Structural Mapping of Mutations in Spike, RdRp and Orf3a Genes of SARS-CoV-2 in Influenza Like Illness (ILI) Patients
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethical Considerations
2.2. Study Specimens
2.3. Laboratory Investigations
2.4. Real-Time Reverse Transcription PCR for Detection of Influenza and SARS-CoV-2
2.5. Surveillance of Influenza in Saudi Arabia
2.6. Genome Amplification, Sequencing and Phylogenetic Analysis
2.7. Statistical Analysis
3. Results
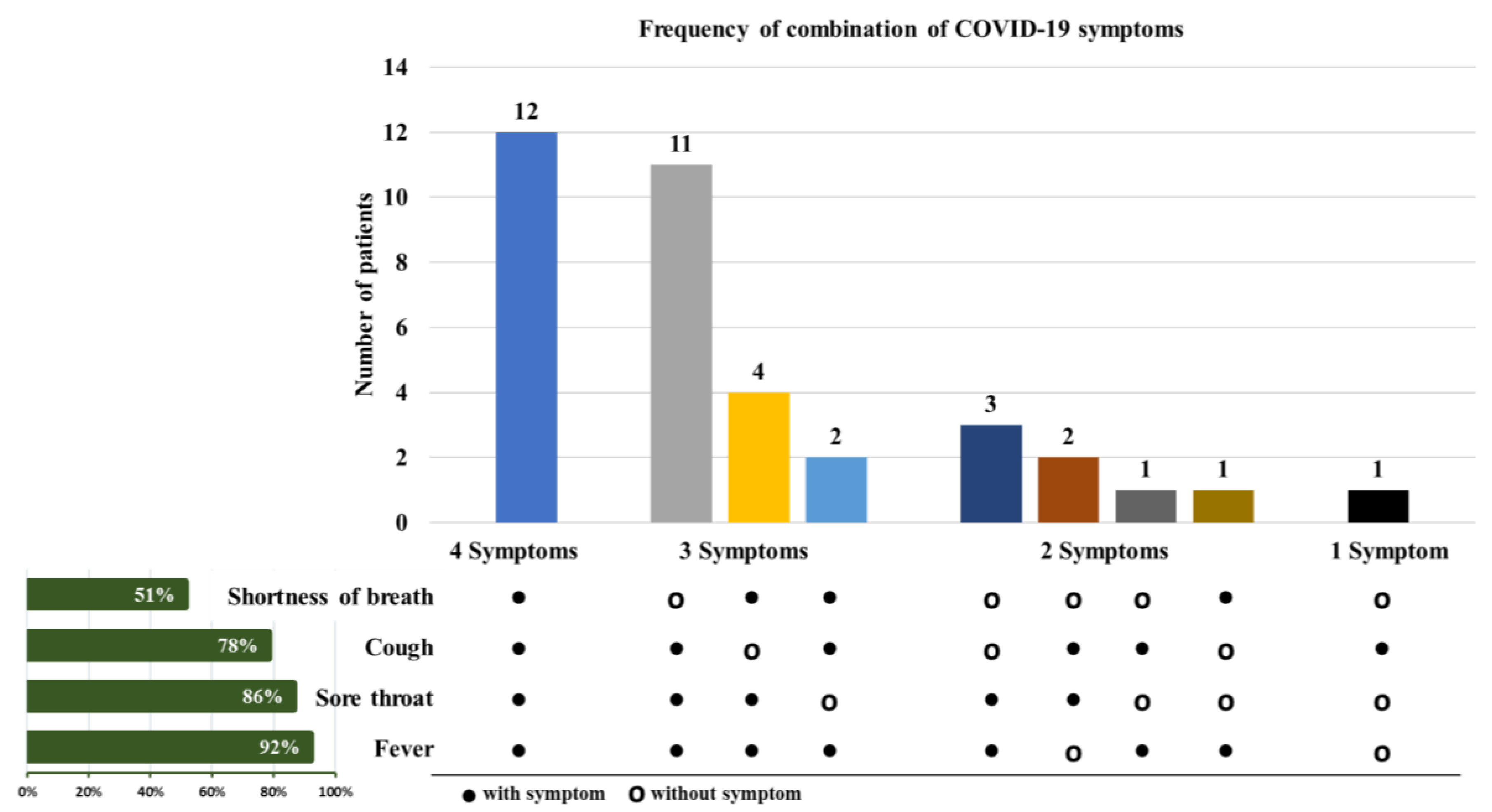
3.1. Shortness of Breath in ILI Patients
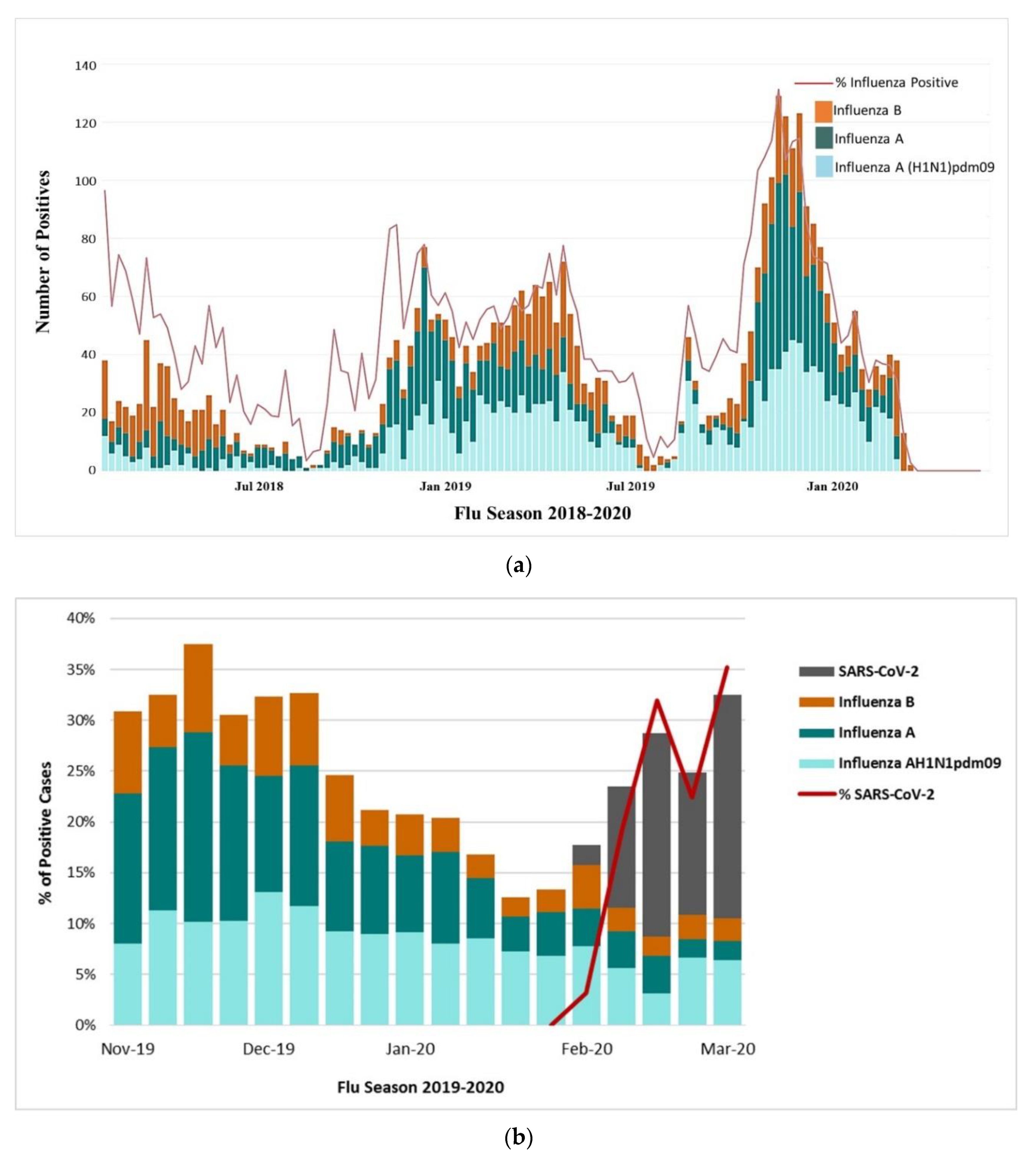
3.2. Surveillance of Influenza in Saudi Arabia
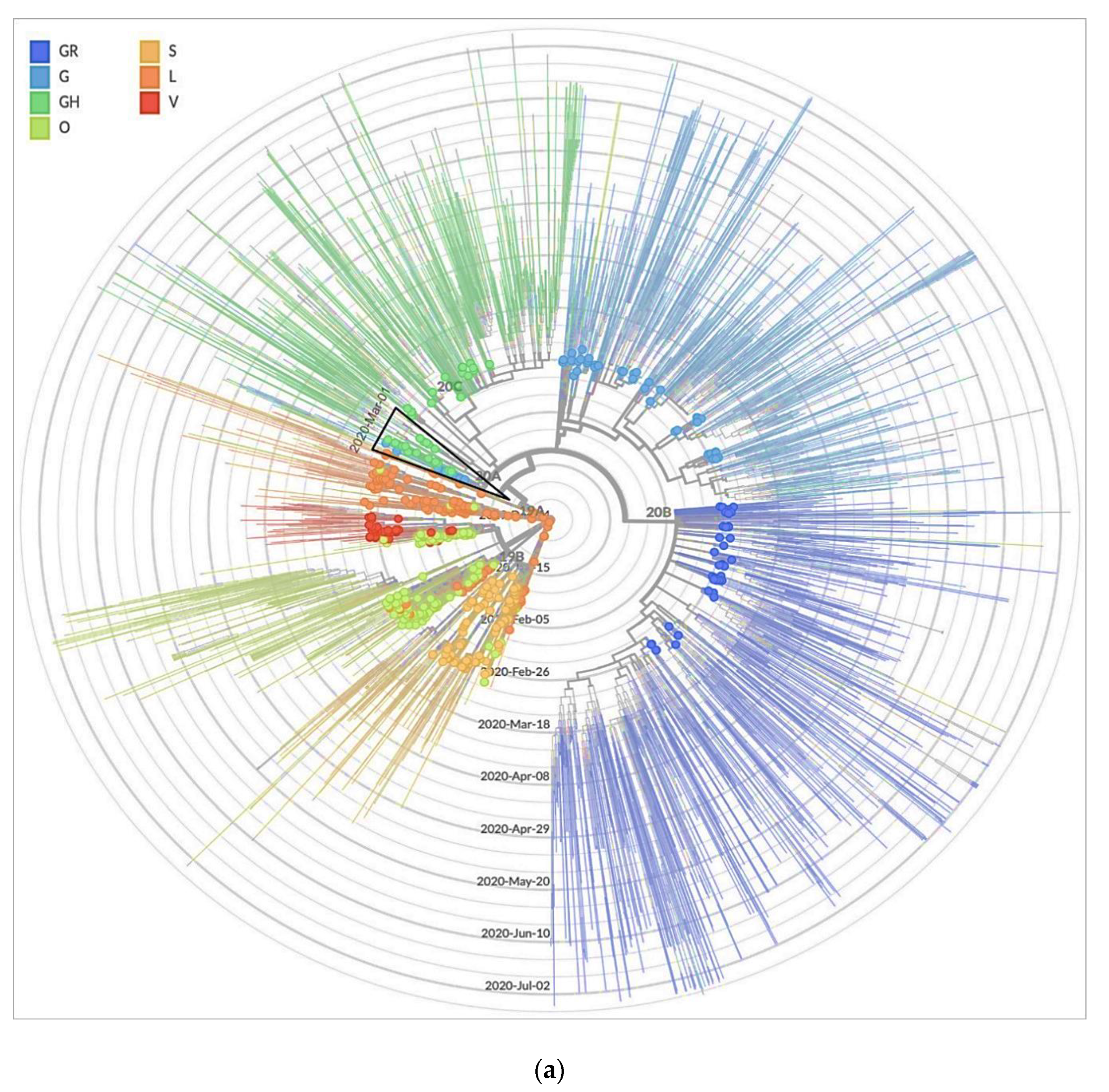
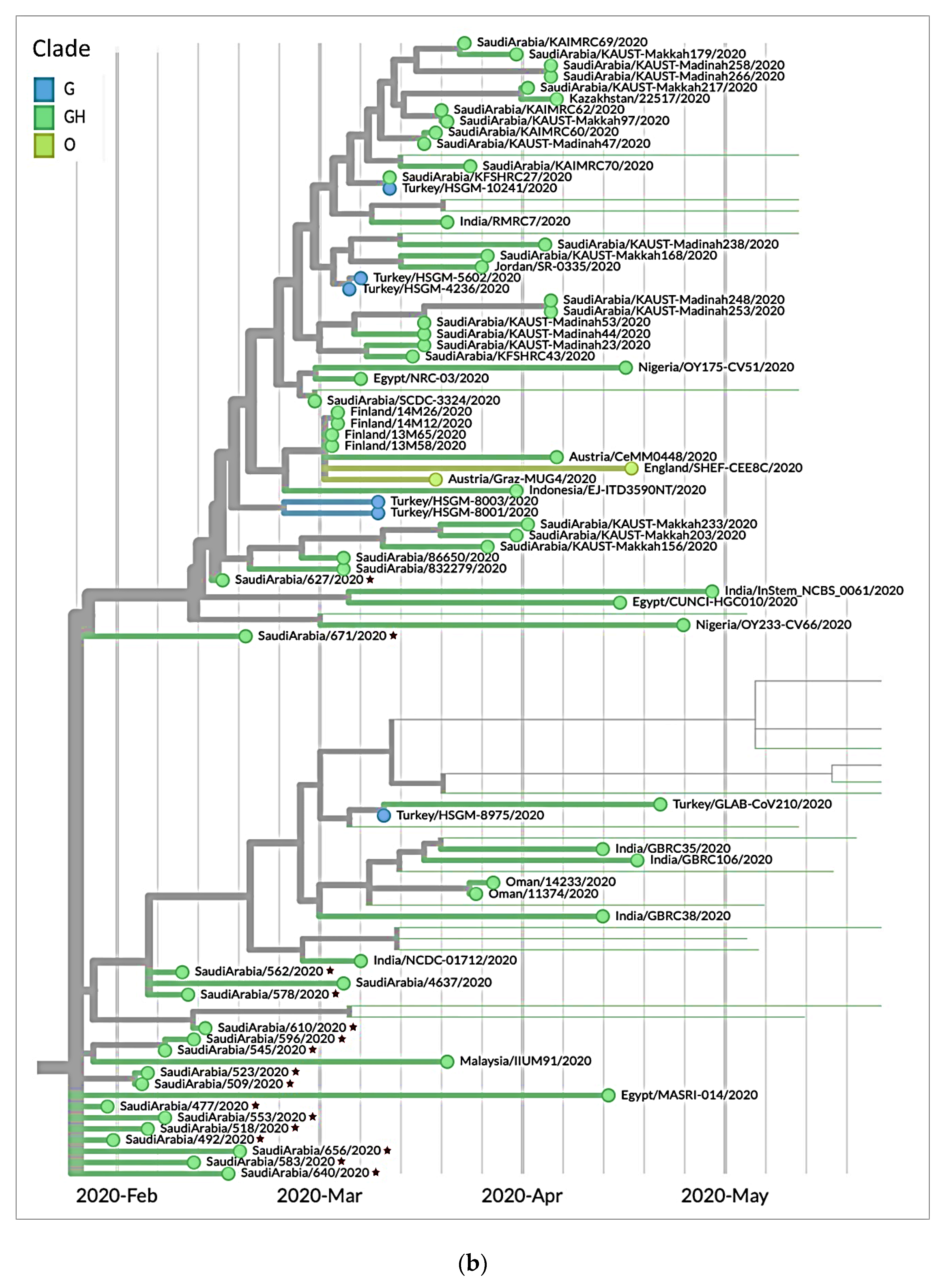
3.3. Genetic Sequencing of SARS-CoV-2 Isolates
3.4. Structural Predictions of the Proteins
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Coronavirus Disease 2019 (COVID-19): Situation Report, 33. Available online: https://apps.who.int/iris/handle/10665/331219 (accessed on 22 December 2020).
- Perlman, S. Another Decade, Another Coronavirus. N. Engl. J. Med. 2020, 382, 760–762. [Google Scholar] [CrossRef] [PubMed]
- Dong, E.; Du, H.; Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Inf. Dis. 2020, 20, 533–534. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.D.; Wang, Z.Y.; Zhang, S.F.; Li, X.; Li, L.; Li, C.; Cui, Y.; Fu, R.B.; Dong, Y.Z.; Chi, X.Y.; et al. Aerosol and Surface Distribution of Severe Acute Respiratory Syndrome Coronavirus 2 in Hospital Wards, Wuhan, China, 2020. Emerg. Infect. Dis. 2020, 26, 1583–1591. [Google Scholar] [CrossRef] [PubMed]
- Correia, G.; Rodrigues, L.; Gameiro da Silva, M.; Gonçalves, T. Airborne route and bad use of ventilation systems as non-negligible factors in SARS-CoV-2 transmission. Med. Hypotheses 2020, 141, 109781. [Google Scholar] [CrossRef]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus-Infected Pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef]
- Maugeri, A.; Barchitta, M.; Battiato, S.; Agodi, A. Estimation of Unreported Novel Coronavirus (SARS-CoV-2) Infections from Reported Deaths: A Susceptible-Exposed-Infectious-Recovered-Dead Model. J. Clin. Med. 2020, 9, 1350. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control. Novel Coronavirus Disease 2019 (COVID-19) Pandemic: Increased Transmission in the EU/EEA and the UK—Sixth update—Stockholm: ECDC. 2020. Available online: https://www.ecdc.europa.eu/sites/default/files/documents/RRA-sixth-update-Outbreak-of-novel-coronavirus-disease-2019-COVID-19.pdf (accessed on 22 December 2020).
- Kimball, A.; Hatfield, K.M.; Arons, M.; James, A.; Taylor, J.; Spicer, K.; Bardossy, A.C.; Oakley, L.P.; Tanwar, S.; Chisty, Z.; et al. Asymptomatic and Presymptomatic SARS-CoV-2 Infections in Residents of a Long-Term Care Skilled Nursing Facility—King County, Washington, March 2020. MMWR Morb. Mortal Wkly. Rep. 2020, 69, 377–381. [Google Scholar] [CrossRef] [Green Version]
- Li, R.; Pei, S.; Chen, B.; Song, Y.; Zhang, T.; Yang, W.; Shaman, J. Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2). Science 2020, 368, 489–493. [Google Scholar] [CrossRef] [Green Version]
- Wei, W.E.; Li, Z.; Chiew, C.J.; Yong, S.E.; Toh, M.P.; Lee, V.J. Presymptomatic Transmission of SARS-CoV-2—Singapore, 23 January–16 March 2020. MMWR Morb. Mortal Wkly. Rep. 2020, 69, 411–415. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Du, Z.; Wang, L.; Cauchemez, S.; Xu, X.; Wang, X.; Cowling, B.J.; Meyers, L.A. Risk for Transportation of Coronavirus Disease from Wuhan to Other Cities in China. Emerg. Infect. Dis. 2020, 26, 1049–1052. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oran, D.P.; Topol, E.J. Prevalence of Asymptomatic SARS-CoV-2 Infection: A Narrative Review. Ann. Intern. Med. 2020, 173, 362–367. [Google Scholar] [CrossRef]
- Zou, L.; Ruan, F.; Huang, M.; Liang, L.; Huang, H.; Hong, Z.; Yu, J.; Kang, M.; Song, Y.; Xia, J.; et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N. Engl. J. Med. 2020, 382, 1177–1179. [Google Scholar] [CrossRef] [PubMed]
- Perkins, T.A.; Cavany, S.M.; Moore, S.M.; Oidtman, R.J.; Lerch, A.; Poterek, M. Estimating unobserved SARS-CoV-2 infections in the United States. Proc. Natl. Acad. Sci. USA 2020, 117, 22597–22602. [Google Scholar] [CrossRef]
- Sutton, D.; Fuchs, K.; D’Alton, M.; Goffman, D. Universal Screening for SARS-CoV-2 in Women Admitted for Delivery. N. Engl. J. Med. 2020, 382, 2163–2164. [Google Scholar] [CrossRef]
- Gianino, M.M.; Politano, G.; Scarmozzino, A.; Stillo, M.; Amprino, V.; Di Carlo, S.; Benso, A.; Zotti, C.M. Cost of Sickness Absenteeism during Seasonal Influenza Outbreaks of Medium Intensity among Health Care Workers. Int. J. Environ. Res. Public Health 2019, 16, 747. [Google Scholar] [CrossRef] [Green Version]
- Musher, D.M. How contagious are common respiratory tract infections? N. Engl. J. Med. 2003, 348, 1256–1266. [Google Scholar] [CrossRef]
- World Health Organization. Coronavirus Disease 2019 (COVID-19): WHO COVID-19 Case Definition. 2020. Available online: https://www.who.int/publications/i/item/WHO-2019-nCoV-Surveillance_Case_Definition-2020.1 (accessed on 22 December 2020).
- Kong, W.H.; Li, Y.; Peng, M.W.; Kong, D.G.; Yang, X.B.; Wang, L.; Liu, M.Q. SARS-CoV-2 detection in patients with influenza-like illness. Nat. Microbiol. 2020, 5, 675–678. [Google Scholar] [CrossRef] [Green Version]
- Epidemiology Working Group for NCIP Epidemic Response; Chinese Center for Disease Control and Prevention. The epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19) in China. Zhonghua Liu Xing Bing Xue Za Zhi 2020, 41, 145–151. (In Chinese) [Google Scholar]
- WHO. Draft Landscape of COVID-19 Candidate Vaccines. Available online: https://www.who.int/publications/m/item/draft-landscape-of-covid-19-candidate-vaccines (accessed on 22 December 2020).
- Krammer, F. SARS-CoV-2 vaccines in development. Nature 2020, 586, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Laine, C.; Cotton, D.; Moyer, D.V. COVID-19 Vaccine: Promoting Vaccine Acceptance. Ann. Intern. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Gimferrer, L.; Andrés, C.; Rando, A.; Piñana, M.; Codina, M.G.; del Carmen Martin, M.; Fuentes, F.; Rubio, S.; Alcubilla, P.; Pumarola, T.; et al. Evaluation of Seegene Allplex Respiratory Panel 1 kit for the detection of influenza virus and human respiratory syncytial virus. J. Clin. Virol. 2018, 105, 31–34. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef] [Green Version]
- Abdalla, O.; Mohammed, M.; Hakawi, A.M.; Aljifri, A.; Abdalla, M.; Eltigani, S.; Mujib, S.A.; Assiri, A. Hospital-based surveillance of influenza A(H1N1)pdm09 virus in Saudi Arabia, 2010–2016. Ann. Saudi Med. 2020, 40, 1–6. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S. Fast gapped-read alignment with Bowtie 2. Nature Methods. 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The Sequence alignment/map (SAM) format and SAMtools. Bioinformatics. 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [Green Version]
- Rambaut, A.; Holmes, E.C.; O’Toole, Á.; Hill, V.; McCrone, J.T.; Ruis, C.; du Plessis, L.; Pybus, O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 2020. [Google Scholar] [CrossRef]
- Rozewicki, J.; Li, S.; Amada, K.M.; Standley, D.M.; Katoh, K. MAFFT-DASH: Integrated protein sequence and structural alignment. Nucleic Acids Res. 2019, 47, W5–W10. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization EMRO. C Influenza (Seasonal). 2020. Available online: http://www.emro.who.int/health-topics/influenza/influenza-updates.html (accessed on 16 May 2020).
- Hadfield, J.; Megill, C.; Bell, S.M.; Huddleston, J.; Potter, B.; Callender, C.; Sagulenko, P.; Bedford, T.; Neher, R.A. Nextstrain: Real-time tracking of pathogen evolution. Bioinformatics 2018, 34, 4121–4123. [Google Scholar] [CrossRef] [PubMed]
- Rhizobium, G.E. Complete Genome Sequence of the Sesbania Symbiont and Rice. Nucleic Acids Res. 2013, 1, 13–14. [Google Scholar] [CrossRef]
- Naeem, A.; Elbakkouri, K.; Alfaiz, A.; Hamed, M.E.; Alsaran, H.; AlOtaiby, S.; Enani, M.; Alosaimi, B. Antigenic drift of hemagglutinin and neuraminidase in seasonal H1N1 influenza viruses from Saudi Arabia in 2014 to 2015. J. Med. Virol. 2020, 11. [Google Scholar] [CrossRef]
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.Y.; Chen, L.; Wang, M. Presumed Asymptomatic Carrier Transmission of COVID-19. JAMA 2020, 323, 1406–1407. [Google Scholar] [CrossRef] [Green Version]
- Stringhini, S.; Wisniak, A.; Piumatti, G.; Azman, A.S.; Lauer, S.A.; Baysson, H.; De Ridder, D.; Petrovic, D.; Schrempft, S.; Marcus, K.; et al. Seroprevalence of anti-SARS-CoV-2 IgG antibodies in Geneva, Switzerland (SEROCoV-POP): A population-based study. Lancet 2020, 396, 313–319. [Google Scholar] [CrossRef]
- Rothe, C.; Schunk, M.; Sothmann, P.; Bretzel, G.; Froeschl, G.; Wallrauch, C.; Zimmer, T.; Thiel, V.; Janke, C.; Guggemos, W.; et al. Transmission of 2019-nCoV Infection from an Asymptomatic Contact in Germany. N. Engl. J. Med. 2020, 382, 970–971. [Google Scholar] [CrossRef] [Green Version]
- Sakurai, A.; Sasaki, T.; Kato, S.; Hayashi, M.; Tsuzuki, S.I.; Ishihara, T.; Iwata, M.; Morise, Z.; Doi, Y. Natural History of Asymptomatic SARS-CoV-2 Infection. N. Engl. J. Med. 2020, 383, 885–886. [Google Scholar] [CrossRef]
- Sood, N.; Simon, P.; Ebner, P.; Eichner, D.; Reynolds, J.; Bendavid, E.; Bhattacharya, J. Seroprevalence of SARS-CoV-2-Specific Antibodies Among Adults in Los Angeles County, California, on 10–11 April 2020. JAMA 2020, 323, 2425–2427. [Google Scholar] [CrossRef]
- Pollán, M.; Pérez-Gómez, B.; Pastor-Barriuso, R.; Oteo, J.; Hernán, M.A.; Pérez-Olmeda, M.; Sanmartín, J.L.; Fernández-García, A.; Cruz, I.; de Larrea, N.F.; et al. Prevalence of SARS-CoV-2 in Spain (ENE-COVID): A nationwide, population-based seroepidemiological study. Lancet 2020, 396, 535–544. [Google Scholar] [CrossRef]
- Wu, Z.; McGoogan, J.M. Characteristics of and Important Lessons from the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72,314 Cases from the Chinese Center for Disease Control and Prevention. JAMA 2020, 323, 1239–1242. [Google Scholar] [CrossRef] [PubMed]
- Uyoga, S.; Adetifa, I.; Karanja, H.; Nyagwange, J.; Tuju, J.; Wanjiku, P.; Aman, R.; Mwangangi, M.; Amoth, P.; Kasera, K.; et al. Seroprevalence of Anti-SARS-CoV-2 IgG Antibodies in Kenyan Blood Donors. Science 2021, 371, 79–82. [Google Scholar] [CrossRef] [PubMed]
- Spellberg, B.; Haddix, M.; Lee, R.; Butler-Wu, S.; Holtom, P.; Yee, H.; Gounder, P. Community Prevalence of SARS-CoV-2 Among Patients with Influenza like Illnesses Presenting to a Los Angeles Medical Center in March 2020. JAMA 2020, 323, 1966–1967. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Delaney, M.; Shah, R.K.; Campos, J.M.; Wessel, D.L.; DeBiasi, R.L. Comparison of Clinical Features of COVID-19 vs Seasonal Influenza A and B in US Children. JAMA Netw. Open 2020, 3, e2020495. [Google Scholar] [CrossRef] [PubMed]
- Cuadrado-Payán, E.; Montagud-Marrahi, E.; Torres-Elorza, M.; Bodro, M.; Blasco, M.; Poch, E.; Soriano, A.; Piñeiro, G.J. SARS-CoV-2 and influenza virus co-infection. Lancet 2020, 395, e84. [Google Scholar] [CrossRef]
- Lessler, J.; Salje, H.; Van Kerkhove, M.D.; Ferguson, N.M.; Cauchemez, S.; Rodriquez-Barraquer, I.; Hakeem, R.; Jombart, T.; Aguas, R.; Al-Barrak, A.; et al. Estimating the Severity and Subclinical Burden of Middle East Respiratory Syndrome Coronavirus Infection in the Kingdom of Saudi Arabia. Am. J. Epidemiol. 2016, 183, 657–663. [Google Scholar] [CrossRef] [Green Version]
- Yu, X.; Wei, D.; Chen, Y.; Zhang, D.; Zhang, X. Retrospective detection of SARS-CoV-2 in hospitalized patients with influenza-like illness. Emerg. Microbes Infect. 2020, 9, 1470–1473. [Google Scholar] [CrossRef]
- Nakagawa, S.; Miyazawa, T. Genome evolution of SARS-CoV-2 and its virological characteristics. Inflamm. Regen. 2020, 40, 17. [Google Scholar] [CrossRef]
- Gámbaro, F.; Behillil, S.; Baidaliuk, A.; Donati, F.; Albert, M.; Alexandru, A.; Vanpeene, M.; Bizard, M.; Brisebarre, A.; Barbet, M.; et al. Introductions and early spread of SARS-CoV-2 in France, 24 January to 23 March 2020. Eurosurveillance 2020, 25, 2001200. [Google Scholar] [CrossRef]
- Deslandes, A.; Berti, V.; Tandjaoui-Lambotte, Y.; Alloui, C.; Carbonnelle, E.; Zahar, J.R.; Brichler, S.; Cohen, Y. SARS-CoV-2 was already spreading in France in late December 2019. Int. J. Antimicrob. Agents 2020, 55, 106006. [Google Scholar] [CrossRef]
- Liu, M.; Gu, C.; Wu, J.; Zhu, Y. Amino acids 1 to 422 of the spike protein of SARS associated coronavirus are required for induction of cyclooxygenase-2. Virus Genes 2006, 33, 309–317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell 2020, 182, 812–827.e19. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wu, J.; Nie, J.; Zhang, L.; Hao, H.; Liu, S.; Zhao, C.; Zhang, Q.; Liu, H.; Nie, L.; et al. The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity. Cell 2020, 182, 1284–1294.e9. [Google Scholar] [CrossRef] [PubMed]
- Kannan, S.R.; Spratt, A.N.; Quinn, T.P.; Heng, X.; Lorson, C.L.; Sönnerborg, A.; Byrareddy, S.N.; Singh, K. Infectivity of SARS-CoV-2: There Is Something More than D614G? J. Neuroimmune. Pharmacol. 2020, 15, 574–577. [Google Scholar] [CrossRef]
- Julias, J.G.; Boyer, P.L.; McWilliams, M.J.; Alvord, W.G.; Hughes, S.H. Mutations at position 184 of human immunodeficiency virus type-1 reverse transcriptase affect virus titer and viral DNA synthesis. Virology 2004, 322, 13–21. [Google Scholar] [CrossRef] [Green Version]
- Fan, N.; Evans, D.B.; Rank, K.B.; Thomas, R.C.; Tarpley, W.G.; Sharma, S.K. Mechanism of resistance to U-90152S and sensitization to L-697,661 by a proline to leucine change at residue 236 of human immunodeficiency virus type 1 (HIV-1) reverse transcriptase. FEBS Lett. 1995, 359, 233–238. [Google Scholar] [CrossRef] [Green Version]
- Nelson, C.W.; Ardern, Z.; Goldberg, T.L.; Meng, C.; Kuo, C.H.; Ludwig, C.; Kolokotronis, S.O.; Wei, X. Dynamically evolving novel overlapping gene as a factor in the SARS-CoV-2 pandemic. eLife 2020, 9, e59633. [Google Scholar] [CrossRef]

| Baseline Variables | All Patients | SARS-CoV-2 Pos | SARS-CoV-2 Neg | p-Value |
|---|---|---|---|---|
| n = 739 | n = 37 (5%) | n = 702 (95%) | ||
| Characteristics | ||||
| Median Age | 33.4 ± 8.5 | 33.9 ± 9.6 | 33.4 ± 8.5 | |
| Gender | ||||
| Men | 242 (33%) | 8 (3.3%) | 234 (96.6%) | p = 0.13 * |
| Women | 497 (67%) | 29 (5.8%) | 468 (94.1%) | |
| Flu Vaccination | ||||
| Vaccinated | 470 (64%) | 25 (68%) | 445 (63%) | p = 0.60 * |
| Non-vaccinated | 269 (36%) | 12 (32%) | 257 (67%) | |
| SARS-CoV-2 Positives (n = 37) | |||||
|---|---|---|---|---|---|
| Vaccinated | Non-Vaccinated | ||||
| Symptoms | With | Without | With | Without | |
| Fever | 22 (59%) | 3 (8%) | 11 (30%) | 1 (3%) | p = 0.73 |
| Sore throat | 22 (59%) | 3 (8%) | 10 (27%) | 2 (6%) | p = 0.69 |
| Cough | 20 (54%) | 5 (14%) | 9 (24%) | 3 (8%) | p = 0.72 |
| Shortness of breath | 10 (27%) | 15 (41%) | 9 (24%) | 3 (8%) | p = 0.04 * |
| Co-infection | 7 (19%) | 18 (49%) | 3 (8%) | 9 (24%) | p = 0.84 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alosaimi, B.; Naeem, A.; Alghoribi, M.F.; Okdah, L.; Hamed, M.E.; AlYami, A.S.; Alotaibi, A.; Enani, M. Structural Mapping of Mutations in Spike, RdRp and Orf3a Genes of SARS-CoV-2 in Influenza Like Illness (ILI) Patients. Viruses 2021, 13, 136. https://0-doi-org.brum.beds.ac.uk/10.3390/v13010136
Alosaimi B, Naeem A, Alghoribi MF, Okdah L, Hamed ME, AlYami AS, Alotaibi A, Enani M. Structural Mapping of Mutations in Spike, RdRp and Orf3a Genes of SARS-CoV-2 in Influenza Like Illness (ILI) Patients. Viruses. 2021; 13(1):136. https://0-doi-org.brum.beds.ac.uk/10.3390/v13010136
Chicago/Turabian StyleAlosaimi, Bandar, Asif Naeem, Majed F. Alghoribi, Lilian Okdah, Maaweya E. Hamed, Ahmad S. AlYami, Athari Alotaibi, and Mushira Enani. 2021. "Structural Mapping of Mutations in Spike, RdRp and Orf3a Genes of SARS-CoV-2 in Influenza Like Illness (ILI) Patients" Viruses 13, no. 1: 136. https://0-doi-org.brum.beds.ac.uk/10.3390/v13010136






