Towards a New Understanding of Decision-Making by Hematopoietic Stem Cells
Abstract
:1. Introduction
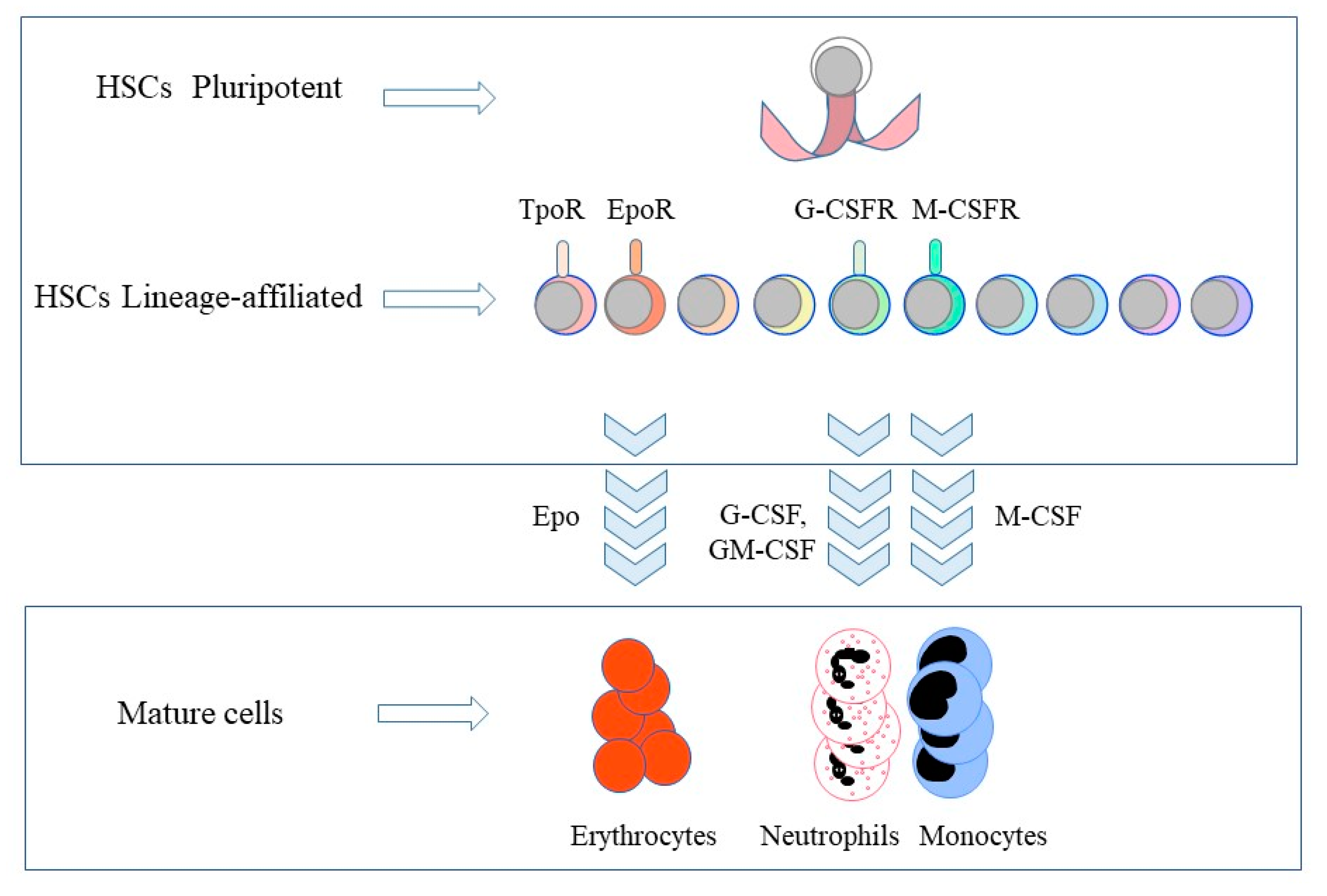
2. The Selective Expression of Cytokine Receptors by Hematopoietic Stem Cells
3. The Instructive Action of Hematopoietic Cytokines
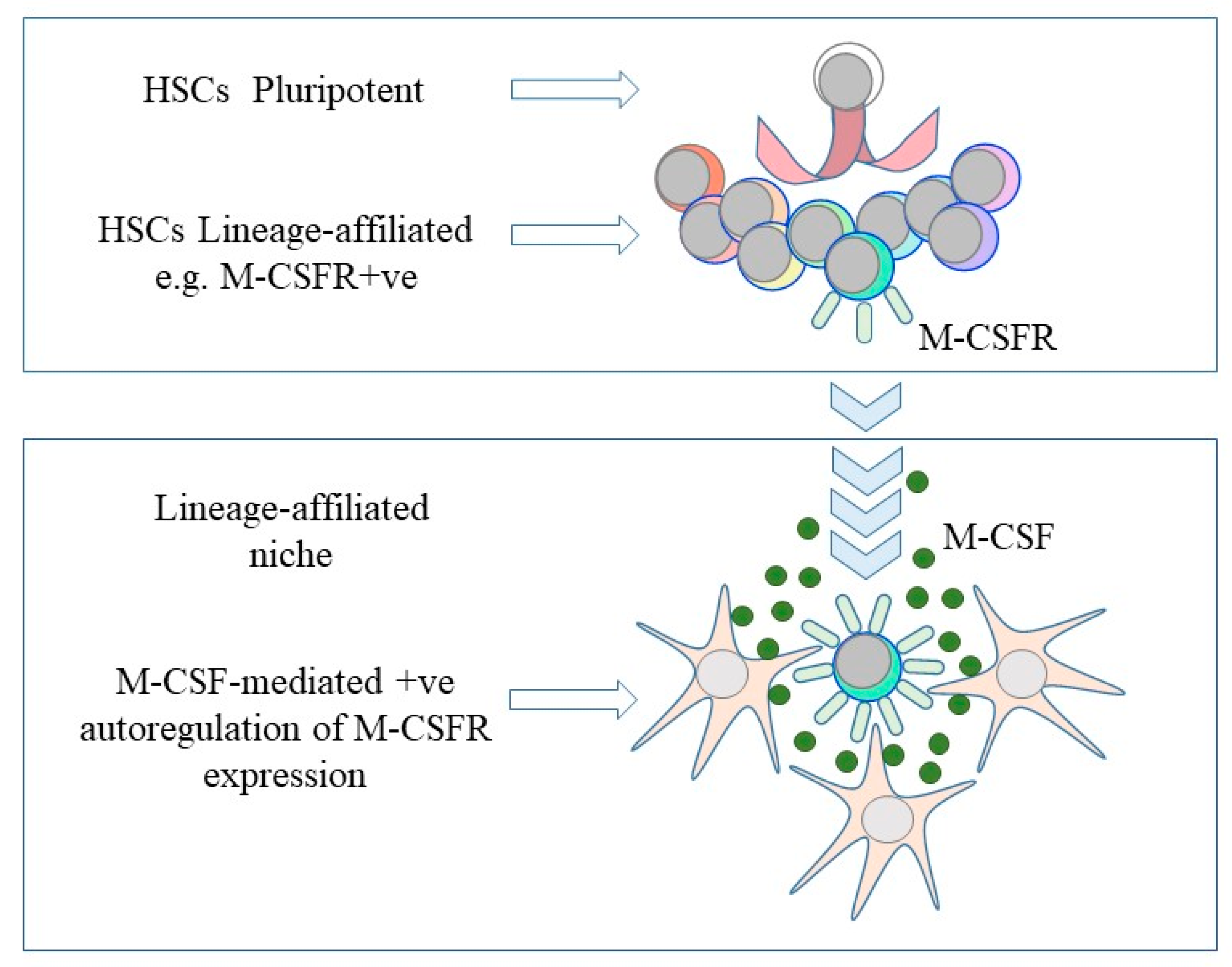
4. HSC Commitment to Differentiation and Finding a New Niche
5. Natural Variation within Hematopoietic Stem Cells
6. A Natural Selection Model of Hematopoiesis
7. Concluding Remarks
Funding
Acknowledgments
Conflicts of Interest
References
- Schofield, R. The relationship between the spleen colony-forming cell and the haematopoietic stem cell. Blood Cells 1978, 41, 7–25. [Google Scholar]
- Pinho, S.; Frenette, P.S. Haematopoietic stem cell activity and interactions with the niche. Nat. Rev. Mol. Cell Biol. 2019, 20, 303–320. [Google Scholar] [CrossRef] [PubMed]
- Okumura, N.; Ebihara, Y.; Tanaka, I.; Koike, K.; Komiyama, A.; Nakahata, T. Chemotactic and chemokinetic activities of stem cell factor on murine hematopoietic progenitor cells. Blood 1996, 87, 4100–4108. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Driessen, R.L.; Johnson, H.M.; Nilsson, S.K. Membrane-bound stem cell factor is a key regulator in the initial lodgement of stem cells within the endosteal marrow region. Exp. Hematol. 2003, 31, 1284–1291. [Google Scholar] [CrossRef] [PubMed]
- Ashman, L.K. The biology of stem cell factor and its receptor C-kit. Int. J. Biochem. Cell. Biol. 1999, 31, 1037–1051. [Google Scholar] [CrossRef]
- Sharma, S.; Gurudutta, G.U.; Satija, N.K.; Pati, S.; Gupta, P.; Verma, Y.K.; Singh, V.K.; Tripathi, R.P. Stem cell c-KIT and HOXB4 genes: Critical roles and mechanisms in self-renewal, proliferation, and differentiation. Stem Cells Dev. 2006, 15, 755–778. [Google Scholar] [CrossRef]
- Ceredig, R.; Rolink, A.G.; Brown, G. Models of haematopoiesis: Seeing the wood for the trees. Nat. Rev. Immunol. 2009, 9, 293–300. [Google Scholar] [CrossRef]
- Brown, G.; Ceredig, R. Modelling the hematopoietic landscape. Front. Cell Dev. Biol. 2019, 7, 104. [Google Scholar] [CrossRef]
- Hume, D.A.; MacDonald, K.P. Therapeutic applications of macrophage colony-stimulating factor (CSF-1) and antagonists of CSF-1 receptor (CSF-1R) signaling. Blood 2012, 119, 1810–1811, 1820. [Google Scholar] [CrossRef]
- Koury, M.J.; Bondurant, M.C. Erythropoietin retards DNA breakdown and prevents programmed death in erythroid progenitor cells. Science 1990, 248, 378–381. [Google Scholar] [CrossRef]
- Demetri, G.D.; Griffin, J.D. Granulocyte colony-stimulating factor and its receptor. Blood 1991, 78, 2791–2808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Metcalf, D. Hematopoietic regulators: Redundancy or subtlety? Blood 1993, 82, 3515–3523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gasson, J.C. Molecular physiology of granulocyte-macrophage colony stimulating factor. Blood 1991, 77, 1131–1145. [Google Scholar] [CrossRef] [PubMed]
- Kaushansky, K. Thrombopoietin: The primary regulator of megakaryocyte and platelet production. Thromb. Haemost. 1995, 74, 521–525. [Google Scholar] [CrossRef]
- Mooney, C.J.; Cunningham, A.; Tsapogas, P.; Toellner, K.-M.; Brown, G. Selective expression of Flt3 within the mouse hematopoietic stem cell compartment. Int. J. Mol. Sci. 2017, 18, 1037. [Google Scholar] [CrossRef]
- Osawa, M.; Hanada, K.; Hamada, H.; Nakauchi, H. Long-term lymphohematopoietic reconstitution by a single CD34-low/negative hematopoietic stem cell. Science 1996, 273, 242–245. [Google Scholar] [CrossRef] [Green Version]
- Shinjo, K.; Takeshita, A.; Higuchi, M.; Ohnishi, K.; Ohno, R. Erythropoietin receptor expression on human bone marrow erythroid precursor cells by a newly-devised quantitative flow-cytometric assay. Br. J. Haematol. 1997, 96, 551–558. [Google Scholar] [CrossRef]
- Zriwil, A.; Bolers, C.; Wittmann, L.; Green, J.C.A.; Woll, P.S.; Jacobsen, S.E.W.; Sitnicka, E. Macrophage colony-stimulating factor receptor marks and regulates a fetal myeloid-primed B-cell progenitor in mice. Blood 2016, 128, 217–226. [Google Scholar] [CrossRef] [Green Version]
- Mossadegh-Keller, N.; Sarrazin, S.; Kandalla, P.K.; Espinosa, L.; Stanley, E.R.; Nutt, S.L.; Moore, J.; Sieweke, M.H. M-CSF instructs myeloid lineage fate in single haematopoietic stem cells. Nature 2013, 497, 239–243. [Google Scholar] [CrossRef]
- Stoffel, R.; Ziegler, S.; Ghilhardi, N.; Ledermann, B.; de Sauvage, F.J.; Skoda, R.C. Permissive role of thrombopoietin and granulocyte colony-stimulating factor receptors in hematopoietic cell fate decisions in vivo. Proc. Natl. Acad Sci. USA 1999, 96, 698–702. [Google Scholar] [CrossRef] [Green Version]
- Mansson, R.; Hultquist, A.; Luc, S.; Yang, L.; Anderson, K.; Kharazi, S.; Al-Hashmi, S.; Liuba, K.; Thoren, L.; Adolffsson, J.; et al. Molecular evidence for hierarchical transcriptional lineage priming in fetal and adult stem cells and multipotent progenitors. Immunity 2007, 26, 407–419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Graaf, C.; Metcalf, D. Thrombopoietin and hematopoietic stem cells. Cell Cycle 2011, 10, 1582–1589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshihara, H.; Arai, F.; Hosokawa, K.; Hagiwara, T.; Takubo, K.; Nakamura, Y.; Gomei, Y.; Iwasaki, H.; Matsuoka, S.; Miyanoto, K.; et al. Thrombopoietin/MPL signaling regulates hematopoietic stem cell quiescence and interaction with the osteoblastic niche. Cell Stem Cell 2007, 1, 685–697.22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morrison, S.J.; Wright, D.E.; Weissman, I.L. Cyclophosphamide/granulocyte colony-stimulating factor induces hematopoietic stem cells to proliferate prior to mobilization. Proc. Natl. Acad. Sci. USA 1997, 94, 1908–1913. [Google Scholar] [CrossRef] [Green Version]
- Kondo, M.; Scherer, D.C.; Miyamoto, T.; King, A.G.; Akashi, K.; Sugamura, K.; Weissman, I.L. Cell-fate conversion of lymphoid-committed progenitors by instructive actions of cytokines. Nature 2000, 407, 383–386. [Google Scholar] [CrossRef] [PubMed]
- Tsapogas, P.; Mooney, C.J.; Brown, G.; Rolink, A. The cytokine Flt3-ligand in normal and malignant hematopoiesis. Int. J. Mol. Sci. 2017, 18, 1115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamamoto, R.; Morita, Y.; Ooehara, J.; Hamanaka, S.; Onodera, M.; Rudolph, K.L.; Ema, H.; Nakauchi, H. Clonal analysis unveils self-renewing lineage-restricted progenitors generated directly from hematopoietic stem cells. Cell 2013, 154, 1112–1126. [Google Scholar] [CrossRef] [Green Version]
- Shin, J.Y.; Hu, W.; Naramura, M.; Park, C.Y. High c-Kit expression identifies hematopoietic stem cells with impaired self-renewal and megakaryocytic bias. J. Exp. Med. 2014, 211, 217–231. [Google Scholar] [CrossRef] [Green Version]
- Sanjuan-Pla, A.; Macaulay, I.C.; Jensen, C.T.; Woll, P.S.; Luis, T.C.; Mead, A.; Moore, S.; Carella, C.; Bouriez Jones, T.; Chowdhury, O.; et al. Platelet-biased stem cells reside at the apex of the hematopoietic stem-cell hierarchy. Nature 2013, 502, 232–236. [Google Scholar] [CrossRef]
- Grover, A.; Mancini, E.; Moore, S.; Mead, A.J.; Atkinson, D.; Rasmussen, K.D.; O’Carroll, D.; Jacobsen, S.E.; Nerlove, C. Erythropoietin guides multipotent hematopoietic progenitor cells towards an erythroid fate. J. Exp. Med. 2014, 211, 181–188. [Google Scholar] [CrossRef] [Green Version]
- Metcalf, D.; Burgess, A.W. Clonal analysis of progenitor cell commitment of granulocyte or macrophage production. J. Cell Physiol. 1982, 111, 275–283. [Google Scholar] [CrossRef] [PubMed]
- Rieger, M.A.; Hoppe, P.S.; Smejkal, B.M.; Eitelhuber, A.C.; Schroeder, T. Hematopoietic cytokines can instruct lineage choice. Science 2009, 325, 217–218. [Google Scholar] [CrossRef] [PubMed]
- Tsapogas, P.; Swee, L.K.; Nusser, A.; Nuber, N.; Kreuzaler, M.; Capoferri, G.; Rolink, H.; Ceredig, R.; Rolink, A. In vivo evidence for an instructive role of fms-like tyrosine kinase-3 (FLT3) ligand in hematopoietic development. Haematologica 2014, 99, 638–646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Colmore, A.; Amorim, M.; Pontier, A.L.; Wang, S.; Jablonski, E.; Sipkins, D.A. Leukaemia cells create bone marrow niches that disrupt the behaviour of normal hematopoietic progenitor cells. Science 2008, 322, 1861–1865. [Google Scholar] [CrossRef] [Green Version]
- Chasis, J.A.; Mohandas, N. Erythroblastic islands: Niches for erythropoiesis. Blood 2008, 112, 470–478. [Google Scholar] [CrossRef] [Green Version]
- Comazzetto, S.; Murphy, M.M.; Berto, S.; Jeffery, E.; Zhao, Z.; Morrison, S.J. Restricted hematopoietic progenitors and reythropoiesis require SCF from leptin receptor+ niche cells in bone marrow. Cell Stem Cell 2019, 245, 477–486. [Google Scholar] [CrossRef] [Green Version]
- Zwezdaryk, K.J.; Coffelt, S.B.; Figueroa, Y.G.; Liu, J.; Phinney, D.G.; LaMarca, H.L.; Florez, L.; Morris, C.B.; Hoyle, G.W.; Scandurro, A.B. Erythropoietin, a hypoxia-regulated factor, elicits a pro-angiogenic program in human mesenchymal stem cells. Exp. Hemato. 2007, 35, 640–652. [Google Scholar] [CrossRef]
- Anagnostou, A.; Lee, E.S.; Keissiman, N.; Levinson, R.; Steiner, M. Erytrhopoietin has a mitogenic and postive chemotactic effect on endothelial cells. Proc. Natl. Acad. Sci. USA 1990, 87, 5978–5982. [Google Scholar] [CrossRef] [Green Version]
- Poniewierska-Baran, A.; Rajewska, J.R.; Ratajczak, M.Z. Erythropoietin enhances migration of human nuroblastoma cells: In vitro studies and potential therapeutic implication. J. Cancer Stem Cell Res. 2017, 5, e1003. [Google Scholar]
- Gomez-Cambronero, J.; Horn, J.; Paul, C.C.; Baumann, M.A. Granulocyte-macrophage colony-stimulating factor is a chemoattractant cytokine for human neutrophils: Involvement of the ribosomal p70 S6 kinase signalling pathway. J. Immunol. 2003, 171, 6846–6855. [Google Scholar] [CrossRef] [Green Version]
- Bussolino, F.; Wang, J.M.; Defilippi, P.; Turrini, F.; Sanavio, F.; Edgell, J.; Aglietta, M.; Arese, P.; Mantovani, A. Granulocyte- and granulocyte-macrophage-colony stimulating factors induce human endothelial cells to migrateand proliferate. Nature 1989, 337, 471. [Google Scholar] [CrossRef] [PubMed]
- Vaillant, P.; Muller, V.; Martinet, Y.; Martinet, N. Human granulocyte and granulocyte-macrophage-colony stimulating factors are chemotactic and “competence” growth factors for human mesenchymal cells. Biochem. Biophys. Res. Commun. 1993, 192, 879–885. [Google Scholar] [CrossRef] [PubMed]
- Fuller, K.; Owens, J.M.; Jagger, A.; Wilson, A.; Moss, R.; Chambers, T.J. Macrophage colony-stimulating factors stimulates survival and chemotactic behaviour in isolated osteoclasts. J. Exp. Med. 1993, 178, 1733–1744. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.M.; Griffin, J.D.; Rambaldi, A.; Chen, Z.D.; Mantovani, A. Induction of monocyte migration by recombinant macrophage colony-stimulating factor. J. Immunol. 1988, 141, 575–579. [Google Scholar]
- Vedham, V.; Phee, H.; Coggeshall, K.M. Vav activation and function as a Rac guanine nucleotide exchange factor in macrophage colony-stimulating factor-induced macrophage chemotaxis. Mol Cell Biol. 2005, 25, 4211–4220. [Google Scholar] [CrossRef] [Green Version]
- Pierce, J.H.; Di Marco, E.; Cox, G.W.; Lombardi, D.; Ruggiero, M.; Varesio, L.; Wang, L.M.; Choudhury, G.C.; Sakaguchi, A.Y.; Di Fiore, P.P. Macrophage-colony-stimulating factor (CSF-1) induces proliferation, chemotaxis, and reversible monocytic differentiation in myeloid progenitor cells transfected with human c-fms/CSF-1 receptor cDNA. Proc. Natl. Acad. Sci. USA 1990, 87, 5613–5617. [Google Scholar] [CrossRef] [Green Version]
- Pollard, J.W.; Bartocci, A.; Arceci, R.; Orlofsky, A.; Ladner, M.B.; Stanley, E.R. Apparent role of the macrophage growth factor, CSF-1, in placental development. Nature 1987, 330, 484–486. [Google Scholar] [CrossRef]
- Filderman, A.E.; Bruckner, A.; Kacinski, B.M.; Deng, N.; Remold, H.G. Macrophage colony-stimulating factor (CSF-1) enhances invasiveness in CSF-1 receptor-positive carcinoma cell lines. Cancer Res. 1992, 52, 3661–3666. [Google Scholar]
- Alberti-Servera, L.; von Muenchow, L.; Tsapogas, P.; Capoferri, G.; Eschbach, K.; Beisel, C. Single-cell RNA sequencing reveals developmental heterogeneity among early lymphoid progenitors. EMBO J. 2017, 36, 3619–3633. [Google Scholar] [CrossRef]
- Weinreb, C.; Rodriguez-Fraticelli, A.R.; Camargo, F.; Klein, A.M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 2020, 367, pii: eaaw3381. [Google Scholar] [CrossRef]
- Nestorowa, S.; Hamey, F.K.; Pijuan Sala, B.; Diamanti, E.; Shepherd, M.; Laurenti, E.; Wilson, N.K.; Kent, D.G.; Gottens, B. A single-cell resolution map of mouse hematopoietic stem and progenitor cell differentiation. Blood 2016, 128, e20–e31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Furusawa, C.; Kaneko, K. A dynamical-systems view of stem cell biology. Science 2012, 338, 215–217. [Google Scholar] [CrossRef] [PubMed]
- Raser, J.M.; O’Shea, E.K. Noise in gene expression: Origins, consequences, and control. Science 2005, 309, 2010–2013. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lund, R.J.; Narva, E.; Lahesmaa, R. Genetic and epigenetic stability of human pluripotent stem cells. Nat. Rev. Genet. 2012, 13, 732–744. [Google Scholar] [CrossRef]
- Barroso, G.V.; Puzovic, N.; Dutheil, J.Y. The evolution of gene-specific transcriptional noise is driven by selection at the pathway level. Genetics 2018, 208, 1173–1189. [Google Scholar] [CrossRef] [Green Version]
- Doi, Y.; Yokota, Y.; Satoh, Y.; Okuzaki, D.; Tokunaga, M.; Ishibashi, T.; Sudo, T.; Ueda, T.; Shingai, Y.; Ichi, M. Variable SATB1 levels regulate hematopoietic stem cell heterogeneity with distinct lineage fate. Cell Rep. 2018, 23, 3223–3235. [Google Scholar] [CrossRef]
- Chang, H.H.; Hemberg, M.; Barahana, M.; Ingber, D.E.; Huang, S. Transcriptome-wide noise controls lineage choice in mammalian progenitor cells. Nature 2008, 453, 544–547. [Google Scholar] [CrossRef] [Green Version]
- Yokota, T. “Hierarchy” and “Holoaciary”: A paradigm of the hematopoietic system. Cells 2019, 8, pii E1138. [Google Scholar] [CrossRef] [Green Version]
- Lloyd, A.M.; Schena, M.; Walbot, V.; Davis, R.W. Epidermal cell fate determination in Arabidopsis: Patterns defined by a steroid-inducible regulator. Science 1994, 266, 436–439. [Google Scholar] [CrossRef]
- Yu, Q.; Li, P.; Liang, N.; Wang, H.; Xu, M.; Wu, S. Cell-fate specification in Arabidopsis roots requires coordinative action of lineage instruction and positional reprogramming. Plant Physiol. 2017, 175, 816–827. [Google Scholar] [CrossRef] [Green Version]
- Szyf, M.; McGowan, P.M.; Meaney, M.J. The social environment and the epigenome. Environ. Mol. Mutagenesis 2008, 49, 46–60. [Google Scholar] [CrossRef] [PubMed]
- Bloom, A.B.; Zaman, M.H. Influence of the microenvironment on cell fate determination and migration. Physiol. Genomics 2014, 46, 309–314. [Google Scholar] [CrossRef] [PubMed] [Green Version]
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brown, G. Towards a New Understanding of Decision-Making by Hematopoietic Stem Cells. Int. J. Mol. Sci. 2020, 21, 2362. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21072362
Brown G. Towards a New Understanding of Decision-Making by Hematopoietic Stem Cells. International Journal of Molecular Sciences. 2020; 21(7):2362. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21072362
Chicago/Turabian StyleBrown, Geoffrey. 2020. "Towards a New Understanding of Decision-Making by Hematopoietic Stem Cells" International Journal of Molecular Sciences 21, no. 7: 2362. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21072362



