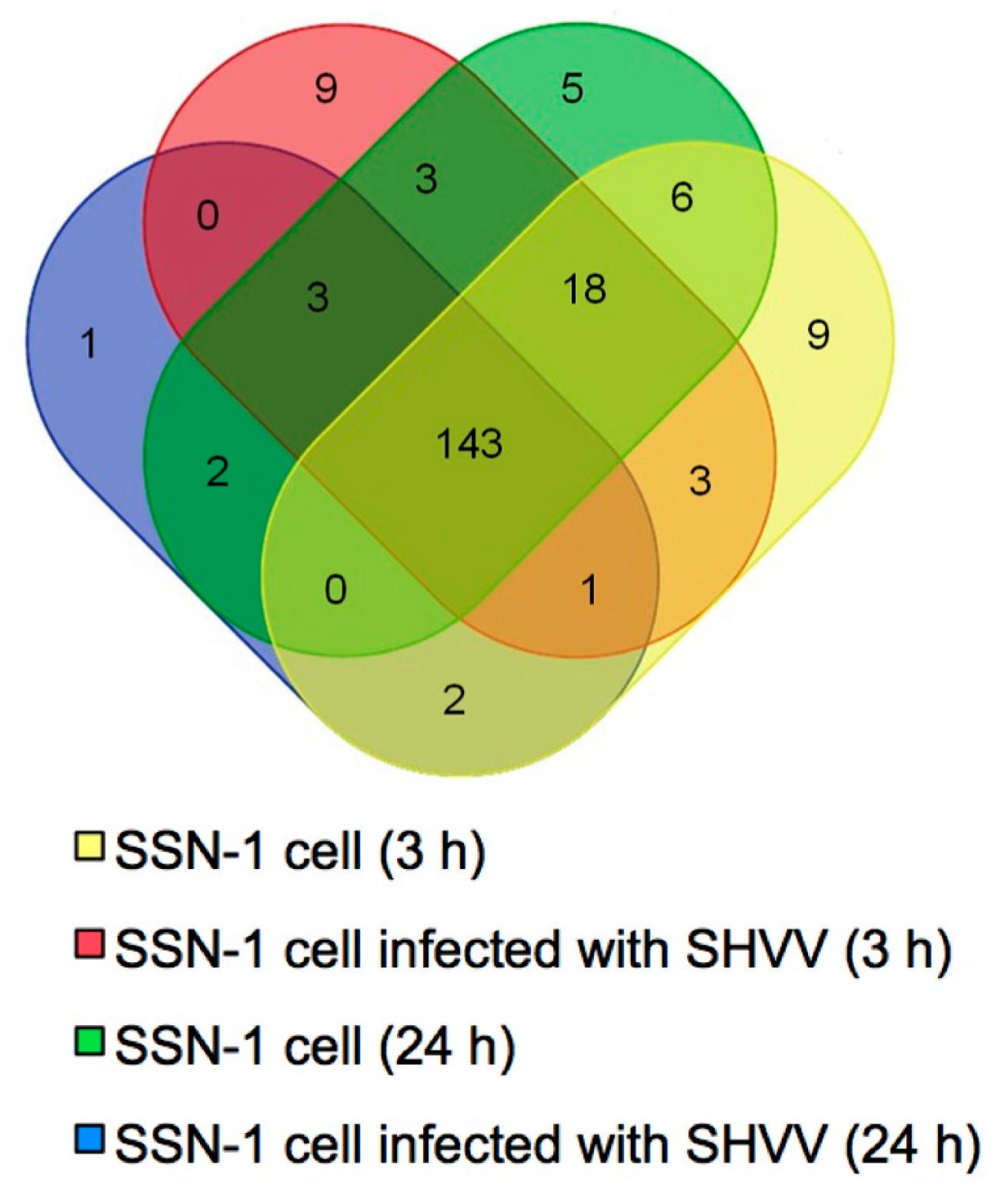
2.3. Prediction and Validation of Novel miRNAs
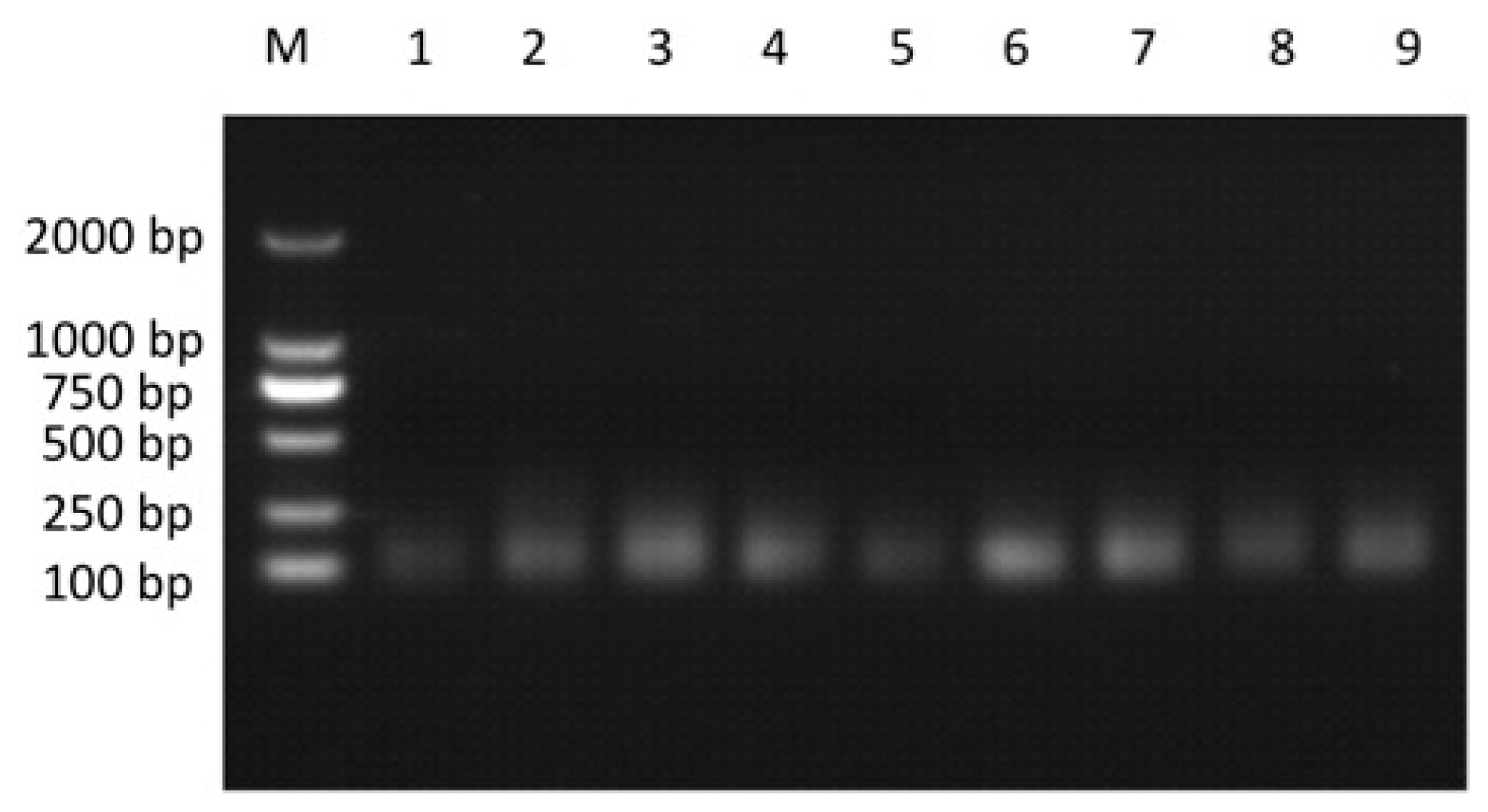
| Sequence (5′→3′) | Mature Reads | Fold Change (Log2 I24/C24) * | p-Value | |
|---|---|---|---|---|
| 1 | UCCAUCAGUCACGUGACCUAC | 26 | −2.15182 | 2.03 × 10−16 |
| 2 | UCGGGUCGCUAAUGACGUCACC | 30 | −3.57979 | 4.64 × 10−21 |
| 3 | ACCAGGUGCUGUAAGCUU | 67 | <1 | – |
| 4 | CUUUUAAUCUGAGGGUCCA | 12 | <1 | – |
| 5 | AUGACUCGAACCCGAGGACUCG | 13 | 1.57979 | 0.019847 |
| 6 | AUCCGGCUCGAAGGACCAA | 157 | −4.36217 | 0 |
| 7 | AAACACUGCCAGCUGCCACA | 5 | −3.67933 | 0.00165 |
| 8 | GGGGCCUGAGUCCUUCUG | 17 | 13.75742 | 0 |
| 9 | ACCCCACUCCUGGUACCA | 51 | −4.59089 | 8.16 × 10−197 |
2.4. Differentially-Expressed miRNAs and Validation of the miRNAs by qRT-PCR
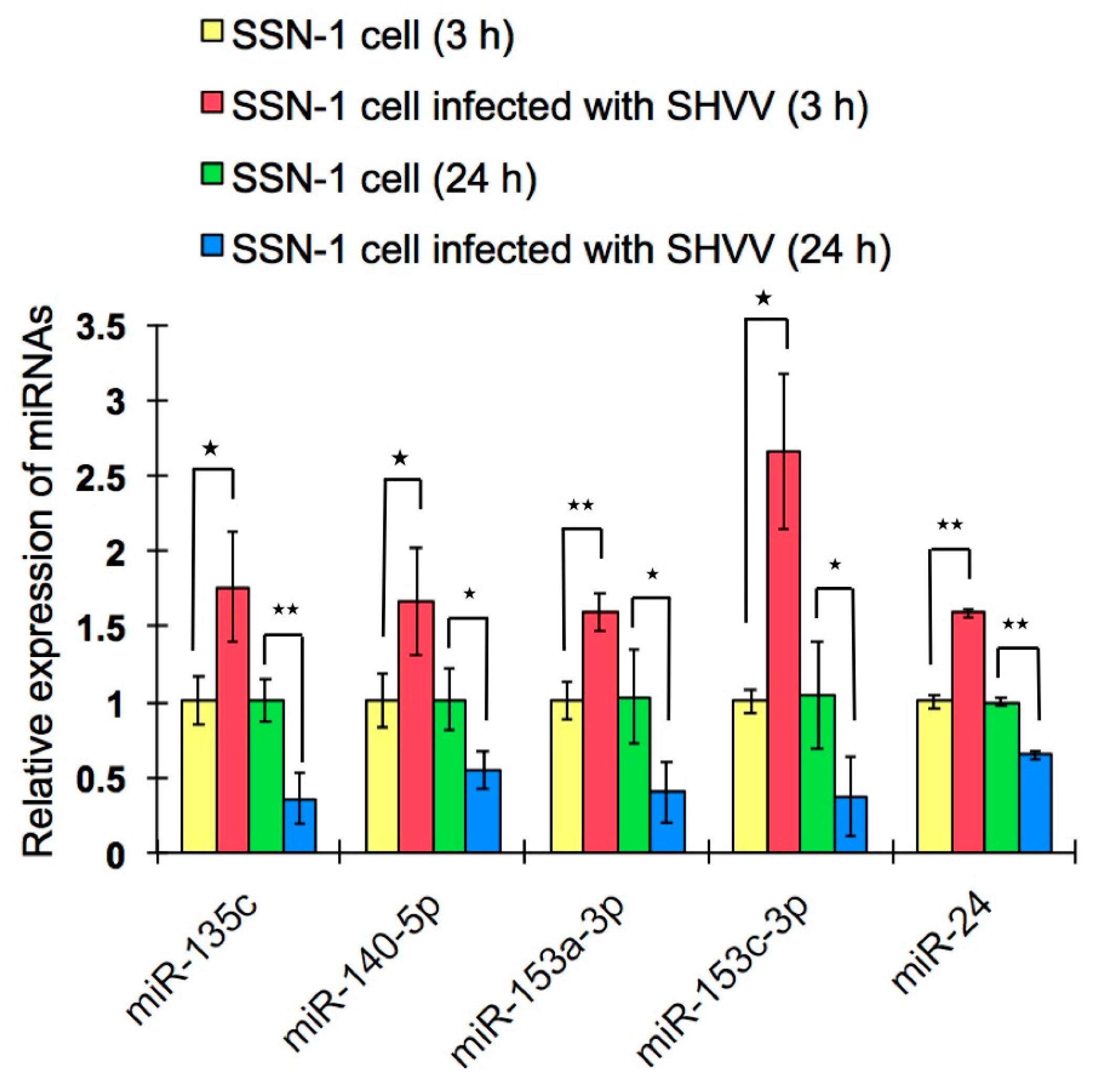
2.5. miRNA Targets Prediction
| Name | Sequence (5′→3′) | Target | Fold Change (Log2 I24/C24) * |
|---|---|---|---|
| miR-23a-3-5p | GGAUUCCUGGCAGAGUGAUUU | N, L | −7.92953 |
| miR-199-5p | CCCAGUGUUCAGACUACCUGUUC | N, L | −3.96545 |
| miR-338 | UCCAGCAUCAGUGAUUUUGUUG | N, L | −4.16476 |
| miR-145-3p | GGAUUCCUGGAAAUACUGUUCU | N, L | −6.05784 |
| miR-100-3p | CAAGCUUGUAUCUAUAGGUAUC | N, G | −3.61793 |
| miR-216b | UAAUCUCUGCAGGCAACUGUGA | N, G, L | −6.73689 |
| miR-130c-5p | GCCCUUUUUCUGUUGUACUACU | N, G, L | −4.79246 |
| miR-214 | ACAGCAGGCACAGACAGGCAG | N, P | −4.27387 |
| miR-731 | AAUGACACGUUUUCUCCCGGAUCG | N | −3.18425 |
| miR-29b | UAGCACCAUUUGAAAUCAGUGU | P | −3.20471 |
| miR-29a | UAGCACCAUUUGAAAUCGGUUA | P | −3.25812 |
| miR-135c | UAUGGCUUUCUAUUCCUAUGUG | M | −1.6469 |
| miR-145-5p | GUCCAGUUUUCCCAGGAAUCCC | G | −7.017 |
| miR-92b-3p | UAUUGCACUCGUCCCGGCCUCC | G | −4.75081 |
| miR-153b-3p | UUGCAUAGUCACAAAAAUGAGC | G, L | −2.41897 |
| miR-7147 | UGUACCAUGCUGGUAGCCAGU | G | −4.7948 |
| miR-92a-3p | UAUUGCACUUGUCCCGGCCUGU | G | −3.97976 |
| miR-184 | UGGACGGAGAACUGAUAAGGGC | G | −2.16476 |
| miR-301c-3p | CAGUGCAAUAGUAUUGUCAUAG | G | −2.45693 |
| miR-363-3p | AAUUGCACGGUAUCCAUCUGUA | G | −3.26429 |
| miR-454b | UAGUGCAAUAUUGCUUAUAGGG | G | −2.98676 |
| miR-301a | CAGUGCAAUAGUAUUGUCAAAG | G | −2.1425 |
| miR-130a | CAGUGCAAUGUUAAAAGGGCAU | G | −3.11229 |
| miR-23a-5p | GAAUUCCUGGCAGAGUGAUUU | G, L | −6.09946 |
| miR-153a-3p | UUGCAUAGUCACAAAAGUGAUC | G, L | −2.95278 |
| miR-25-3p | CAUUGCACUUGUCUCGGUCUGA | G | −2.31239 |
| miR-730 | UCCUCAUUGUGCAUGCUGUGUGU | G | −4.60532 |
| miR-204-5p | UUCCCUUUGUCAUCCUAUGCCU | G, L | −3.14817 |
| miR-301b-3p | CAGUGCAAUAGUAUUGUCAUUG | G | −4.92954 |
| miR-724 | UUAAAGGGAAUUUGCGACUGUU | L | −2.63081 |
| miR-181c-5p | CACAUUCAUUGCUGUCGGUGGG | L | −1.84696 |
| miR-125a | UCCCUGAGACCCUUAACCUGUG | L | −3.16476 |
| miR-183-5p | UAUGGCACUGGUAGAAUUCACUG | L | −1.57979 |
| miR-199-3p | UACAGUAGUCUGCACAUUGGUU | L | −2.97786 |
| miR-26a-2-3p | CCUAUUCAUGAUUACUUGCACU | L | −3.8599 |
| miR-140-3p | UACCACAGGGUAGAACCACGGAC | L | −3.03951 |
| miR-146a | UGAGAACUGAAUUCCAUAGAUGG | L | −1.98658 |
| miR-150 | UCUCCCAAUCCUUGUACCAGUG | L | −2.63751 |
| miR-221-3p | AGCUACAUUGUCUGCUGGGUUUC | L | −2.48955 |
| miR-34a | UGGCAGUGUCUUAGCUGGUUGU | L | −2.5756 |
| miR-101b | UACAGUACUAUGAUAACUGAAG | L | −2.73747 |
| miR-9-5p | UCUUUGGUUAUCUAGCUGUAUGA | L | −1.54182 |
| miR-181b-5p | AACAUUCAUUGCUGUCGGUGGG | L | −1.75554 |
| miR-107a-3p | AGCAGCAUUGUACAGGGCUAUCA | L | −2.81985 |
| miR-101a | UACAGUACUGUGAUAACUGAAG | L | −3.05599 |
| miR-181c-3p | CUCGCCGGACAAUGAAUGAGAA | L | −5.01275 |
| miR-103 | AGCAGCAUUGUACAGGGCUAUGA | L | −2.61915 |
| miR-146b | UGAGAACUGAAUUCCAAGGGUG | L | −3.91539 |
| miR-138-5p | AGCUGGUGUUGUGAAUCAGGCC | L | −4.94236 |
| miR-429a | UAAUACUGUCUGGUAAUGCCGU | L | −2.44487 |
| miR-153b-5p | GUCAUUUUUGUGGUUUGCAGCU | L | −3.67933 |
| miR-125c-5p | UCCCUGAGACCCUAACUCGUGA | L | −5.07926 |
| miR-181a-5p | AACAUUCAACGCUGUCGGUGAGU | L | −1.97076 |
| miR-199-3-3p | ACAGUAGUCCGCACAUUGGUU | L | −3.81683 |
| miR-125b-5p | UCCCUGAGACCCUAACUUGUGA | L | −3.43276 |
| miR-26a-5p | UUCAAGUAAUCCAGGAUAGGCU | L | −3.35436 |
| miR-722 | UUUUUUGCAGAAACGUUUCAGAUU | L | −6.25146 |
| miR-222a-3p | AGCUACAUCUGGCUACUGGGUCUC | L | −3.55456 |
2.6. Effects of Three Selected miRNAs on SHVV Multiplication
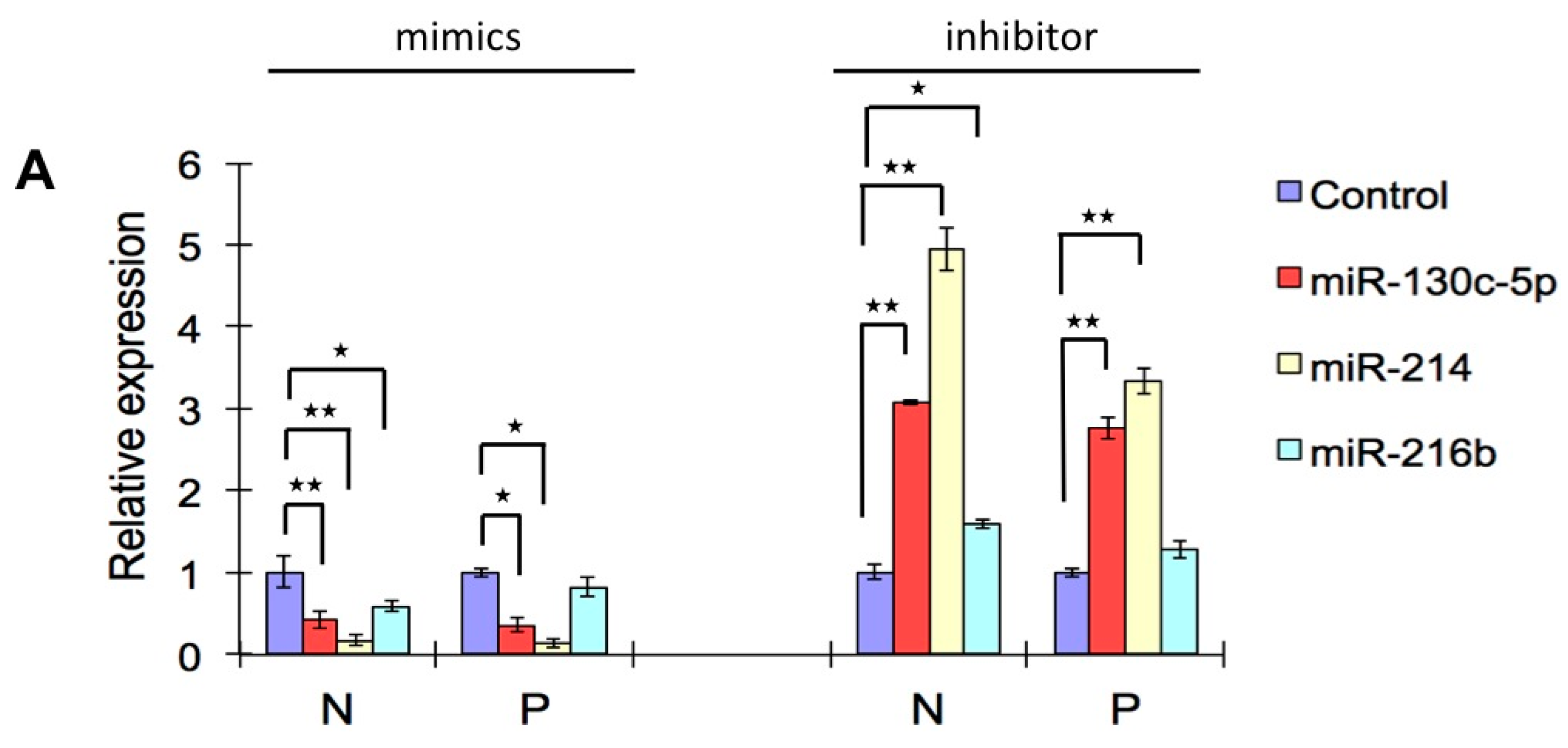
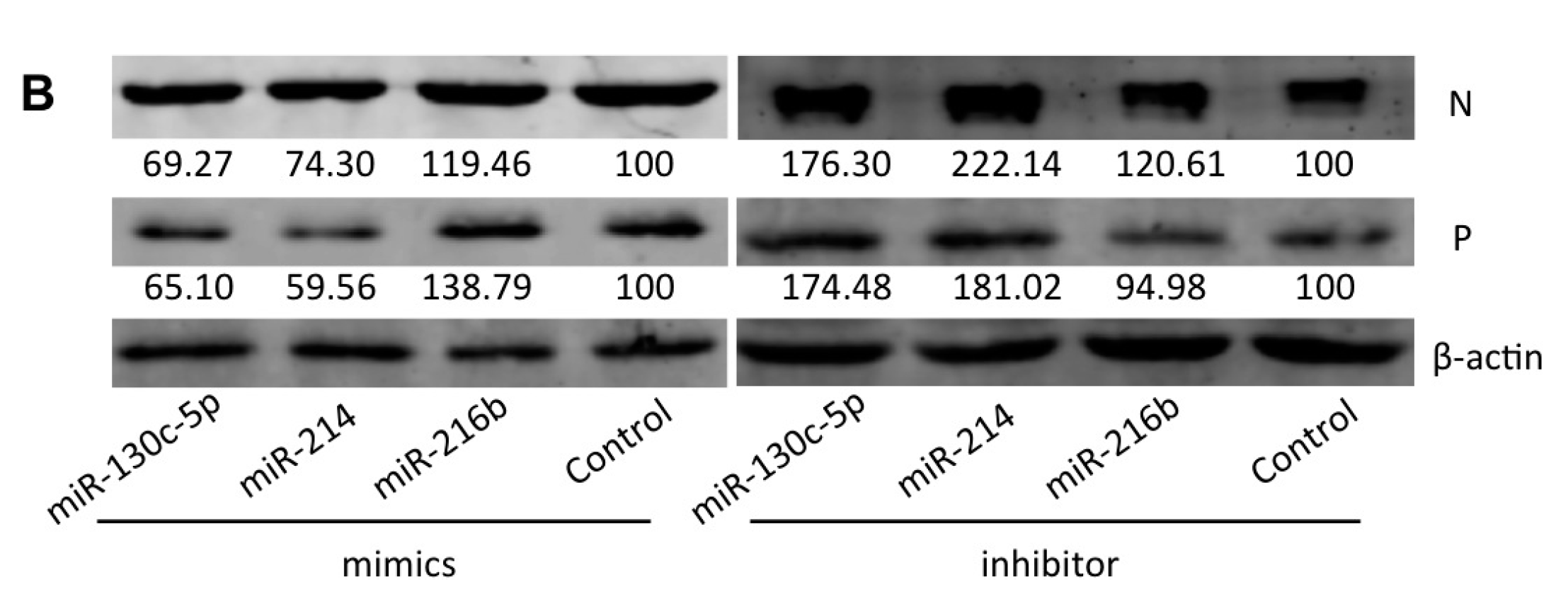
3. Discussion
4. Experimental Section
4.1. Virus, Cells and RNA Preparation
4.2. Small RNA Library Construction
4.3. Sequencing Data Analysis
4.4. Identification of Novel miRNAs
4.5. Analysis of Differentially-Expressed miRNAs
4.6. Prediction of miRNA Targets on the SHVV Genome
4.7. RT-PCR and qRT-PCR Analysis of miRNAs
4.8. Effects of Three Randomly-Selected Differentially-Expressed miRNAs on Virus Multiplication
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Fan, H.X.; Tang, H. Complex interactions between microRNAs and hepatitis B/C viruses. World J. Gastroenterol. 2014, 20, 13477–13492. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.G.; Yu, J.F.; Zhang, Y.; Gong, D.Q.; Gu, Z.L. Identification and characterization of microRNA from chicken adipose tissue and skeletal muscle. Poult. Sci. 2012, 91, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Djuranovic, S.; Nahvi, A.; Green, R. miRNA-mediated gene silencing by translational repression followed by mRNA deadenylation and decay. Science 2012, 336, 237–240. [Google Scholar] [CrossRef] [PubMed]
- Trakooljul, N.; Hicks, J.A.; Liu, H.C. Identification of target genes and pathways associated with chicken microRNA miR-143. Anim. Genet. 2010, 41, 357–364. [Google Scholar] [PubMed]
- Bartel, D.P. MicroRNAs, target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Doench, J.G.; Sharp, P.A. Specificity of microRNA target selection in translational repression. Genes Dev. 2004, 18, 504–511. [Google Scholar] [CrossRef] [PubMed]
- Cullen, B.R. Five questions about viruses and microRNAs. PLoS Pathog. 2010, 6, e1000787. [Google Scholar] [CrossRef] [PubMed]
- Skalsky, R.L.; Cullen, B.R. Viruses, microRNAs, and host interactions. Annu. Rev. Microbiol. 2010, 64, 123–141. [Google Scholar] [CrossRef] [PubMed]
- Cullen, B.R. Viruses and microRNAs, RISCy interactions with serious consequences. Genes Dev. 2011, 25, 1881–1894. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Calado, D.P.; Galler, G.; Thai, T.H.; Patterson, H.C.; Wang, J.; Rajewsky, N.; Bender, T.P.; Rajewsky, K. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 2007, 131, 146–159. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Vernooy, S.Y.; Guo, M.; Hay, B.A. The Drosophila microRNA Mir-14 suppresses cell death and is required for normal fat metabolism. Curr. Biol. 2003, 13, 790–795. [Google Scholar] [CrossRef]
- Ou, J.; Meng, Q.; Li, Y.; Xiu, Y.; Du, J.; Gu, W.; Wu, T.; Li, W.; Ding, Z.; Wang, W. Identification and comparative analysis of the Eriocheir sinensis microRNA transcriptome response to Spiroplasma eriocheiris infection using a deep sequencing approach. Fish Shellfish Immunol. 2012, 32, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Trobaugh, D.W.; Gardner, C.L.; Sun, C.; Haddow, A.D.; Wang, E.; Chapnik, E.; Mildner, A.; Weaver, S.C.; Ryman, K.D.; Klimstra, W.B. RNA viruses can hijack vertebrate microRNAs to suppress innate immunity. Nature 2014, 506, 245–248. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Liu, L.; Zohaib, A.; Lin, L.; Yuan, J.; Wang, M.; Liu, X. MicroRNA profile analysis of Epithelioma papulosum cyprini cell line before and after SVCV infection. Dev. Comp. Immunol. 2015, 48, 124–128. [Google Scholar] [CrossRef] [PubMed]
- Betts, A.M.; Stone, D.M.; Way, K.; Torhy, C.; Chilmonczyk, S.; Benmansour, A.; de Kinkelin, P. Emerging vesiculo-type virus infections of freshwater fishes in Europe. Dis. Aquat. Organ. 2003, 57, 201–212. [Google Scholar] [CrossRef] [PubMed]
- Crane, M.; Hyatt, A. Viruses of fish, an overview of significant pathogens. Viruses 2011, 3, 2025–2046. [Google Scholar] [CrossRef] [PubMed]
- McFee, R.B. Global infections—Avian influenza and other significant emerging pathogens: An overview. Dis. Mon. 2007, 53, 343–347. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wen, Y.; Hu, X.; Wang, W.; Liang, X.; Li, J.; Vakharia, V.; Lin, L. Breaking the host range: Mandarin fish is susceptible to a vesiculovirus derived from snakehead fish. J. Gen. Virol. 2015, 96, 775–781. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.; Wang, Q.; Wang, Y.; Liu, C.; Liang, H.; Fang, X.; Wu, S. Genomic characterization and taxonomic position of a rhabdovirus from a hybrid snakehead. Arch. Virol. 2014, 159, 2469–2473. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Asim, M.; Yi, L.; Hegazy, A.M.; Hu, X.; Zhou, Y.; Ai, T.; Lin, L. Abortive infection of snakehead fish vesiculovirus in ZF4 cells was associated with the RLRs pathway activation by viral replicative intermediates. Int. J. Mol. Sci. 2015, 16, 6235–6250. [Google Scholar] [CrossRef] [PubMed]
- Landgraf, P.; Rusu, M.; Sheridan, R.; Sewer, A.; Iovino, N.; Aravin, A.; Pfeffer, S.; Rice, A.; Kamphorst, A.O.; Landthaler, M.; et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007, 129, 1401–1414. [Google Scholar] [CrossRef] [PubMed]
- Cullen, B.R. How do viruses avoid inhibition by endogenous cellular microRNAs? PLoS Pathog. 2013, 9, e1003694. [Google Scholar] [CrossRef] [PubMed]
- Chi, W.; Tong, C.; Gan, X.; He, S. Characterization and comparative profiling of MiRNA transcriptomes in bighead carp and silver carp. PLoS ONE 2011, 6, e23549. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Ding, L.; Li, Y.; Zhang, X.; Liang, Y.; Sun, X.; Teng, C.B. Identification and profiling of microRNAs from skeletal muscle of the common carp. PLoS ONE 2012, 7, e30925. [Google Scholar] [CrossRef] [PubMed]
- Yi, S.; Gao, Z.X.; Zhao, H.; Zeng, C.; Luo, W.; Chen, B.; Wang, W.M. Identification and characterization of microRNAs involved in growth of blunt snout bream (Megalobrama amblycephala) by Solexa sequencing. BMC Genom. 2013, 14, 754. [Google Scholar] [CrossRef] [PubMed]
- Frerichs, G.N.; Morgan, D.; Hart, D.; Skerrow, C.; Roberts, R.J.; Onions, D.E. Spontaneously productive C-type retrovirus infection of fish cell lines. J. Gen. Virol. 1991, 72, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.J.; Zhang, Y.P.; Li, Y.; Zheng, H.W.; Zheng, Y.S.; Liu, C.J. Distinct expression pattern of miRNAs in Marek’s disease virus infected-chicken splenic tumors and non-tumorous spleen tissues. Res. Vet. Sci. 2014, 97, 156–161. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Zhu, J.; Zhou, H.; Zhao, Z.; Zou, Z.; Liu, X.; Lin, X.; Zhang, X.; Deng, X.; Wang, R.; et al. Identification of cellular microRNA-136 as a dual regulator of RIG-I-mediated innate immunity that antagonizes H5N1 IAV replication in A549 cells. Sci. Rep. 2015, 5, 14991. [Google Scholar] [CrossRef] [PubMed]
- Taganov, K.D.; Boldin, M.P.; Chang, K.J.; Baltimore, D. NF-ĸB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA 2006, 103, 12481–12486. [Google Scholar] [CrossRef] [PubMed]
- Henke, J.I.; Goergen, D.; Zheng, J.; Song, Y.; Schuttler, C.G.; Fehr, C.; Junemann, C.; Niepmann, M. microRNA-122 stimulates translation of hepatitis C virus RNA. EMBO J. 2008, 27, 3300–3310. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.K.; Zhang, Q.; Gao, L.; Li, N.; Chen, X.X.; Feng, W.H. Increasing expression of microRNA 181 inhibits porcine reproductive and respiratory syndrome virus replication and has implications for controlling virus infection. J. Virol. 2013, 87, 1159–1171. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.J.; Yang, J.; Fan, X.L.; Zhao, H.B.; Hu, W.; Li, Z.P.; Yu, G.C.; Ding, X.R.; Wang, J.Z.; Bo, X.C.; et al. Cellular microRNA let-7c inhibits M1 protein expression of the H1N1 influenza A virus in infected human lung epithelial cells. J. Cell. Mol. Med. 2012, 16, 2539–2546. [Google Scholar] [CrossRef] [PubMed]
- miRBase. Available online: http://www.mirbase.org (accessed on 1 January 2016).
- Miranda. Available online: http://www.microrna.org/microrna/home.do (accessed on 1 January 2016).
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Tu, J.; Yuan, J.; Liu, X.; Zhao, L.; Dawar, F.U.; Khattak, M.N.K.; Hegazy, A.M.; Chen, N.; Vakharia, V.N.; et al. Identification and Characterization of MicroRNAs in Snakehead Fish Cell Line upon Snakehead Fish Vesiculovirus Infection. Int. J. Mol. Sci. 2016, 17, 154. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms17020154
Liu X, Tu J, Yuan J, Liu X, Zhao L, Dawar FU, Khattak MNK, Hegazy AM, Chen N, Vakharia VN, et al. Identification and Characterization of MicroRNAs in Snakehead Fish Cell Line upon Snakehead Fish Vesiculovirus Infection. International Journal of Molecular Sciences. 2016; 17(2):154. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms17020154
Chicago/Turabian StyleLiu, Xiaodan, Jiagang Tu, Junfa Yuan, Xueqin Liu, Lijuan Zhao, Farman Ullah Dawar, Muhammad Nasir Khan Khattak, Abeer M. Hegazy, Nan Chen, Vikram N. Vakharia, and et al. 2016. "Identification and Characterization of MicroRNAs in Snakehead Fish Cell Line upon Snakehead Fish Vesiculovirus Infection" International Journal of Molecular Sciences 17, no. 2: 154. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms17020154






