Chlamydia psittaci PmpD-N Modulated Chicken Macrophage Function by Triggering Th2 Polarization and the TLR2/MyD88/NF-κB Signaling Pathway
Abstract
:1. Introduction
2. Results
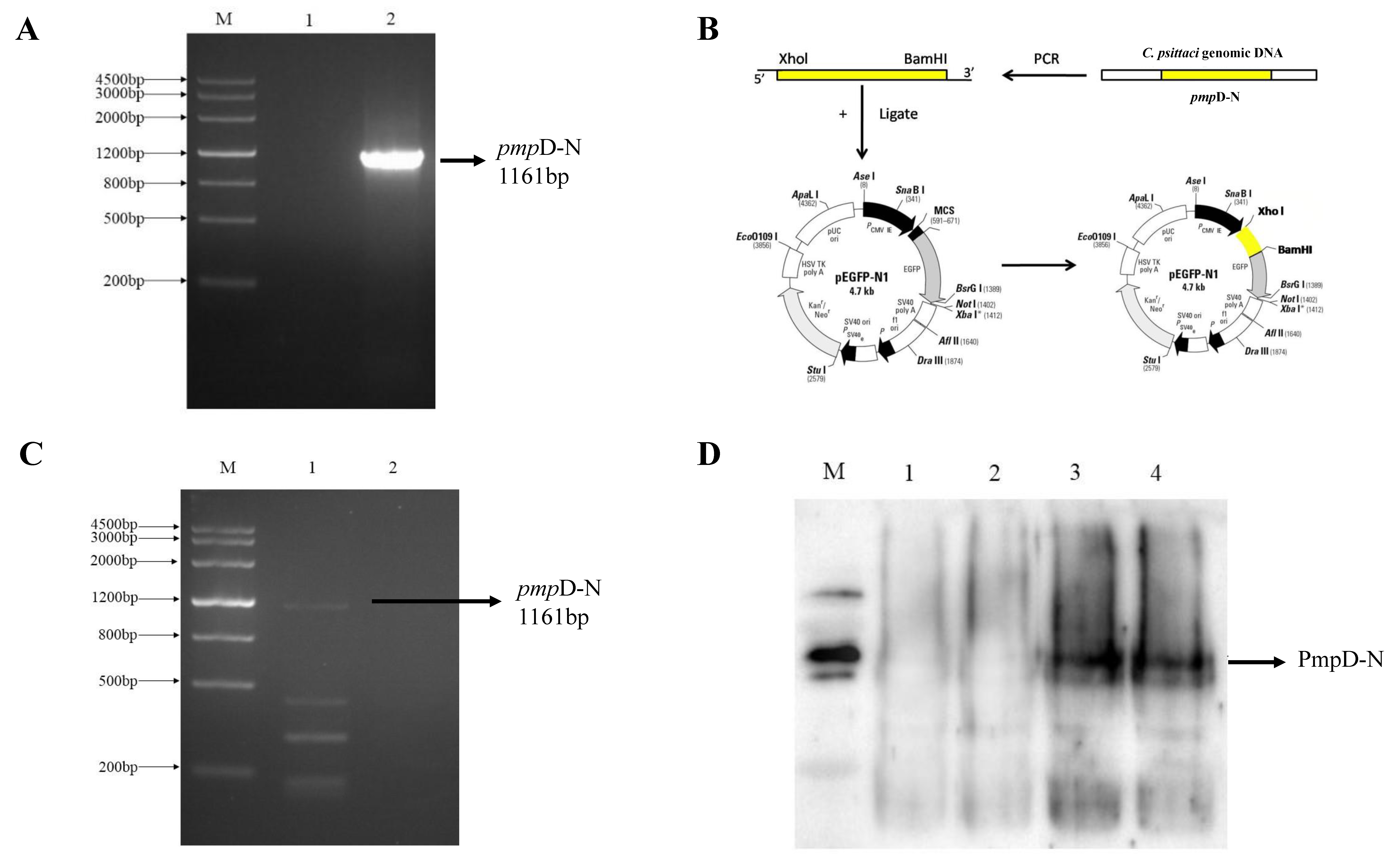
2.1. Endotoxin Removal of the Recombinant PmpD-N and Construction of pEGFP-N1-pmpD-N Plasmids
2.2. The Effect of PmpD-N and pEGFP-N1-pmpD-N on the Replication of C. Psittaci in HD11 Cells
2.3. The Effect of PmpD-N on the Phagocytic Function of HD11 Cells
2.4. The Effect of PmpD-N on the Release of Nitric Oxide in HD11 Cells
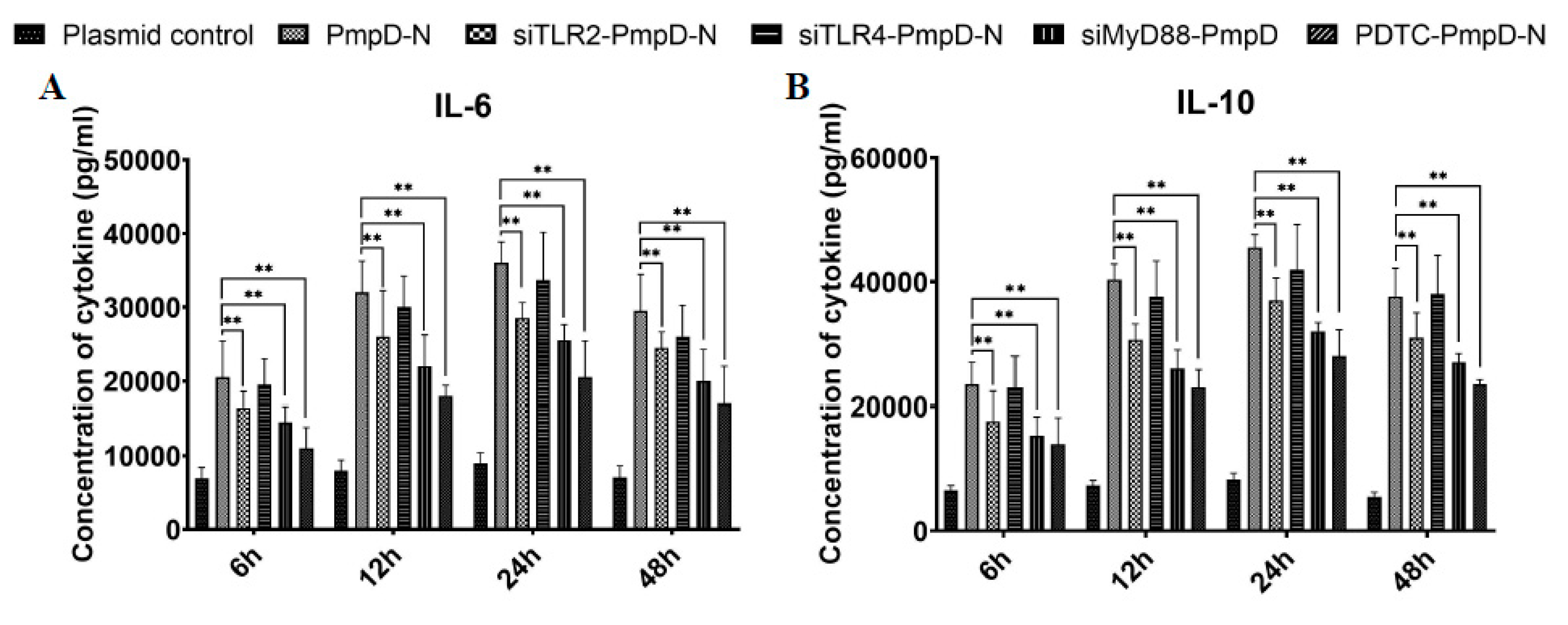
2.5. The Effect of PmpD-N on Cytokine Secretions in HD11 Cells
2.6. The Effect of PmpD-N on the mRNA Expression of Toll-like Receptors (TLRs) and Myeloid Differentiation Factor 88 (MyD88) and Nuclear Factor kappa B (NF-κB) in HD11 Cells
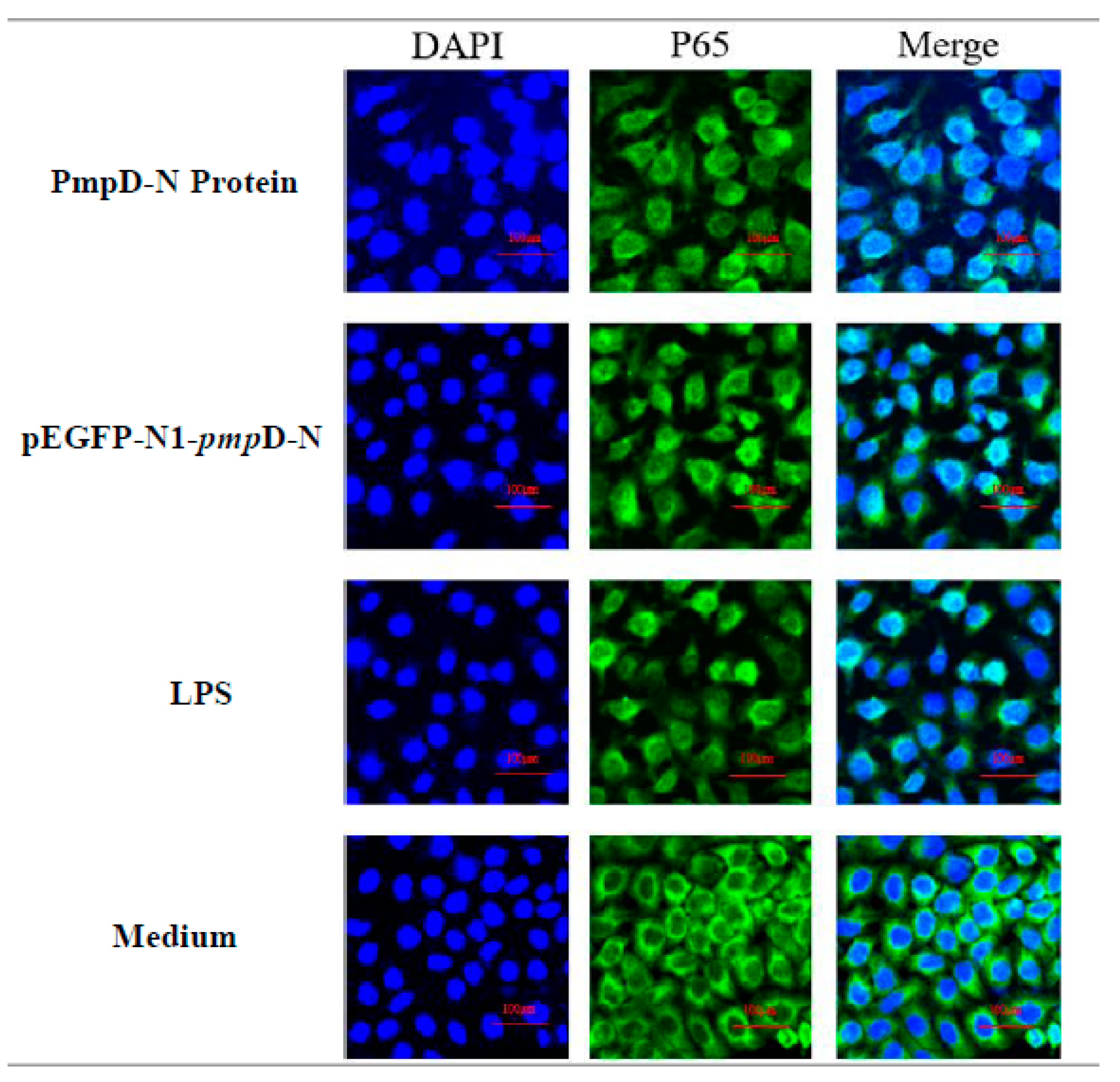
2.7. The Effect of PmpD-N on the Nuclear Translocation of NF-κB in the Treated HD11
2.8. The Effect of PmpD-N on the Binding of the Transcription Factor NF-κB protein to the NF-κB Promoter DNA
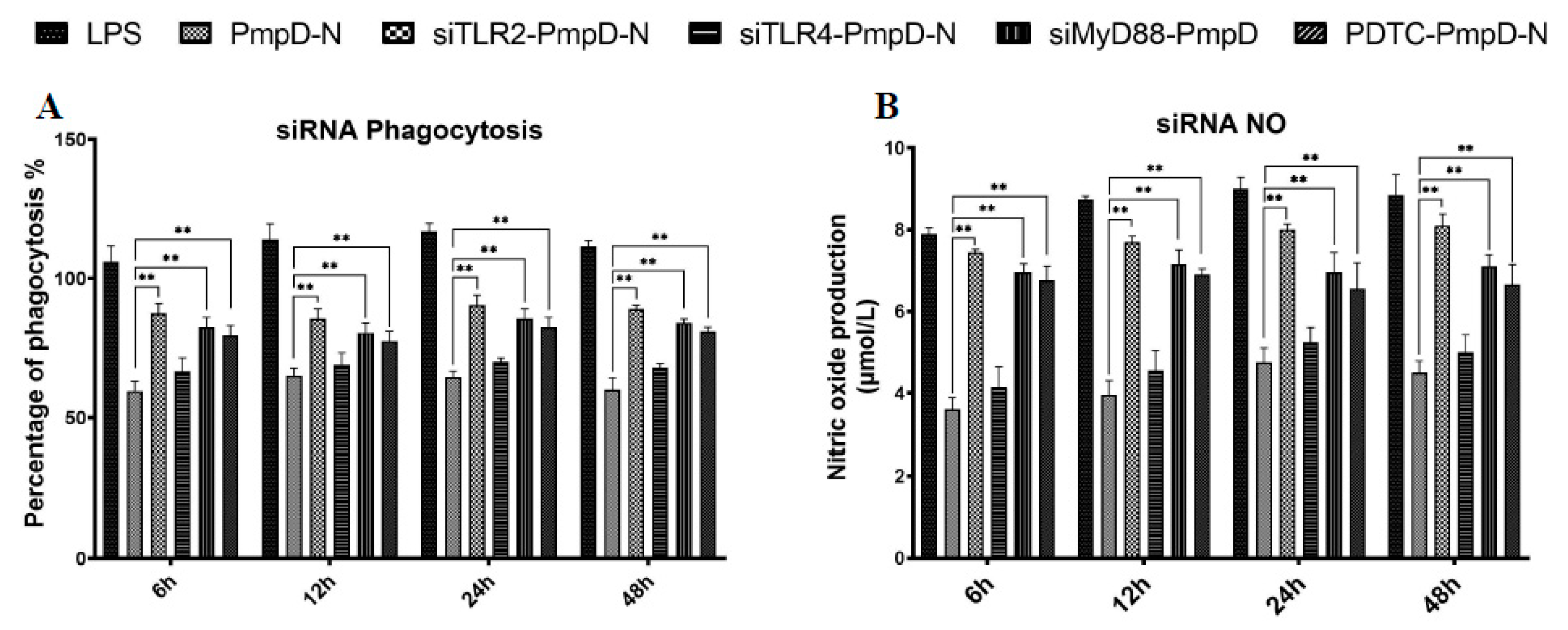
2.9. The Effect of Short Interfering RNA (siRNA) or Treatment with NF-κB Inhibitor on Cytokine Secretion, Phagocytic Function, and NO Production
3. Materials and Methods
3.1. Quantitative Endotoxin of the Recombinant PmpD-N
3.2. Cell Culture and Bacterial Strain
3.3. The Culture of HD11 Cells with PmpD-N, pEGFP-N1-pmpD-N, or Plasmids
3.4. C. psittaci Infection of HD11 Cells Treated with PmpD-N, pEGFP-N1-pmpD-N, or Plasmids
4. Western Blotting Assay
4.1. The Effect of PmpD-N or pEGFP-N1-pmpD-N on the Phagocytic Function of Macrophages
4.2. Nitric Oxide Determination by C.-psittaci-Infected Macrophages
4.3. Cytokine and Toll-like Receptor Analysis
4.4. Confocal Microscopy
4.5. Electrophoretic Mobility Shift Assay (EMSA)
4.6. Luciferase Reporter Assay
4.7. Short Interfering RNA (siRNA)
5. Statistical Analysis
6. Discussion
Author Contributions
Funding
Conflicts of Interest
Additional Information
References
- Rodolakis, A.; Yousef Mohamad, K. Zoonotic potential of Chlamydophila. Vet. Microbiol. 2010, 140, 382–391. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plummer, F.A.; Simonsen, J.N.; Cameron, D.W.; Ndinya-Achola, J.O.; Kreiss, J.K.; Gakinya, M.N.; Waiyaki, P.; Cheang, M.; Piot, P.; Ronald, A.R.; et al. Cofactors in male-female sexual transmission of human immunodeficiency virus type 1. J. Infect. Dis. 1991, 163, 233–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dickx, V.; Geens, T.; Deschuyffeleer, T.; Tyberghien, L.; Harkinezhad, T.; Beeckman, D.S.; Braeckman, L.; Vanrompay, D. Chlamydophila psittaci zoonotic risk assessment in a chicken and turkey slaughterhouse. J. Clin. Microbiol. 2010, 48, 3244–3250. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lagae, S.; Kalmar, I.; Laroucau, K.; Vorimore, F.; Vanrompay, D. Emerging Chlamydia psittaci infections in chickens and examination of transmission to humans. J. Med. Microbiol. 2014, 63, 399–407. [Google Scholar] [CrossRef] [Green Version]
- Horn, M.; Collingro, A.; Schmitz-Esser, S.; Beier, C.L.; Purkhold, U.; Fartmann, B.; Brandt, P.; Nyakatura, G.J.; Droege, M.; Frishman, D.; et al. Illuminating the evolutionary history of chlamydiae. Science 2004, 304, 728–730. [Google Scholar] [CrossRef]
- Vandahl, B.B.; Birkelund, S.; Christiansen, G. Genome and proteome analysis of Chlamydia. Proteomics 2004, 4, 2831–2842. [Google Scholar] [CrossRef]
- Voigt, A.; Schofl, G.; Saluz, H.P. The Chlamydia psittaci genome: A comparative analysis of intracellular pathogens. PLoS ONE 2012, 7, e35097. [Google Scholar] [CrossRef] [Green Version]
- Grimwood, J.; Stephens, R.S. Computational analysis of the polymorphic membrane protein superfamily of Chlamydia trachomatis and Chlamydia pneumoniae. Microb. Comp. Genom. 1999, 4, 187–201. [Google Scholar] [CrossRef]
- Van Lent, S.; De Vos, W.H.; Huot Creasy, H.; Marques, P.X.; Ravel, J.; Vanrompay, D.; Bavoil, P.; Hsia, R.C. Analysis of polymorphic membrane protein expression in cultured cells identifies PmpA and PmpH of Chlamydia psittaci as candidate factors in pathogenesis and immunity to infection. PLoS ONE 2016, 11, e0162392. [Google Scholar] [CrossRef] [Green Version]
- Moulder, J.W.; Hatch, T.P.; Byrne, G.I.; Kellogg, K.R. Immediate toxicity of high multiplicities of Chlamydia psittaci for mouse fibroblasts (L cells). Infect. Immun. 1976, 14, 277–289. [Google Scholar] [CrossRef] [Green Version]
- Todd, W.J.; Storz, J. Ultrastructural cytochemical evidence for the activation of lysosomes in the cytocidal effect of Chlamydia psittaci. Infect. Immun. 1975, 12, 638–646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lépinay, A.; Orfila, J.; Anteunis, A.; Boutry, J.; Orme-Roselli, L.; Robineaux, R. Study with electron microscopy of the development and morphology of Chlamydia psittaci in mouse macrophages. In Annales de l’Institut Pasteur; 1970; p. 222. [Google Scholar]
- Kiselev, A.O.; Stamm, W.E.; Yates, J.R.; Lampe, M.F. Expression, processing, and localization of PmpD of Chlamydia trachomatis serovar L2 during the chlamydial developmental cycle. PLoS ONE 2007, 2, e568. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wehrl, W.; Brinkmann, V.; Jungblut, P.R.; Meyer, T.F.; Szczepek, A.J. From the inside out—Processing of the chlamydial autotransporter PmpD and its role in bacterial adhesion and activation of human host cells. Mol. Microbiol. 2004, 51, 319–334. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.S.; Chu, J.; Zhang, Q.; Sun, W.; Zhang, T.Y.; He, C. Development of a novel PmpD-N ELISA for Chlamydia psittaci infection. Biomed. Environ. Sci. 2016, 29, 315–322. [Google Scholar] [PubMed]
- Liu, S.; Sun, W.; Huang, X.; Zhang, W.; Jia, C.; Luo, J.; Shen, Y.; El-Ashram, S.; He, C. A promising recombinant herpesvirus of turkeys vaccine expressing PmpD-N of Chlamydia psittaci based on elongation factor-1 alpha promoter. Front. Vet. Sci. 2017, 4, 221. [Google Scholar] [CrossRef] [Green Version]
- Niessner, A.; Kaun, C.; Zorn, G.; Speidl, W.; Turel, Z.; Christiansen, G.; Pedersen, A.S.; Birkelund, S.; Simon, S.; Georgopoulos, A.; et al. Polymorphic membrane protein (PMP) 20 and PMP 21 of Chlamydia pneumoniae induce proinflammatory mediators in human endothelial cells In Vitro by activation of the nuclear factor-kappaB pathway. J. Infect. Dis. 2003, 188, 108–113. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kari, L.; Southern, T.R.; Downey, C.J.; Watkins, H.S.; Randall, L.B.; Taylor, L.D.; Sturdevant, G.L.; Whitmire, W.M.; Caldwell, H.D. Chlamydia trachomatis polymorphic membrane protein D is a virulence factor involved in early host-cell interactions. Infect. Immun. 2014, 82, 2756–2762. [Google Scholar] [CrossRef] [Green Version]
- Brunner, M.; Gruber, M.; Schmid, D.; Baran, H.; Moeslinger, T. Proliferation of macrophages due to the inhibition of inducible nitric oxide synthesis by oxidized low-density lipoproteins. EXCLI J. 2015, 14, 439–451. [Google Scholar]
- Koziel, D.; Michaelis, U.; Kruse, T. Broad application and optimization of a single wash-step for integrated endotoxin depletion during protein purification. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2018, 1091, 101–107. [Google Scholar] [CrossRef]
- Beug, H.; von Kirchbach, A.; Doderlein, G.; Conscience, J.F.; Graf, T. Chicken hematopoietic cells transformed by seven strains of defective avian leukemia viruses display three distinct phenotypes of differentiation. Cell 1979, 18, 375–390. [Google Scholar] [CrossRef]
- Vanrompay, D.; Charlier, G.; Ducatelle, R.; Haesebrouck, F. Ultrastructural changes in avian Chlamydia psittaci serovar A-, B-, and D-infected buffalo green monkey cells. Infect. Immun. 1996, 64, 1265–1271. [Google Scholar] [CrossRef] [Green Version]
- Chu, J.; Zhang, Q.; Zhang, T.; Han, E.; Zhao, P.; Khan, A.; He, C.; Wu, Y. Chlamydia psittaci infection increases mortality of avian influenza virus H9N2 by suppressing host immune response. Sci. Rep. 2016, 6, 29421. [Google Scholar] [CrossRef]
- MacMicking, J.; Xie, Q.W.; Nathan, C. Nitric oxide and macrophage function. Annu. Rev. Immunol. 1997, 15, 323–350. [Google Scholar] [CrossRef]
- Beeckman, D.S.; Rothwell, L.; Kaiser, P.; Vanrompay, D.C. Differential cytokine expression in Chlamydophila psittaci genotype A-, B- or D-infected chicken macrophages after exposure to Escherichia coli O2:K1 LPS. Dev. Comp. Immunol. 2010, 34, 812–820. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, H.; Ren, J.; Tang, X.; Jing, Y.; Xing, D.; Zhao, G.; Yao, Z.; Yang, X.; Bai, H. IL-17A synergizes with IFN-gamma to upregulate iNOS and No production and inhibit chlamydial growth. PLoS ONE 2012, 7, e39214. [Google Scholar]
- Rajaram, K.; Nelson, D.E. Chlamydia muridarum infection of macrophages elicits bactericidal nitric oxide production via reactive oxygen species and cathepsin B. Infect. Immun. 2015, 83, 3164–3175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joyee, A.G.; Yang, X. Role of toll-like receptors in immune responses to chlamydial infections. Curr. Pharm. Des. 2008, 14, 593–600. [Google Scholar] [PubMed]
- Hodgson, A.; Wan, F. Interference with nuclear factor kappaB signaling pathway by pathogen-encoded proteases: Global and selective inhibition. Mol. Microbiol. 2016, 99, 439–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pan, Q.; Zhang, Q.; Chu, J.; Pais, R.; Liu, S.; He, C.; Eko, F.O. Chlamydia abortus Pmp18.1 induces IL-1beta secretion by TLR4 activation through the MyD88, NF-kappaB, and caspase-1 signaling pathways. Front. Cell. Infect. Microbiol. 2017, 7, 514. [Google Scholar] [CrossRef] [PubMed]
- Bas, S.; Neff, L.; Vuillet, M.; Spenato, U.; Seya, T.; Matsumoto, M.; Gabay, C. The proinflammatory cytokine response to Chlamydia trachomatis elementary bodies in human macrophages is partly mediated by a lipoprotein, the macrophage infectivity potentiator, through TLR2/TLR1/TLR6 and CD14. J. Immunol. 2008, 180, 1158–1168. [Google Scholar] [CrossRef]
- Omosun, Y.; McKeithen, D.; Ryans, K.; Kibakaya, C.; Blas-Machado, U.; Li, D.; Singh, R.; Inoue, K.; Xiong, Z.G.; Eko, F.; et al. Interleukin-10 modulates antigen presentation by dendritic cells through regulation of NLRP3 inflammasome assembly during Chlamydia infection. Infect. Immun. 2015, 83, 4662–4672. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nau, G.J.; Richmond, J.F.; Schlesinger, A.; Jennings, E.G.; Lander, E.S.; Young, R.A. Human macrophage activation programs induced by bacterial pathogens. Proc. Natl. Acad. Sci. USA 2002, 99, 1503–1508. [Google Scholar] [CrossRef] [PubMed] [Green Version]




© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chu, J.; Li, X.; Qu, G.; Wang, Y.; Li, Q.; Guo, Y.; Hou, L.; Liu, J.; Eko, F.O.; He, C. Chlamydia psittaci PmpD-N Modulated Chicken Macrophage Function by Triggering Th2 Polarization and the TLR2/MyD88/NF-κB Signaling Pathway. Int. J. Mol. Sci. 2020, 21, 2003. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21062003
Chu J, Li X, Qu G, Wang Y, Li Q, Guo Y, Hou L, Liu J, Eko FO, He C. Chlamydia psittaci PmpD-N Modulated Chicken Macrophage Function by Triggering Th2 Polarization and the TLR2/MyD88/NF-κB Signaling Pathway. International Journal of Molecular Sciences. 2020; 21(6):2003. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21062003
Chicago/Turabian StyleChu, Jun, Xiaohui Li, Guanggang Qu, Yihui Wang, Qiang Li, Yongxia Guo, Lei Hou, Jue Liu, Francis O. Eko, and Cheng He. 2020. "Chlamydia psittaci PmpD-N Modulated Chicken Macrophage Function by Triggering Th2 Polarization and the TLR2/MyD88/NF-κB Signaling Pathway" International Journal of Molecular Sciences 21, no. 6: 2003. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21062003




