Isolation and Characterization of a Novel Four-Transmembrane Protein PMP22CD Specifically Expressed in the Testis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cloning of the gene PMP22CD
2.2. In silico analysis
2.3. Expression pattern analysis of PMP22CD
2.4. Analysis of subcellular localization
2.5.Cell transfection and western blotting
3. Results
3.1. Human PMP22CD gene and its genomic organization
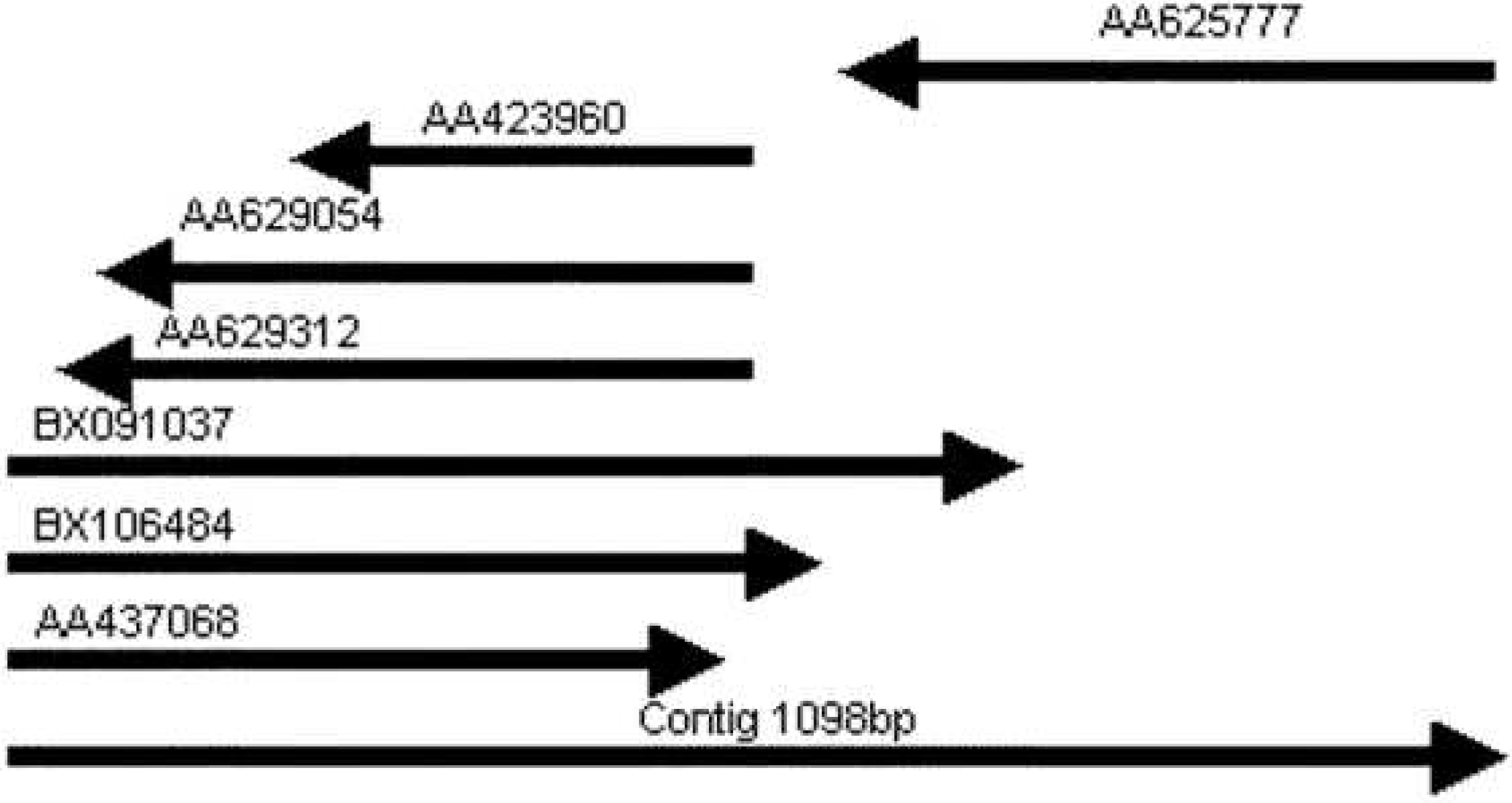
 , Polyadenylation signals;
, Polyadenylation signals;  , in-frame stop codon.
, in-frame stop codon.
 , Polyadenylation signals;
, Polyadenylation signals;  , in-frame stop codon.
, in-frame stop codon.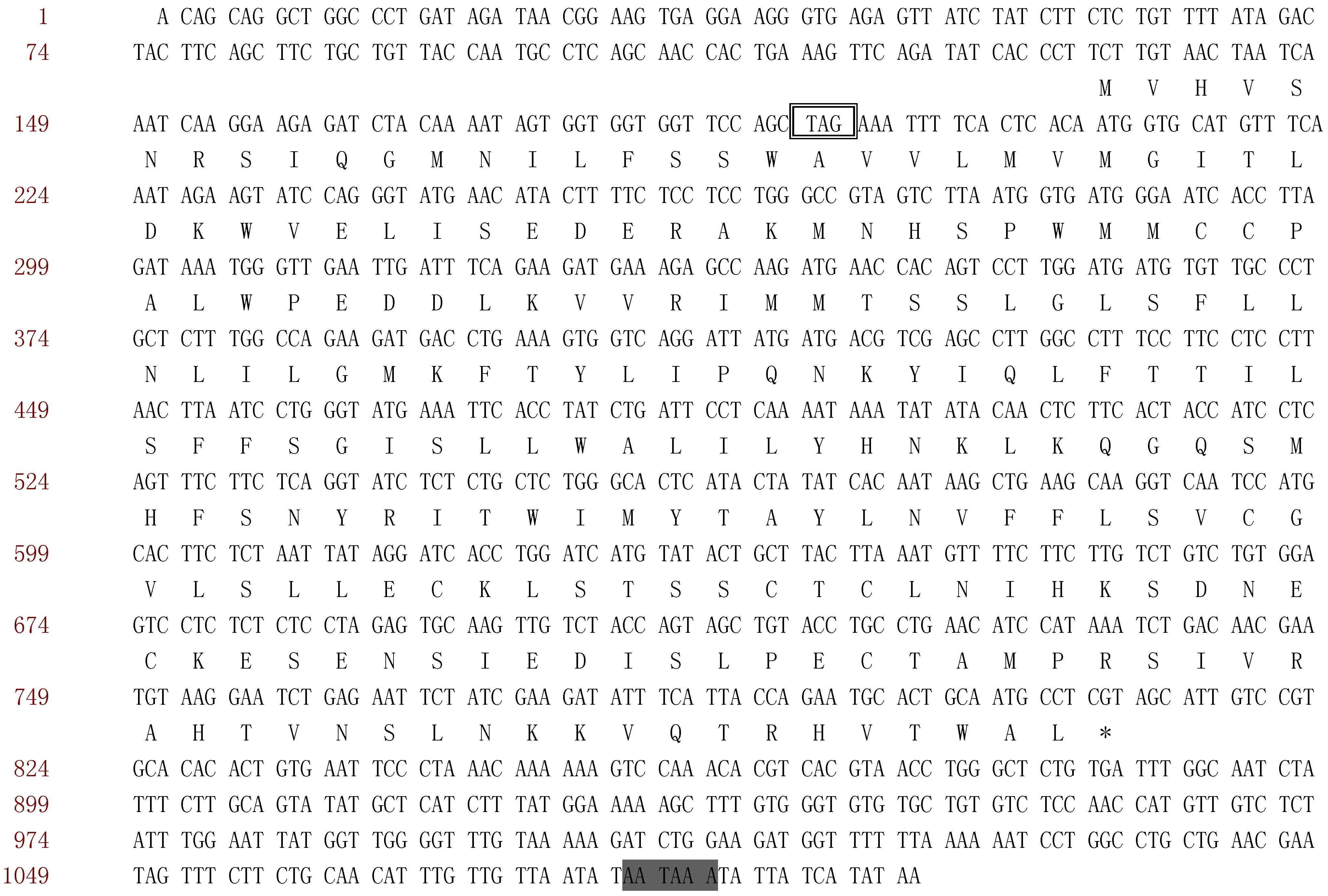
| 3’Splice acceptor | Exon | Size(bp) | 5’ Splice donor | Intron | Size(bp) |
|---|---|---|---|---|---|
| cDNA end ACAGCAGGCT | 1 | 389 | TGGCCAGAAGgtcagaactg | 1 | 608 |
| tttccaacagATGACCTGAA | 2 | 147 | TTCTTCTCAGgtaacttcct | 2 | 280 |
| tggacatcagGTATCTCTCT | 3 | 135 | TCTGTCTGTGgtgagtgtct | 3 | 722 |
3.2. In silico analysis of PMP22CD gene

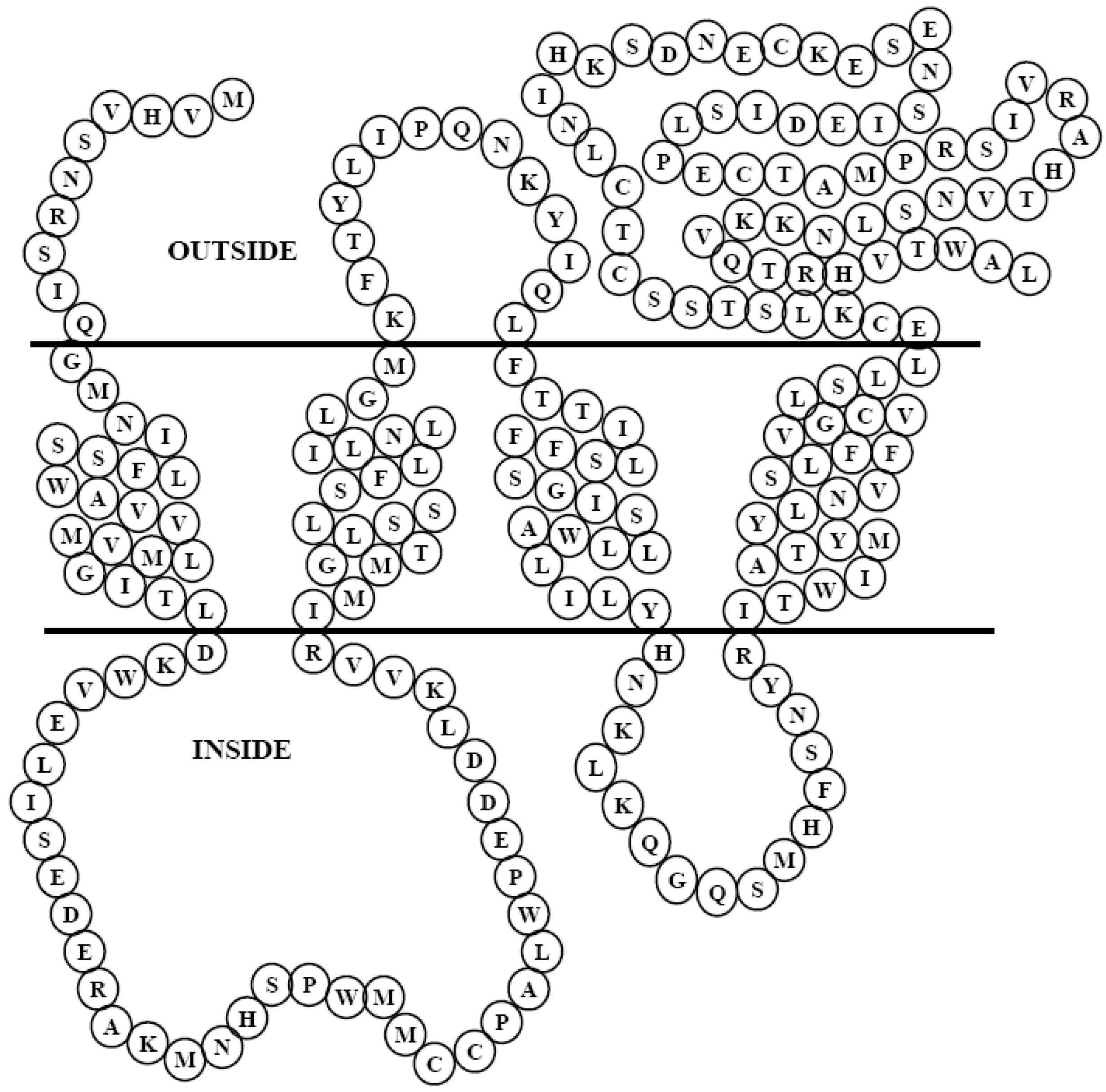


3.3. Expression pattern of PMP22CD gene
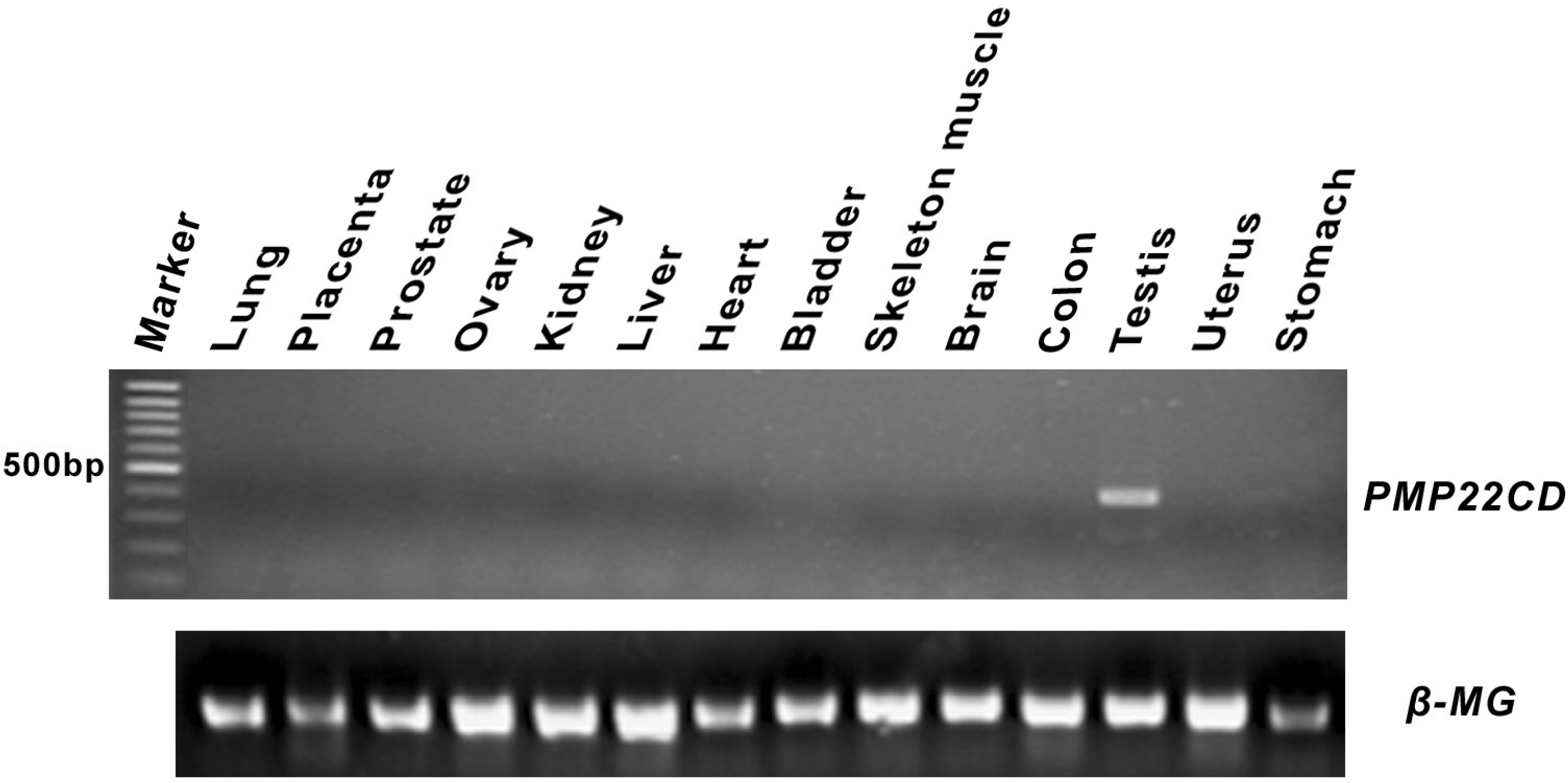

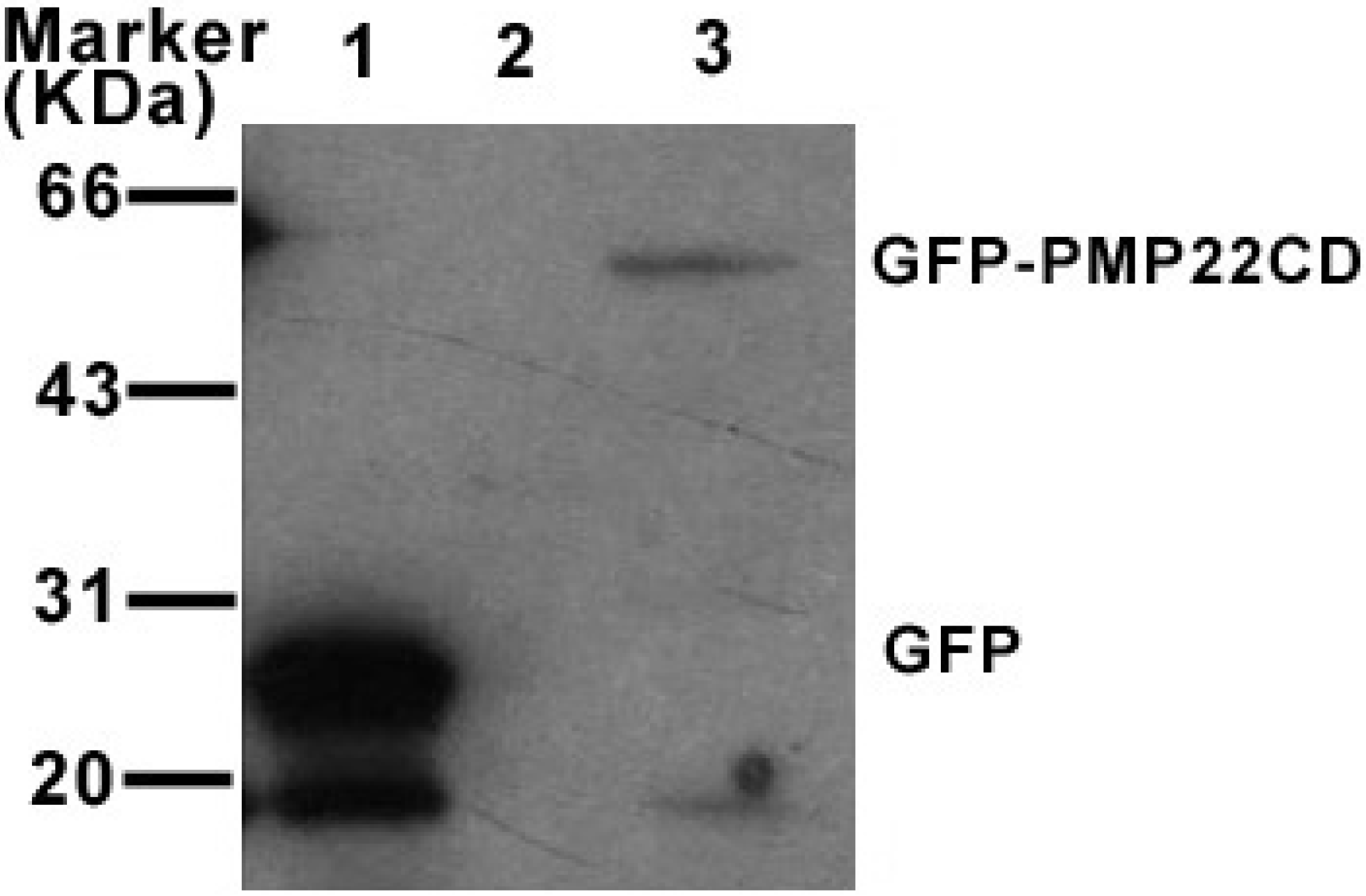
3.4. PMP22CD protein expression and its subcellular localization
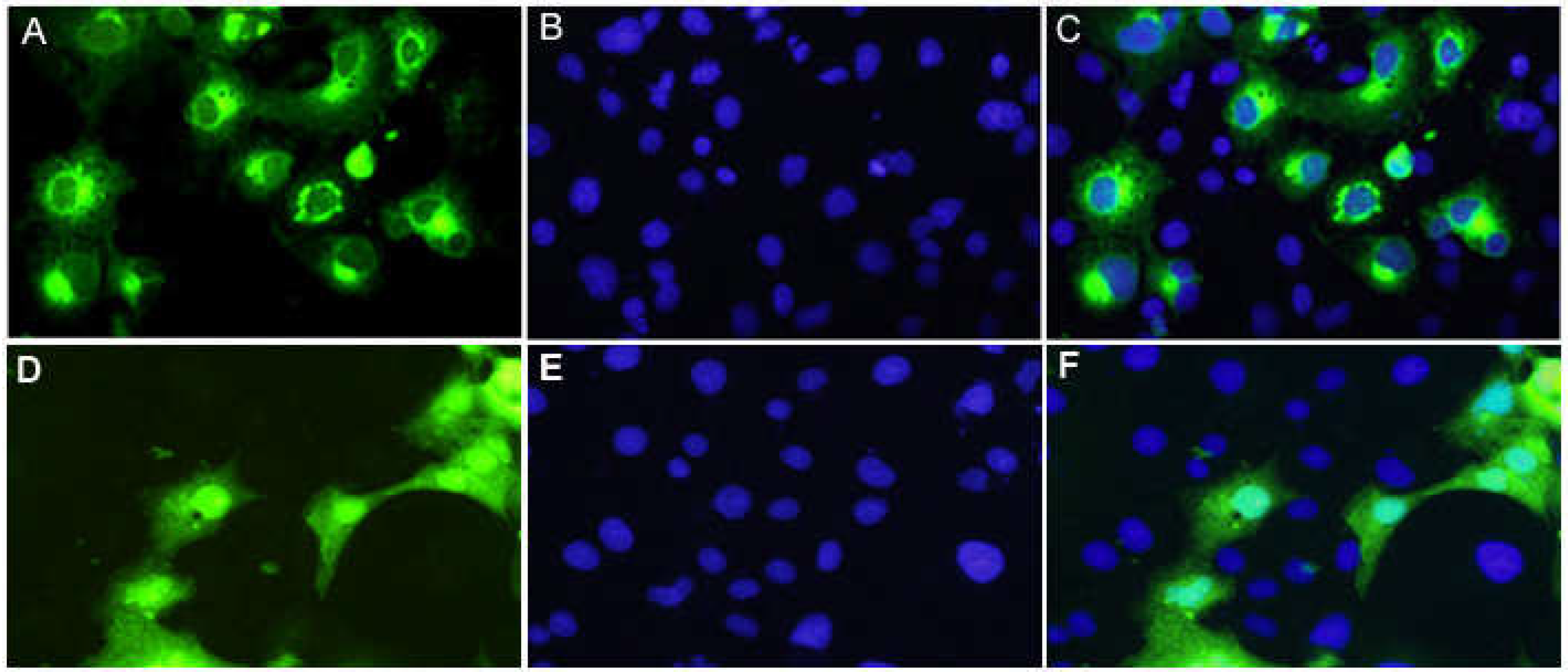
- A:
- Cells transfected with pEGFP-PMP22CD
- B:
- The nucleus of cells transfected with pEGFP-PMP22CD were stained with DAPI
- C:
- Colocalization of A and B
- D:
- Cells transfected with pEGFP-C1
- E:
- The nucleus of cells transfected with pEGFP-C1 were stained with DAPI
- F:
- Colocalization of D and E
4. Discussion
Acknowledgement
References and Notes
- Nagafuchi, A. Molecular architecture of adherens junctions. Curr Opin Cell Biol. 2001, 13, 600–603. [Google Scholar]
- Furuse, M.; Hirase, T.; Itoh, M.; Nagafuchi, A.; Yonemura, S.; Tsukita, S. Occludin: a novel integral membrane protein localizing at tight junctions. J Cell Biol. 1993, 123, 1777–1788. [Google Scholar]
- Muller, H. W. Tetraspan myelin protein PMP22 and demyelinating peripheral neuropathies: new facts and hypotheses. Glia. 2000, 29, 182–185. [Google Scholar]
- Furuse, M.; Fujita, K.; Hiiragi, T.; Fujimoto, K.; Tsukita, S. Claudin-1 and -2: novel integral membrane proteins localizing at tight junctions with no sequence similarity to occludin. J Cell Biol. 1998, 141, 1539–1550. [Google Scholar]
- Tsukita, S.; Furuse, M. Overcoming barriers in the study of tight junction functions: from occludin to claudin. Genes Cells. 1998, 3, 569–573. [Google Scholar] [CrossRef]
- Morita, K.; Furuse, M.; Fujimoto, K.; Tsukita, S. Claudin multigene family encoding four-transmembrane domain protein components of tight junction strands. Proc Natl Acad Sci U S A. 1999, 96, 511–516. [Google Scholar]
- Tsukita, S.; Furuse, M.; Itoh, M. Multifunctional strands in tight junctions. Nat Rev Mol Cell Biol. 2001, 2, 285–293. [Google Scholar] [CrossRef]
- Christophe-Hobertus, C.; Szpirer, C.; Guyon, R.; Christophe, D. Identification of the gene encoding Brain Cell Membrane Protein 1 (BCMP1), a putative four-transmembrane protein distantly related to the Peripheral Myelin Protein 22 / Epithelial Membrane Proteins and the Claudins. BMC Genomics. 2001, 2, 3. [Google Scholar] [CrossRef]
- Kearsey, J.; Petit, S.; De Oliveira, C.; Schweighoffer, F. A novel four transmembrane spanning protein, CLP24. A hypoxically regulated cell junction protein. Eur J Biochem. 2004, 271, 2584–2592. [Google Scholar] [CrossRef]
- Gonzalez-Mariscal, L.; Betanzos, A.; Nava, P.; Jaramillo, B. E. Tight junction proteins. Prog Biophys Mol Biol. 2003, 81, 1–44. [Google Scholar]
- Huang, C.; Guo, J.; Liu, S.; Shan, Y.; Wu, S.; Cai, Y.; Yu, L. Isolation, tissue distribution and prokaryotic expression of a novel human X-linked gene LHFPL1. DNA Seq. 2004, 15, 299–302. [Google Scholar]
- Huang, C.; Zhou, J.; Wu, S.; Shan, Y.; Teng, S.; Yu, L. Cloning and tissue distribution of the human B3GALT7 gene, a member of the beta1,3-Glycosyltransferase family. Glycoconj J. 2004, 21, 267–273. [Google Scholar] [CrossRef]
- Altschul, S. F.; Madden, T. L.; Schaffer, A. A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D. J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Tusnady, G. E.; Simon, I. The HMMTOP transmembrane topology prediction server. Bioinformatics. 2001, 17, 849–850. [Google Scholar] [CrossRef]
- Wang, M.; Wang, J.; Zhang, Z.; Zhao, Z.; Zhang, R.; Hu, X.; Tan, T.; Luo, S.; Luo, Z. Dissecting phenotypic variation among AIS patients. Biochem Biophys Res Commun. 2005, 335, 335–342. [Google Scholar] [CrossRef]
- Inai, T.; Sengoku, A.; Guan, X.; Hirose, E.; Iida, H.; Shibata, Y. Heterogeneity in expression and subcellular localization of tight junction proteins, claudin-10 and -15, examined by RT-PCR and immunofluorescence microscopy. Arch Histol Cytol. 2005, 68, 349–360. [Google Scholar] [CrossRef]
- Lee, N. P.; Tong, M. K.; Leung, P. P.; Chan, V. W.; Leung, S.; Tam, P. C.; Chan, K. W.; Lee, K. F.; Yeung, W. S.; Luk, J. M. Kidney claudin-19: localization in distal tubules and collecting ducts and dysregulation in polycystic renal disease. FEBS Lett. 2006, 580, 923–931. [Google Scholar]
- Li, W. Y.; Huey, C. L.; Yu, A. S. Expression of claudin-7 and -8 along the mouse nephron. Am J Physiol Renal Physiol. 2004, 286, F1063–1071. [Google Scholar] [CrossRef] [Green Version]
- Gye, M. C. Expression of claudin-1 in mouse testis. Arch Androl. 2003, 49, 271–279. [Google Scholar]
- Sengupta, P.; Chou, J. H.; Bargmann, C. I. odr-10 encodes a seven transmembrane domain olfactory receptor required for responses to the odorant diacetyl. Cell. 1996, 84, 899–909. [Google Scholar] [CrossRef] [Green Version]
- Note: The sequence data reported here have been submitted to the GenBank database under accession number AY634366.
© 2006 by MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, M.; Luo, G.; Li, F.; Wu, S.; Jiang, J.; Huang, C. Isolation and Characterization of a Novel Four-Transmembrane Protein PMP22CD Specifically Expressed in the Testis. Int. J. Mol. Sci. 2006, 7, 425-437. https://0-doi-org.brum.beds.ac.uk/10.3390/i7100425
Wang M, Luo G, Li F, Wu S, Jiang J, Huang C. Isolation and Characterization of a Novel Four-Transmembrane Protein PMP22CD Specifically Expressed in the Testis. International Journal of Molecular Sciences. 2006; 7(10):425-437. https://0-doi-org.brum.beds.ac.uk/10.3390/i7100425
Chicago/Turabian StyleWang, Minghua, Guanmei Luo, Fang Li, Shiliang Wu, Juxiang Jiang, and Chaoqun Huang. 2006. "Isolation and Characterization of a Novel Four-Transmembrane Protein PMP22CD Specifically Expressed in the Testis" International Journal of Molecular Sciences 7, no. 10: 425-437. https://0-doi-org.brum.beds.ac.uk/10.3390/i7100425




