1. Introduction
The adsorption and penetration of myophage T4 is a fascinating example of a self-triggered multistep event in molecular biology. The process is divided into five steps: (1) the adsorption of long tail fibres; (2) the binding of short tail fibres; (3) tail-sheath contraction; (4) tail-tube movement through the outer membrane and periplasm; and (5) the contact of the tail-tube tip with the inner membrane to initiate the phage DNA ejection process.
Bacteriophage T4 contacts its host cells by initially stochastic low-affinity binding six long tail fibres to the outer membrane [
1,
2,
3]. This binding triggers a conformational change in the geometry of the baseplate that results in stretching out six short tail fibres (gp12), leading to an irreversible and firm binding of the phage to the bacterial lipopolysaccharide moiety of the cell [
4,
5]. The conformational transition of the baseplate to a starlike configuration then starts the contraction of the tail structure [
6]. Thereby, the gp18 proteins of the tail sheath rearrange their intersubunit contacts, leading to wavelike shortening of the sheath [
7,
8]. Consequently, the tail sheath is shortened from 92.5 to 42 nm, whereas the inner tail tube stays in its stiff cylinder-like structure that is mechanically forced to move through the centre of the baseplate.
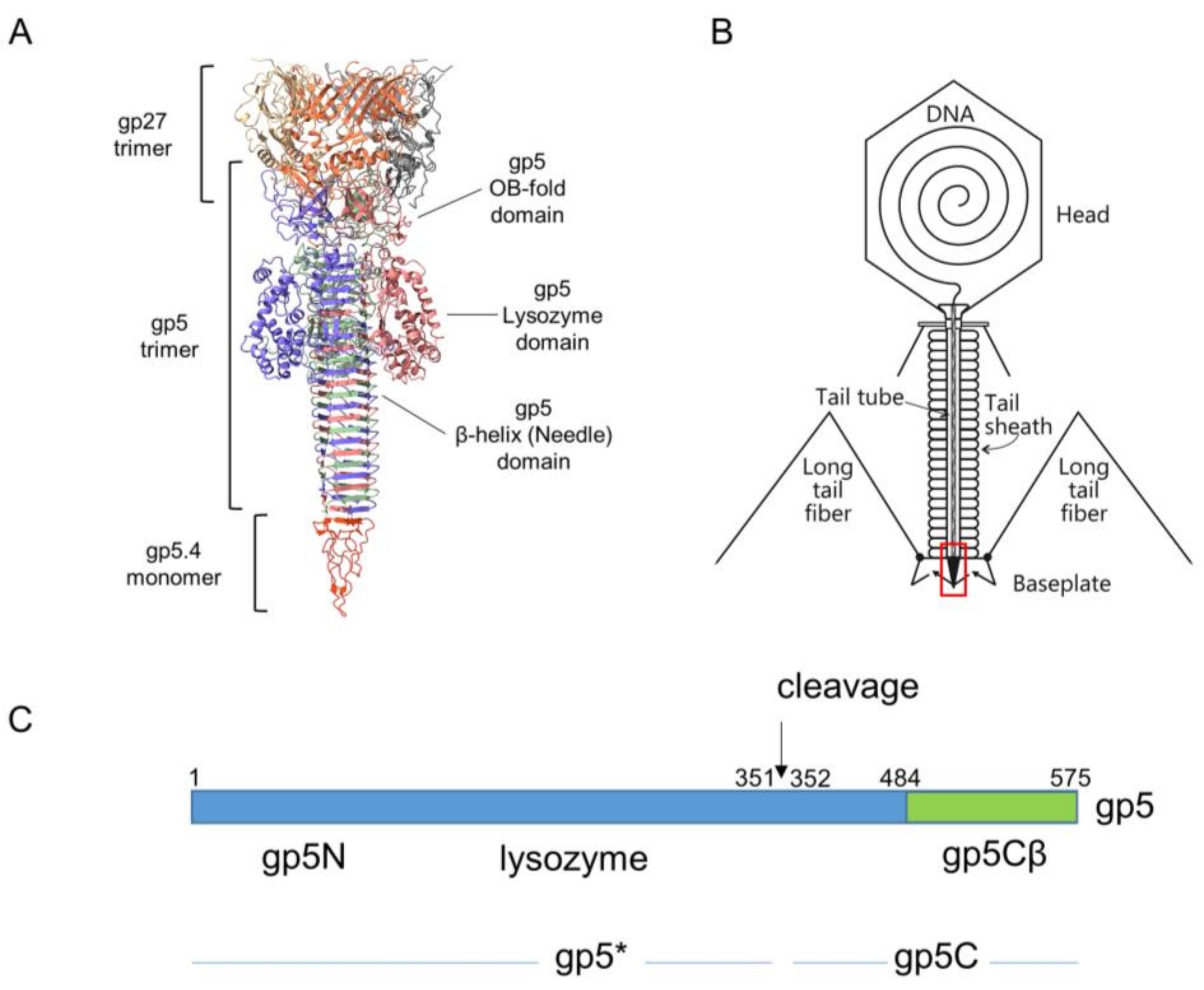
The movement of the tail tube towards the cell surface takes baseplate components gp5.4, gp5, gp27, gp48, and gp54 along at the leading end of the tube (gp19); gp5.4, at the tip of this structure, is tightly connected to the C-terminal part of gp5, which forms a stable triple helix, and is called the gp5C-gp5.4 central spike complex (
Figure 1A) [
9,
10].
Gp5 exists in the phage as a cleaved protein, with the N-terminal part (gp5*) bound to gp27 and the C-terminal part tightly bound to gp5.4. The penetration of the outer membrane is most likely executed by mechanical force generated by the sheath contraction and applied to the central spike structure. In the periplasm, the tip of the tail moves through the peptidoglycan layer, which is facilitated by the mureolytic activity of the lysozyme domain of gp5.
The molecular details that lead to contact with the inner membrane and to the translocation of the phage DNA through the membrane into the cytoplasm are still unknown. Here, a number of scenarios are discussed. The needle at the tip of the tail tube may come into contact with the inner membrane and initiate an opening of a DNA-conductive channel. Second, the central spike complex dissociates from the tail tube in the periplasm, leaving the N-terminal part of gp5 exposed on the tube tip, which could then come into contact with the inner membrane. Furthermore, it is possible that, in the periplasm, the entire T4 tip structure dissociates from the tube, and gp27 or gp48 comes into contact with the inner membrane for DNA translocation.
To shed some light on these interesting events, we purified the gp5Cβ–gp5.4 complex, a tip fragment of the central spike complex, and tested its binding to liposomes and spheroplasts. We found that this spike fragment binds to periplasmic protein PpiD that is anchored to the inner membrane. PpiD is a periplasmic chaperone [
12] that is connected to the SecYEG translocon [
13]. A chromosomal deletion of PpiD resulted in a decrease of T4 plating efficiency on this strain, suggesting that PpiD might play an important role in T4 infection.
2. Materials and Methods
2.1. Bacterial Strains and Phage
T4 was propagated on
E. coli B and on BW25113 (parental strain of
ΔppiD; Keio collection) in Luria broth (LB medium). Media preparation and bacterial growth were performed according to standard methods [
14].
ppiD deletion strain JW0431 was cultured in LB medium containing 25 µg/mL kanamycin at 37 °C.
E. coli BL21 (DE3) [
15], transformed with plasmid pEEva2 encoding the gp5Cβ–5.4 complex or plasmid pASK-IBA3C encoding PpiD, respectively, was grown in LB in the presence of ampicillin (100 µg/mL, final concentration).
2.2. Purification of gp5Cβ–gp5.4 Complex
E. coli BL21 (DE) harbouring plasmid pEEva2 encoding the His–gp5Cβ–gp5.4 complex was grown in a 2xYT medium containing 100 µg/mL ampicillin at 37 °C until the culture had reached OD600 of 0.6. The culture was cooled down to 18 °C. Gp5Cβ–5.4 complex expression was induced by the addition of isopropyl-β-D-thiogalactopyranoside (IPTG) to 0.5 mM, and the culture was grown at 18 °C overnight. Cells were harvested by centrifugation at 8000× g at 4 °C for 15 min. Cell pellets were resuspended in a lysis buffer (20 mM Tris-HCl (pH 8), 300 mM NaCl, 5 mM imidazole) on ice and lysed using the OneShot (Constant Systems) at 1.23 kbar. Before cell disruption, 1 mM phenylmethylsulfanyl-fluoride (PMSF) was added. The lysate was then centrifuged at 35,000× g and 4 °C for 20 min. The supernatant was loaded into a 2 mL Ni-Sepharose 6 Fast Flow matrix. After consecutively washing the column with 10 mL of each of the two wash buffers (20 mM Tris-HCl (pH 8), 300 mM NaCl, 37.5 mM imidazole (Wash Buffer 1) or 50 mM imidazole (Wash Buffer 2)), the complex was eluted with 20 mL elution buffer (20 mM Tris-HCl (pH 8), 150 mM NaCl, 300 mM imidazole) in 2 mL fractions. The elution fractions containing the target protein were further purified by ion-exchange chromatography performed with a Resource Q 1 mL column connected to an ÄKTA purifier 10 system. The sample was loaded onto the column that was equilibrated with Buffer A (20 mM Tris-HCl (pH 8)) and eluted with a linear gradient against Buffer B (20 mM Tris-HCl (pH 8), 1 M NaCl). Relevant fractions were combined and concentrated using Sartorius Vivaspin centrifugal concentrators with a molecular weight cut-off of 10 kDa. This sample was loaded onto a Superdex 75 10/300 GL column connected to an ÄKTA purifier 10 system, and purified in a gel filtration buffer (10 mM Tris-HCl (pH 8), 150 mM NaCl).
2.3. Size-Exclusion Chromatography–Multiple-Angle Light Scattering (SEC-MALS)
Protein samples were injected into a Superdex 200 increase 10/300 GL column, equilibrated with a gel filtration buffer (10 mM Tris-HCl (pH 8), 150 mM NaCl). The SEC column was linked to a static 3-angle light-scattering detector (miniDAWN Treos II) and a refractive-index detector (Optilab T-rEX) (Wyatt Technology, Santa Barbara, CA, USA). Data were collected every second at a flow rate of 0.5 mL/min. Data analysis was carried out using ASTRA VII software, yielding the molar mass for each fraction. In addition, protein-conjugate analysis was performed to calculate the molar mass of each component of a protein complex. The light-scattering detectors were normalised, and data quality was assessed by a BSA standard (Thermo, Waltham, MA, USA).
2.4. PpiD Purification
E. coli PpiD was expressed from plasmid pASK-IBA3C encoding PipD-Strep (kindly provided by HG Koch, Freiburg) in BL21 (DE3) cells in LB medium containing 25 µg/mL chloramphenicol at 37 °C until the culture had reached an OD600 of 0.6. The culture was induced by the addition of anhydrotetracycline to a final concentration of 200 µg/L for 2 h at 37 °C. Cells were harvested by centrifugation at 10,000× g and 4 °C for 15 min. Cell pellets were resuspended in Buffer A (50 mM Tris-HCl (pH 7.4), 50 mM NaCl, 1 mM Ethylen-diamino-tetra-acidic acid (EDTA)) on ice and lysed using the OneShot instrument at 1.23 kbar. Before cell disruption, 1 mM PMSF was added. The lysate was then centrifuged at 35,000× g and 4 °C for 45 min. The pellet was resuspended in Buffer A containing 1% N-lauroylsarcosine to solubilise the inner membrane at 4 °C overnight. The solubilised membrane fraction was collected by ultracentrifugation at 110,000× g at 4 °C for 40 min and loaded onto 1 mL of Strep-Tactin XT matrix. After washing the matrix with 5 mL of Buffer W (100 mM Tris-HCl (pH 8), 150 mM NaCl, 1 mM EDTA), the protein was eluted with 3 mL of Buffer BXT (100 mM Tris-HCl (pH 8), 150 mM NaCl, 1 mM EDTA, 50 mM biotin) in 500 µL fractions. The elution fractions containing the target protein were further purified by gel filtration chromatography performed with a Superdex 200 increase 10/300 GL column connected to an ÄKTA purifier 10 system in Buffer A.
2.5. Spheroplast Preparation
For cross-linking and binding assays, spheroplasts were generated after centrifugation of
E. coli BL21 (DE3) harbouring plasmid pMAL-p5X (NEB) at 5000× g and 4 °C for 15 min. The cell pellet was resuspended in an ice-cold spheroplast buffer (20 mM Tris-HCl (pH 8), 0.5 M sucrose) and lysozyme (20 µg/mL), and incubated for 5 min at room temperature (RT). Then, 1.6 mM EDTA was added, and the sample was incubated for 7 min at RT [
16]. Finally, 10 mM MgCl
2 was added to the spheroplasts for stabilization.
2.6. Liposomes and Proteoliposomes
E. coli lipids (Avanti) 1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-ethanolamine (POPE) and 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoglycerol (POPG) were dissolved in dichloromethane and evaporated under a stream of nitrogen. The dry lipid film was dissolved in Buffer A (50 mM Tris-HCl (pH 7.4), 50 mM NaCl, 1 mM EDTA). A 7:3 mixture of PE and PG was used, mimicking the membrane-lipid composition of E. coli. Liposomes with an average diameter of 250 nm were generated by extrusion (Mini extruder, Avanti Polar Lipids. Inc.) 31 times through a 0.4 µm polycarbonate membrane (Whatman). For preparing the proteoliposomes, purified PpiD protein was mixed with a lipid/protein ratio of at least 10,000:1 (mol/mol) and extruded as described above.
4. Discussion
The infection process of myophage T4 involves an astounding cascade of conformational events, and this is exemplary in protein biochemistry. Remarkably, the self-triggered multistep process is independent of any outside energy input, similar to the mechanism of an old-fashioned time watch driven to operate for days by a key-loaded spring. The first steps of the phage adsorption and interaction of tail fibres with the receptors of the host cell [
3], the movement of the short fibres, and the contraction of the tail sheath [
4,
5] are quite well-understood, whereas the molecular events that occur when the phage is penetrating the outer and inner host membrane are still unknown.
When the tail tube, driven by sheath contraction, moves through the baseplate, it takes with itself the central hub proteins of the baseplate, gp5, gp5.4, and gp27, together with gp48 and gp54. Therefore, the structure that enters the host periplasm is a capped tail tube structure, as depicted in
Figure 1. In the present study, we followed the fate of the gp5 and gp5.4 proteins at the leading end of the periplasm-entering tail tube. In the phage particle, gp5 is a cleaved protein that is present as two distinct fragments, gp5* and gp5C [
17]. Gp5C is folded into a triple helix that tightly binds 5.4 and withholds its triple-helical structure [
9], even after denaturing treatments on SDS-PAGE giving rise to an apparent 48 kDa protein band (
Figure 7A). The stability of the complex allowed for purifying a fragment of gp5C (484–575), termed gp5Cβ, with gp5.4 from
E. coli cells encoded on a plasmid [
10]. Expressed from the
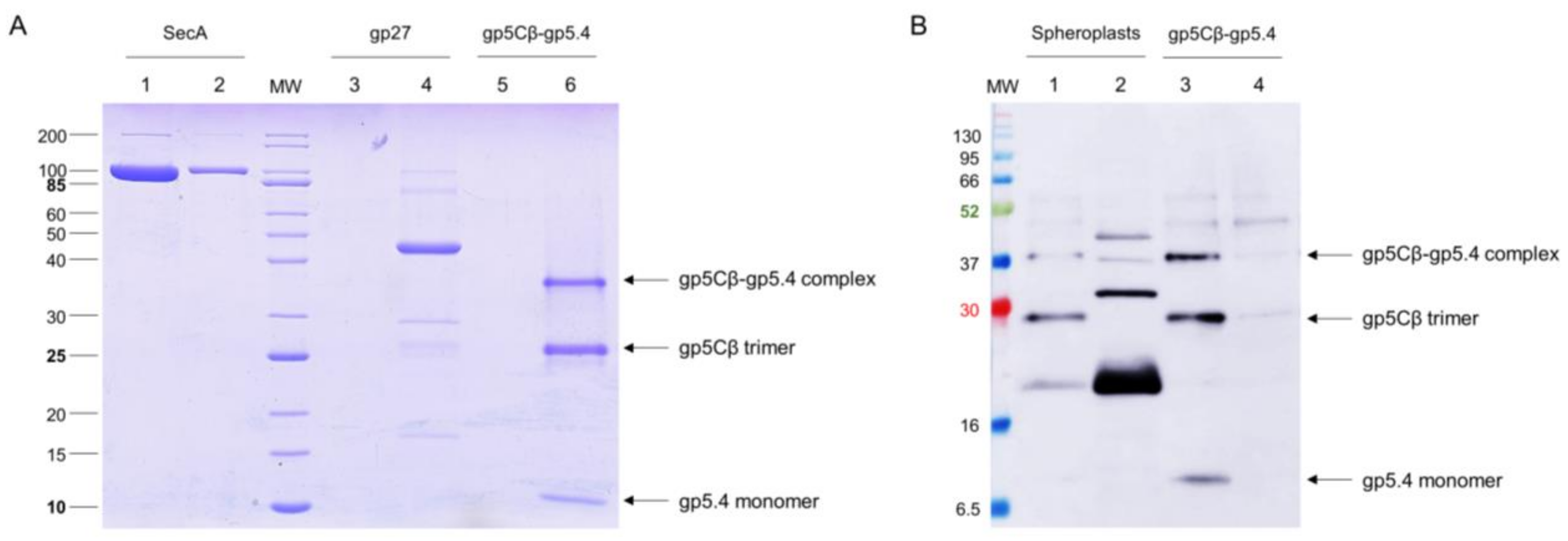
E. coli cells, the two proteins formed a complex, gp5Cβ–gp5.4, which was partially stable on SDS-PAGE (
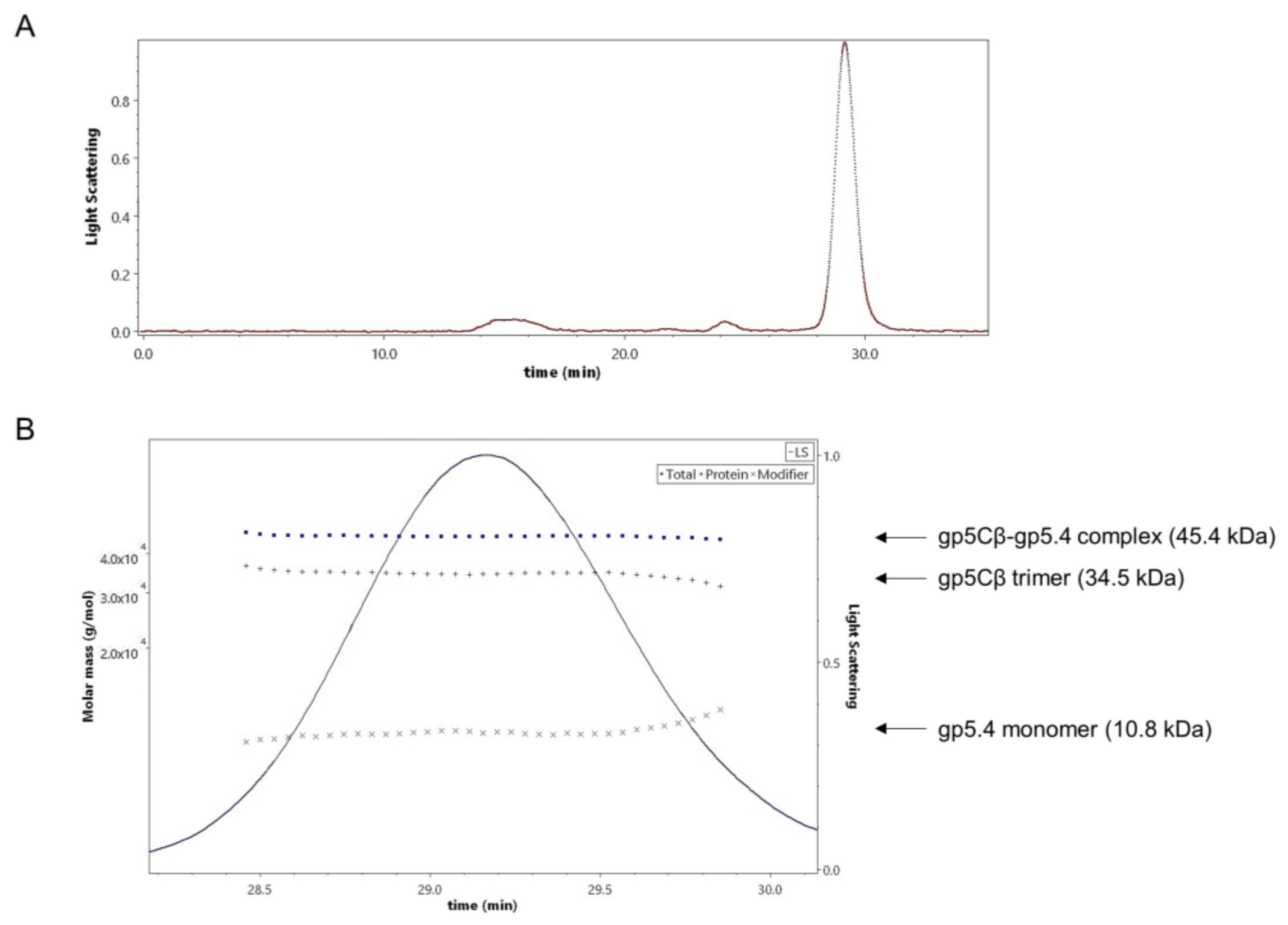
Figure 2B), and encompassed 3 copies of gp5Cβ and one copy of gp5.4, as was verified by MALS analysis (
Figure 3).
The purified complex was then used for binding experiments in vitro to uncover possible interacting partner proteins. First, the addition to spheroplasts and liposomes did not show strong interaction (
Figure 4), suggesting that the central spike complex freely diffuses in the periplasm, as was suggested earlier [
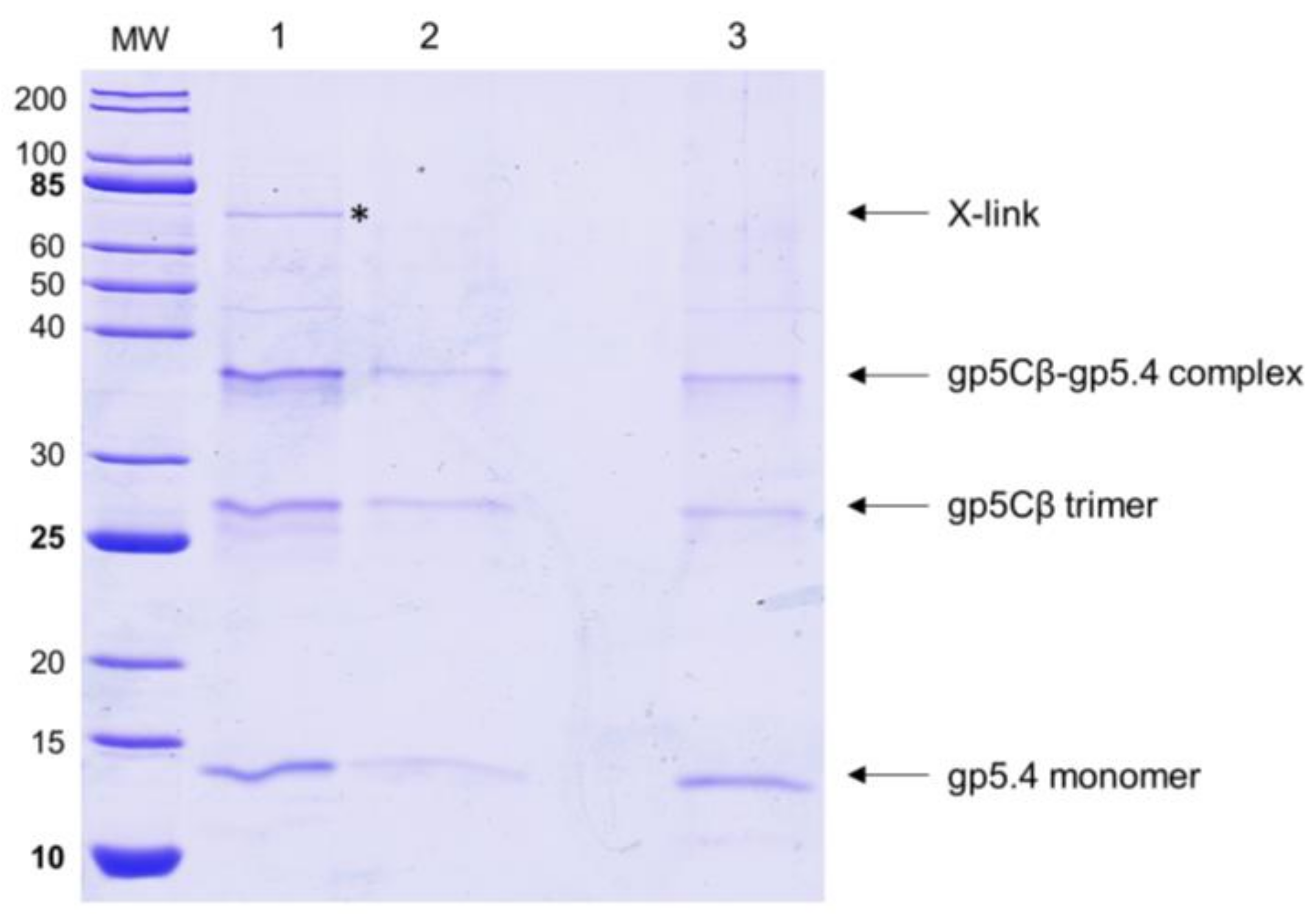
19]. However, when we used formaldehyde as a chemical cross-linker to stabilise possible interactions with spheroplasts, we found that PpiD copurified with the gp5Cβ–5.4 complex (
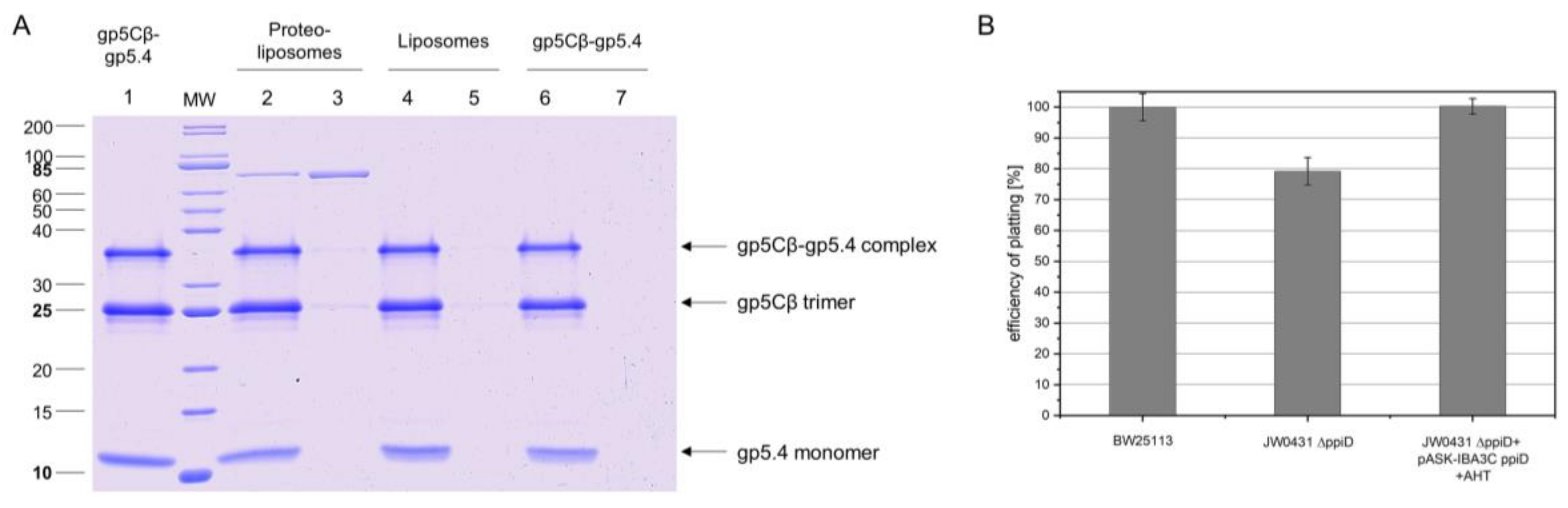
Figure 5). This interaction was verified with proteoliposomes that were reconstituted with purified PpiD (
Figure 6A). PpiD proteoliposomes showed enhanced binding of the added central spike complex when compared with that of pure liposomes. PpiD is a periplasmic chaperone, anchored to the inner membrane that is devoted to binding and stabilising proteins. Regarding its function for T4 infection, it could either catalyse the interaction of the tail tube with the inner membrane for penetration or simply bind the already dissociated gp5Cβ–gp5.4 complex to prevent an inhibitory role for efficient tail tube–inner membrane interaction. Additional experiments are required to clarify these points.
The involvement of PpiD in the T4 infection and propagation process was then studied with efficiency of plating (EOP). This assay quantified the proportion of infecting phage that successfully led to the formation of a plaque. When we compared the parental strain with the deletion mutant of PpiD, a reduction in EOP to less than 80% was determined. This points to a nonessential but supporting role of PpiD in the T4 infection process. Alternatively, other periplasmic chaperones such as SurA or Skp [
20] might step in when
ppiD is deleted and compensate for the loss of PpiD.
When we followed the gp5C-containing complex in phage particles, the major portion was found in a 48 kDa protein band (
Figure 7A,B). Minor bands migrated at 65 and 52 kDa, respectively. The known cleavage site at amino acid residue 351 generates a triple helix of gp5C with a theoretical molecular weight of 59 kDa. Previously, it was shown that gp5C runs with a lower apparent MW [
11], which might be caused by protein folding of the triple helix. After infection, the 48 kDa protein complex remained mainly with the spheroplasts (
Figure 7B). Therefore, the central spike complex might indeed come into contact with the cytoplasmic membrane. However, in our analysis with the purified gp5Cβ–gp5.4 complex, we did not find a strong binding to spheroplasts. It is, therefore, possible that additional factors are required to support the binding of the T4 tail tube to the membrane surface.
Taken together, our results suggest that the T4 central spike complex enters the periplasm and interacts with low affinity to PpiD. In our analysis, we did not observe stable interaction with the inner membrane, which is required for the phage to establish a DNA-translocating complex.