PCR Detection and Phylogenetic Analysis of Megalocytivirus Isolates in Farmed Giant Sea Perch Lates calcarifer in Southern Taiwan
Abstract
:1. Introduction
2. Materials and Methods
2.1. Fish and Virus
2.2. Viral DNA Extraction
2.3. PCR Assays for the Identification and Differentiation of GSPIV Isolates from Giant Sea Perch
2.4. Viral MCP Gene Cloning and Sequencing
2.5. Phylogenetic Analysis
3. Results
3.1. PCR Identification of GSPIV Isolates from Giant Sea Perch
3.1.1. The OIE-Registered Primer Sets
3.1.2. The Megalocytivirus Universal Primer Set
3.1.3. Primer Sets Respectively Specific to RSIV, ISKNV and TRBIV
3.1.4. The Hemi-Nested (hn) Primer Set
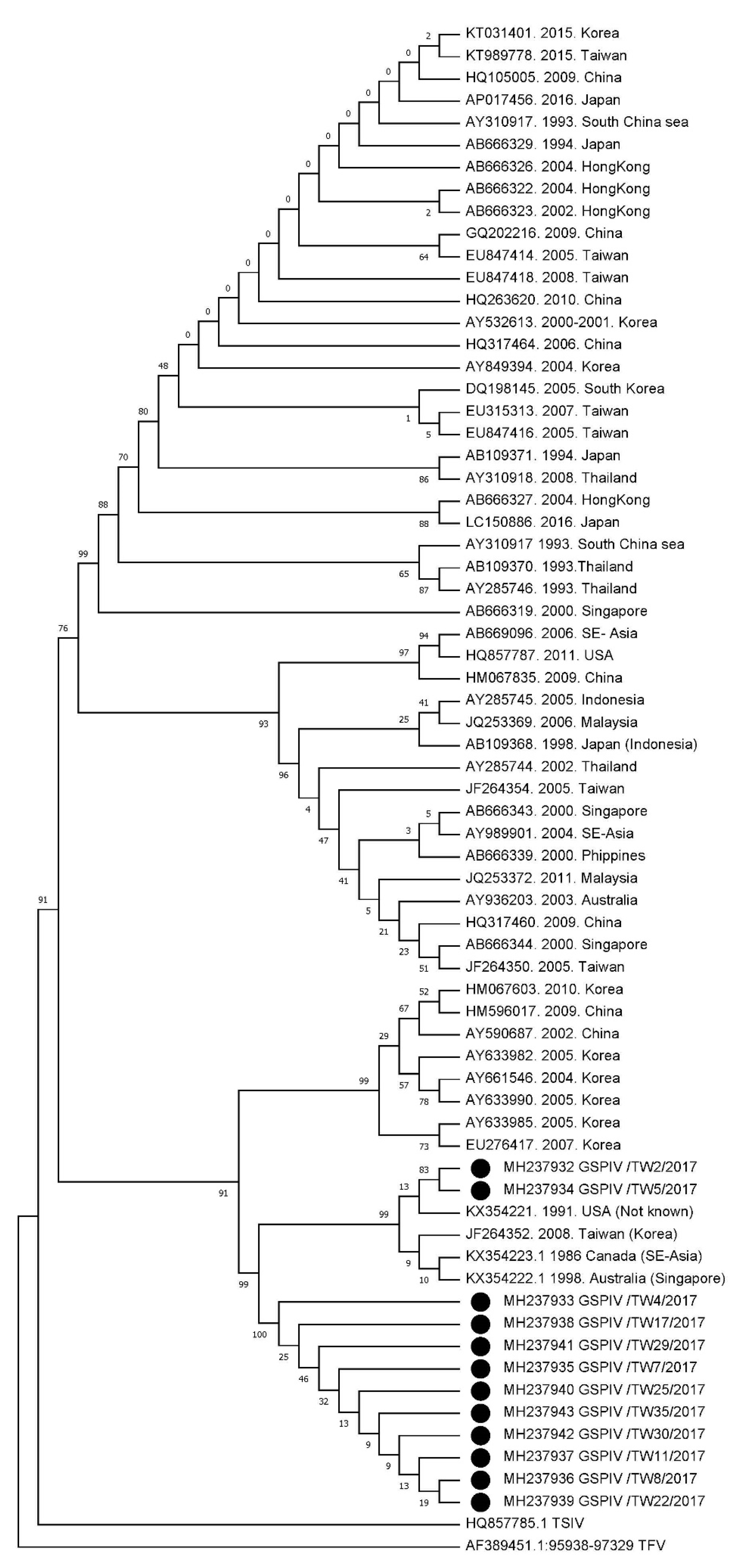
3.2. Phylogenetic Analysis of GSPIV Isolates from Giant Sea Perch
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gibson-Kueh, S.; Netto, P.; Ngoh-Lim, G.H.; Chang, S.F.; Ho, L.L.; Qin, Q.W.; Chua, F.H.; Ng, M.L.; Ferguson, H.W. The pathology of systemic iridoviral disease in fish. J. Comp. Pathol. 2003, 129, 111–119. [Google Scholar] [CrossRef]
- Chinchar, V.G.; Hyatt, A.; Miyazaki, T.; Williams, T. Family Iridoviridae: Poor viral relations no longer. Curr. Top Microbiol. Immunol. 2009, 328, 123–170. [Google Scholar] [CrossRef] [PubMed]
- Williams, T.; Barbosa-Solomieu, V.; Chinchar, V.G. A decade of advances in iridovirus research. Adv Virus Res 2005, 65, 173–248. [Google Scholar] [CrossRef]
- Chinchar, V.G.; Essbayer, S.; He, J.G.; Hyatt, A.; Miyazaki, T.; Seligy, V.; Williams, T. Family Iridoviridae. In Virus Taxonomy, 8th ed.; Fauquet, C.M., Mayo, M.A., Maniloff, J., Desselberger, U., Ball, L.A., Eds.; Elsevier Academic Press: San Diego, CA, USA, 2005; pp. 145–162. [Google Scholar]
- Eaton, H.E.; Ring, B.A.; Brunetti, C.R. The genomic diversity and phylogenetic relationship in the family iridoviridae. Viruses 2010, 2, 1458–1475. [Google Scholar] [CrossRef] [Green Version]
- Kurita, J.; Nakajima, K. Megalocytiviruses. Viruses 2012, 4, 521–538. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, C.S.; Chao, S.Y.; Ku, C.C.; Wen, C.M.; Shih, H.H. PCR amplification and sequence analysis of the major capsid protein gene of megalocytiviruses isolated in Taiwan. J. Fish Dis. 2009, 32, 543–550. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.M.; Tu, C.; Tseng, C.H.; Huang, C.C.; Chou, C.C.; Kuo, H.C.; Chang, S.K. Genetic analysis of fish iridoviruses isolated in Taiwan during 2001-2009. Arch. Virol. 2011, 156, 1505–1515. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Inouye, K.; Yamano, K.; Maeno, Y.; Nakajima, K.; Matsuoka, M.; Wada, Y.; Sorimachi, M. Iridovirus infection of cultured red-sea bream, Pagrus major. Fish Pathol. 1992, 27, 19–27. [Google Scholar] [CrossRef]
- He, J.G.; Deng, M.; Weng, S.P.; Li, Z.; Zhou, S.Y.; Long, Q.X.; Wang, X.Z.; Chan, S.M. Complete genome analysis of the mandarin fish infectious spleen and kidney necrosis iridovirus. Virology 2001, 291, 126–139. [Google Scholar] [CrossRef] [Green Version]
- He, J.G.; Zeng, K.; Weng, S.P.; Chan, S.M. Experimental transmission, pathogenicity and physical-chemical properties of infectious spleen and kidney necrosis virus (ISKNV). Aquaculture 2002, 204, 11–24. [Google Scholar] [CrossRef]
- Shi, C.Y.; Wang, Y.G.; Yang, S.L.; Huang, J.; Wang, Q.Y. The first report of an iridovirus-like agent infection in fanned turbot, Scophthalmus maximus, in China. Aquaculture 2004, 236, 11–25. [Google Scholar] [CrossRef]
- Do, J.W.; Cha, S.J.; Kim, J.S.; An, E.J.; Lee, N.S.; Choi, H.J.; Lee, C.H.; Park, M.S.; Kim, J.W.; Kim, Y.C.; et al. Phylogenetic analysis of the major capsid protein gene of iridovirus isolates from cultured flounders Paralichthys olivaceus in Korea. Dis. Aquat. Organ. 2005, 64, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Do, J.W.; Cha, S.J.; Kim, J.S.; An, E.J.; Park, M.S.; Kim, J.W.; Kim, Y.C.; Park, M.A.; Park, J.W. Sequence variation in the gene encoding the major capsid protein of Korean fish iridoviruses. Arch. Virol. 2005, 150, 351–359. [Google Scholar] [CrossRef]
- Nakajima, K.; Kunita, J. Red sea bream iridoviral disease. Uirusu 2005, 55, 115–125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sawayama, E.; Tanizawa, S.; Kitamura, S.I.; Nakayama, K.; Ohta, K.; Ozaki, A.; Takagi, M. Identification of quantitative trait loci for resistance to RSIVD in red sea bream (Pagrus major). Mar. Biotechnol. (NY) 2017, 19, 601–613. [Google Scholar] [CrossRef]
- Dong, C.; Xiong, X.; Luo, Y.; Weng, S.; Wang, Q.; He, J. Efficacy of a formalin-killed cell vaccine against infectious spleen and kidney necrosis virus (ISKNV) and immunoproteomic analysis of its major immunogenic proteins. Vet. Microbiol. 2013, 162, 419–428. [Google Scholar] [CrossRef]
- Wang, Q.; Ji, W.; Xu, Z. Current use and development of fish vaccines in China. Fish Shellfish Immunol. 2020, 96, 223–234. [Google Scholar] [CrossRef]
- Subramaniam, K.; Gotesman, M.; Smith, C.E.; Steckler, N.K.; Kelley, K.L.; Groff, J.M.; Waltzek, T.B. Megalocytivirus infection in cultured Nile tilapia Oreochromis niloticus. Dis. Aquat. Organ. 2016, 119, 253–258. [Google Scholar] [CrossRef]
- Jeong, J.B.; Kim, H.Y.; Jun, L.J.; Lyu, J.H.; Park, N.G.; Kim, J.K.; Jeong, H.D. Outbreaks and risks of infectious spleen and kidney necrosis virus disease in freshwater ornamental fishes. Dis. Aquat. Organ. 2008, 78, 209–215. [Google Scholar] [CrossRef]
- Subramaniam, K.; Shariff, M.; Omar, A.R.; Hair-Bejo, M.; Ong, B.L. Detection and molecular characterization of infectious spleen and kidney necrosis virus from major ornamental fish breeding states in Peninsular Malaysia. J. Fish Dis. 2014, 37, 609–618. [Google Scholar] [CrossRef]
- Shi, C.Y.; Jia, K.T.; Yang, B.; Huang, J. Complete genome sequence of a Megalocytivirus (family Iridoviridae) associated with turbot mortality in China. Virol. J. 2010, 7, 159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Won, K.M.; Cho, M.Y.; Park, M.A.; Jee, B.Y.; Myeong, J.I.; Kim, J.W. The first report of a megalocytivirus infection in farmed starry flounder, Platichthys stellatus, in Korea. Fish. Aquat. Sci. 2013, 16, 93–99. [Google Scholar] [CrossRef]
- Go, J.; Waltzek, T.B.; Subramaniam, K.; Yun, S.C.; Groff, J.M.; Anderson, I.G.; Chong, R.; Shirley, I.; Schuh, J.C.; Handlinger, J.H.; et al. Detection of infectious spleen and kidney necrosis virus (ISKNV) and turbot reddish body iridovirus (TRBIV) from archival ornamental fish samples. Dis. Aquat. Organ. 2016, 122, 105–123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koda, S.A.; Subramaniam, K.; Francis-Floyd, R.; Yanong, R.P.; Frasca, S., Jr.; Groff, J.M.; Popov, V.L.; Fraser, W.A.; Yan, A.; Mohan, S.; et al. Phylogenomic characterization of two novel members of the genus Megalocytivirus from archived ornamental fish samples. Dis. Aquat. Organ. 2018, 130, 11–24. [Google Scholar] [CrossRef]
- Wang, C.S.; Shih, H.H.; Ku, C.C.; Chen, S.N. Studies on epizootic iridovirus infection among red sea bream, Pagrus major (Temminck & Schlegel), cultured in Taiwan. J. Fish Dis. 2003, 26, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Paria, A.; Dong, J.; Babu, P.P.S.; Makesh, M.; Chaudhari, A.; Thirunavukkarasu, A.R.; Purushothaman, C.S.; Rajendran, K.V. Evaluation of candidate reference genes for quantitative expression studies in Asian seabass (Lates calcarifer) during ontogenesis and in tissues of healthy and infected fishes. Indian J. Exp. Biol. 2016, 54, 597–605. [Google Scholar]
- Kurita, J.; Nakajima, K.; Hirono, I.; Aoki, T. Polymerase chain reaction (PCR) amplification of DNA of red sea bream iridovirus (RSIV). Fish Pathol. 1998, 33, 17–23. [Google Scholar] [CrossRef] [Green Version]
- Oseko, N.; Chuah, T.T.; Palamisamy, V.; Maeno, Y.; Kurita, J. Iridovirus isolated from diseased sea bass Lates calcarifer and red drum Sciaenops ocellatus causing mass mortality in Malaysia. In Proceedings of the 7th Asian Fisheries Forum 04 Abstracts, Penang, Malaysia, 30 November–4 December 2004; Volume 127. [Google Scholar]
- 2019 OIE-Manual of Diagnostic Tests for Aquatic Animals-14/11/2019 (Chapter 2.3.8.). Available online: https://www.oie.int/standard-setting/aquatic-manual/access-online/ (accessed on 14 May 2020).
- Jung-Schroers, V.; Adamek, M.; Wohlsein, P.; Wolter, J.; Wedekind, H.; Steinhagen, D. First outbreak of an infection with infectious spleen and kidney necrosis virus (ISKNV) in ornamental fish in Germany. Dis. Aquat. Organ. 2016, 119, 239–244. [Google Scholar] [CrossRef] [Green Version]
- Minosse, C.; Selleri, M.; Zaniratti, M.S.; Lauria, F.N.; Puro, V.; Carletti, F.; Cappiello, G.; Gualano, G.; Bevilacqua, N.; Capobianchi, M.R. Improved detection of human influenza A and B viruses in respiratory tract specimens by hemi-nested PCR. J. Virol. Methods 2007, 141, 225–228. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic. Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Gias, E.; Johnston, C.; Keeling, S.; Spence, R.P.; McDonald, W.L. Development of real-time PCR assays for detection of megalocytiviruses in imported ornamental fish. J. Fish Dis. 2011, 34, 609–618. [Google Scholar] [CrossRef] [PubMed]
- Rodgers, C.J.; Mohan, C.V.; Peeler, E.J. The spread of pathogens through trade in aquatic animals and their products. Rev. Sci. Tech. Off. Int. Epiz. 2011, 30, 241–256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rimmer, A.E.; Whittington, R.J.; Tweedie, A.; Becker, J.A. Susceptibility of a number of Australian freshwater fishes to dwarf gourami iridovirus (Infectious spleen and kidney necrosis virus). J. Fish Dis. 2017, 40, 293–310. [Google Scholar] [CrossRef]
- Go, J.; Whittington, R. Experimental transmission of infectious spleen and kidney necrosis virus (ISKNV) from freshwater ornamental fish to silver sweep Scorpis lineolata, an Australian marine fish. Dis. Aquat. Organ. 2019, 137, 1–21. [Google Scholar] [CrossRef]

| Isolate a | Date (yr/mo) of Isolation | Fish Farm District b |
|---|---|---|
| GSPIV/TW1 | 2017/04 | PT |
| GSPIV/TW2 | 2017/04 | PT |
| GSPIV/TW3 | 2017/04 | PT |
| GSPIV/TW4 | 2017/05 | K |
| GSPIV/TW5 | 2017/05 | PT |
| GSPIV/TW6 | 2017/05 | PT |
| GSPIV/TW7 | 2017/06 | PT |
| GSPIV/TW8 | 2017/06 | K |
| GSPIV/TW9 | 2017/06 | K |
| GSPIV/TW10 | 2017/06 | K |
| GSPIV/TW11 | 2017/06 | K |
| GSPIV/TW12 | 2017/06 | PT |
| GSPIV/TW13 | 2017/06 | K |
| GSPIV/TW14 | 2017/07 | PT |
| GSPIV/TW15 | 2017/07 | PT |
| GSPIV/TW16 | 2017/07 | PT |
| GSPIV/TW17 | 2017/07 | PT |
| GSPIV/TW18 | 2017/08 | PT |
| GSPIV/TW19 | 2017/08 | PT |
| GSPIV/TW20 | 2017/08 | PT |
| GSPIV/TW21 | 2017/08 | PT |
| GSPIV/TW22 | 2017/08 | PT |
| GSPIV/TW23 | 2017/08 | PT |
| GSPIV/TW24 | 2017/08 | PT |
| GSPIV/TW25 | 2017/09 | PT |
| GSPIV/TW26 | 2017/09 | PT |
| GSPIV/TW27 | 2017/09 | PT |
| GSPIV/TW28 | 2017/09 | PT |
| GSPIV/TW29 | 2017/09 | PT |
| GSPIV/TW30 | 2017/09 | PT |
| GSPIV/TW31 | 2017/09 | PT |
| GSPIV/TW32 | 2017/09 | PT |
| GSPIV/TW33 | 2017/09 | PT |
| GSPIV/TW34 | 2017/09 | PT |
| GSPIV/TW35 | 2017/09 | PT |
| GSPIV/TW36 | 2017/09 | PT |
| GSPIV/TW37 | 2017/09 | PT |
| GSPIV/TW38 | 2017/09 | PT |
| GSPIV/TW39 | 2017/10 | PT |
| Purpose | Primer | Primer Sequence (5′-3′) | PCR Conditions | Product | Target Gene | Reference |
|---|---|---|---|---|---|---|
| Detection | OIE RSIV/ISKNV | |||||
| 1-F | CTCAAACACTCTGGCTCATC | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min; 72 °C, 1 min; 72 °C, 5 min | 570 bp | Pst I fragment | [28] | |
| 1-R | GCACCAACACATCTCCTATC | |||||
| OIE RSIV | ||||||
| 4-F | CGGGGGCAATGACGACTACA | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min; 72°C, 1 min; 72 °C, 5 min | 568 bp | DNA Polymerase | [29] | |
| 4-R | CCGCCTGTGCCTTTTCTGGA | |||||
| RSIV/ISKNV/TRBIV | ||||||
| MCP- uni332- F3 | AGGTGTCGGTGTCATTAACGACCT G | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min; 72 °C, 1 min; 72 °C, 5 min | 777 bp | MCP | [6,31] | |
| MCP- uni1108- R8 | TCTCAGGCATGCTGGGCGCAAAG | |||||
| RSIV | ||||||
| MCP- specR674- F4 | CCCGCACTGACCAACGTGTCC | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min; 72 °C, 1 min; 72 °C, 5 min | 191 bp | MCP | [6] | |
| MCP- specR888- R6 | CACAGGGTGACTGAACTCAGG TCG | |||||
| ISKNV | ||||||
| MCP- specI465- F3 | GGTGGCCGGCATCACCAACGG C | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min; 72 °C, 1 min; 72 °C, 5 min | 413 bp | MCP | [6] | |
| MCP- specI879- R3 | CACGGGGTGACTGAACCTG | |||||
| TRBIV | ||||||
| MCP- specT37- F1 | TTC ATC GAC ATC TCC GCT TTC | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min; 72 °C, 1 min; 72 °C, 5 min | 453 bp | MCP | [6] | |
| MCP-specT490- R1 | TST GAC CGT TGG TGA TAC CGG AG | |||||
| hn-TRBIV | ||||||
| MCP- specT37- F1 | TTC ATC GAC ATC TCC GCT TTC | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min; 72 °C, 1 min; 72 °C, 5 min | 306 bp | MCP | Newly designed | |
| hnMCP-specT342-R2 | CAC CGA CAC CTC CTC AAC C | |||||
| β−actin | ||||||
| β−actin−F | TAC CAC CGG TAT CGT CAT GGA | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 60 °C, 1 min; 72 °C, 1 min; 72 °C, 5 min | 150 bp | β−actin | [27] | |
| β−actin−R | CCA CGC TCT GTC AGG ATC TTC | |||||
| Cloning | JM-MCP | |||||
| JM-MCP-F | AGG TGC GAA CGT AAC CAG T | 94 °C, 5 min; 30 cycles: 95 °C, 30 s; 58 °C, 1 min 30 s; 72 °C, 1 min; 72 °C, 8 min | 1348 bp | MCP | Newly designed | |
| JM-MCP-R | TTA CAG GAT AGG GAA GCC TGC |
| Virus Name | Genus | MCP Accession No. | Country (origin) | Year | Host |
|---|---|---|---|---|---|
| Giant sea perch iridovirus (GSPIV/TW2/PT) | Megalocytivirus | MH237932 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW4/K) | Megalocytivirus | MH237933 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW5/PT) | Megalocytivirus | MH237934 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW7/PT) | Megalocytivirus | MH237935 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW8/K) | Megalocytivirus | MH237936 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW11/K) | Megalocytivirus | MH237937 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW17/PT) | Megalocytivirus | MH237938 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW22/PT) | Megalocytivirus | MH237939 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW25/PT) | Megalocytivirus | MH237940 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW29/PT) | Megalocytivirus | MH237941 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW30/PT) | Megalocytivirus | MH237942 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Giant sea perch iridovirus (GSPIV/TW35/PT) | Megalocytivirus | MH237943 | Taiwan | 2017 | Giant sea perch (Lates calcarifer) |
| Rock bream iridovirus (RBIV-HD) | Megalocytivirus | KT031401 | Korea | 2015 | Rock bream (Oplegnathus fasciatus) |
| Grouper iridovirus | Megalocytivirus | KT989778 | Taiwan | 2015 | Grouper (Epinephelus sp.) |
| Rock bream iridovirus (RBIV-C1) | Megalocytivirus | HQ105005 | China | 2009 | Rock bream (Oplegnathus fasciatus) |
| Red sea bream iridovirus (RIE12-1) | Megalocytivirus | AP017456 | Japan | 2016 | Red sea bream (Chrysophrys major) |
| Sea bass iridovirus | Megalocytivirus | AY310917 | China | 1993 | Sea bass (Lateolabrax sp) |
| Red sea bream iridovirus (RSIV-2) | Megalocytivirus | AB666329 | Japan | 1994 | Greater amberjack (Seriola dumerili) |
| Red sea bream iridovirus (3GG1) | Megalocytivirus | AB666326 | Hong Kong | 2004 | Orange-spotted grouper (Epinephelus coioides) |
| Red sea bream iridovirus (7GG) | Megalocytivirus | AB666322 | Hong Kong | 2004 | Orange-spotted grouper (Epinephelus coioides) |
| Red sea bream iridovirus (GIG42) | Megalocytivirus | AB666323 | Hong Kong | 2002 | Giant grouper (Epinephelus lanceolatus) |
| Spotted knifejaw iridovirus (SKIV-ZJ07) | Megalocytivirus | GQ202216 | China | 2009 | Spotted knifejaw (Oplegnathus punctatus) |
| King grouper iridovirus (KGIV-05) | Megalocytivirus | EU847414 | Taiwan | 2005 | Giant grouper (Epinephelus lanceolatus) |
| Barramundi perch iridovirus (BPIV-08) | Megalocytivirus | EU847418 | Taiwan | 2008 | Barramundi perch (Lates calcarifer) |
| Stone flounder iridovirus (724) | Megalocytivirus | HQ263620 | China | 2010 | Stone flounder (Kareius bicoloratus) |
| Rock bream iridovirus (SBIV-KOR-TY) | Megalocytivirus | AY532613 | Korea | 2000-2001 | Sea bass (Lateolabrax japonicus) |
| Infectious spleen and kidney necrosis virus (ISKNV-HT) | Megalocytivirus | HQ317464 | China | 2006 | Mandarin fish (Siniperca chuatsi) |
| Rock bream iridovirus (CNU-2) | Megalocytivirus | AY849394 | South Korea | 2004 | Rock bream (Oplegnathus fasciatus) |
| Olive flounder iridovirus | Megalocytivirus | DQ198145 | South Korea | 2005 | Bastard halibut (Paralichthys olivaceus) |
| Giant seaperch iridovirus (GSIV-K1) | Megalocytivirus | EU315313 | Taiwan | 2007 | Giant sea perch (Lates calcarifer) |
| Barramundi perch iridovirus (BPIV-05) | Megalocytivirus | EU847416 | Taiwan | 2005 | Barramundi perch (Lates calcarifer) |
| Red sea bream iridovirus | Megalocytivirus | AB109371 | Japan | 1994 | Red sea bream (Chrysophrys major) |
| Red sea bream iridovirus | Megalocytivirus | AY310918 | Japan/Thailand | 2008 | Red sea bream (Chrysophrys major) |
| Red sea bream iridovirus (6SB) | Megalocytivirus | AB666327 | Hong Kong | 2004 | Yellowfin sea bream (Acanthopagrus latus) |
| Red seabream iridovirus (HyoDS-13) | Megalocytivirus | LC150886 | Japan | 2016 | Devil stinger (Inimicus japonicus) |
| Sea bass iridovirus | Megalocytivirus | AY310917 | China | 1993 | Sea bass (Lateolabrax japonicus) |
| Grouper sleepy disease iridovirus | Megalocytivirus | AB109370 | Thailand | 1993 | Brown spotted grouper (Epinephelus malabaricus) |
| Grouper sleepy disease iridovirus | Megalocytivirus | AY285746 | Thailand | 1993 | Brown spotted grouper (Epinephelus malabaricus) |
| Red seabream iridovirus (TGA12) | Megalocytivirus | AB666319 | Singapore | 2000 | Brown-marbled grouper (Epinephelus fuscoguttatus) |
| Infectious spleen and kidney necrosis virus | Megalocytivirus | AB669096 | Southeast Asia/Japan | 2006 | Banggai cardinalfish (Pterapogon kauderni) |
| Banggai cardinalfish iridovirus | Megalocytivirus | HQ857787 | USA | 2011 | Banggai cardinalfish (Pterapogon kauderni) |
| Marble sleepy goby iridovirus | Megalocytivirus | HM067835 | China | 2009 | Marble goby (Oxyeleotris marmorata) |
| African lampeye iridovirus | Megalocytivirus | AY285745 | Indonesia | 2005 | African lampeye (Aplocheilichthys normani) |
| Megalocytivirus Sabah/RAA/2012 (HGIV67) | Megalocytivirus | JQ253369 | Malaysia | 2006 | Barramundi cod (Cromileptes altivelis) |
| African lampeye iridovirus | Megalocytivirus | AB109368 | Indonesia/Japan | 1998 | African lampeye (Aplocheilichthys normani) |
| Dwarf gourami iridovirus | Megalocytivirus | AY285744 | Thailand | 2002 | Dwarf gourami (Trichogaster lalius) |
| Giant sea perch iridovirus (GSIV_Pt_843_05) | Megalocytivirus | JF264354 | Taiwan | 2005 | Giant sea perch (Lates calcarifer) |
| Mullet iridovirus MA5/5 | Megalocytivirus | AB666343 | Singapore | 2000 | Fathead mullet (Mugil cephalus) |
| Dwarf gourami iridovirus | Megalocytivirus | AY989901 | Southeast Asia | 2004 | Dwarf gourami (Trichogaster lalius) |
| Infectious spleen and kidney necrosis virus (lapulapu) | Megalocytivirus | AB666339 | Philippines | 2000 | Orange-spotted grouper (Epinephelus coioides) |
| Megalocytivirus Sabah/RAA/2012 (OSGIV75) | Megalocytivirus | JQ253372 | Malaysia | 2011 | Orange-spotted grouper (Epinephelus coioides) |
| Murray cod iridovirus | Megalocytivirus | AY936203 | Australia | 2003 | Murray cod (Maccullochella peeli) |
| Infectious spleen and kidney necrosis virus (ISKNV-QY) | Megalocytivirus | HQ317460 | China | 2009 | Mandarin fish (Siniperca chuatsi) |
| Dwarf gourami iridovirus DGA4/6K | Megalocytivirus | AB666344 | Singapore | 2000 | Dwarf gourami (Trichogaster lalius) |
| Giant sea perch iridovirus (GSIV_Pt_836_05) | Megalocytivirus | JF264350 | Taiwan | 2005 | Giant sea perch (Lates calcarifer) |
| Sea perch iridovirus (CH-1) | Megalocytivirus | HM067603 | South Korea | 2010 | Sea perch (Lateolabrax sp.) |
| Turbot iridovirus isolate (R-603) | Megalocytivirus | HM596017 | China | 2009 | Turbot (Scophthalmus maximus) |
| Turbot reddish body iridovirus | Megalocytivirus | AY590687 | China | 2002 | Turbot (Scophthalmus maximus) |
| Korean flounder iridovirus (FLIV-MI) | Megalocytivirus | AY633982 | Korea | 2005 | Olive flounder (Paralichthys olivaceus) |
| Olive flounder iridovirus | Megalocytivirus | AY661546 | Korea | 2004 | Olive flounder (Paralichthys olivaceus) |
| Korean flounder iridovirus (FLIV-JJY) | Megalocytivirus | AY633990 | Korea | 2005 | Olive flounder (Paralichthys olivaceus) |
| Korean flounder iridovirus (FLIV-WD2) | Megalocytivirus | AY633985 | Korea | 2005 | Olive flounder (Paralichthys olivaceus) |
| Olive flounder iridovirus (OFLIV-1) | Megalocytivirus | EU276417 | Korea | 2007 | Olive flounder (Paralichthys olivaceus) |
| Turbot reddish body iridovirus (case3) | Megalocytivirus | KX354221 | USA | 1991 | Oscar (Astronotus ocellatus) |
| Rock bream iridovirus (RBIV_Tp_45_08) | Megalocytivirus | JF264352 | Taiwan | 2008 | Rock bream (Oplegnathus fasciatus) |
| Turbot reddish body iridovirus (case1) | Megalocytivirus | KX354223 | Canada/Southeast Asia | 1986 | Angelfish (Pterophyllum scalare) |
| Turbot reddish body iridovirus (case2) | Megalocytivirus | KX354222 | Australia/ Singapore | 1988 | Dwarf gourami (Trichogaster lalius) |
| Threespine stickleback iridovirus | Megalocytivirus | HQ857785 | Canada | 2012 | Threespine stickleback (Gasterosteus aculeatus) |
| Tiger frog virus | Ranavirus | AF389451 | China | 2002 | Tiger frog (Rana tigrina rugulosa) |
| Isolate | OIE Primer Sets a | Universal Primer Set b | Primer Sets Respectively Specific to c | hn-TRBIV d | |||
| 1-F/1-R | 4-F/4-R | RSIV/ISKNV/TRBIV | RSIV | ISKNV | TRBIV | ||
| GSPIV/TW1 | – | – | – | – | – | – | + |
| GSPIV/TW2 | – | – | + | – | – | + | + |
| GSPIV/TW3 | – | – | – | – | – | – | + |
| GSPIV/TW4 | – | – | + | – | – | + | + |
| GSPIV/TW5 | – | – | + | – | – | + | + |
| GSPIV/TW6 | – | – | – | – | – | – | + |
| GSPIV/TW7 | – | – | + | – | – | + | + |
| GSPIV/TW8 | – | – | + | – | – | + | + |
| GSPIV/TW9 | – | – | + | – | – | + | + |
| GSPIV/TW10 | – | – | – | – | – | – | + |
| GSPIV/TW11 | – | – | + | – | – | + | + |
| GSPIV/TW12 | – | – | – | – | – | – | – |
| GSPIV/TW13 | – | – | + | – | – | + | + |
| GSPIV/TW14 | – | – | – | – | – | – | + |
| GSPIV/TW15 | – | – | – | – | – | – | + |
| GSPIV/TW16 | – | – | – | – | – | – | + |
| GSPIV/TW17 | – | – | + | – | – | + | + |
| GSPIV/TW18 | – | – | – | – | – | – | + |
| GSPIV/TW19 | – | – | – | – | – | – | + |
| GSPIV/TW20 | – | – | – | – | – | – | + |
| GSPIV/TW21 | – | – | + | – | – | + | + |
| GSPIV/TW22 | – | – | + | – | – | + | + |
| GSPIV/TW23 | – | – | – | – | – | – | + |
| GSPIV/TW24 | – | – | – | – | – | – | + |
| GSPIV/TW25 | – | – | + | – | – | + | + |
| GSPIV/TW26 | – | – | + | – | – | + | + |
| GSPIV/TW27 | – | – | – | – | – | – | + |
| GSPIV/TW28 | – | – | + | – | – | + | + |
| GSPIV/TW29 | – | – | + | – | – | + | + |
| GSPIV/TW30 | – | – | + | – | – | + | + |
| GSPIV/TW31 | – | – | – | – | – | – | + |
| GSPIV/TW32 | – | – | – | – | – | – | + |
| GSPIV/TW33 | – | – | + | – | – | + | + |
| GSPIV/TW34 | – | – | + | – | – | + | + |
| GSPIV/TW35 | – | – | + | – | – | + | + |
| GSPIV/TW36 | – | – | + | – | + | – | – |
| GSPIV/TW37 | – | – | – | – | – | – | – |
| GSPIV/TW38 | – | – | – | – | – | – | + |
| GSPIV/TW39 | + | + | + | + | – | – | – |
| Virus Isolate a | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1. MH237932 | 100.00 | |||||||||||||||
| 2. MH237933 | 98.01 | 100.00 | ||||||||||||||
| 3. MH237934 | 99.85 | 98.01 | 100.00 | |||||||||||||
| 4. MH237935 | 97.86 | 99.85 | 97.86 | 100.00 | ||||||||||||
| 5. MH237936 | 97.78 | 99.77 | 97.78 | 99.62 | 100.00 | |||||||||||
| 6. MH237937 | 97.78 | 99.77 | 97.78 | 99.62 | 99.54 | 100.00 | ||||||||||
| 7. MH237938 | 98.01 | 100.00 | 98.01 | 99.85 | 99.77 | 99.77 | 100.00 | |||||||||
| 8. MH237939 | 97.71 | 99.69 | 97.71 | 99.54 | 99.46 | 99.46 | 99.69 | 100.00 | ||||||||
| 9. MH237940 | 97.86 | 99.85 | 97.86 | 99.69 | 99.62 | 99.62 | 99.85 | 99.54 | 100.00 | |||||||
| 10. MH237941 | 97.94 | 99.92 | 97.94 | 99.77 | 99.69 | 99.69 | 99.92 | 99.62 | 99.77 | 100.00 | ||||||
| 11. MH237942 | 97.86 | 99.85 | 97.86 | 99.69 | 99.62 | 99.62 | 99.85 | 99.54 | 99.69 | 99.77 | 100.00 | |||||
| 12. MH237943 | 97.86 | 99.85 | 97.86 | 99.69 | 99.62 | 99.62 | 99.85 | 99.54 | 99.69 | 99.77 | 99.69 | 100.00 | ||||
| 13. JF264352 | 99.85 | 98.01 | 99.85 | 97.86 | 97.78 | 97.78 | 98.01 | 97.71 | 97.86 | 97.94 | 97.86 | 97.86 | 100.00 | |||
| 14. KX354221 | 99.85 | 98.01 | 99.85 | 97.86 | 97.78 | 97.78 | 98.01 | 97.71 | 97.86 | 97.94 | 97.86 | 97.86 | 99.93 | 100.00 | ||
| 15. KX354222 | 99.85 | 98.01 | 99.85 | 97.86 | 97.78 | 97.78 | 98.01 | 97.71 | 97.86 | 97.94 | 97.86 | 97.86 | 99.93 | 100.00 | 100.00 | |
| 16. KX354223 | 99.85 | 98.01 | 99.85 | 97.86 | 97.78 | 97.78 | 98.01 | 97.71 | 97.86 | 97.94 | 97.86 | 97.86 | 99.93 | 100.00 | 100.00 | 100.00 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsai, J.-M.; Huang, S.-L.; Yang, C.-D. PCR Detection and Phylogenetic Analysis of Megalocytivirus Isolates in Farmed Giant Sea Perch Lates calcarifer in Southern Taiwan. Viruses 2020, 12, 681. https://0-doi-org.brum.beds.ac.uk/10.3390/v12060681
Tsai J-M, Huang S-L, Yang C-D. PCR Detection and Phylogenetic Analysis of Megalocytivirus Isolates in Farmed Giant Sea Perch Lates calcarifer in Southern Taiwan. Viruses. 2020; 12(6):681. https://0-doi-org.brum.beds.ac.uk/10.3390/v12060681
Chicago/Turabian StyleTsai, Jia-Ming, Song-Lang Huang, and Chung-Da Yang. 2020. "PCR Detection and Phylogenetic Analysis of Megalocytivirus Isolates in Farmed Giant Sea Perch Lates calcarifer in Southern Taiwan" Viruses 12, no. 6: 681. https://0-doi-org.brum.beds.ac.uk/10.3390/v12060681





