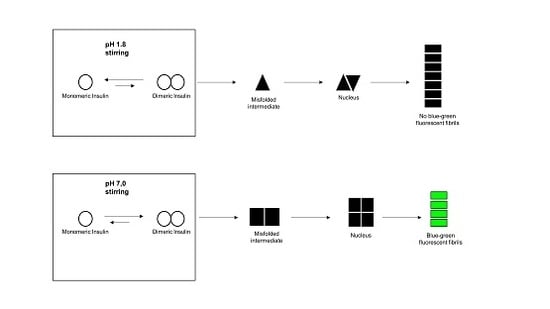
Insights into Insulin Fibril Assembly at Physiological and Acidic pH and Related Amyloid Intrinsic Fluorescence
Abstract
:1. Introduction
2. Results
3. Discussion
4. Material and Methods
4.1. Materials
4.2. Insulin Preparation and Amyloid Fibril Formation
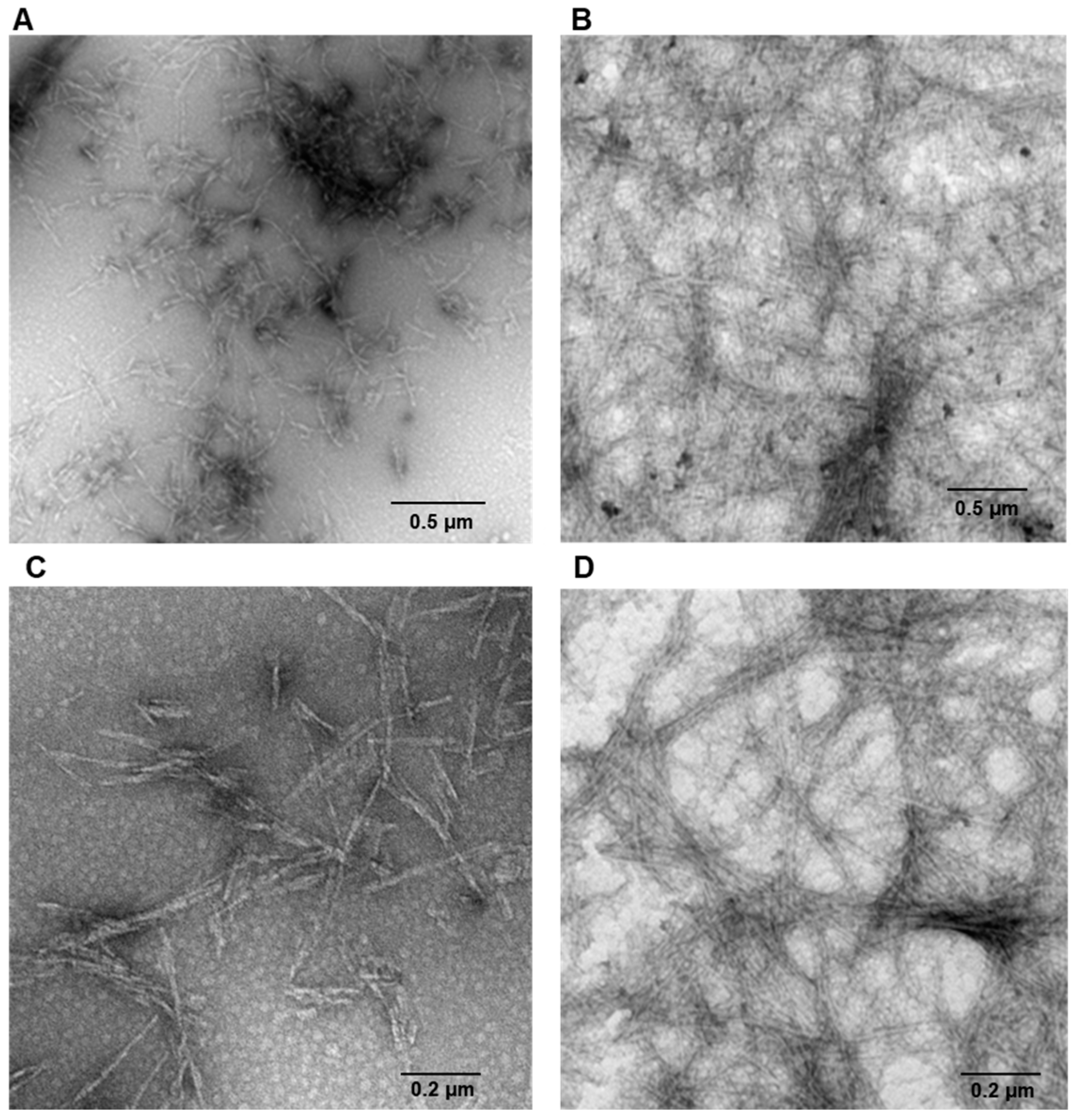
4.3. Transmission Electronic Microscopy (TEM)
4.4. Fluorescence Measurements
4.5. Two Photons Microscopy Measurements
4.6. Circular Dichroism Measurements
4.7. Fourier-Transform Infrared Measurements
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Adams, M.J.; Blundell, T.L.; Dodson, E.J.; Dodson, G.G.; Vijayan, M.; Baker, E.N.; Harding, M.M.; Hodgkin, D.C.; Rimmer, B.; Sheat, S. Structure of rhombohedral 2 zinc insulin crystal. Nature 1969, 224, 491–495. [Google Scholar] [CrossRef]
- Blundell, T.L.; Cutfield, J.F.; Cutfield, S.M.; Dodson, E.J.; Dodson, G.G.; Hodgkin, D.C.; Mercola, D.A.; Vijayan, M. Atomic positions in rhombohedral 2-zinc insulin crystals. Nature 1971, 231, 506–511. [Google Scholar] [CrossRef] [PubMed]
- Dodson, G.; Steiner, D. The role of assembly in insulin’s biosynthesis. Curr. Opin. Struct. Biol. 1998, 8, 189–194. [Google Scholar] [CrossRef]
- Ahmad, A.; Uversky, V.N.; Hong, D.; Fink, A.L. Early events in the fibrillation of monomeric insulin. J. Biol. Chem. 2005, 280, 42669–42675. [Google Scholar] [CrossRef] [PubMed]
- Hong, D.P.; Ahmad, A.; Fink, A.L. Fibrillation of human insulin A and B chains. Biochemistry 2006, 45, 9342–9353. [Google Scholar] [CrossRef] [PubMed]
- Dunn, M.F. Zinc-ligand interactions modulate assembly and stability of the insulin hexamer—A review. Biometals 2005, 18, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Sneideris, T.; Darguzis, D.; Botyriute, A.; Grigaliunas, M.; Winter, R.; Smirnovas, V. pH-Driven polymorphism of insulin amyloid-like fibrils. PLoS ONE 2015, 10, e0136602. [Google Scholar] [CrossRef] [PubMed]
- Rambaran, R.N.; Serpell, L.C. Amyloid fibrils: Abnormal protein assembly. Prion 2008, 2, 112–117. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.C.; Nayak, A.; Sethuraman, A.; Belfort, G.; McRae, G.J. A three-stage kinetic model of amyloid fibrillation. Biophys. J. 2007, 92, 3448–3458. [Google Scholar] [CrossRef] [PubMed]
- Dische, F.E.; Wernstedt, C.; Westermark, G.T.; Westermark, P.; Pepys, M.B.; Rennie, J.A.; Gilbey, S.G.; Watkins, P.J. Insulin as an amyloid-fibril protein at sites of repeated insulin injections in a diabetic patient. Diabetologia 1988, 31, 158–161. [Google Scholar] [CrossRef] [PubMed]
- Swift, B. Examination of insulin injection sites: An unexpected finding of localized amyloidosis. Diabet. Med. 2002, 19, 881–882. [Google Scholar] [CrossRef] [PubMed]
- Shikama, Y.; Kitazawa, J.; Yagihashi, N.; Uehara, O.; Murata, Y.; Yajima, N.; Ryuichi, W.; Soroku, Y. Localized amyloidosis at the site of repeated insulin injection in a diabetic patient. Intern. Med. 2010, 49, 397–401. [Google Scholar] [CrossRef] [PubMed]
- Okamura, S.; Hayashino, Y.; Kore-Eda, S.; Tsujii, S. Localized amyloidosis at the site of repeated insulin injection in a patient with type 2 diabetes. Diabetes Care 2013, 36, e200. [Google Scholar] [CrossRef] [PubMed]
- Nagase, T.; Iwaya, K.; Iwaki, Y.; Kotake, F.; Uchida, R.; Oh-i, T.; Sekine, H.; Miwa, K.; Murakami, S.; Odaka, T.; et al. Insulin-derived amyloidosis and poor glycemic control: A case series. Am. J. Med. 2014, 127, 450–454. [Google Scholar] [CrossRef] [PubMed]
- D’Souza, A.; Theis, J.D.; Vrana, J.A.; Dogan, A. Pharmaceutical amyloidosis associated with subcutaneous insulin and enfuvirtide administration. Amyloid 2014, 21, 71–75. [Google Scholar] [CrossRef] [PubMed]
- Kerr, D.; Wizemann, E.; Senstius, J.; Zacho, M.; Ampudia-Blasco, F.J.J. Stability and performance of rapid-acting insulin analogs used for continuous subcutaneous insulin infusion: A systematic review. J. Diabetes Sci. Technol. 2013, 7, 1595–1606. [Google Scholar] [CrossRef] [PubMed]
- Gupta, Y.; Singla, G.; Singla, R. Insulin-derived amyloidosis. Indian J. Endocrinol. Metab. 2015, 19, 174–177. [Google Scholar] [CrossRef] [PubMed]
- Nettleton, E.J.; Tito, P.; Sunde, M.; Bouchard, M.; Dobson, C.M.; Robinson, C.V. Characterization of the oligomeric states of insulin in self-assembly and amyloid fibril formation by mass spectrometry. Biophys. J. 2000, 79, 1053–1065. [Google Scholar] [CrossRef]
- Bouchard, M.; Zurdo, J.; Nettleton, E.J.; Dobson, C.M.; Robinson, C.V. Formation of insulin amyloid fibrils followed by FTIR simultaneously with CD and electron microscopy. Protein Sci. 2000, 9, 1960–1967. [Google Scholar] [CrossRef] [PubMed]
- Smirnovas, V.; Winter, R. Revealing different aggregation pathways of amyloidogenic proteins by ultrasound velocimetry. Biophys. J. 2008, 94, 3241–3246. [Google Scholar] [CrossRef] [PubMed]
- Vestergaard, B.; Groenning, M.; Roessle, M.; Kastrup, J.S.; van de Weert, M.; Flink, J.M.; Frokjaer, S.; Gajhede, M.; Svergun, D.I. A helical structural nucleus is the primary elongating unit of insulin amyloid fibrils. PLoS Biol. 2007, 5, e134. [Google Scholar] [CrossRef] [PubMed]
- Foderà, V.; Cataldo, S.; Librizzi, F.; Pignataro, B.; Spiccia, P.; Leone, M. Self-organization pathways and spatial heterogeneity in insulin amyloid fibril formation. J. Phys. Chem. B. 2009, 113, 10830–10837. [Google Scholar] [CrossRef] [PubMed]
- Ivanova, M.I.; Sievers, S.A.; Sawaya, M.R.; Wall, J.S.; Eisenberg, D. Molecular basis for insulin fibril assembly. Proc. Natl. Acad. Sci. USA 2009, 106, 18990–18995. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, L.; Khurana, R.; Coats, A.; Frokjaer, S.; Brange, J.; Vyas, S.; Uversky, V.N.; Fink, A.L. Effect of environmental factors on the kinetics of insulin fibril formation: Elucidation of the molecular mechanism. Biochemistry 2001, 40, 6036–6046. [Google Scholar] [CrossRef] [PubMed]
- Tiiman, A.; Noormägi, A.; Friedemann, M.; Krishtal, J.; Palumaa, P.; Tõugu, V. Effect of agitation on the peptide fibrillization: Alzheimer’s amyloid-β peptide 1–42 but not amylin and insulin fibrils can grow under quiescent conditions. J. Pept. Sci. 2013, 19, 386–391. [Google Scholar] [CrossRef] [PubMed]
- Noormägi, A.; Valmsen, K.; Tõugu, V.; Palumaa, P. Insulin Fibrillization at Acidic and Physiological pH Values is Controlled by Different Molecular Mechanisms. Protein J. 2015, 34, 398–403. [Google Scholar] [CrossRef] [PubMed]
- Yoshihara, H.; Saito, J.; Tanabe, A.; Amada, T.; Asakura, T.; Kitagawa, K.; Asada, S. Characterization of Novel Insulin Fibrils That Show Strong Cytotoxicity Under Physiological pH. J. Pharm. Sci. 2016, 105, 1419–1426. [Google Scholar] [CrossRef] [PubMed]
- Surmacz-Chwedoruk, W.; Babenko, V.; Dec, R.; Szymczak, P.; Dzwolak, W. The emergence of superstructural order in insulin amyloid fibrils upon multiple rounds of self-seeding. Sci. Rep. 2016, 6, 32022. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, J.S.; Andersen, C.B.; Otzen, D.E. Amyloid structure—One but not the same: The many levels of fibrillar polymorphism. FEBS J. 2010, 277, 4591–4601. [Google Scholar] [CrossRef] [PubMed]
- Eichner, T.; Radford, S.E. A diversity of assembly mechanisms of a generic amyloid fold. Mol. Cell 2011, 43, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Fändrich, M.; Meinhardt, J.; Grigorieff, N. Structural polymorphism of Alzheimer Aβ and other amyloid fibrils. Prion 2009, 3, 89–93. [Google Scholar] [CrossRef] [PubMed]
- Meinhardt, J.; Sachse, C.; Hortschansky, P.; Grigorieff, N.; Fändrich, M. Aβ(1–40) fibril polymorphism implies diverse interaction patterns in amyloid fibrils. J. Mol. Biol. 2009, 386, 869–877. [Google Scholar] [CrossRef] [PubMed]
- Seilheimer, B.; Bohrmann, B.; Bondolfi, L.; Müller, F.; Stüber, D.; Döbeli, H. The toxicity of the Alzheimer’s β-amyloid peptide correlates with a distinct fiber morphology. J. Struct. Biol. 1997, 119, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Petkova, A.T.; Leapman, R.D.; Guo, Z.; Yau, W.M.; Mattson, M.P.; Tycko, R. Self-propagating, molecular-level polymorphism in Alzheimer’s β-amyloid fibrils. Science 2005, 307, 262–265. [Google Scholar] [CrossRef] [PubMed]
- Aguzzi, A.; Haass, C. Games played by rogue proteins in prion disorders and Alzheimer’s disease. Science 2003, 302, 814–818. [Google Scholar] [CrossRef] [PubMed]
- Chan, F.T.S.; Kaminski, G.S.; Kumita, J.R.; Bertoncini, C.W.; Dobson, C.M.; Kaminski, C.F. Protein amyloids develop an intrinsic fluorescence signature during aggregation. Analyst 2013, 138, 2156–2162. [Google Scholar] [CrossRef] [PubMed]
- Pinotsi, D.; Buell, A.K.; Dobson, C.M.; Kaminski Schierle, G.S.; Kaminski, C.F. A label-free, quantitative assay of amyloid fibril growth based on intrinsic fluorescence. ChemBioChem 2013, 14, 846–850. [Google Scholar] [CrossRef] [PubMed]
- Pinotsi, D.; Kaminski Schierle, G.S.; Kaminski, C.F. Optical Super-Resolution Imaging of β-Amyloid Aggregation in Vitro and in Vivo: Method and Techniques. Methods Mol. Biol. 2016, 1303, 125–141. [Google Scholar] [CrossRef] [PubMed]
- Tcherkasskaya, O. Photo-activity induced by amyloidogenesis. Protein Sci. 2007, 16, 561–571. [Google Scholar] [CrossRef] [PubMed]
- Hanczyc, P.; Samoc, M.; Norden, B. Multiphoton absorption in amyloid protein fibres. Nat. Photonics 2013, 7, 969–972. [Google Scholar] [CrossRef]
- Pinotsi, D.; Grisanti, L.; Mahou, P.; Gebauer, R.; Kaminski, C.F.; Hassanali, A.; Kaminski Schierle, G.S. Proton Transfer and Structure-Specific Fluorescence in Hydrogen Bond-Rich Protein Structures. J. Am. Chem. Soc. 2016, 138, 3046–3057. [Google Scholar] [CrossRef] [PubMed]
- Grisanti, L.; Pinotsi, D.; Gebauer, R.; Kaminski Schierle, G.S.; Hassanali, A.A. A computational study on how structure influences the optical properties in model crystal structures of amyloid fibrils. Phys. Chem. Chem. Phys. 2017, 19, 4030–4040. [Google Scholar] [CrossRef] [PubMed]
- LeVine, H., 3rd. Thioflavine T interaction with synthetic Alzheimer’s disease β-amyloid peptides: Detection of amyloid aggregation in solution. Protein Sci. 1993, 2, 404–410. [Google Scholar] [CrossRef] [PubMed]
- Zandomeneghi, G.; Krebs, M.R.H.; Mccammon, M.G.; Fandrich, M. FTIR reveals differences between native β-sheet proteins and amyloid fibrils. Protein Sci. 2004, 13, 3314–3321. [Google Scholar] [CrossRef] [PubMed]
- Sarroukh, R.; Goormaghtigh, E.; Ruysschaert, J.M.; Raussens, V. ATR-FTIR: A “rejuvenated” tool to investigate amyloid proteins. Biochim. Biophys. Acta 2013, 1828, 2328–2338. [Google Scholar] [CrossRef] [PubMed]
- Natalello, A.; Doglia, S.M. Insoluble protein assemblies characterized by fourier transform infrared spectroscopy. Methods Mol. Biol. 2015, 1258, 347–369. [Google Scholar] [CrossRef] [PubMed]
- Barth, A. Infrared spectroscopy of proteins. Biochim. Biophys. Acta 2007, 1767, 1073–1101. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Yang, S.; Kong, J.; Dong, A.; Yu, S. Obtaining information about protein secondary structures in aqueous solution using Fourier transform IR spectroscopy. Nat. Protoc. 2015, 10, 382–396. [Google Scholar] [CrossRef] [PubMed]
- Baldassarre, M.; Li, C.; Eremina, N.; Goormaghtigh, E.; Barth, A. Simultaneous Fitting of Absorption Spectra and Their Second Derivatives for an Improved Analysis of Protein Infrared Spectra. Molecules 2015, 20, 12599–12622. [Google Scholar] [CrossRef] [PubMed]
- Seshadri, S.; Khurana, R.; Fink, A.L. Fourier transform infrared spectroscopy in analysis of protein deposits. Methods Enzymol. 1999, 309, 559–576. [Google Scholar] [CrossRef] [PubMed]
- Ami, D.; Ricagno, S.; Bolognesi, M.; Bellotti, V.; Doglia, S.M.; Natalello, A. Structure, Stability, and Aggregation of β-2 Microglobulin Mutants: Insights from a Fourier Transform Infrared Study in Solution and in the Crystalline State. Biophys. J. 2012, 102, 1676–1684. [Google Scholar] [CrossRef] [PubMed]
- Kauffmann, E.; Darnton, N.C.; Austin, R.H.; Batt, C.; Gerwert, K. Lifetimes of intermediates in the β-sheet to α-helix transition of β-lactoglobulin by using a diffusional IR mixer. Proc. Natl. Acad. Sci. USA 2001, 98, 6646–6649. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, L.; Frokjaer, S.; Brange, J.; Uversky, V.N.; Fink, A.L. Probing the mechanism of insulin fibril formation with insulin mutants. Biochemistry 2001, 40, 8397–8409. [Google Scholar] [CrossRef] [PubMed]
- Selivanova, O.M.; Galzitskaya, O.V. Structural polymorphism and possible pathways of amyloid fibril formation on the example of insulin protein. Biochemistry 2012, 77, 1237–1247. [Google Scholar] [CrossRef] [PubMed]
- Maity, S.; Hashemi, M.; Lyubchenko, Y.L. Nano-assembly of amyloid β peptide: Role of the hairpin fold. Sci. Rep. 2017, 7, 2344. [Google Scholar] [CrossRef] [PubMed]
- Hudson, S.A.; Ecroyd, H.; Kee, T.W.; Carver, J.A. The thioflavin T fluorescence assay for amyloid fibril detection can be biased by the presence of exogenous compounds. FEBS J. 2009, 276, 5960–5972. [Google Scholar] [CrossRef] [PubMed]
- Del Mercato, L.L.; Pompa, P.P.; Maruccio, G.; Della Torre, A.; Sabella, S.; Tamburro, A.M.; Cingolani, R.; Rinaldi, R. Chargetransport and intrinsicfluorescence in amyloid-likefibrils. Proc. Natl. Acad. Sci. USA 2007, 104, 18019–18024. [Google Scholar] [CrossRef] [PubMed]
- Anand, U.; Mukherjee, M. Exploring the self-assembly of a short aromatic Aβ(16–24) peptide. Langmuir 2013, 29, 2713–2721. [Google Scholar] [CrossRef] [PubMed]
- Sharpe, S.; Simonetti, K.; Yau, J.; Walsh, P. Solid-state NMR characterization of autofluorescent fibrils formed by the elastin-derived peptide GVGVAGVG. Biomacromolecules 2011, 12, 1546–1555. [Google Scholar] [CrossRef] [PubMed]
- Sobczyk, L.; Grabowski, S.J.; Krygowski, T.M. Interrelation between H-Bond and Pi-Electron Delocalization. Chem. Rev. 2005, 105, 3513–3560. [Google Scholar] [CrossRef] [PubMed]
- Portaccio, M.; Gravagnuolo, A.M.; Longobardi, S.; Giardinab, P.; Rea, I.; De Stefanoc, L.; Cammarota, M.; Lepore, M. ATR FT-IR spectroscopy on Vmh2 hydrophobin self-assembled layers for Teflon membrane bio-functionalization. Appl. Surf. Sci. 2015, 351, 673–680. [Google Scholar] [CrossRef]
- Fabian, H.; Mäntele, W. Infrared Spectroscopy of Proteins. In Handbook of Vibrational Spectroscopy; Wiley: Hoboken, NJ, USA, 2006. [Google Scholar] [CrossRef]
- Levenberg, K. A method for the estimation of certain nonlinear problems in least squares. Q. Appl. Math. 1944, 2, 164–168. [Google Scholar] [CrossRef]
- Marquardt, D.W. An algorithm for least squares estimation of nonlinear parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Mei, Y.; Miller, L.; Gao, W.; Gross, R.A. Imaging the distribution and secondary structure of immobilized enzymes using infrared microspectroscopy. Biomacromolecules 2003, 4, 70–74. [Google Scholar] [CrossRef] [PubMed]


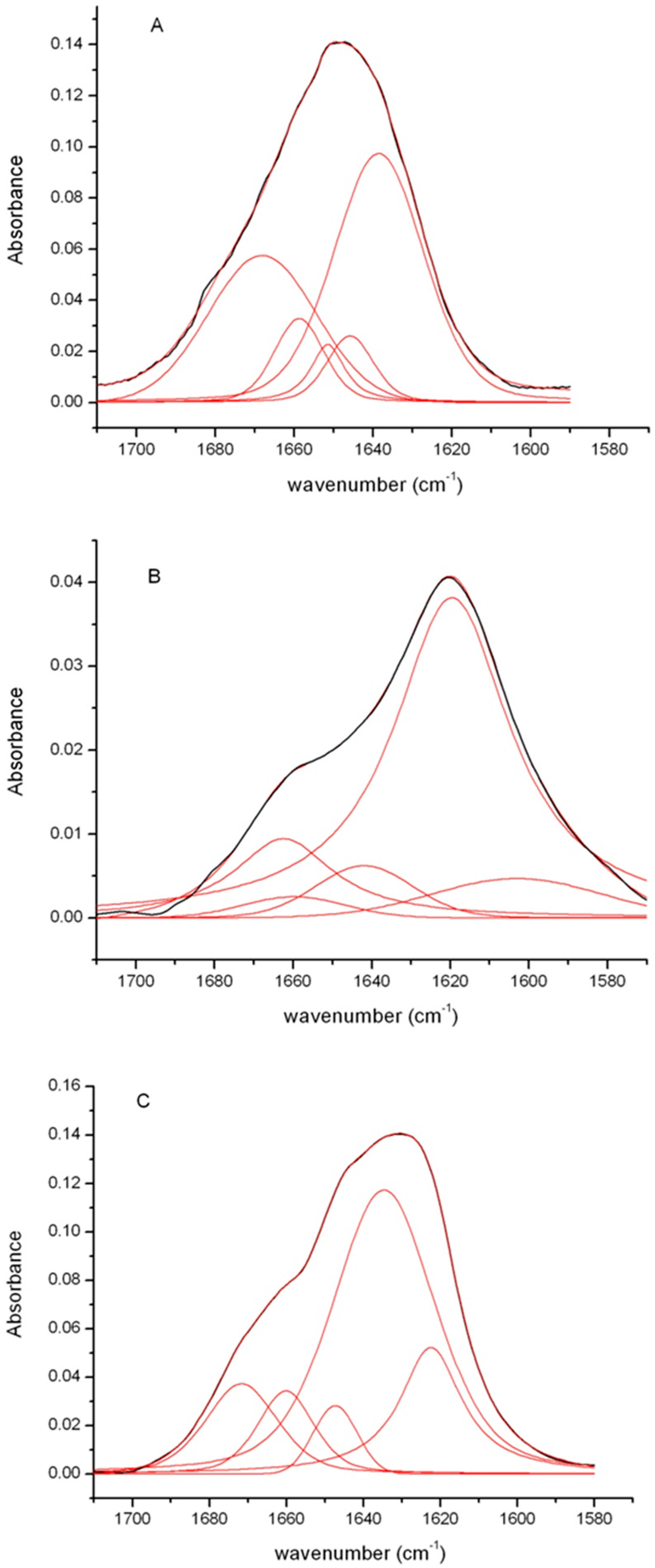
| Protein | Secondary Structure (%) | |||
|---|---|---|---|---|
| Intermolecular β-sheet 1611–1630 cm−1 | Intramolecular β-sheet 1615–1638 cm−1 | α-helix 1640–1655 cm−1 | Disordered/β-turn 1658–1686 cm−1 | |
| Native insulin | 0 | 0 | 60 | 40 |
| Fibrils pH 7.0 | 18 | 56 | 5 | 21 |
| Fibrils pH 1.8 | 76 | 0 | 7 | 17 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iannuzzi, C.; Borriello, M.; Portaccio, M.; Irace, G.; Sirangelo, I. Insights into Insulin Fibril Assembly at Physiological and Acidic pH and Related Amyloid Intrinsic Fluorescence. Int. J. Mol. Sci. 2017, 18, 2551. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18122551
Iannuzzi C, Borriello M, Portaccio M, Irace G, Sirangelo I. Insights into Insulin Fibril Assembly at Physiological and Acidic pH and Related Amyloid Intrinsic Fluorescence. International Journal of Molecular Sciences. 2017; 18(12):2551. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18122551
Chicago/Turabian StyleIannuzzi, Clara, Margherita Borriello, Marianna Portaccio, Gaetano Irace, and Ivana Sirangelo. 2017. "Insights into Insulin Fibril Assembly at Physiological and Acidic pH and Related Amyloid Intrinsic Fluorescence" International Journal of Molecular Sciences 18, no. 12: 2551. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18122551