Defenses against Virus and Vector: A Phloem-Biological Perspective on RTM- and SLI1-Mediated Resistance to Potyviruses and Aphids
Abstract
:1. Introduction
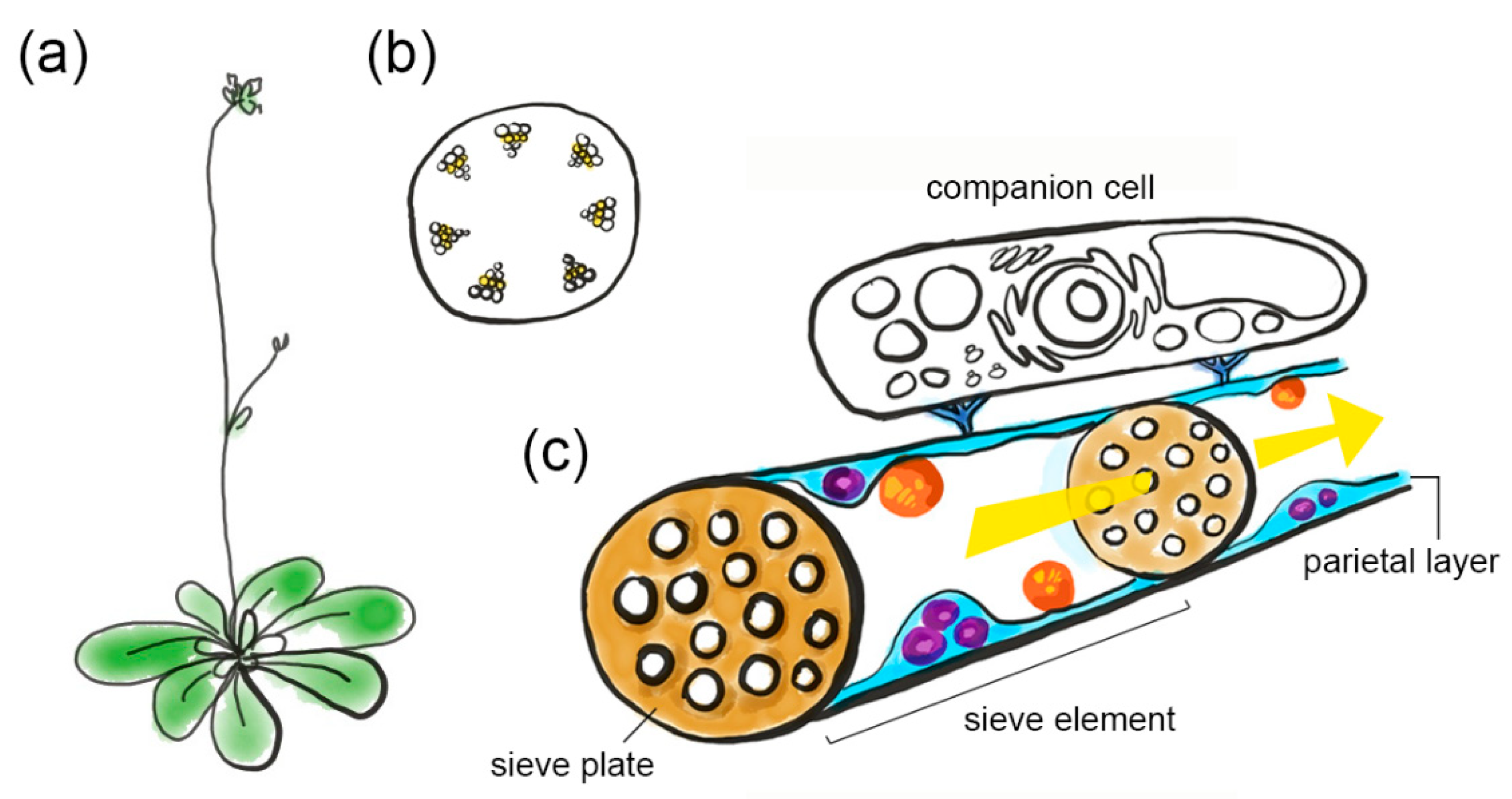
2. Phloem Sap Transport and the Architecture of Sieve Tubes
3. Potyvirus Cell-to-Cell and Long-Distance Movement
4. RTM-Mediated Resistance to Potyviruses
5. SLI1-Mediated Resistance to Myzus persicae Aphids
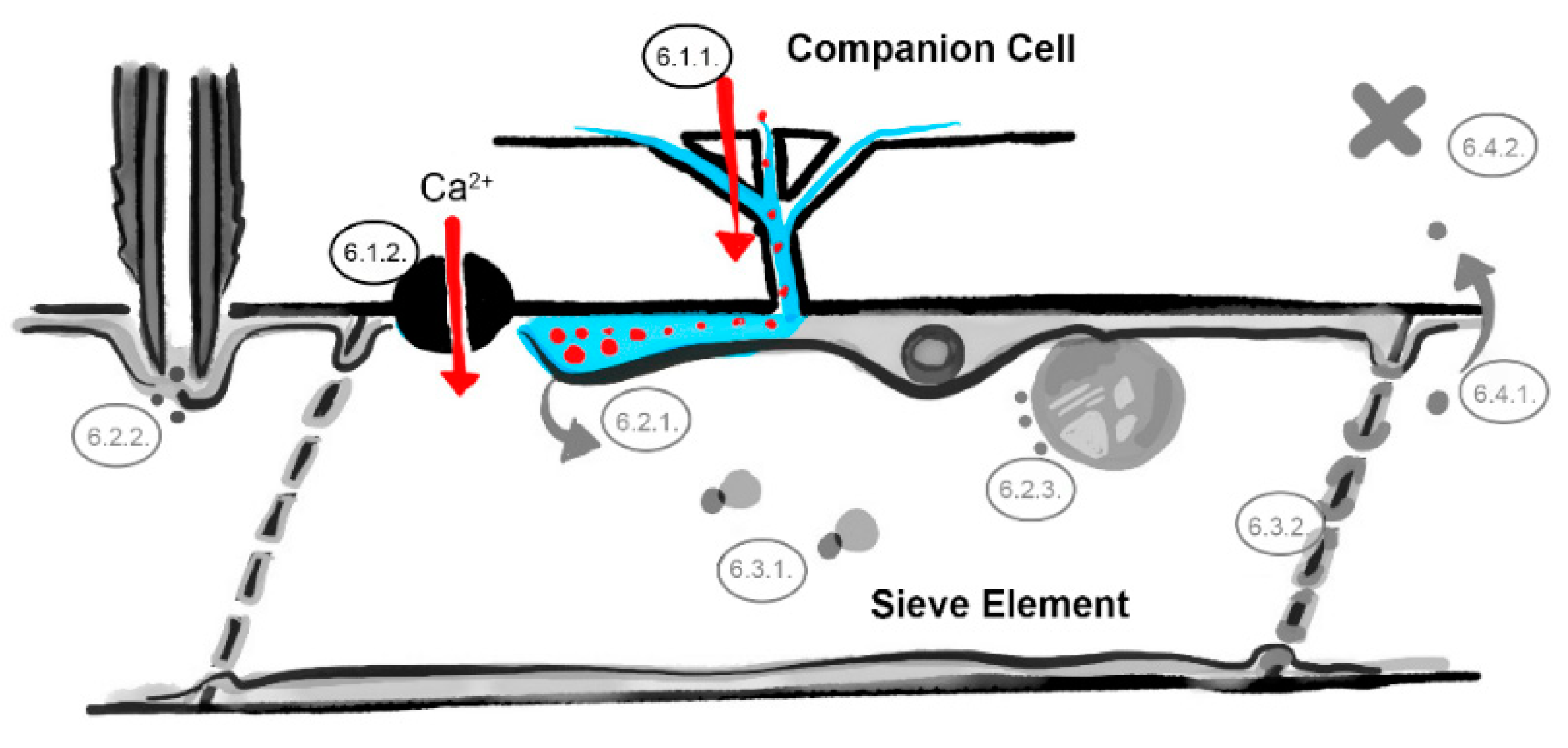
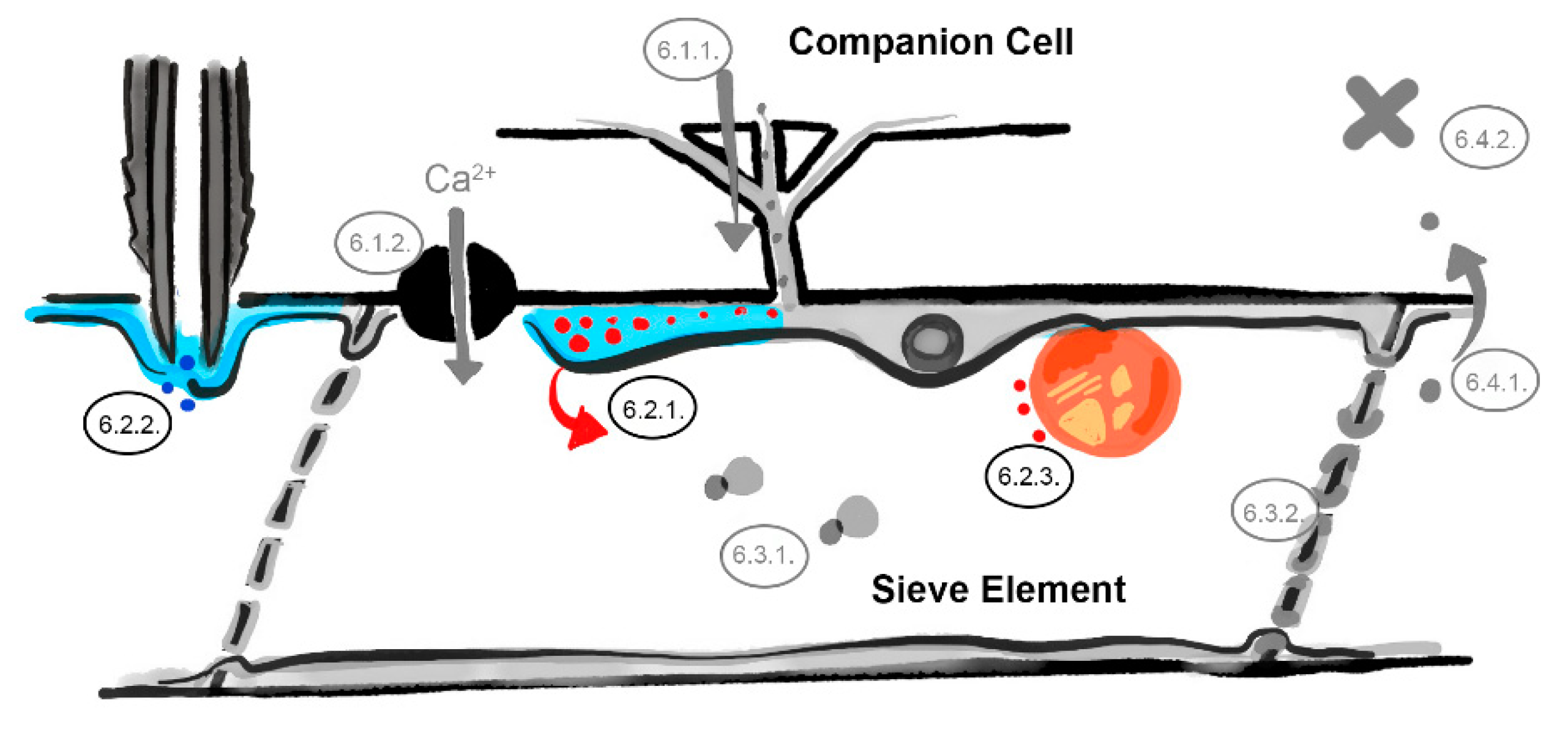
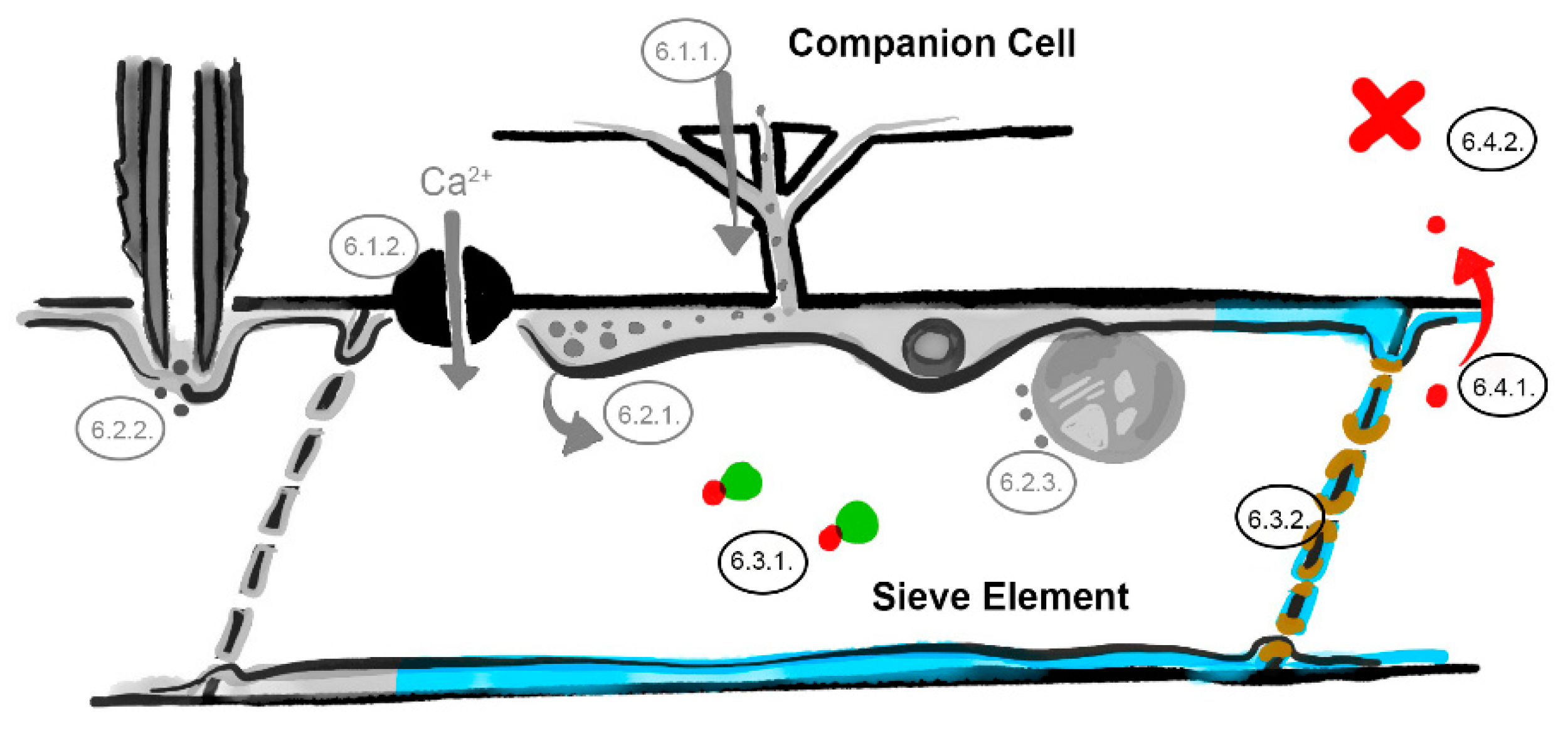
6. What Are the Underlying Mechanisms of RTM- and SLI1-Mediated Resistance?
6.1. The Companion Cell–Sieve Element Interface (Loading)
6.1.1. Restricted Viral Entry into the Sieve Tube
6.1.2. Influx of Ions and Loading of Defense Molecules
6.2. The Parietal Layer–Phloem Sap Interface
6.2.1. Sieve Element Reticulum Defenses against Virus Replication and Assembly
6.2.2. Parietal Occlusion Mechanisms against Aphids
6.2.3. Plastid- and Mitochondria-Induced Defenses Against Viruses and Aphids
6.3. Limiting Factors in the Phloem Sap
6.3.1. Phloem-Mobile Virus Chaperones
6.3.2. Sieve Tube Occlusion
6.4. The Sieve Element–Companion Cell Interface (Unloading)
6.4.1. Restricted Virus Unloading
6.4.2. Unloading of Systemic Resistance Factors
7. Conclusions and Future Outlooks
Funding
Conflicts of Interest
References
- Lecoq, H.; Labonne, G.; Pitrat, M. Specificity of resistance to virus transmission by aphids in Cucumis melo. Ann. Phytopathol. 1980, 12, 139–144. [Google Scholar]
- Pitrat, M.; Lecoq, H. Inheritance of Resistance to Cucumber Mosaic Virus Transmission by Aphis gossypii in Cucumis melo. Phytopathology 1980, 70, 958–961. [Google Scholar] [CrossRef] [Green Version]
- Boissot, N.; Schoeny, A.; Vanlerberghe-Masutti, F. Vat, an Amazing Gene Conferring Resistance to Aphids and Viruses They Carry: From Molecular Structure to Field Effects. Front. Plant Sci. 2016, 7, 1420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bondino, H.G.; Valle, E.M.; ten Have, A. Evolution and functional diversification of the small heat shock protein/α-crystallin family in higher plants. Planta 2012, 235, 1299–1313. [Google Scholar] [CrossRef] [PubMed]
- Whitham, S.A.; Anderberg, R.J.; Chisholm, S.T.; Carrington, J.C. Arabidopsis RTM2 gene is necessary for specific restriction of Tobacco Etch Virus and encodes an unusual small heat shock—Like protein. Plant Cell 2000, 12, 569–582. [Google Scholar]
- Chisholm, S.T.; Mahajan, S.K.; Whitham, S.A.; Yamamoto, M.L.; Carrington, J.C. Cloning of the Arabidopsis RTM1 gene, which controls restriction of long-distance movement of tobacco etch virus. Proc. Natl. Acad. Sci. USA 2000, 97, 489–494. [Google Scholar] [CrossRef] [Green Version]
- Hartig, T. Vergleichende Untersuchungen über die Organisation des Stammes der einheimischen Waldbäume. Jahresber. Fortschr. Forstwissensch. Forstl. Naturkunde 1837, 1, 125–168. [Google Scholar]
- Thompson, G.A.; Schulz, A. Macromolecular trafficking in the phloem. Trends Plant Sci. 1999, 4, 354–360. [Google Scholar] [CrossRef]
- Knoblauch, M.; Peters, W.S. Münch, morphology, microfluidics—Our structural problem with the phloem. Plant Cell Environ. 2010, 33, 1439–1452. [Google Scholar] [CrossRef]
- Lemoine, R.; Camera, S.L.; Atanassova, R.; Dédaldéchamp, F.; Allario, T.; Pourtau, N.; Bonnemain, J.; Laloi, M.; Coutos-Thévenot, P.; Maurousset, L.; et al. Source-to-sink transport of sugar and regulation by environmental factors. Front. Plant Sci. 2013, 4, 1–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- White, R.G. Cell Biology of Sieve Element–Companion Cell Complexes. In Phloem: Molecular Cell Biology, Systemic Communication, Biotic Interactions; Thompson, G.A., Van Bel, A.J.E., Eds.; Wiley-Blackwell: Oxford, UK, 2013; pp. 8–29. [Google Scholar]
- Evert, R.F. Dicotyledons. In Sieve Elements: Comparative Structure, Induction and Development; Behnke, H.D., Sjolund, R.D., Eds.; Springer: Berlin/Heidelberg, Germany, 1990; pp. 103–137. [Google Scholar]
- Van Bel, A.J.E. The phloem, a miracle of ingenuity. Plant Cell Environ. 2003, 26, 125–149. [Google Scholar] [CrossRef]
- Van Bel, A.J.E.; van Rijen, H.V.M. Microelectrode-recorded development of the symplasmic autonomy of the sieve element/companion cell complex in the stem phloem of Lupinus luteus L. Planta 1994, 192, 165–175. [Google Scholar] [CrossRef]
- Oparka, K.J.; Turgeon, R. Sieve elements and companion cells—Traffic control centers of the phloem. Plant Cell 1999, 11, 739–750. [Google Scholar]
- Turgeon, R.; Wolf, S. Phloem Transport: Cellular Pathways and Molecular Trafficking. Annu. Rev. Plant Biol. 2009, 60, 207–221. [Google Scholar] [CrossRef] [Green Version]
- Kehr, J.; Buhtz, A. Long distance transport and movement of RNA through the phloem. J. Exp. Bot. 2008, 59, 85–92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stadler, R.; Lauterbach, C.; Sauer, N. Cell-to-Cell Movement of Green Fluorescent Protein Reveals Post-Phloem Transport in the Outer Integument and Identifies Symplastic Domains in Arabidopsis Seeds and Embryos. Plant Physiol. 2005, 139, 701–712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Balachandran, S.; Xiang, Y.; Schobert, C.; Thompson, G.A.; Lucas, W.J. Phloem sap proteins from Cucurbita maxima and Ricinus communis have the capacity to traffic cell to cell through plasmodesmata. Proc. Natl. Acad. Sci. USA 1997, 94, 14150–14155. [Google Scholar] [CrossRef] [Green Version]
- Ishiwatari, Y.; Fujiwara, T.; McFarland, K.C.; Nemoto, K.; Hayashi, H.; Chino, M.; Lucas, W.J. Rice phloem thioredoxin h has the capacity to mediate its own cell-to-cell transport through plasmodesmata. Planta 1998, 205, 12–22. [Google Scholar] [CrossRef]
- Lough, T.J.; Lucas, W.J. Integrative Plant Biology: Role of Phloem Long-Distance Macromolecular Trafficking. Annu. Rev. Plant Biol. 2006, 57, 203–232. [Google Scholar] [CrossRef] [Green Version]
- Oparka, K.J.; Cruz, S.S. The great escape: Phloem transport and unloading of macromolecules. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2000, 51, 323–347. [Google Scholar] [CrossRef] [Green Version]
- Leisner, S.M.; Turgeon, R. Movement of virus and photoassimilate in the phloem: A comparative analysis. BioEssays 1993, 15, 741–748. [Google Scholar] [CrossRef] [PubMed]
- Revers, F.; García, J.A. Molecular Biology of Potyviruses. In Advances in Virus Research; Maramorosch, K., Mettenleiter, T.C., Eds.; Academic Press: Cambridge, MA, USA, 2015; Volume 92, pp. 101–199. [Google Scholar]
- Harris, K.F. An ingestion-egestion hypothesis of noncirculative virus transmission. In Aphids As Virus Vectors; Harris, K.F., Maramorosch, K., Eds.; Academic Press: New York, NY, USA, 1977; pp. 165–220. [Google Scholar]
- Vuorinen, A.L.; Kelloniemi, J.; Valkonen, J.P.T. Why do viruses need phloem for systemic invasion of plants? Plant Sci. 2011, 181, 355–363. [Google Scholar] [CrossRef] [PubMed]
- Heinlein, M. Plasmodesmata: Channels for Viruses on the Move. In Plasmodesmata. Methods in Molecular Biology (Methods and Protocols); Heinlein, M., Ed.; Springer: New York, NY, USA, 2015; pp. 25–54. [Google Scholar]
- Sevilem, I.; Yadav, S.R.; Helariutta, Y. Plasmodesmata: Channels for Intercellular Signaling During Plant Growth and Development. In Plasmodesmata. Methods in Molecular Biology (Methods and Protocols); Heinlein, M., Ed.; Springer Science & Business Media: New York, NY, USA, 2015; Volume 1217, pp. 3–24. [Google Scholar]
- Ritzenthaler, C.; Hofmann, C. Tubule-Guided Movement of Plant Viruses. In Viral Transport in Plants. Plant Cell Monographs; Waigmann, E., Heinlein, M., Eds.; Springer: Berlin/Heidelberg, Germany, 2007; Volume 7, pp. 63–84. [Google Scholar]
- Waigmann, E.; Ueki, S.; Trutnyeva, K.; Citovsky, V. The Ins and Outs of Nondestructive Cell-to-Cell and Systemic Movement of Plant Viruses. CRC Crit. Rev. Plant Sci. 2004, 23, 195–250. [Google Scholar] [CrossRef]
- Hipper, C.; Brault, V.; Ziegler-Graff, V.; Revers, F. Viral and Cellular Factors Involved in Phloem Transport of Plant Viruses. Front. Plant Sci. 2013, 4, 154. [Google Scholar] [CrossRef] [Green Version]
- Mahajan, S.K.; Chisholm, S.T.; Whitham, S.A.; Carrington, J.C. Identification and characterization of a locus (RTM1) that restricts long-distance movement of tobacco etch virus in Arabidopsis thaliana. Plant J. 1998, 14, 177–186. [Google Scholar] [CrossRef]
- Decroocq, V.; Sicard, O.; Alamillo, J.M.; Lansac, M.; Eyquard, J.P.; García, J.A.; Candresse, T.; Le Gall, O.; Revers, F. Multiple Resistance Traits Control Plum pox virus Infection in Arabidopsis thaliana. MPMI 2006, 19, 541–549. [Google Scholar] [CrossRef] [Green Version]
- Rubio, B.; Cosson, P.; Caballero, M.; Revers, F.; Bergelson, J.; Roux, F.; Schurdi-Levraud, V. Genome-wide association study reveals new loci involved in Arabidopsis thaliana and Turnip mosaic virus (TuMV) interactions in the field. New Phytol. 2019, 221, 2026–2038. [Google Scholar] [CrossRef]
- Chisholm, S.T.; Parra, M.A.; Anderberg, R.J.; Carrington, J.C. Arabidopsis RTM1 and RTM2 genes function in phloem to restrict long-distance movement of Tobacco Etch Virus. Plant Physiol. 2001, 127, 1667–1675. [Google Scholar] [CrossRef]
- Cosson, P.; Sofer, L.; Le, Q.H.; Léger, V.; Schurdi-Levraud, V.; Whitham, S.A.; Yamamoto, M.L.; Gopalan, S.; Le Gall, O.; Candresse, T.; et al. RTM3, which controls long-distance movement of potyviruses, is a member of a new plant gene family encoding a meprin and TRAF homology domain-containing protein. Plant Physiol. 2010, 154, 222–232. [Google Scholar] [CrossRef] [Green Version]
- Whitham, S.A.; Yamamoto, M.L.; Carrington, J.C. Selectable viruses and altered susceptibility mutants in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1999, 96, 772–777. [Google Scholar] [CrossRef] [Green Version]
- Cosson, P.; Schurdi-Levraud, V.; Le, Q.H.; Sicard, O.; Caballero, M.; Roux, F.; Le Gall, O.; Candresse, T.; Revers, F. The RTM resistance to potyviruses in Arabidopsis thaliana: Natural variation of the RTM genes and evidence for the implication of additional genes. PLoS ONE 2012, 7, e39169. [Google Scholar] [CrossRef] [Green Version]
- Esch, L.; Schaffrath, U. An Update on Jacalin-Like Lectins and Their Role in Plant Defense. Int. J. Mol. Sci. 2017, 18, 1592. [Google Scholar]
- Bakthisaran, R.; Tangirala, R.; Rao, C.M. Small heat shock proteins: Role in cellular functions and pathology. Biochim. Biophys. Acta BBA Proteins Proteomics 2015, 1854, 291–319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cosson, P.; Sofer, L.; Schurdi-Levraud, V.; Revers, F. A member of a new plant gene family encoding a meprin and TRAF homology (MATH) domain-containing protein is involved in restriction of long distance movement of plant viruses. Plant Signal Behav. 2010, 5, 1321–1323. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, H.H. Structure of TRAF Family: Current Understanding of Receptor Recognition. Front. Immunol. 2018, 9, 1999. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cayla, T.; Batailler, B.; Le Hir, R.; Revers, F.; Anstead, J.A.; Thompson, G.A.; Grandjean, O.; Dinant, S. Live imaging of companion cells and sieve elements in Arabidopsis leaves. PLoS ONE 2015, 10, e0118122. [Google Scholar] [CrossRef] [PubMed]
- Decroocq, V.; Salvador, B.; Sicard, O.; Glasa, M.; Cosson, P.; Svanella-Dumas, L.; Revers, F.; García, J.A.; Candresse, T. The Determinant of Potyvirus Ability to Overcome the RTM Resistance of Arabidopsis thaliana Maps to the N-Terminal Region of the Coat Protein. MPMI 2009, 22, 1302–1311. [Google Scholar] [CrossRef] [Green Version]
- Sofer, L.; Cabanillas, D.G.; Gayral, M.; Téplier, R.; Pouzoulet, J.; Ducousso, M.; Dufin, L.; Bréhélin, C.; Ziegler-Graff, V.; Brault, V.; et al. Identification of host factors potentially involved in RTM-mediated resistance during potyvirus long distance movement. Arch. Virol. 2017, 162, 1855–1865. [Google Scholar] [CrossRef]
- Kloth, K.J.; Busscher, J.; Wiegers, G.; Kruijer, W.; Buijs, G.; Meyer, R.C.; Albrectsen, B.R.; Bouwmeester, H.J.; Dicke, M.; Jongsma, M.A. Sieve Element-Lining Chaperone 1 restricts aphid feeding on Arabidopsis during heat stress. Plant Cell 2017, 29, 2450–2464. [Google Scholar] [CrossRef] [Green Version]
- Will, T.; Tjallingii, W.F.; Thönnessen, A.; van Bel, A.J.E. Molecular sabotage of plant defense by aphid saliva. Proc. Natl. Acad. Sci. USA 2007, 104, 10536–10541. [Google Scholar] [CrossRef] [Green Version]
- Wan, J.; Cabanillas, D.G.; Zheng, H.Q.; Laliberté, J.F. Turnip mosaic virus Moves Systemically through Both Phloem and Xylem as Membrane-Associated Complexes. Plant Physiol. 2015, 167, 1374–1388. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blackman, L.M.; Boevink, P.; Cruz, S.S.; Palukaitis, P.; Oparka, K.J. The Movement Protein of Cucumber Mosaic Virus Traffics into Sieve Elements in Minor Veins of Nicotiana clevelandii. Plant Cell 1998, 10, 525–537. [Google Scholar] [CrossRef] [Green Version]
- Crawford, K.M.; Zambryski, P.C. Phloem transport: Are you chaperoned? Curr. Biol. 1999, 9, R281–R285. [Google Scholar] [CrossRef] [Green Version]
- Collum, T.D.; Padmanabhan, M.S.; Hsieh, Y.-C.; Culver, J.N. Tobacco mosaic virus-directed reprogramming of auxin/indole acetic acid protein transcriptional responses enhances virus phloem loading. Proc. Natl. Acad. Sci. USA 2016, 113, E2740–E2749. [Google Scholar] [CrossRef] [Green Version]
- Toyota, M.; Spencer, D.; Sawai-Toyota, S.; Jiaqi, W.; Zhang, T.; Koo, A.J.; Howe, G.A.; Gilroy, S. Glutamate triggers long-distance, calcium-based plant defense signaling. Science 2018, 361, 1112–1115. [Google Scholar] [CrossRef]
- Nguyen, C.T.; Kurenda, A.; Stolz, S.; Chételat, A.; Farmer, E.E. Identification of cell populations necessary for leaf-to-leaf electrical signaling in a wounded plant. Proc. Natl. Acad. Sci. USA 2018, 115, 10178–10183. [Google Scholar] [CrossRef] [Green Version]
- Knoblauch, M.; Peters, W.S.; Ehlers, K.; van Bel, A.J.E. Reversible Calcium-Regulated Stopcocks in Legume Sieve Tubes. Plant Cell 2001, 13, 1221–1230. [Google Scholar] [CrossRef] [Green Version]
- Will, T.; Kornemann, S.R.; Furch, A.C.U.; Tjallingii, W.F.; van Bel, A.J.E. Aphid watery saliva counteracts sieve-tube occlusion: A universal phenomenon? JEB 2009, 212, 3305–3312. [Google Scholar] [CrossRef] [Green Version]
- Kettles, G.J.; Drurey, C.; Schoonbeek, H.-J.; Maule, A.J.; Hogenhout, S.A. Resistance of Arabidopsis thaliana to the green peach aphid, Myzus persicae, involves camalexin and is regulated by microRNAs. New Phytol. 2013, 198, 1178–1190. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.H.; Lee, B.W.; Schroeder, F.C.; Jander, G. Identification of indole glucosinolate breakdown products with antifeedant effects on Myzus persicae (green peach aphid). Plant J. 2008, 54, 1015–1026. [Google Scholar] [CrossRef]
- Jørgensen, M.E.; Nour-Eldin, H.H.; Halkier, B.A. Transport of defense compounds from source to sink: Lessons learned from glucosinolates. Trends Plant Sci. 2015, 20, 508–514. [Google Scholar] [CrossRef] [PubMed]
- Andersen, T.G.; Nour-Eldin, H.H.; Fuller, V.L.; Olsen, C.E.; Burow, M.; Halkier, B.A. Integration of Biosynthesis and Long-Distance Transport Establish Organ-Specific Glucosinolate Profiles in Vegetative Arabidopsis. Plant Cell 2013, 25, 3133–3145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sjolund, R.D.; Shih, C.Y. Freeze-fracture analysis of phloem structure in plant tissue cultures: I. The sieve element reticulum. J. Ultrastruct. Res. 1983, 82, 111–121. [Google Scholar] [CrossRef]
- Callaway, A.; Liu, W.; Andrianov, V.; Stenzler, L.; Zhao, J.; Wettlaufer, S.; Jayakumar, P.; Howell, S.H. Characterization of cauliflower mosaic virus (CaMV) resistance in virus-resistant ecotypes of Arabidopsis. MPMI 1996, 9, 810–818. [Google Scholar] [CrossRef]
- D’Arcy, C.J.; Zoeten, G.A. Beet Western Yellows virus in phloem tissue of Thlapsi arvense. Phytopathology 1979, 69, 1194–1198. [Google Scholar] [CrossRef]
- Léonard, S.; Viel, C.; Beauchemin, C.; Daigneault, N.; Fortin, M.G.; Laliberté, J.-F. Interaction of VPg-Pro of Turnip mosaic virus with the translation initiation factor 4E and the poly(A)-binding protein in planta. J. Gen. Virol. 2004, 85, 1055–1063. [Google Scholar] [CrossRef]
- Cotton, S.; Grangeon, R.; Thivierge, K.; Mathieu, I.; Ide, C.; Wei, T.; Wang, A.; Laliberté, J.-F. Turnip Mosaic Virus RNA Replication Complex Vesicles Are Mobile, Align with Microfilaments, and Are Each Derived from a Single Viral Genome. J. Virol. 2009, 83, 10460–10471. [Google Scholar] [CrossRef] [Green Version]
- Sasaki, T.; Chino, M.; Hayashi, H.; Fujiwara, T. Detection of Several mRNA Species in Rice Phloem Sap. Plant Cell Physiol. 1998, 39, 895–897. [Google Scholar] [CrossRef]
- Doering-Saad, C.; Newbury, H.J.; Bale, J.S.; Pritchard, J. Use of aphid stylectomy and RT-PCR for the detection of transporter mRNAs in sieve elements. J. Exp. Bot. 2002, 53, 631–637. [Google Scholar] [CrossRef]
- Kim, Y.J.; Maizel, A.; Chen, X. Traffic into silence: Endomembranes and post-transcriptional RNA silencing. EMBO J. 2014, 33, 968–980. [Google Scholar] [CrossRef] [Green Version]
- Garcia-Ruiz, H.; Takeda, A.; Chapman, E.J.; Sullivan, C.M.; Fahlgren, N.; Brempelis, K.J.; Carrington, J.C. Arabidopsis RNA-Dependent RNA Polymerases and Dicer-Like Proteins in Antiviral Defense and Small Interfering RNA Biogenesis during Turnip Mosaic Virus Infection. Plant Cell 2010, 22, 481–496. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- van Emden, H.F.; Harrington, R. Aphids As Crop Pests; CABI: Oxford, UK, 2007. [Google Scholar]
- Will, T.; Furch, A.C.U.; Zimmermann, M.R. How phloem-feeding insects face the challenge of phloem-located defenses. Front. Plant Sci. 2013, 4, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhattacharyya, D.; Chakraborty, S. Chloroplast: The Trojan horse in plant–virus interaction. Mol. Plant Pathol. 2018, 19, 504–518. [Google Scholar] [CrossRef] [Green Version]
- Behnke, H.D. Sieve-element characters. Nord. J. Bot. 1981, 1, 381–400. [Google Scholar] [CrossRef]
- Knoblauch, M.; Van Bel, A.J.E. Sieve tubes in action. Plant Cell 1998, 10, 35–50. [Google Scholar] [CrossRef] [Green Version]
- Ruan, L.; Zhou, C.; Jin, E.; Kucharavy, A.; Zhang, Y.; Wen, Z.; Florens, L.; Li, R. Cytosolic proteostasis through importing of misfolded proteins into mitochondria. Nature 2017, 543, 443. [Google Scholar] [CrossRef]
- Misawa, T.; Takahama, M.; Saitoh, T. Mitochondria–Endoplasmic Reticulum Contact Sites Mediate Innate Immune Responses. In Organelle Contact Sites: From Molecular Mechanism to Disease; Tagaya, M., Simmen, T., Eds.; Springer: Singapore, 2017; pp. 187–197. [Google Scholar]
- Fisher, D.B.; Wu, Y.; Ku, M.S.B. Turnover of Soluble Proteins in the Wheat Sieve Tube. Plant Physiol. 1992, 100, 1433–1441. [Google Scholar] [CrossRef] [Green Version]
- Gaupels, F.; Buhtz, A.; Knauer, T.; Deshmukh, S.; Waller, F.; van Bel, A.J.E.; Kogel, K.-H.; Kehr, J. Adaptation of aphid stylectomy for analyses of proteins and mRNAs in barley phloem sap. J. Exp. Bot. 2008, 59, 3297–3306. [Google Scholar] [CrossRef] [Green Version]
- Navarro, J.A.; Sanchez-Navarro, J.A.; Pallas, V. Key checkpoints in the movement of plant viruses through the host. In Advances in Virus Research; Kielian, M., Mettenleiter, T.C., Roossinck, M.J., Eds.; Academic Press: Cambridge, MA, USA, 2019; Volume 104, pp. 1–64. [Google Scholar]
- Gómez, G.; Pallás, V. A long-distance translocatable phloem protein from cucumber forms a ribonucleoprotein complex in vivo with Hop Stunt Viroid RNA. J. Virol. 2004, 78, 10104–10110. [Google Scholar] [CrossRef] [Green Version]
- Requena, A.; Simón-Buela, L.; Salcedo, G.; García-Arenal, F. Potential Involvement of a Cucumber Homolog of Phloem Protein 1 in the Long-Distance movement of Cucumber mosaic virus Particles. MPMI 2006, 19, 734–746. [Google Scholar] [CrossRef]
- Shalitin, D.; Wolf, S. Interaction between phloem proteins and viral movement proteins. Aust. J. Plant Physiol. 2000, 27, 801–806. [Google Scholar] [CrossRef]
- Knoblauch, M.; Mullendore, D. Sieve element occlusion. In Phloem: Molecular Cell Biology, Systemic Communication, Biotic Interactions; Thompson, G.A., van Bel, A.J.E., Eds.; Wiley-Blackwell: Oxford, UK, 2013; pp. 141–153. [Google Scholar]
- Esau, K.; Cheadle, V.I. Size of pores and their contents in sieve elements of Dicotyledons. Proc. Natl. Acad. Sci. USA 1959, 45, 156–162. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kalmbach, L.; Helariutta, Y. Sieve Plate Pores in the Phloem and the Unknowns of Their Formation. Plants 2019, 8, 25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Furch, A.C.U.; Hafke, J.B.; Schulz, A.; van Bel, A.J.E. Ca2+-mediated remote control of reversible sieve tube occlusion in Vicia faba. J. Exp. Bot. 2007, 58, 2827–2838. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knoblauch, M.; Froelich, D.R.; Pickard, W.F.; Peters, W.S. SEORious business: Structural proteins in sieve tubes and their involvement in sieve element occlusion. J. Exp. Bot. 2014, 65, 1879–1893. [Google Scholar] [CrossRef]
- Ernst, A.M.; Jekat, S.B.; Zielonka, S.; Müller, B.; Neumann, U.; Rüping, B.; Twyman, R.M.; Krzyzanek, V.; Prüfer, D.; Noll, G.A. Sieve element occlusion (SEO) genes encode structural phloem proteins involved in wound sealing of the phloem. Proc. Natl. Acad. Sci. USA 2012, 109, E1980–E1989. [Google Scholar] [CrossRef] [Green Version]
- Froelich, D.R.; Mullendore, D.L.; Jensen, K.H.; Ross-Elliott, T.J.; Anstead, J.A.; Thompson, G.A.; Pélissier, H.C.; Knoblauch, M. Phloem ultrastructure and pressure flow: Sieve-element-occlusion-related agglomerations do not affect translocation. Plant Cell 2011, 23, 4428–4445. [Google Scholar] [CrossRef] [Green Version]
- Tjallingii, W.F. Salivary secretions by aphids interacting with proteins of phloem wound responses. J. Exp. Bot. 2006, 57, 739–745. [Google Scholar] [CrossRef]
- Will, T.; Vilcinskas, A. The structural sheath protein of aphids is required for phloem feeding. Insect Biochem. Mol. Biol. 2015, 57, 34–40. [Google Scholar] [CrossRef]
- Esau, K.; Cronshaw, J. Endoplasmic reticulum in the sieve element of Cucurbita. J. Ultrastruct. Res. 1968, 23, 1–14. [Google Scholar] [CrossRef]
- Ruffet, E.; Paquet, N.; Frutiger, S.; Hughes, G.J.; Jaton, J.C. Structural and electron-microscopic studies of jacalin from jackfruit (Artocarpus integrifolia) show that this lectin is a 65 kDa tetramer. Biochem. J. 1992, 286, 131–134. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haslbeck, M.; Franzmann, T.; Weinfurtner, D.; Buchner, J. Some like it hot: The structure and function of small heat-shock proteins. Nat. Struct. Mol. Biol. 2005, 12, 842. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.S.; Wellink, J.; Goldbach, R.W.; van Lent, J.W.M. Phloem loading and unloading of Cowpea mosaic virus in Vigna unguiculata. J. Gen. Virol. 2002, 83, 1493–1504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, F.; D’Auria, J.C.; Tholl, D.; Ross, J.R.; Gershenzon, J.; Noel, J.P.; Pichersky, E. An Arabidopsis thaliana gene for methylsalicylate biosynthesis, identified by a biochemical genomics approach, has a role in defense. Plant J. 2003, 36, 577–588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, M.; Ye, X.; Willie, K.; Lin, J.; Zhang, X.; Redinbaugh, M.G.; Simon, A.E.; Morris, T.J.; Qu, F. The Capsid Protein of Turnip Crinkle Virus Overcomes Two Separate Defense Barriers To Facilitate Systemic Movement of the Virus in Arabidopsis. J. Virol. 2010, 84, 7793–7802. [Google Scholar] [CrossRef] [Green Version]
- Gill, C.C.; Chong, J. Development of the infection in oat leaves inoculated with barley yellow dwarf virus. Virology 1975, 66, 440–453. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kloth, K.J.; Kormelink, R. Defenses against Virus and Vector: A Phloem-Biological Perspective on RTM- and SLI1-Mediated Resistance to Potyviruses and Aphids. Viruses 2020, 12, 129. https://0-doi-org.brum.beds.ac.uk/10.3390/v12020129
Kloth KJ, Kormelink R. Defenses against Virus and Vector: A Phloem-Biological Perspective on RTM- and SLI1-Mediated Resistance to Potyviruses and Aphids. Viruses. 2020; 12(2):129. https://0-doi-org.brum.beds.ac.uk/10.3390/v12020129
Chicago/Turabian StyleKloth, Karen J., and Richard Kormelink. 2020. "Defenses against Virus and Vector: A Phloem-Biological Perspective on RTM- and SLI1-Mediated Resistance to Potyviruses and Aphids" Viruses 12, no. 2: 129. https://0-doi-org.brum.beds.ac.uk/10.3390/v12020129




