Tongue Sole CD209: A Pattern-Recognition Receptor that Binds a Broad Range of Microbes and Promotes Phagocytosis
Abstract
:1. Introduction
2. Results
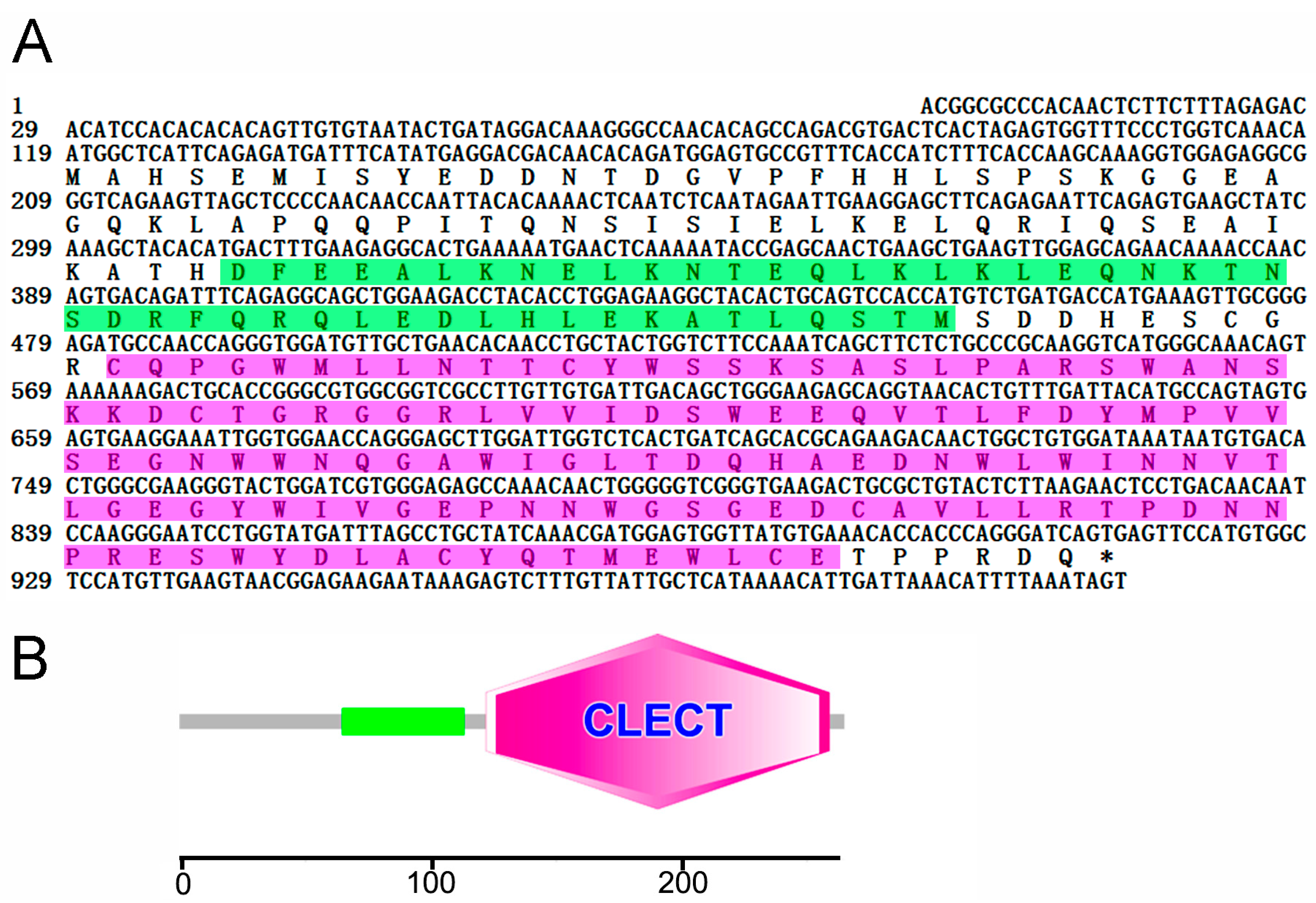
2.1. Sequence Analysis of CsCD209
2.2. Three-Dimensional Structure Characteristics of CsCD209
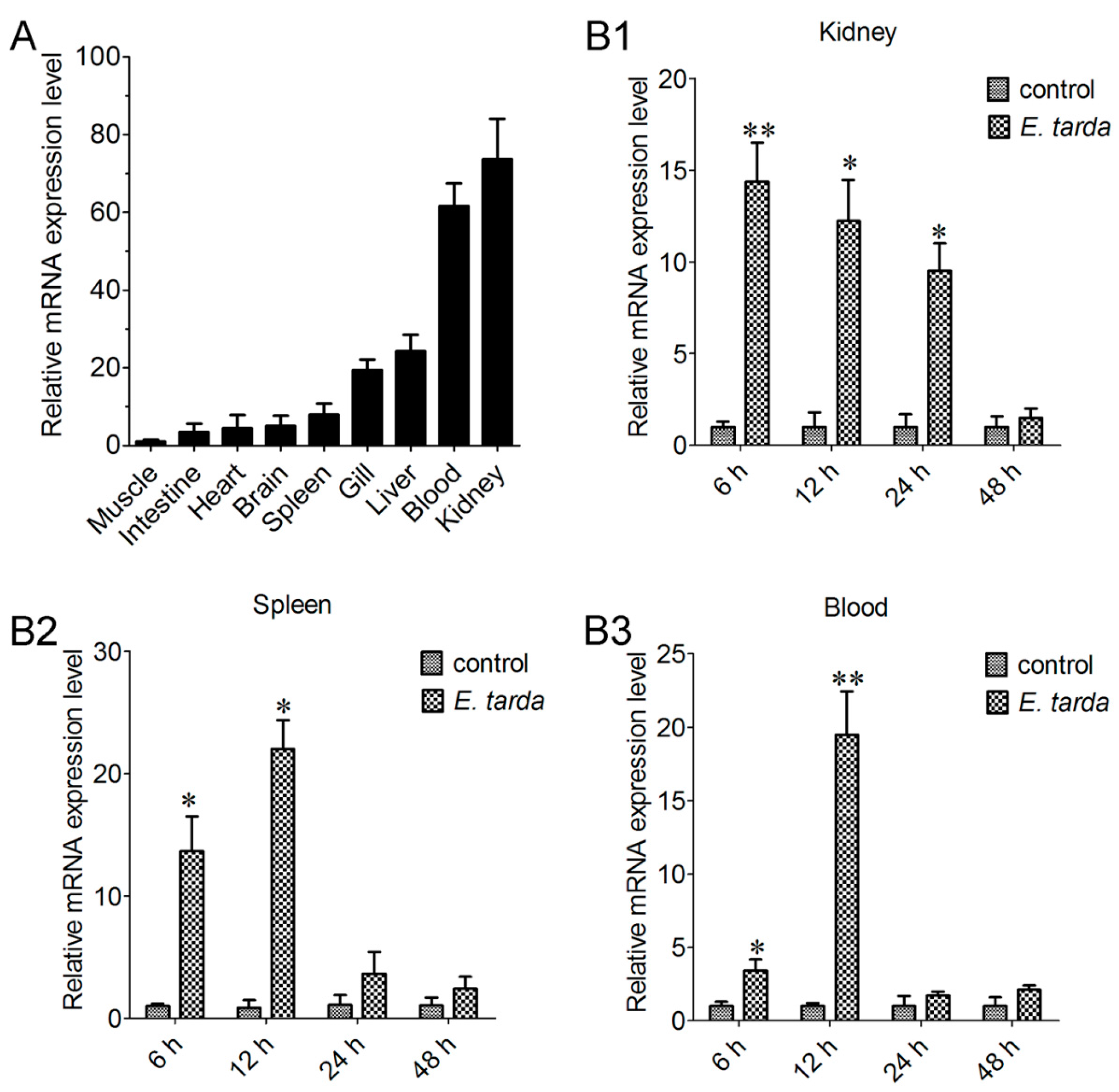
2.3. Expression of CsCD209 in Fish Tissues
2.4. Expression of CsCD209 in Head Kidney Leukocytes and Blood
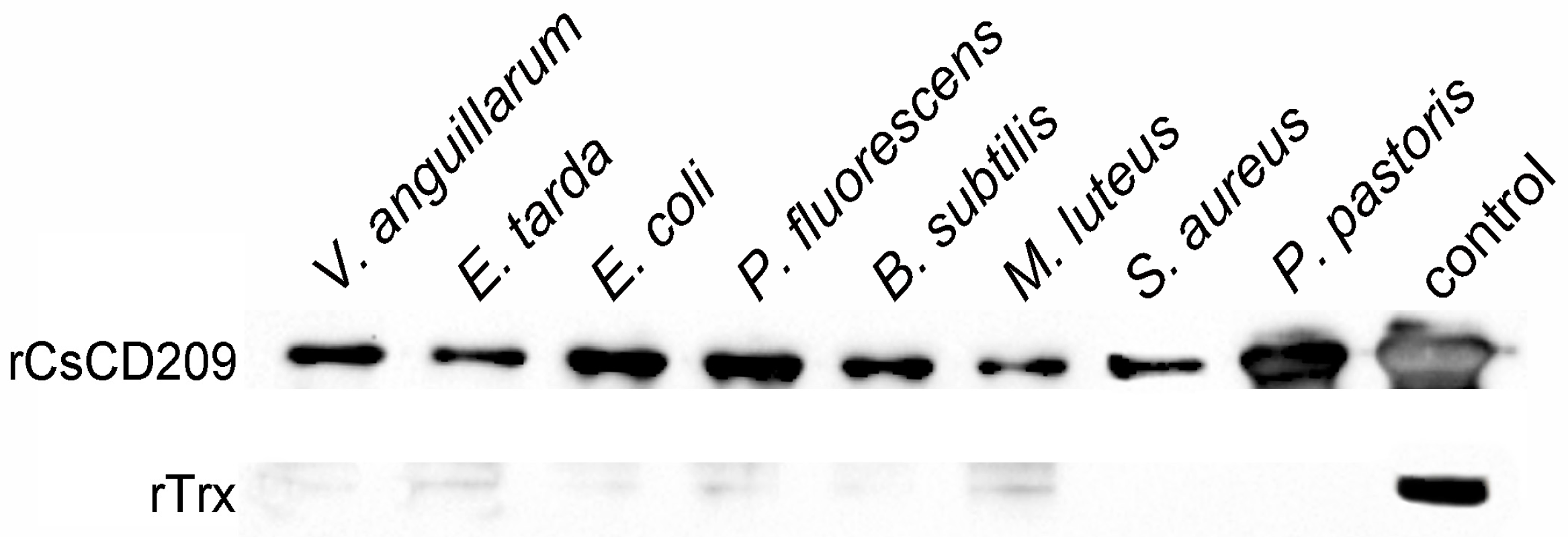
2.5. Microbial Binding Activity of rCsCD209
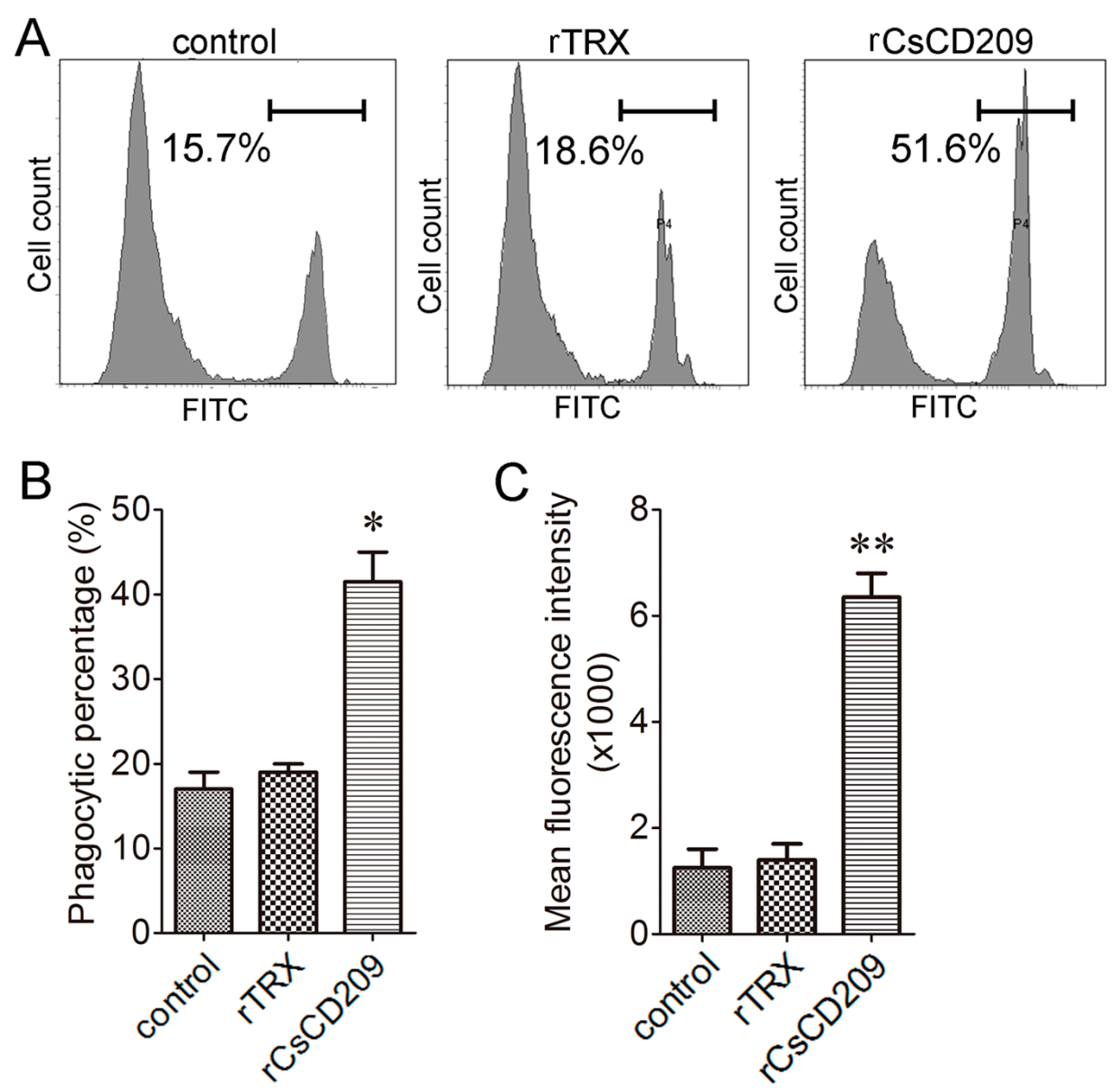
2.6. Effect of rCsCD209 on Phagocytosis Towards Bacteria
3. Discussion
4. Materials and Methods
4.1. Fish
4.2. Bacterial Strains and Culture Conditions
4.3. Sequence Analysis
4.4. Quantitative Real Time Reverse Transcription-PCR (qRT-PCR)
4.4.1. qRT-PCR Analysis of CsCD209 Expression in Different Fish Tissues under Normal Physiological Conditions
4.4.2. qRT-PCR Analysis of CsCD209 Expression in Different Fish Tissues during Pathogen Infection
4.5. Prokaryotic Expression and Purification of Recombinant Proteins
4.6. Generation of Mouse Polyclonal Antibody Against rCsCD209
4.7. Western Blot Analysis
4.7.1. Western Blot Analysis of CsCD209 in Head Kidney Leukocytes and Serum
4.7.2. Western Blot Analysis of rCsCD209 Binding to Microbes
4.8. Flow Cytometry
4.8.1. Flow Cytometry Analysis of CsCD209 Expression
4.8.2. Flow Cytometry Analysis of rCsCD209 Binding to Bacteria
4.8.3. Flow Cytometry Analysis of Phagocytosis
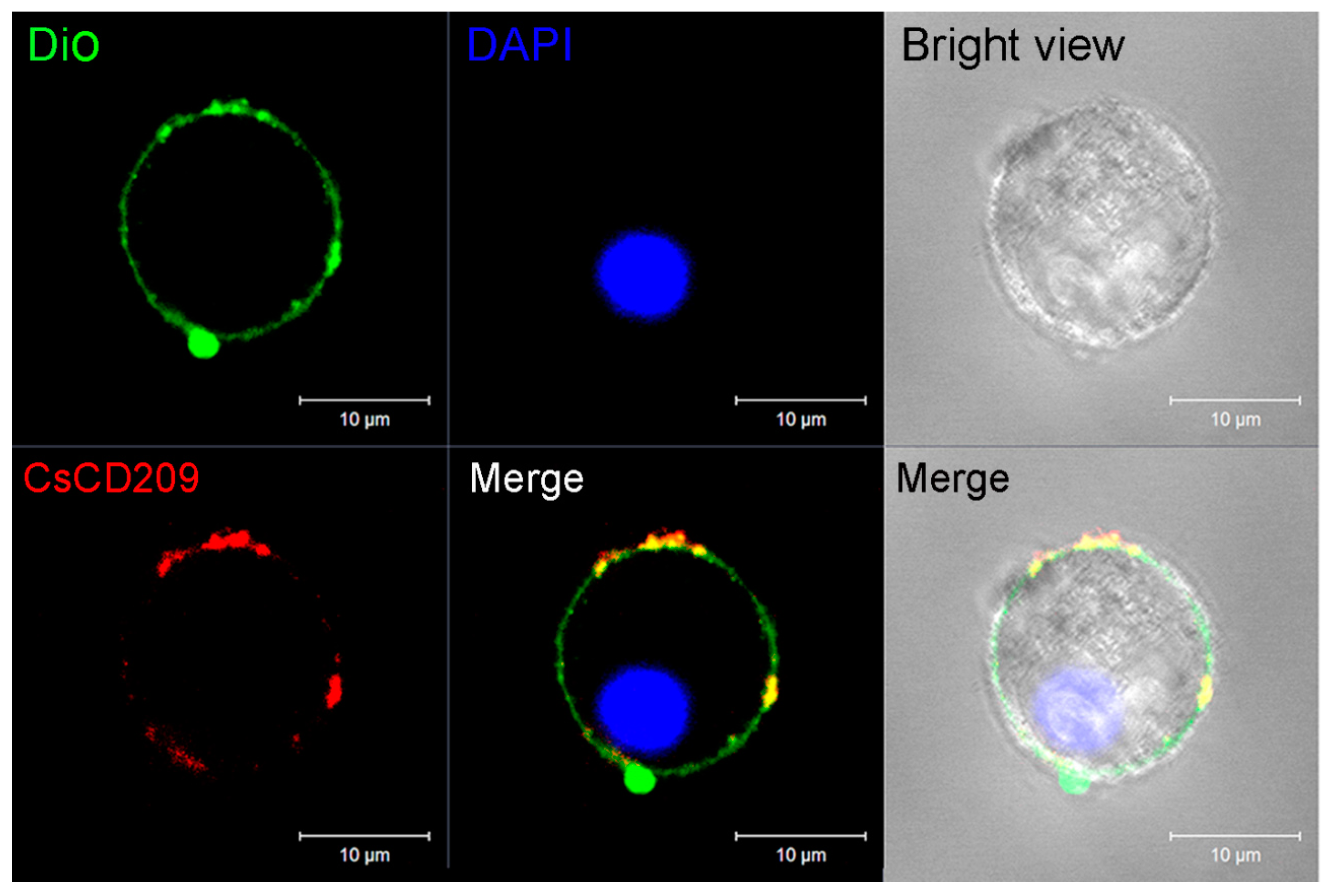
4.9. Microscopic Analysis of the Subcellular Distribution of CsCD209
4.10. Statistical Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wu, J.; Chen, Z.J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 2014, 32, 461–488. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, A.; Medzhitov, R. Regulation of adaptive immunity by the innate immune system. Science 2010, 327, 291–295. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.; Mazmanian, S.K. Innate immune recognition of the microbiota promotes host-microbial symbiosis. Nat. Immunol. 2013, 14, 668–675. [Google Scholar] [CrossRef] [PubMed]
- Abraham, C.; Medzhitov, R. Interactions between the host innate immune system and microbes in inflammatory bowel disease. Gastroenterology 2011, 140, 1729–1737. [Google Scholar] [CrossRef] [PubMed]
- Seimon, T.A.; Obstfeld, A.; Moore, K.J.; Golenbock, D.T.; Tabas, I. Combinatorial pattern recognition receptor signaling alters the balance of life and death in macrophages. Proc. Natl. Acad. Sci. USA 2006, 103, 19794–19799. [Google Scholar] [CrossRef] [PubMed]
- Tassaneetrithep, B.; Burgess, T.H.; Granelli-Piperno, A.; Trumpfheller, C.; Finke, J.; Sun, W.; Eller, M.A.; Pattanapanyasat, K.; Sarasombath, S.; Birx, D.L.; et al. DC-SIGN (CD209) mediates dengue virus infection of human dendritic cells. J. Exp. Med. 2003, 197, 823–839. [Google Scholar] [CrossRef] [PubMed]
- Davis, C.W.; Mattei, L.M.; Nguyen, H.Y.; Ansarah-Sobrinho, C.; Doms, R.W.; Pierson, T.C. The location of asparagine-linked glycans on West Nile virions controls their interactions with CD209 (dendritic cell-specific ICAM-3 grabbing nonintegrin). J. Biol. Chem. 2006, 281, 37183–37194. [Google Scholar] [CrossRef] [PubMed]
- Ortiz, M.; Kaessmann, H.; Zhang, K.; Bashirova, A.; Carrington, M.; Quintana-Murci, L.; Telenti, A. The evolutionary history of the CD209 (DC-SIGN) family in humans and non-human primates. Genes Immun. 2008, 9, 483–492. [Google Scholar] [CrossRef] [PubMed]
- Montoya, D.; Cruz, D.; Teles, R.M.; Lee, D.J.; Ochoa, M.T.; Krutzik, S.R.; Chun, R.; Schenk, M.; Zhang, X.; Ferguson, B.G.; et al. Divergence of macrophage phagocytic and antimicrobial programs in leprosy. Cell. Host Microbe 2009, 6, 343–353. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Gomez, D.; Dominguez-Soto, A.; Ancochea, J.; Jimenez-Heffernan, J.A.; Leal, J.A.; Corbi, A.L. Dendritic cell-specific intercellular adhesion molecule 3-grabbing nonintegrin mediates binding and internalization of Aspergillus fumigatus conidia by dendritic cells and macrophages. J. Immunol. 2004, 173, 5635–5643. [Google Scholar] [CrossRef] [PubMed]
- Cambi, A.; Gijzen, K.; de Vries, I.J.M.; Torensma, R.; Joosten, B.; Adema, G.J.; Netea, M.G.; Kullberg, B.J.; Romani, L.; Figdor, C.G. The C-type lectin DC-SIGN (CD209) is an antigen-uptake receptor for Candida albicans on dendritic cells. Eur. J. Immunol. 2003, 33, 532–538. [Google Scholar] [CrossRef] [PubMed]
- Gunther, P.S.; Mikeler, E.; Hamprecht, K.; Schneider-Schaulies, J.; Jahn, G.; Dennehy, K.M. CD209/DC-SIGN mediates efficient infection of monocyte-derived dendritic cells by clinical adenovirus 2C isolates in the presence of bovine lactoferrin. J. Gen. Virol. 2011, 92, 1754–1759. [Google Scholar] [CrossRef] [PubMed]
- Soilleux, E.J.; Morris, L.S.; Leslie, G.; Chehimi, J.; Luo, Q.; Levroney, E.; Trowsdale, J.; Montaner, L.J.; Doms, R.W.; Weissman, D.; et al. Constitutive and induced expression of DC-SIGN on dendritic cell and macrophage subpopulations in situ and in vitro. J. Leukoc. Biol. 2002, 71, 445–457. [Google Scholar] [PubMed]
- Wichukchinda, N.; Kitamura, Y.; Rojanawiwat, A.; Nakayama, E.E.; Song, H.; Pathipvanich, P.; Auwanit, W.; Sawanpanyalert, P.; Iwamoto, A.; Shioda, T.; et al. The polymorphisms in DC-SIGNR affect susceptibility to HIV type 1 infection. AIDS Res. Hum. Retroviruses 2007, 23, 686–692. [Google Scholar] [CrossRef] [PubMed]
- Krishnadas, D.K.; Ahn, J.S.; Han, J.; Kumar, R.; Agrawal, B. Immunomodulation by hepatitis C virus-derived proteins: Targeting human dendritic cells by multiple mechanisms. Int. Immunol. 2010, 22, 491–502. [Google Scholar] [CrossRef] [PubMed]
- Saitoh, T.; Komano, J.; Saitoh, Y.; Misawa, T.; Takahama, M.; Kozaki, T.; Uehata, T.; Iwasaki, H.; Omori, H.; Yamaoka, S.; et al. Neutrophil extracellular traps mediate a host defense response to human immunodeficiency virus-1. Cell Host Microbe 2012, 12, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Lugo-Villarino, G.; Balla, K.M.; Stachura, D.L.; Banuelos, K.; Werneck, M.B.F.; Traver, D. Identification of dendritic antigen-presenting cells in the zebrafish. P Natl. Acad. Sci. USA 2010, 107, 15850–15855. [Google Scholar] [CrossRef] [PubMed]
- Lin, A.F.; Xiang, L.X.; Wang, Q.L.; Dong, W.R.; Gong, Y.F.; Shao, J.Z. The DC-SIGN of zebrafish: Insights into the existence of a CD209 homologue in a lower vertebrate and its involvement in adaptive immunity. J. Immunol. 2009, 183, 7398–7410. [Google Scholar] [CrossRef] [PubMed]
- Bassity, E.; Clark, T.G. Functional identification of dendritic cells in the teleost model, rainbow trout (Oncorhynchus mykiss). PLoS ONE 2012, 7, e33196. [Google Scholar] [CrossRef] [PubMed]
- Johansson, P.; Corripio-Miyar, Y.; Wang, T.; Collet, B.; Secombes, C.J.; Zou, J. Characterisation and expression analysis of the rainbow trout (Oncorhynchus mykiss) homologue of the human dendritic cell marker CD208/lysosomal associated membrane protein 3. Dev. Comp. Immunol. 2012, 37, 402–413. [Google Scholar] [CrossRef] [PubMed]
- Wittamer, V.; Bertrand, J.Y.; Gutschow, P.W.; Traver, D. Characterization of the mononuclear phagocyte system in zebrafish. Blood 2011, 117, 7126–7135. [Google Scholar] [CrossRef] [PubMed]
- Feinberg, H.; Mitchell, D.A.; Drickamer, K.; Weis, W.I. Structural basis for selective recognition of oligosaccharides by DC-SIGN and DC-SIGNR. Science 2001, 294, 2163–2166. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Feinberg, H.; Conroy, E.; Mitchell, D.A.; Alvarez, R.; Blixt, O.; Taylor, M.E.; Weis, W.I.; Drickamer, K. Structural basis for distinct ligand-binding and targeting properties of the receptors DC-SIGN and DC-SIGNR. Nat. Struct. Mol. Biol. 2004, 11, 591–598. [Google Scholar] [CrossRef] [PubMed]
- Van Liempt, E.; Bank, C.M.; Mehta, P.; Garcia-Vallejo, J.J.; Kawar, Z.S.; Geyer, R.; Alvarez, R.A.; Cummings, R.D.; Kooyk, Y.; van Die, I. Specificity of DC-SIGN for mannose- and fucose-containing glycans. FEBS Lett. 2006, 580, 6123–6131. [Google Scholar] [CrossRef] [PubMed]
- Feinberg, H.; Powlesland, A.S.; Taylor, M.E.; Weis, W.I. Trimeric structure of langerin. J. Biol. Chem. 2010, 285, 13285–13293. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Chen, Y.; Hao, L.; Zhang, Y. DC-SIGN and immunoregulation. Cell. Mol. Immunol. 2006, 3, 279–283. [Google Scholar] [PubMed]
- Van Gisbergen, K.P.J.M.; Sanchez-Hernandez, M.; Geijtenbeek, T.B.H.; van Kooyk, Y. Neutrophils mediate immune modulation of dendritic cells through glycosylation-dependent interactions between Mac-1 and DC-SIGN. J. Exp. Med. 2005, 201, 1281–1292. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Geijtenbeek, T.B.; Krooshoop, D.J.; Bleijs, D.A.; van Vliet, S.J.; van Duijnhoven, G.C.; Grabovsky, V.; Alon, R.; Figdor, C.G.; van Kooyk, Y. DC-SIGN-ICAM-2 interaction mediates dendritic cell trafficking. Nat. Immunol. 2000, 1, 353–357. [Google Scholar] [CrossRef] [PubMed]
- Granja, A.G.; Leal, E.; Pignatelli, J.; Castro, R.; Abos, B.; Kato, G.; Fischer, U.; Tafalla, C. Identification of Teleost Skin CD8α+ Dendritic-like Cells, Representing a potential common ancestor for mammalian cross-presenting dendritic cells. J. Immunol. 2015, 195, 1825–1837. [Google Scholar] [CrossRef] [PubMed]
- Mummidi, S.; Catano, G.; Lam, L.; Hoefle, A.; Telles, V.; Begum, K.; Jimenez, F.; Ahuja, S.S.; Ahuja, S.K. Extensive repertoire of membrane-bound and soluble dendritic cell-specific ICAM-3-grabbing nonintegrin 1 (DC-SIGN1) and DC-SIGN2 isoforms. Inter-individual variation in expression of DC-SIGN transcripts. J. Biol. Chem. 2001, 276, 33196–33212. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.; Torensma, R.; van Vliet, S.J.; van Duijnhoven, G.C.; Adema, G.J.; van Kooyk, Y.; Figdor, C.G. Identification of DC-SIGN, a novel dendritic cell-specific ICAM-3 receptor that supports primary immune responses. Cell. 2000, 100, 575–585. [Google Scholar] [CrossRef]
- Svajger, U.; Anderluh, M.; Jeras, M.; Obermajer, N. C-type lectin DC-SIGN: An adhesion, signalling and antigen-uptake molecule that guides dendritic cells in immunity. Cell. Signal. 2010, 22, 1397–1405. [Google Scholar] [CrossRef] [PubMed]
- Van Kooyk, Y.; Geijtenbeek, T.B. DC-SIGN: Escape mechanism for pathogens. Nat. Rev. Immunol. 2003, 3, 697–709. [Google Scholar] [CrossRef] [PubMed]
- Tailleux, L.; Schwartz, O.; Herrmann, J.L.; Pivert, E.; Jackson, M.; Amara, A.; Legres, L.; Dreher, D.; Nicod, L.P.; Gluckman, J.C.; et al. DC-SIGN is the major Mycobacterium tuberculosis receptor on human dendritic cells. J. Exp. Med. 2003, 197, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Geijtenbeek, T.B.; Kwon, D.S.; Torensma, R.; van Vliet, S.J.; van Duijnhoven, G.C.; Middel, J.; Cornelissen, I.L.; Nottet, H.S.; KewalRamani, V.N.; Littman, D.R.; et al. DC-SIGN, a dendritic cell-specific HIV-1-binding protein that enhances trans-infection of T cells. Cell. 2000, 100, 587–597. [Google Scholar] [CrossRef]
- Aderem, A.; Underhill, D.M. Mechanisms of phagocytosis in macrophages. Annu. Rev. Immunol. 1999, 17, 593–623. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, S.; Grinstein, S. Phagocytosis and innate immunity. Curr. Opin. Immunol. 2002, 14, 136–145. [Google Scholar] [CrossRef]
- Zamze, S.; Martinez-Pomares, L.; Jones, H.; Taylor, P.R.; Stillion, R.J.; Gordon, S.; Wong, S.Y. Recognition of bacterial capsular polysaccharides and lipopolysaccharides by the macrophage mannose receptor. J. Biol. Chem. 2002, 277, 41613–41623. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Skurnik, M.; Zhang, S.S.; Schwartz, O.; Kalyanasundaram, R.; Bulgheresi, S.; He, J.J.; Klena, J.D.; Hinnebusch, B.J.; Chen, T. Human dendritic cell-specific intercellular adhesion molecule-grabbing nonintegrin (CD209) is a receptor for Yersinia pestis that promotes phagocytosis by dendritic cells. Infect. Immun. 2008, 76, 2070–2079. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Snyder, S.; Feng, P.; Azadi, P.; Zhang, S.S.; Bulgheresi, S.; Sanderson, K.E.; He, J.; Klena, J.; Chen, T. Role of N-acetylglucosamine within core lipopolysaccharide of several species of gram-negative bacteria in targeting the DC-SIGN (CD209). J. Immunol. 2006, 177, 4002–4011. [Google Scholar] [CrossRef] [PubMed]
- Cormier, E.G.; Durso, R.J.; Tsamis, F.; Boussemart, L.; Manix, C.; Olson, W.C.; Gardner, J.P.; Dragic, T. L-SIGN (CD209L) and DC-SIGN (CD209) mediate transinfection of liver cells by hepatitis C virus. Proc. Natl. Acad. Sci. USA 2004, 101, 14067–14072. [Google Scholar] [CrossRef] [PubMed]
- Do Vale, A.; Silva, M.T.; dos Santos, N.M.; Nascimento, D.S.; Reis-Rodrigues, P.; Costa-Ramos, C.; Ellis, A.E.; Azevedo, J.E. AIP56, a novel plasmid-encoded virulence factor of Photobacterium damselae subsp. piscicida with apoptogenic activity against sea bass macrophages and neutrophils. Mol. Microbiol. 2005, 58, 1025–1038. [Google Scholar] [CrossRef] [PubMed]
- Li, M.F.; Li, J.; Sun, L. CsMAP34, a teleost MAP with dual role: A promoter of MASP-assisted complement activation and a regulator of immune cell activity. Sci. Rep. 2016, 6, 39287. [Google Scholar] [CrossRef] [PubMed]
- Long, H.; Chen, C.; Zhang, J.; Sun, L. Antibacterial and antiviral properties of tongue sole (Cynoglossus semilaevis) high mobility group B2 protein are largely independent on the acidic C-terminal domain. Fish. Shellfish Immunol. 2014, 37, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.C.; Sun, L. Tongue sole (Cynoglossus semilaevis) prothymosin alpha: Cytokine-like activities associated with the intact protein and the C-terminal region that lead to antiviral immunity via Myd88-dependent and -independent pathways respectively. Dev. Comp. Immunol. 2015, 53, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Chen, Y.; Wang, M.; Yin, Y.; Pan, Y.; Gu, B.; Yu, G.; Li, Y.; Wong, B.H.; Liang, Y.; et al. A novel lectin from Agrocybe aegerita shows high binding selectivity for terminal N-acetylglucosamine. Biochem. J. 2012, 443, 369–378. [Google Scholar] [CrossRef] [PubMed]
- Voullaire, L.; Ioannou, P.; Nouri, S.; Williamson, R. Fetal nucleated red blood cells from CVS washings: An aid to development of first trimester non-invasive prenatal diagnosis. Prenat. Diagn. 2001, 21, 827–834. [Google Scholar] [CrossRef] [PubMed]
- Kurucz, E.; Markus, R.; Zsamboki, J.; Folkl-Medzihradszky, K.; Darula, Z.; Vilmos, P.; Udvardy, A.; Krausz, I.; Lukacsovich, T.; Gateff, E.; et al. Nimrod, a putative phagocytosis receptor with EGF repeats in Drosophila plasmatocytes. Curr. Biol. 2007, 17, 649–654. [Google Scholar] [CrossRef] [PubMed]




© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, S.; Sun, L. Tongue Sole CD209: A Pattern-Recognition Receptor that Binds a Broad Range of Microbes and Promotes Phagocytosis. Int. J. Mol. Sci. 2017, 18, 1848. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18091848
Jiang S, Sun L. Tongue Sole CD209: A Pattern-Recognition Receptor that Binds a Broad Range of Microbes and Promotes Phagocytosis. International Journal of Molecular Sciences. 2017; 18(9):1848. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18091848
Chicago/Turabian StyleJiang, Shuai, and Li Sun. 2017. "Tongue Sole CD209: A Pattern-Recognition Receptor that Binds a Broad Range of Microbes and Promotes Phagocytosis" International Journal of Molecular Sciences 18, no. 9: 1848. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18091848



