Analysis of Astroglial Secretomic Profile in the Mecp2-Deficient Male Mouse Model of Rett Syndrome
Abstract
:1. Introduction
2. Results
2.1. Proteomic Analysis of Astroglial Secretome
2.2. Functional and Network Analyses of Secreted Proteins Deregulated upon Mecp2 Deficiency
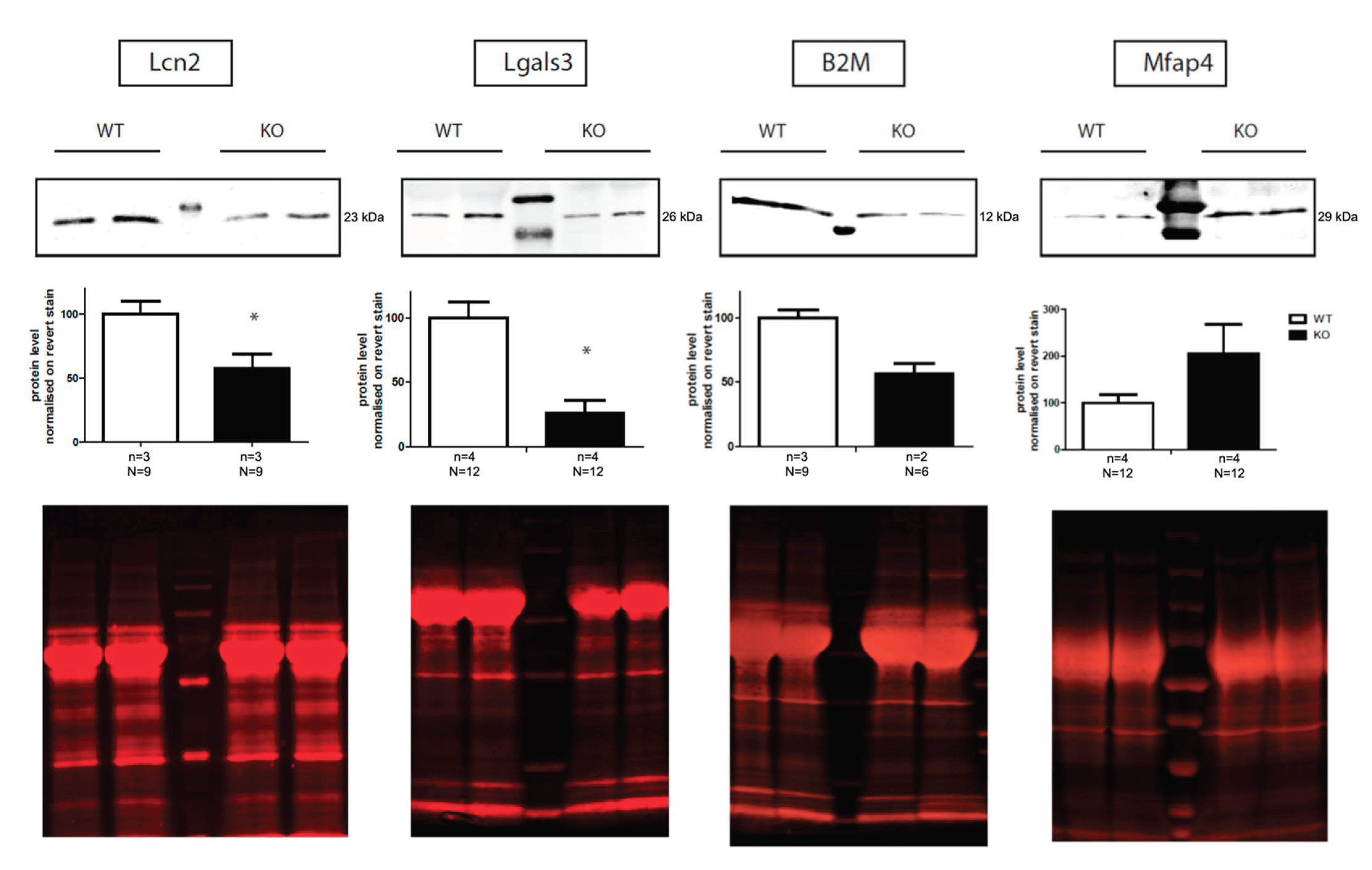
2.3. Validation of Proteomic Data by Western Blot
2.4. Lcn2 and Lgals3 Supplementation Improves Dendritic Arborization of Mecp2 KO Neurons
3. Discussion
3.1. Limitations
3.2. GO Analysis Reveals Enriched Biological ProcessesiIn the RTT Astroglial Secretome
3.3. Network Analysis Identifies Two Proteins of Interest: Lcn2 and Lgals3
3.4. A Well-Known Candidate, Lcn2
3.5. A Multifunction Protein, Lgals3
3.6. Lcn2, Lgals3 and BDNF
4. Materials and Methods
4.1. Animals
4.2. Reagents
4.3. Primary Astroglial Culture and Medium Collection
4.4. Secretomics
4.4.1. Sample Purification
4.4.2. Protein Digestion and iTRAQ Labelling
4.4.3. Peptide Separation by OFFGEL Isoelectrofocusing and Nano LC-MS/MS Analysis
4.4.4. iTRAQ Data Analysis
4.4.5. Gene Ontology Analysis
4.5. Western Blotting
4.6. Primary Neuronal Cultures and Treatments
4.7. Immunofluorescence
4.8. Sholl Aanalysis
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Amir, R.E.; Van den Veyver, I.B.; Wan, M.; Tran, C.Q.; Francke, U.; Zoghbi, H.Y. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat. Genet. 1999, 23, 185–188. [Google Scholar] [CrossRef]
- Hagberg, B.; Aicardi, J.; Dias, K.; Ramos, O. A progressive syndrome of autism, dementia, ataxia, and loss of purposeful hand use in girls: Rett’s syndrome: Report of 35 cases. Ann. Neurol. 1983, 14, 471–479. [Google Scholar] [CrossRef]
- Ip, J.P.K.; Mellios, N.; Sur, M. Rett syndrome: Insights into genetic, molecular and circuit mechanisms. Nat. Rev. Neurosci. 2018, 19, 368–382. [Google Scholar] [CrossRef] [PubMed]
- Kishi, N.; Macklis, J.D. MECP2 is progressively expressed in post-migratory neurons and is involved in neuronal maturation rather than cell fate decisions. Mol. Cell. Neurosci. 2004, 27, 306–321. [Google Scholar] [CrossRef] [PubMed]
- Mullaney, B.C.; Johnston, M.V.; Blue, M.E. Developmental expression of methyl-CpG binding protein 2 is dy-namically regulated in the rodent brain. Neuroscience 2004, 123, 939–949. [Google Scholar] [CrossRef]
- Barres, B.A. The mystery and magic of glia: A perspective on their roles in health and disease. Neuron 2008, 60, 430–440. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Durkee, C.A.; Araque, A. Diversity and specificity of astrocyte-neuron communication. Neuroscience 2019, 396, 73–78. [Google Scholar] [CrossRef]
- Santello, M.; Toni, N.; Volterra, A. Astrocyte function from information processing to cognition and cognitive impairment. Nat. Neurosci. 2019, 22, 154–166. [Google Scholar] [CrossRef] [Green Version]
- Shahbazian, M.D.; Antalffy, B.; Armstrong, D.L.; Zoghbi, H.Y. Insight into Rett syndrome: MeCP2 levels dis-play tissue- and cell-specific differences and correlate with neuronal maturation. Hum. Mol. Genet. 2002, 11, 115–124. [Google Scholar] [CrossRef]
- Ballas, N.; Lioy, D.T.; Grunseich, C.; Mandel, G. Non-cell autonomous influence of MeCP2-deficient glia on neuronal dendritic morphology. Nat. Neurosci. 2009, 12, 311–317. [Google Scholar] [CrossRef]
- Maezawa, I.; Swanberg, S.; Harvey, D.; LaSalle, J.M.; Jin, L.W. Rett syndrome astrocytes are abnormal and spread MeCP2 deficiency through gap junctions. J. Neurosci. 2009, 29, 5051–5061. [Google Scholar] [CrossRef] [PubMed]
- Lioy, D.T.; Garg, S.K.; Monaghan, C.E.; Raber, J.; Foust, K.D.; Kaspar, B.K.; Hirrlinger, P.G.; Kirchhoff, F.; Bissonnette, J.M.; Ballas, N.; et al. A role for glia in the progression of Rett’s syndrome. Nature 2011, 475, 497–500. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.; Singh, J.; Pillai, P.P.; Frost, E.E. Involvement of MeCP2 in regulation of myelin-related gene ex-pression in cultured rat oligodendrocytes. J. Mol. Neurosci. 2015, 57, 176–184. [Google Scholar] [CrossRef] [PubMed]
- Tochiki, K.K.; Cunningham, J.; Hunt, S.P.; Geranton, S.M. The expression of spinal methyl-CpG-binding protein 2, DNA methyltransferases and histone deacetylases is modulated in persistent pain states. Mol. Pain. 2012, 8, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garg, S.K.; Lioy, D.T.; Knopp, S.J.; Bissonnette, J.M. Conditional depletion of methyl-CpG-binding protein 2 in astrocytes depresses the hypercapnic ventilatory response in mice. J. Appl. Physiol. 2015, 119, 670–676. [Google Scholar] [CrossRef] [Green Version]
- Turovsky, E.; Karagiannis, A.; Abdala, A.P.; Gourine, A.V. Impaired CO2 sensitivity of astrocytes in a mouse model of Rett syndrome. J. Physiol. 2015, 593, 3159–3168. [Google Scholar] [CrossRef] [Green Version]
- Delepine, C.; Nectoux, J.; Letourneur, F.; Baud, V.; Chelly, J.; Billuart, P.; Bienvenu, T. Astrocyte transcriptome from the Mecp2(308)-truncated mouse model of rett syndrome. Neuromol. Med. 2015, 17, 353–363. [Google Scholar] [CrossRef]
- Rakela, B.; Brehm, P.; Mandel, G. Astrocytic modulation of excitatory synaptic signaling in a mouse model of Rett syndrome. Elife 2018, 7, e31629. [Google Scholar] [CrossRef]
- Chenau, J.; Michelland, S.; de Fraipont, F.; Josserand, V.; Coll, J.L.; Favrot, M.C.; Seve, M. The cell line secre-tome, a suitable tool for investigating proteins released in vivo by tumors: Application to the study of p53-modulated proteins secreted in lung cancer cells. J. Proteome. Res. 2009, 8, 4579–4591. [Google Scholar] [CrossRef]
- Kim, J.H.; Afridi, R.; Lee, W.H.; Suk, K. Proteomic examination of the neuroglial secretome: Lessons for the clinic. Expert Rev. Proteom. 2020, 17, 207–220. [Google Scholar] [CrossRef]
- Song, P.; Kwon, Y.; Joo, J.Y.; Kim, D.G.; Yoon, J.H. Secretomics to discover regulators in diseases. Int. J. Mol. Sci. 2019, 20, 3893. [Google Scholar] [CrossRef] [Green Version]
- Williams, E.C.; Zhong, X.; Mohamed, A.; Li, R.; Liu, Y.; Dong, Q.; Ananiev, G.E.; Mok, J.C.; Lin, B.R.; Lu, J.; et al. Mutant astrocytes differentiated from Rett syndrome pa-tients-specific iPSCs have adverse effects on wild-type neurons. Hum. Mol. Genet. 2014, 23, 2968–2980. [Google Scholar] [CrossRef]
- Goron, A.; Breuillard, C.; Cunin, V.; Bourgoin-Voillard, S.; Seve, M.; Moinard, C. Modulation of muscle protein synthesis by amino acids: What consequences for the secretome? A preliminary in vitro study. Amino Acids 2019, 51, 1681–1688. [Google Scholar] [CrossRef]
- Vizcaino, J.A.; Deutsch, E.W.; Wang, R.; Csordas, A.; Reisinger, F.; Rios, D.; Dianes, J.A.; Sun, Z.; Farrah, T.; Bandeira, N.; et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 2014, 32, 223–226. [Google Scholar] [CrossRef]
- Zoghbi, H.Y. Postnatal neurodevelopmental disorders: Meeting at the synapse? Science 2003, 302, 826–830. [Google Scholar] [CrossRef]
- Armstrong, D.D. Neuropathology of Rett syndrome. J. Child. Neurol. 2005, 20, 747–753. [Google Scholar] [CrossRef]
- van den Pol, A.N.; Spencer, D.D. Differential neurite growth on astrocyte substrates: Interspecies facilitation in green fluorescent protein-transfected rat and human neurons. Neuroscience 2000, 95, 603–616. [Google Scholar] [CrossRef]
- Ferreira, A.C.; Pinto, V.; Da Mesquita, S.; Novais, A.; Sousa, J.C.; Correia-Neves, M.; Sousa, N.; Palha, J.A.; Marques, F. Lipocalin-2 is involved in emotional behaviors and cognitive function. Front. Cell Neurosci. 2013, 7, 122. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferreira, A.C.; Santos, T.; Sampaio-Marques, B.; Novais, A.; Mesquita, S.D.; Ludovico, P.; Bernardino, L.; Cor-reia-Neves, M.; Sousa, N.; Palha, J.A.; et al. Lipocalin-2 regulates adult neurogenesis and contextual discriminative behaviours. Mol. Psychiatry 2018, 23, 1031–1039. [Google Scholar] [CrossRef] [PubMed]
- Yoo, H.I.; Kim, E.G.; Lee, E.J.; Hong, S.Y.; Yoon, C.S.; Hong, M.J.; Park, S.J.; Woo, R.S.; Baik, T.K.; Song, D.Y. Neuroanatomical distribution of galectin-3 in the adult rat brain. J. Mol. Histol. 2017, 48, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Al-Dalahmah, O.; Campos Soares, L.; Nicholson, J.; Draijer, S.; Mundim, M.; Lu, V.M.; Sun, B.; Tyler, T.; Ador-jan, I.; O’Neill, E.; et al. Galectin-3 modulates postnatal subventricular zone gliogenesis. Glia 2020, 68, 435–450. [Google Scholar] [CrossRef] [Green Version]
- Kimpinski, K.; Campenot, R.B.; Mearow, K. Effects of the neurotrophins nerve growth factor, neurotrophin-3, and brain-derived neurotrophic factor (BDNF) on neurite growth from adult sensory neurons in compartmented cultures. J. Neurobiol. 1997, 33, 395–410. [Google Scholar] [CrossRef]
- Verkhratsky, A.; Matteoli, M.; Parpura, V.; Mothet, J.P.; Zorec, R. Astrocytes as secretory cells of the central nervous system: Idiosyncrasies of vesicular secretion. EMBO J. 2016, 35, 239–257. [Google Scholar] [CrossRef] [Green Version]
- Bayraktar, O.A.; Bartels, T.; Holmqvist, S.; Kleshchevnikov, V.; Martirosyan, A.; Polioudakis, D.; Ben Haim, L.; Young, A.M.H.; Batiuk, M.Y.; Prakash, K.; et al. Astrocyte layers in the mammalian cerebral cortex revealed by a single-cell in situ transcriptomic map. Nat. Neurosci. 2020, 23, 500–509. [Google Scholar] [CrossRef]
- Boutej, H.; Rahimian, R.; Thammisetty, S.S.; Beland, L.C.; Lalancette-Hebert, M.; Kriz, J. Diverging mRNA and protein networks in activated microglia reveal srsf3 suppresses translation of highly upregulated innate immune transcripts. Cell Rep. 2017, 21, 3220–3233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flo, T.H.; Smith, K.D.; Sato, S.; Rodriguez, D.J.; Holmes, M.A.; Strong, R.K.; Akira, S.; Aderem, A. Lipocalin 2 mediates an innate immune response to bacterial infection by sequestrating iron. Nature 2004, 432, 917–921. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Jha, M.K.; Suk, K. Lipocalin-2 in the inflammatory activation of brain astrocytes. Crit. Rev. Immunol. 2015, 35, 77–84. [Google Scholar] [CrossRef]
- Lee, S.; Lee, J.; Kim, S.; Park, J.Y.; Lee, W.H.; Mori, K.; Kim, S.H.; Kim, I.K.; Suk, K. A dual role of lipocalin 2 in the apoptosis and deramification of activated microglia. J. Immunol. 2007, 179, 3231–3241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, S.; Park, J.Y.; Lee, W.H.; Kim, H.; Park, H.C.; Mori, K.; Suk, K. Lipocalin-2 is an autocrine mediator of re-active astrocytosis. J. Neurosci. 2009, 29, 234–249. [Google Scholar] [CrossRef]
- Burguillos, M.A.; Svensson, M.; Schulte, T.; Boza-Serrano, A.; Garcia-Quintanilla, A.; Kavanagh, E.; Santiago, M.; Viceconte, N.; Oliva-Martin, M.J.; Osman, A.M.; et al. Microglia-secreted galectin-3 acts as a toll-like receptor 4 ligand and contributes to microglial activation. Cell Rep. 2015, 10, 1626–1638. [Google Scholar] [CrossRef] [Green Version]
- Lalancette-Hebert, M.; Swarup, V.; Beaulieu, J.M.; Bohacek, I.; Abdelhamid, E.; Weng, Y.C.; Sato, S.; Kriz, J. Galectin-3 is required for resident microglia activation and proliferation in response to ischemic injury. J. Neurosci. 2012, 32, 10383–10395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Derecki, N.C.; Cronk, J.C.; Lu, Z.; Xu, E.; Abbott, S.B.; Guyenet, P.G.; Kipnis, J. Wild-type microglia arrest pa-thology in a mouse model of Rett syndrome. Nature 2012, 484, 105–109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Wegener, J.E.; Huang, T.W.; Sripathy, S.; De Jesus-Cortes, H.; Xu, P.; Tran, S.; Knobbe, W.; Leko, V.; Britt, J.; et al. Wild-type microglia do not reverse pathology in mouse models of Rett syndrome. Nature 2015, 521, E1–E4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Periyasamy, P.; Thangaraj, A.; Guo, M.L.; Hu, G.; Callen, S.; Buch, S. Epigenetic promoter DNA methylation of miR-124 promotes HIV-1 tat-mediated microglial activation via MECP2-STAT3 axis. J. Neurosci. 2018, 38, 5367–5383. [Google Scholar] [CrossRef] [PubMed]
- Squillaro, T.; Alessio, N.; Cipollaro, M.; Melone, M.A.; Hayek, G.; Renieri, A.; Giordano, A.; Galderisi, U. Re-duced expression of MECP2 affects cell commitment and maintenance in neurons by triggering senescence: New perspective for Rett syndrome. Mol. Biol. Cell. 2012, 23, 1435–1445. [Google Scholar] [CrossRef]
- Dumic, J.; Dabelic, S.; Flogel, M. Galectin-3: An open-ended story. Biochim. Biophys. Acta 2006, 1760, 616–635. [Google Scholar] [CrossRef] [PubMed]
- Jha, M.K.; Lee, S.; Park, D.H.; Kook, H.; Park, K.G.; Lee, I.K.; Suk, K. Diverse functional roles of lipocalin-2 in the central nervous system. Neurosci. Biobehav. Rev. 2015, 49, 135–156. [Google Scholar] [CrossRef]
- Krzeslak, A.; Lipinska, A. Galectin-3 as a multifunctional protein. Cell Mol. Biol. Lett. 2004, 9, 305–328. [Google Scholar] [PubMed]
- Xu, X.; Miller, E.C.; Pozzo-Miller, L. Dendritic spine dysgenesis in Rett syndrome. Front. Neuroanat. 2014, 8, 97. [Google Scholar] [CrossRef] [Green Version]
- Berard, J.L.; Zarruk, J.G.; Arbour, N.; Prat, A.; Yong, V.W.; Jacques, F.H.; Akira, S.; David, S. Lipocalin 2 is a novel immune mediator of experimental autoimmune encephalomyelitis pathogenesis and is modulated in mul-tiple sclerosis. Glia 2012, 60, 1145–1159. [Google Scholar] [CrossRef]
- Bi, F.; Huang, C.; Tong, J.; Qiu, G.; Huang, B.; Wu, Q.; Li, F.; Xu, Z.; Bowser, R.; Xia, X.G.; et al. Reactive as-trocytes secrete lcn2 to promote neuron death. Proc. Natl. Acad. Sci. USA 2013, 110, 4069–4074. [Google Scholar] [CrossRef] [Green Version]
- Ip, J.P.; Nocon, A.L.; Hofer, M.J.; Lim, S.L.; Muller, M.; Campbell, I.L. Lipocalin 2 in the central nervous sys-tem host response to systemic lipopolysaccharide administration. J Neuroinflammation 2011, 8, 124. [Google Scholar] [CrossRef] [Green Version]
- Jang, E.; Kim, J.H.; Lee, S.; Kim, J.H.; Seo, J.W.; Jin, M.; Lee, M.G.; Jang, I.S.; Lee, W.H.; Suk, K. Phenotypic polarization of activated astrocytes: The critical role of lipocalin-2 in the classical inflammatory activation of astrocytes. J. Immunol. 2013, 191, 5204–5219. [Google Scholar] [CrossRef] [Green Version]
- Jang, E.; Lee, S.; Kim, J.H.; Kim, J.H.; Seo, J.W.; Lee, W.H.; Mori, K.; Nakao, K.; Suk, K. Secreted protein lipocalin-2 promotes microglial M1 polarization. FASEB J. 2013, 27, 1176–1190. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yasui, D.H.; Xu, H.; Dunaway, K.W.; Lasalle, J.M.; Jin, L.W.; Maezawa, I. MeCP2 modulates gene expression pathways in astrocytes. Mol. Autism 2013, 4, 3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ben-Shachar, S.; Chahrour, M.; Thaller, C.; Shaw, C.A.; Zoghbi, H.Y. Mouse models of MeCP2 disorders share gene expression changes in the cerebellum and hypothalamus. Hum. Mol. Genet. 2009, 18, 2431–2442. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Colantuoni, C.; Jeon, O.H.; Hyder, K.; Chenchik, A.; Khimani, A.H.; Narayanan, V.; Hoffman, E.P.; Kaufmann, W.E.; Naidu, S.; Pevsner, J. Gene expression profiling in postmortem Rett Syndrome brain: Differential gene expression and patient classification. Neurobiol. Dis. 2001, 8, 847–865. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Banerjee, A.; Miller, M.T.; Li, K.; Sur, M.; Kaufmann, W.E. Towards a better diagnosis and treatment of Rett syndrome: A model synaptic disorder. Brain 2019, 142, 239–248. [Google Scholar] [CrossRef]
- Wesley, U.V.; Vemuganti, R.; Ayvaci, E.R.; Dempsey, R.J. Galectin-3 enhances angiogenic and migratory po-tential of microglial cells via modulation of integrin linked kinase signaling. Brain Res. 2013, 1496, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Pecorelli, A.; Cervellati, C.; Cordone, V.; Hayek, J.; Valacchi, G. Compromised immune/inflammatory responses in Rett syndrome. Free Radic. Biol. Med. 2020, 152, 100–106. [Google Scholar] [CrossRef]
- Chen, Y.C.; Ma, Y.L.; Lin, C.H.; Cheng, S.J.; Hsu, W.L.; Lee, E.H. Galectin-3 negatively regulates hippo-campus-dependent memory formation through inhibition of integrin signaling and galectin-3 phosphorylation. Front. Mol. Neurosci. 2017, 10, 217. [Google Scholar] [CrossRef]
- Pasquini, L.A.; Millet, V.; Hoyos, H.C.; Giannoni, J.P.; Croci, D.O.; Marder, M.; Liu, F.T.; Rabinovich, G.A.; Pasquini, J.M. Galectin-3 drives oligodendrocyte differentiation to control myelin integrity and function. Cell Death Differ. 2011, 18, 1746–1756. [Google Scholar] [CrossRef] [PubMed]
- Sartim, A.G.; Sartim, M.A.; Cummings, R.D.; Dias-Baruffi, M.; Joca, S.R. Impaired emotional response to stress in mice lacking galectin-1 or galectin-3. Physiol. Behav. 2020, 220, 112862. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Pozzo-Miller, L. BDNF deregulation in Rett syndrome. Neuropharmacology 2014, 76, 737–746. [Google Scholar] [CrossRef] [Green Version]
- Ehinger, Y.; Bruyere, J.; Panayotis, N.; Abada, Y.S.; Borloz, E.; Matagne, V.; Scaramuzzino, C.; Vitet, H.; De-latour, B.; Saidi, L.; et al. Huntingtin phosphorylation governs BDNF homeostasis and improves the phenotype of Mecp2 knockout mice. EMBO Mol. Med. 2020, 12, e10889. [Google Scholar] [CrossRef] [PubMed]
- Xing, C.; Wang, X.; Cheng, C.; Montaner, J.; Mandeville, E.; Leung, W.; van Leyen, K.; Lok, J.; Wang, X.; Lo, E.H. Neuronal production of lipocalin-2 as a help-me signal for glial activation. Stroke 2014, 45, 2085–2092. [Google Scholar] [CrossRef]
- Stajic, D.; Selakovic, D.; Jovicic, N.; Joksimovic, J.; Arsenijevic, N.; Lukic, M.L.; Rosic, G. The role of galectin-3 in modulation of anxiety state level in mice. Brain Behav. Immun. 2019, 78, 177–187. [Google Scholar] [CrossRef]
- Lehmann, S.G.; Seve, M.; Vanwonterghem, L.; Michelland, S.; Cunin, V.; Coll, J.L.; Hurbin, A.; Bourgo-in-Voillard, S. A large scale proteome analysis of the gefitinib primary resistance overcome by KDAC inhibition in KRAS mutated adenocarcinoma cells overexpressing amphiregulin. J. Proteomics 2019, 195, 114–124. [Google Scholar] [CrossRef]
- Lehmann, S.G.; Bourgoin-Voillard, S.; Seve, M.; Rachidi, W. Tubulin beta-3 chain as a new candidate protein biomarker of human skin aging: A preliminary study. Oxid. Med. Cell Longev. 2017, 2017, 5140360. [Google Scholar] [CrossRef]
- Breitwieser, F.P.; Muller, A.; Dayon, L.; Kocher, T.; Hainard, A.; Pichler, P.; Schmidt-Erfurth, U.; Superti-Furga, G.; Sanchez, J.C.; Mechtler, K.; et al. General statistical modeling of data from protein rel-ative expression isobaric tags. J. Proteome Res. 2011, 10, 2758–2766. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehen-sive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DA-VID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Doncheva, N.T.; Morris, J.H.; Gorodkin, J.; Jensen, L.J. Cytoscape StringApp: Network analysis and visualiza-tion of proteomics data. J Proteome Res. 2019, 18, 623–632. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferreira, T.A.; Blackman, A.V.; Oyrer, J.; Jayabal, S.; Chung, A.J.; Watt, A.J.; Sjostrom, P.J.; van Meyel, D.J. Neuronal morphometry directly from bitmap images. Nat. Methods 2014, 11, 982–984. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Accession | Name | Gene Name | Peptides (>95%) | Mean of iTRAQ Ratio KO/WT | p Value Rat. KO/WT |
|---|---|---|---|---|---|
| P16045 | Galectin-1 | Lgals1 | 9 | 0.14 | 1.73 × 10−2 |
| P11672 | Neutrophil Gelatinase-Associated Lipocalin | Lcn2 | 93 | 0.25 | 1.12 × 10−5 |
| P10148 | C-C Motif Chemokine 2 | Ccl2 | 3 | 0.35 | 8.05 × 10−4 |
| P16110 | Galectin-3 | Lgals3 | 6 | 0.35 | 1.10 × 10−2 |
| P24452 | Macrophage-Capping Protein | Capg | 6 | 0.41 | 2.06 × 10−4 |
| Q61362 | Chitinase-3-Like Protein 1 | Chi3l1 | 100 | 0.46 | 1.68 × 10−6 |
| P21460 | Cystatin-C | Cst3 | 69 | 0.48 | 4.76 × 10−2 |
| P01887 | Beta-2-Microglobulin | B2m | 24 | 0.52 | 4.01 × 10−4 |
| Q9CXI5 | Mesencephalic Astrocyte-Derived Neurotrophic Factor | Manf | 2 | 0.54 | 7.21 × 10−4 |
| Q8VCT3 | Aminopeptidase B | Rnpep | 7 | 0.59 | 5.69 × 10−3 |
| Q91WP6 | Serine Protease Inhibitor A3N | Serpina3n | 25 | 0.60 | 1.71 × 10−2 |
| Q06890 | Clusterin | Clu | 92 | 1.33 | 8.28 × 10−3 |
| P01027 | Complement C3 | C3 | 766 | 1.46 | 1.9 × 10−2 |
| Q91ZJ9 | Hyaluronidase-1 | Hyal1 | 3 | 1.56 | 1.46 × 10−3 |
| Q9CQ01 | Ribonuclease T2 | Rnaset2 | 15 | 1.62 | 1.03 × 10−3 |
| P01029 | Complement C4-B | C4b C4 | 141 | 1.85 | 1.86 × 10−3 |
| O88998 | Noelin | Olfm1 | 6 | 1.89 | 1.57 × 10−3 |
| Q61220 | Protein Kinase C-Binding Protein NELL2 | Nell2 | 3 | 1.90 | 1.58 × 10−3 |
| O35684 | Neuroserpin | Serpini1 | 4 | 2.03 | 1.38 × 10−4 |
| P97298 | Pigment Epithelium-Derived Factor | Serpinf1 | 24 | 2.04 | 4.77 × 10−3 |
| B9EKR1 | Receptor-Type Tyrosine-Protein Phosphatase Zeta | Ptprz1 | 13 | 2.06 | 6.80 × 10−3 |
| O54782 | Epididymis-Specific Alpha-Mannosidase | Man2b2 | 46 | 2.11 | 2.72 × 10−5 |
| Q07797 | Galectin-3-Binding Protein | Lgals3bp | 26 | 2.11 | 8.30 × 10−4 |
| O09114 | Prostaglandin-H2 D-Isomerase | Ptgds | 22 | 2.17 | 2.02 × 10−3 |
| P50608 | Fibromodulin | Fmod | 19 | 2.37 | 1.86 × 10−4 |
| Q99LS0 | Augurin | Ecrg4 | 8 | 2.53 | 2.49 × 10−6 |
| Q9D2L1 | Arylsulfatase K | Arsk | 5 | 2.68 | 1.78 × 10−2 |
| Q9D1H9 | Microfibril-Associated Glycoprotein 4 | Mfap4 | 2 | 2.75 | 4.80 × 10−4 |
| Q9EQG7 | Ectonucleotide Pyrophosphatase/Phosphodiesterase Family Member 5 | Enpp5 | 13 | 2.88 | 3.05 × 10−7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ehinger, Y.; Matagne, V.; Cunin, V.; Borloz, E.; Seve, M.; Bourgoin-Voillard, S.; Borges-Correia, A.; Villard, L.; Roux, J.-C. Analysis of Astroglial Secretomic Profile in the Mecp2-Deficient Male Mouse Model of Rett Syndrome. Int. J. Mol. Sci. 2021, 22, 4316. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22094316
Ehinger Y, Matagne V, Cunin V, Borloz E, Seve M, Bourgoin-Voillard S, Borges-Correia A, Villard L, Roux J-C. Analysis of Astroglial Secretomic Profile in the Mecp2-Deficient Male Mouse Model of Rett Syndrome. International Journal of Molecular Sciences. 2021; 22(9):4316. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22094316
Chicago/Turabian StyleEhinger, Yann, Valerie Matagne, Valérie Cunin, Emilie Borloz, Michel Seve, Sandrine Bourgoin-Voillard, Ana Borges-Correia, Laurent Villard, and Jean-Christophe Roux. 2021. "Analysis of Astroglial Secretomic Profile in the Mecp2-Deficient Male Mouse Model of Rett Syndrome" International Journal of Molecular Sciences 22, no. 9: 4316. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22094316






