Gibberellic Acid: A Key Phytohormone for Spikelet Fertility in Rice Grain Production
Abstract
:1. Introduction
2. Gibberellic Acid (GA) Signal Transduction in Rice Development
3. Roles of GA Signaling in Male Organ Development in Rice
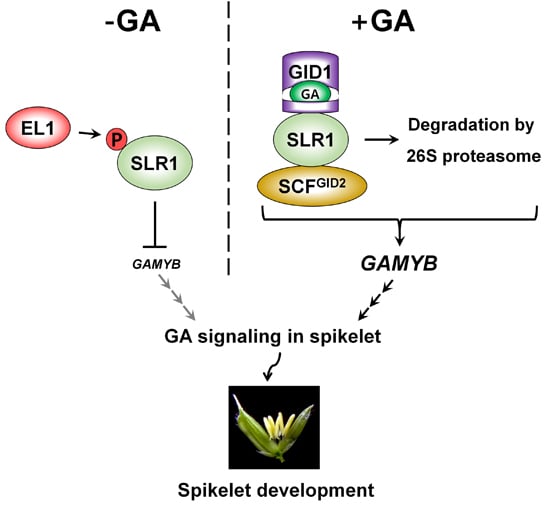
4. The Role of GA in Rice Spikelet Fertility
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| GA | Gibberellic acid |
| SLR1 | Slender rice1 |
| GID1 | GA INSENSITIVE DWARF1 |
| GID2 | GA INSENSITIVE DWARF2 |
| SCF | Skp, Cullin, F-box |
| OsSPY | Rice SPINDLY |
| EL1 | Early flowering1 |
| PMC | Pollen mother cell |
| OsCPS | Rice ent-copalyl diphosphate synthase |
| CYP703 | Cytochrome P450 hydroxylase 703 |
| KAR | β-Ketoacyl reductase |
| TDR | TAPETUM DEGENERATION RETARDATION |
| OsCP | Rice cysteine protease |
| Col-0 | Columbia |
| Ler | Landsberg erecta |
| PIF | Phytochrome-interacting factor |
| JAZ | Jasmonate ZIM-domain |
| EIN3 | Ethylene insensitive3 |
References
- Yabuta, T.; Sumiki, Y. On the crystal of gibberellin, a substance to promote plant growth. J. Agric. Chem. Soc. Jpn. 1938, 14, 1526. [Google Scholar]
- Schwechheimer, C. Gibberellin signaling in plants—The extended version. Front. Plant Sci. 2012, 2. [Google Scholar] [CrossRef] [PubMed]
- Davière, J.M.; Achard, P. Gibberellin signaling in plants. Development 2013, 140, 1147–1151. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, S. Gibberellin metabolism and its regulation. Annu. Rev. Plant Biol. 2008, 59, 225–251. [Google Scholar] [CrossRef] [PubMed]
- Nester, J.E.; Zeevaart, J.A.D. Flower development in normal tomato and a gibberellin-deficient (ga-2) mutant. Am. J. Bot. 1988, 75, 45–55. [Google Scholar] [CrossRef]
- Ritchie, S.; Gilroy, S. Gibberellins: Regulating genes and germination. New Phytol. 1998, 140, 363–383. [Google Scholar] [CrossRef]
- Goto, N.; Pharis, R.P. Role of gibberellins in the development of floral organs of the gibberellin-deficient mutant, ga1-1, of Arabidopsis thaliana. Can. J. Bot. 1999, 77, 944–954. [Google Scholar] [CrossRef]
- Woodger, F.J.; Millar, A.; Murray, F.; Jacobsen, J.V.; Gubler, F. The role of GAMYB transcription factors in GA-regulated gene expression. J. Plant Growth Regul. 2003, 22, 176–184. [Google Scholar] [CrossRef]
- Singh, D.P.; Jermakow, A.M.; Swain, S.M. Gibberellins are required for seed development and pollen tube growth in Arabidopsis. Plant Cell 2002, 14, 3133–3147. [Google Scholar] [CrossRef] [PubMed]
- Chhun, T.; Aya, K.; Asano, K.; Yamamoto, E.; Morinaka, Y.; Watanabe, M.; Kitano, H.; Ashikari, M.; Matsuoka, M.; Ueguchi-Tanaka, M. Gibberellin regulates pollen viability and pollen tube growth in rice. Plant Cell 2007, 19, 3876–3888. [Google Scholar] [CrossRef] [PubMed]
- Kwon, C.T.; Kim, S.H.; Kim, D.; Paek, N.C. The rice floral repressor Early flowering1 affects spikelet fertility by modulating gibberellin signaling. Rice 2015, 8. [Google Scholar] [CrossRef] [PubMed]
- Aya, K.; Ueguchi-Tanaka, M.; Kondo, M.; Hamada, K.; Yano, K.; Nishimura, M.; Matsuoka, M. Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB. Plant Cell 2009, 21, 1453–1472. [Google Scholar] [CrossRef] [PubMed]
- Hedden, P.; Phillips, A.L. Gibberellin metabolism: New insights revealed by the genes. Trends Plant Sci. 2000, 5, 523–530. [Google Scholar] [CrossRef]
- Fleet, C.M.; Sun, T.P. A DELLAcate balance: The role of gibberellin in plant morphogenesis. Curr. Opin. Plant Biol. 2005, 8, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.B.; Xu, Y.Y.; Ma, H.; Chong, K. Cell biological characterization of male meiosis and pollen development in rice. J. Integr. Plant Biol. 2005, 47, 734–744. [Google Scholar] [CrossRef]
- Itoh, J.; Nonomura, K.; Ikeda, K.; Yamaki, S.; Inukai, Y.; Yamagishi, H.; Kitano, H.; Nagato, Y. Rice plant development: From zygote to spikelet. Plant Cell Physiol. 2005, 46, 23–47. [Google Scholar] [CrossRef] [PubMed]
- Wilson, Z.A.; Zhang, D.B. From Arabidopsis to rice: Pathways in pollen development. J. Exp. Bot. 2009, 60, 1479–1492. [Google Scholar] [CrossRef] [PubMed]
- Plackett, A.R.G.; Thomas, S.G.; Wilson, Z.A.; Hedden, P. Gibberellin control of stamen development: A fertile field. Trends Plant Sci. 2011, 16, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Ueguchi-Tanaka, M.; Nakajima, M.; Motoyuki, A.; Matsuoka, M. Gibberellin receptor and its role in gibberellin signaling in plants. Annu. Rev. Plant Biol. 2007, 58, 183–198. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Genschik, P. Releasing the brakes of plant growth: How GAs shutdown DELLA proteins. J. Exp. Bot. 2009, 60, 1085–1092. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, A.; Ueguchi-Tanaka, M.; Sonoda, Y.; Kitano, H.; Koshioka, M.; Futsuhara, Y.; Matsuoka, M.; Yamaguchi, J. slender rice, a constitutive gibberellin response mutant, is caused by a null mutation of the SLR1 gene, an ortholog of the height-regulating gene GAI/RGA/RHT/D8. Plant Cell 2001, 13, 999–1010. [Google Scholar] [CrossRef] [PubMed]
- Itoh, H.; Ueguchi-Tanaka, M.; Sato, Y.; Ashikari, M.; Matsuoka, M. The gibberellin signaling pathway is regulated by the appearance and disappearance of SLENDER RICE1 in nuclei. Plant Cell 2002, 14, 57–70. [Google Scholar] [CrossRef] [PubMed]
- Ueguchi-Tanaka, M.; Ashikari, M.; Nakajima, M.; Itoh, H.; Katoh, E.; Kobayashi, M.; Chow, T.Y.; Hsing, Y.I.C.; Kitano, H.; Yamaguchi, I.; et al. GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 2005, 437, 693–698. [Google Scholar] [CrossRef] [PubMed]
- Ueguchi-Tanaka, M.; Nakajima, M.; Katoh, E.; Ohmiya, H.; Asano, K.; Saji, S.; Xiang, H.Y.; Ashikari, M.; Kitano, H.; Yamaguchi, I.; et al. Molecular interactions of a soluble gibberellin receptor, GID1, with a rice DELLA protein, SLR1, and gibberellin. Plant Cell 2007, 19, 2140–2155. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Itoh, H.; Gomi, K.; Ueguchi-Tanaka, M.; Ishiyama, K.; Kobayashi, M.; Jeong, D.H.; An, G.; Kitano, H.; Ashikari, M.; et al. Accumulation of phosphorylated repressor for gibberellin signaling in an F-box mutant. Science 2003, 299, 1896–1898. [Google Scholar] [CrossRef] [PubMed]
- Gomi, K.; Sasaki, A.; Itoh, H.; Ueguchi-Tanaka, M.; Ashikari, M.; Kitano, H.; Matsuoka, M. GID2, an F-box subunit of the SCF E3 complex, specifically interacts with phosphorylated SLR1 protein and regulates the gibberellin-dependent degradation of SLR1 in rice. Plant J. 2004, 37, 626–634. [Google Scholar] [CrossRef] [PubMed]
- Itoh, H.; Sasaki, A.; Ueguchi-Tanaka, M.; Ishiyama, K.; Kobayashi, M.; Hasegawa, Y.; Minami, E.; Ashikari, M.; Matsuoka, M. Dissection of the phosphorylation of rice DELLA protein, SLENDER RICE1. Plant Cell Physiol. 2005, 46, 1392–1399. [Google Scholar] [CrossRef] [PubMed]
- Shimada, A.; Ueguchi-Tanaka, M.; Sakamoto, T.; Fujioka, S.; Takatsuto, S.; Yoshida, S.; Sazuka, T.; Ashikari, M.; Matsuoka, M. The rice SPINDLY gene functions as a negative regulator of gibberellin signaling by controlling the suppressive function of the DELLA protein, SLR1, and modulating brassinosteroid synthesis. Plant J. 2006, 48, 390–402. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Xue, H.W. Rice early flowering1, a CKI, phosphorylates DELLA protein SLR1 to negatively regulate gibberellin signalling. EMBO J. 2010, 29, 1916–1927. [Google Scholar] [CrossRef] [PubMed]
- Kwon, C.T.; Yoo, S.C.; Koo, B.H.; Cho, S.H.; Park, J.W.; Zhang, Z.Y.; Li, J.J.; Li, Z.C.; Paek, N.C. Natural variation in Early flowering1 contributes to early flowering in japonica rice under long days. Plant Cell Environ. 2014, 37, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Gubler, F.; Kalla, R.; Roberts, J.K.; Jacobsen, J.V. Gibberellin-regulated expression of a myb gene in barley aleurone cells: Evidence for Myb transactivation of a high-pl α-amylase gene promoter. Plant Cell 1995, 7, 1879–1891. [Google Scholar] [CrossRef] [PubMed]
- Gubler, F.; Watts, R.J.; Kalla, R.; Matthews, P.; Keys, M.; Jacobsen, J.V. Cloning of a rice cDNA encoding a transcription factor homologous to barley GAMyb. Plant Cell Physiol. 1997, 38, 362–365. [Google Scholar] [CrossRef] [PubMed]
- Gubler, F.; Raventos, N.; Keys, M.; Watts, R.; Mundy, J.; Jacobsen, J.V. Target genes and regulatory domains of the GAMYB transcriptional activator in cereal aleurone. Plant J. 1999, 17, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Gubler, F.; Chandler, P.M.; White, R.G.; Llewellyn, D.J.; Jacobsen, J.V. Gibberellin signaling in barley aleurone cells. Control of SLN1 and GAMYB expression. Plant Physiol. 2002, 129, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, M.; Inukai, Y.; Ueguchi-Tanaka, M.; Itoh, H.; Izawa, T.; Kobayashi, Y.; Hattori, T.; Miyao, A.; Hirochika, H.; Ashikari, M.; et al. Loss-of-function mutations of the rice GAMYB gene impair α-amylase expression in aleurone and flower development. Plant Cell 2004, 16, 33–44. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Herr, A.; Baulcombe, D.C.; Harberd, N.P. Modulation of floral development by a gibberellin-regulated microRNA. Development 2004, 131, 3357–3365. [Google Scholar] [CrossRef] [PubMed]
- Tsuji, H.; Aya, K.; Ueguchi-Tanaka, M.; Shimada, Y.; Nakazono, M.; Watanabe, R.; Nishizawa, N.K.; Gomi, K.; Shimada, A.; Kitano, H.; et al. GAMYB controls different sets of genes and is differentially regulated by microRNA in aleurone cells and anthers. Plant J. 2006, 47, 427–444. [Google Scholar] [CrossRef] [PubMed]
- Lanahan, M.B.; Ho, T.H.D. Slender barley—A constitutive gibberellin-response mutant. Planta 1988, 175, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Jacobsen, S.E.; Olszewski, N.E. Characterization of the arrest in anther development associated with gibberellin deficiency of the gib-1 mutant of tomato. Plant Physiol. 1991, 97, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Evans, M.M.; Poethig, R.S. Gibberellins promote vegetative phase change and reproductive maturity in maize. Plant Physiol. 1995, 108, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Izhaki, A.; Borochov, A.; Zamski, E.; Weiss, D. Gibberellin regulates post-microsporogenesis processes in petunia anthers. Physiol. Plant 2002, 115, 442–447. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Qin, L.J.; Lee, S.C.; Fu, X.D.; Richards, D.E.; Cao, D.N.; Luo, D.; Harberd, N.P.; Peng, J.R. Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 2004, 131, 1055–1064. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, T.; Miura, K.; Itoh, H.; Tatsumi, T.; Ueguchi-Tanaka, M.; Ishiyama, K.; Kobayashi, M.; Agrawal, G.K.; Takeda, S.; Abe, K.; et al. An overview of gibberellin metabolism enzyme genes and their related mutants in rice. Plant Physiol. 2004, 134, 1642–1653. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Vriezen, W.H.; van Der Straeten, D.; Harberd, N.P. Ethylene regulates Arabidopsis development via the modulation of DELLA protein growth repressor function. Plant Cell 2003, 15, 2816–2825. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Cheng, H.; de Grauwe, L.; Decat, J.; Schoutteten, H.; Moritz, T.; van Der Straeten, D.; Peng, J.; Harberd, N.P. Integration of plant responses to environmentally activated phytohormonal signals. Science 2006, 311, 91–94. [Google Scholar] [CrossRef] [PubMed]
- Navarro, L.; Bari, R.; Achard, P.; Lisón, P.; Nemri, A.; Harberd, N.P.; Jones, J.D. DELLAs control plant immune responses by modulating the balance and salicylic acid signaling. Curr. Biol. 2008, 18, 650–655. [Google Scholar] [CrossRef] [PubMed]
- McCormick, S. Control of male gametophyte development. Plant Cell 2004, 16, S142–S153. [Google Scholar] [CrossRef] [PubMed]
- Scott, R.J.; Spielman, M.; Dickinson, H.G. Stamen structure and function. Plant Cell 2004, 16, S46–S60. [Google Scholar] [CrossRef] [PubMed]
- Ma, H. Molecular genetic analyses of microsporogenesis and microgametogenesis in flowering plants. Annu. Rev. Plant Biol. 2005, 56, 393–434. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.X.; Liu, Y.G. Molecular control of male reproductive development and pollen fertility in rice. J. Integr. Plant Biol. 2012, 54, 967–978. [Google Scholar] [CrossRef] [PubMed]
- Hirano, K.; Aya, K.; Hobo, T.; Sakakibara, H.; Kojima, M.; Shim, R.A.; Hasegawa, Y.; Ueguchi-Tanaka, M.; Matsuoka, M. Comprehensive transcriptome analysis of phytohormone biosynthesis and signaling genes in microspore/pollen and tapetum of rice. Plant Cell Physiol. 2008, 49, 1429–1450. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.H.; Mitchum, M.G.; Barnaby, N.; Ayele, B.T.; Ogawa, M.; Nam, E.; Lai, W.C.; Hanada, A.; Alonso, J.M.; Ecker, J.R.; et al. Potential sites of bioactive gibberellin production during reproductive growth in Arabidopsis. Plant Cell 2008, 20, 320–336. [Google Scholar] [CrossRef] [PubMed]
- Aya, K.; Suzuki, G.; Suwabe, K.; Hobo, T.; Takahashi, H.; Shiono, K.; Yano, K.; Tsutsumi, N.; Nakazono, M.; Nagamura, Y.; et al. Comprehensive network analysis of anther-expressed genes in rice by the combination of 33 laser microdissection and 143 spatiotemporal microarrays. PLoS ONE 2011, 6, e26162. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Zhang, D.S.; Liu, H.S.; Yin, C.S.; Li, X.X.; Liang, W.Q.; Yuan, Z.; Xu, B.; Chu, H.W.; Wang, J.; et al. The rice tapetum degeneration retardation gene is required for tapetum degradation and anther development. Plant Cell 2006, 18, 2999–3014. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.S.; Liang, W.; Yin, C.; Zong, J.; Gu, F.; Zhang, D. OsC6, encoding a lipid transfer protein, is required for postmeiotic anther development in rice. Plant Physiol. 2010, 154, 149–162. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Zhang, D.C.; Tang, M.F.; Li, D.Y.; Zhu, Y.X.; Zhu, L.H.; Chen, C.Y. THIS1 is a putative lipase that regulates tillering, plant height, and spikelet fertility in rice. J. Exp. Bot. 2013, 64, 4389–4402. [Google Scholar] [CrossRef] [PubMed]
- Plackett, A.R.G.; Ferguson, A.C.; Powers, S.J.; Wanchoo-Kohli, A.; Phillips, A.L.; Wilson, Z.A.; Hedden, P.; Thomas, S.G. DELLA activity is required for successful pollen development in the Columbia ecotype of Arabidopsis. New Phytol. 2014, 201, 825–836. [Google Scholar] [CrossRef] [PubMed]
- Murray, F.; Kalla, R.; Jacobsen, J.; Gubler, F. A role for HvGAMYB in anther development. Plant J. 2003, 33, 481–491. [Google Scholar] [CrossRef] [PubMed]
- Batch, J.J.; Morgan, D.G. Male-sterility induced in barley by photoperiod. Nature 1974, 250, 165–167. [Google Scholar] [CrossRef]
- Higashitani, A. High temperature injury and auxin biosynthesis in microsporogenesis. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Sakata, T.; Oda, S.; Tsunaga, Y.; Shomura, H.; Kawagishi-Kobayashi, M.; Aya, K.; Saeki, K.; Endo, T.; Nagano, K.; Kojima, M.; et al. Reduction of gibberellin by low temperature disrupts pollen development in rice. Plant Physiol. 2014, 164, 2011–2019. [Google Scholar] [CrossRef] [PubMed]
- Park, J.H.; Halitschke, R.; Kim, H.B.; Baldwin, I.T.; Feldmann, K.A.; Feyereisen, R. A knock-out mutation in allene oxide synthase results in male sterility and defective wound signal transduction in Arabidopsis due to a block in jasmonic acid biosynthesis. Plant J. 2002, 31, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Cecchetti, V.; Altamura, M.M.; Falasca, G.; Costantino, P.; Cardarelli, M. Auxin regulates Arabidopsis anther dehiscence, pollen maturation, and filament elongation. Plant Cell 2008, 20, 1760–1774. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Yuan, Z.; Chen, M.; Yin, C.; Luo, Z.; Zhao, X.; Liang, W.; Hu, J.; Zhang, D. Jasmonic acid regulates spikelet development in rice. Nat. Commun. 2014, 19, 3476. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Sakuraba, Y.; Lee, T.; Kim, K.W.; An, G.; Lee, H.Y.; Paek, N.C. Mutation of Oryza sativa CORONATINE INSENSITIVE 1b (OsCOI1b) delays leaf senescence. J. Integr. Plant Biol. 2015, 57, 562–576. [Google Scholar] [CrossRef] [PubMed]
- De Lucas, M.; Davière, J.M.; Rodríguez-Falcón, M.; Pontin, M.; Iglesias-Pedraz, J.M.; Lorrain, S.; Fankhauser, C.; Blázquez, M.A.; Titarenko, E.; Prat, S. A molecular framework for light and gibberellin control of cell elongation. Nature 2008, 451, 480–484. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Lee, L.Y.; Xia, K.; Yan, Y.; Yu, H. DELLAs modulate jasmonate signaling via competitive binding to JAZs. Dev. Cell 2010, 19, 884–894. [Google Scholar] [CrossRef] [PubMed]
- An, F.; Zhang, X.; Zhu, Z.; Ji, Y.; He, W.; Jiang, Z.; Li, M.; Guo, H. Coordinated regulation of apical hook development by gibberellins and ethylene in etiolated Arabidopsis seedlings. Cell Res. 2012, 22, 915–927. [Google Scholar] [CrossRef] [PubMed]
- Wild, M.; Davière, J.M.; Cheminant, S.; Regnault, T.; Baumberger, N.; Heintz, D.; Baltz, R.; Genschik, P.; Achard, P. The Arabidopsis DELLA RGA-LIKE3 is a direct target of MYC2 and modulates jasmonate signaling responses. Plant Cell 2012, 24, 3307–3319. [Google Scholar] [CrossRef] [PubMed]
- Davière, J.M.; Achard, P. A pivotal role of DELLAs in regulating multiple hormone signals. Mol. Plant 2016, 9, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Dill, A.; Sun, T.P. Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics 2001, 159, 777–785. [Google Scholar] [PubMed]
- Fuentes, S.; Ljung, K.; Sorefan, K.; Alvey, E.; Harberd, N.P.; Ostergaard, L. Fruit growth in Arabidopsis occurs via DELLA-dependent and DELLA-independent gibberellin responses. Plant Cell 2012, 24, 3982–3996. [Google Scholar] [CrossRef] [PubMed]

| Gene | MSU ID a | RAP ID b | Function | Conserved Domain | Polypeptide | Reference |
|---|---|---|---|---|---|---|
| SLR1 | Os03g49990 | Os03g0707600 | Transcription regulation | DELLA, GRAS | 625 aa | [21] |
| GID1 | Os05g33730 | Os05g0407500 | GA receptor | Hormone sensitive lipase (HSL) | 354 aa | [23] |
| GID2 | Os02g36974 | Os02g0580300 | DELLA degradation | F-box | 212 aa | [25] |
| EL1 | Os03g57940 | Os03g0793500 | Phosphorylation | Ser/Thr kinase | 707 aa | [29] |
| GAMYB | Os01g59660 | Os01g0812000 | Transcription factor | Myb | 553 aa | [32] |
| OsSPY | Os08g44510 | Os08g0559300 | O-GlcNAcylation | O-GlcNAc transferase | 927aa | [28] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kwon, C.-T.; Paek, N.-C. Gibberellic Acid: A Key Phytohormone for Spikelet Fertility in Rice Grain Production. Int. J. Mol. Sci. 2016, 17, 794. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms17050794
Kwon C-T, Paek N-C. Gibberellic Acid: A Key Phytohormone for Spikelet Fertility in Rice Grain Production. International Journal of Molecular Sciences. 2016; 17(5):794. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms17050794
Chicago/Turabian StyleKwon, Choon-Tak, and Nam-Chon Paek. 2016. "Gibberellic Acid: A Key Phytohormone for Spikelet Fertility in Rice Grain Production" International Journal of Molecular Sciences 17, no. 5: 794. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms17050794