Impact of the Lipopolysaccharide Chemotype of Salmonella Enterica Serovar Typhimurium on Virulence in Gnotobiotic Piglets
Abstract
:1. Introduction
2. Results
2.1. Clinical Signs
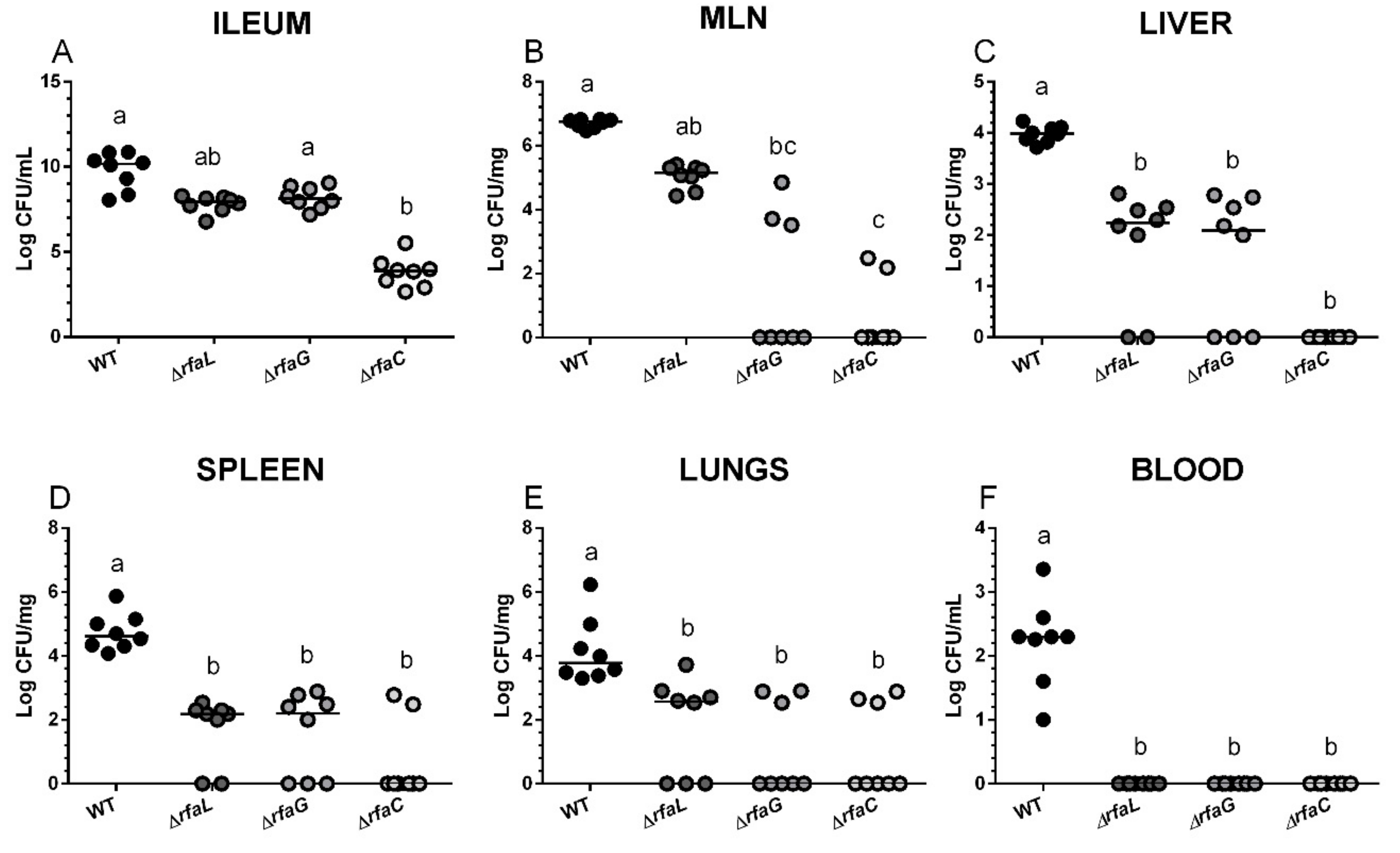
2.2. Salmonella Colonization of the Small Intestine and Its Translocation
2.3. Histopathological Changes in the Ileum
2.4. Expression of Tight Junction Proteins Claudin-1, Claudin-2, and Occludin in the Ileum
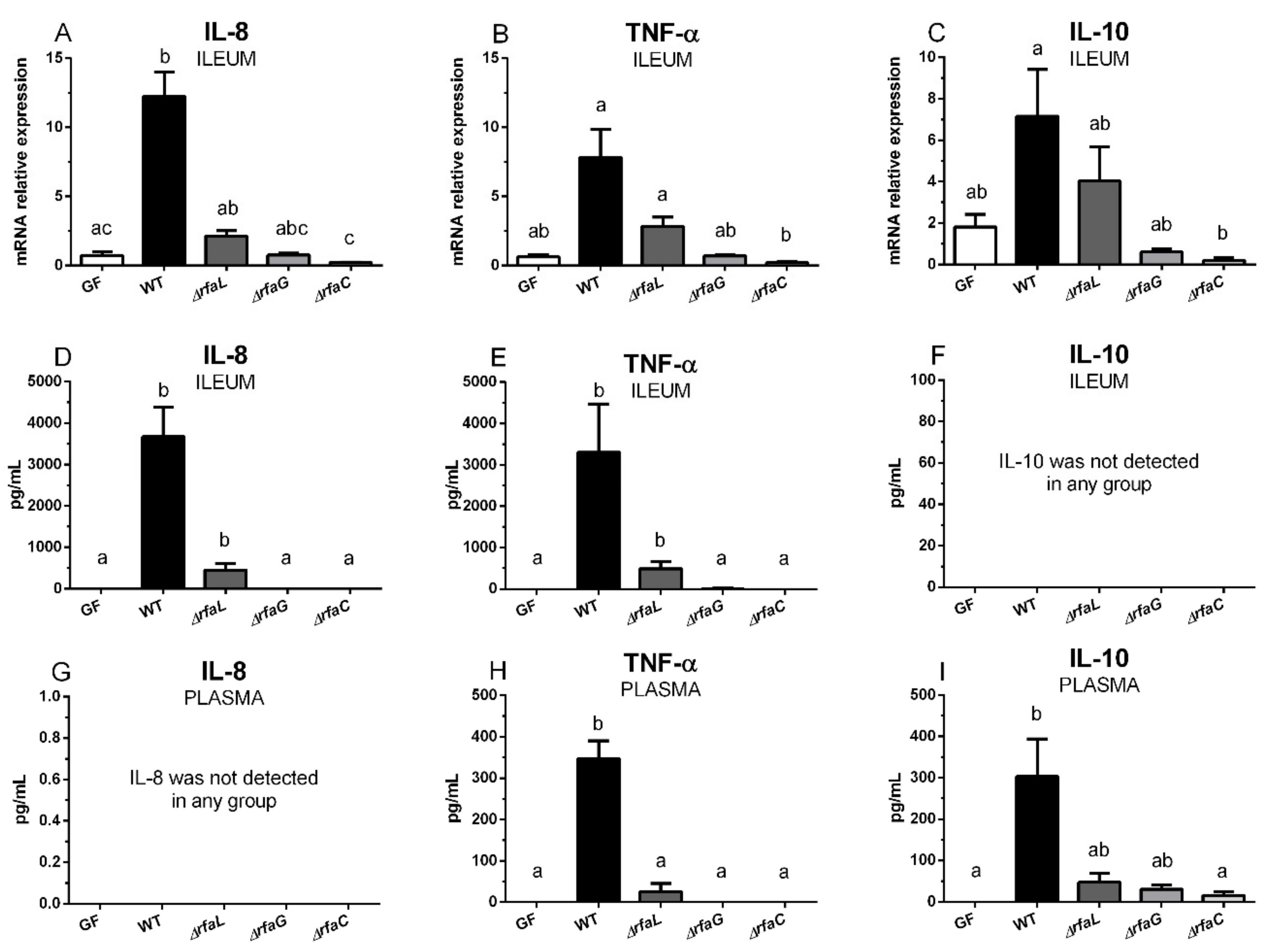
2.5. Transcriptions of IL-8, TNF-α, and IL-10 in the Ileum, and Their Local and Systemic Levels
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Ethical Statement
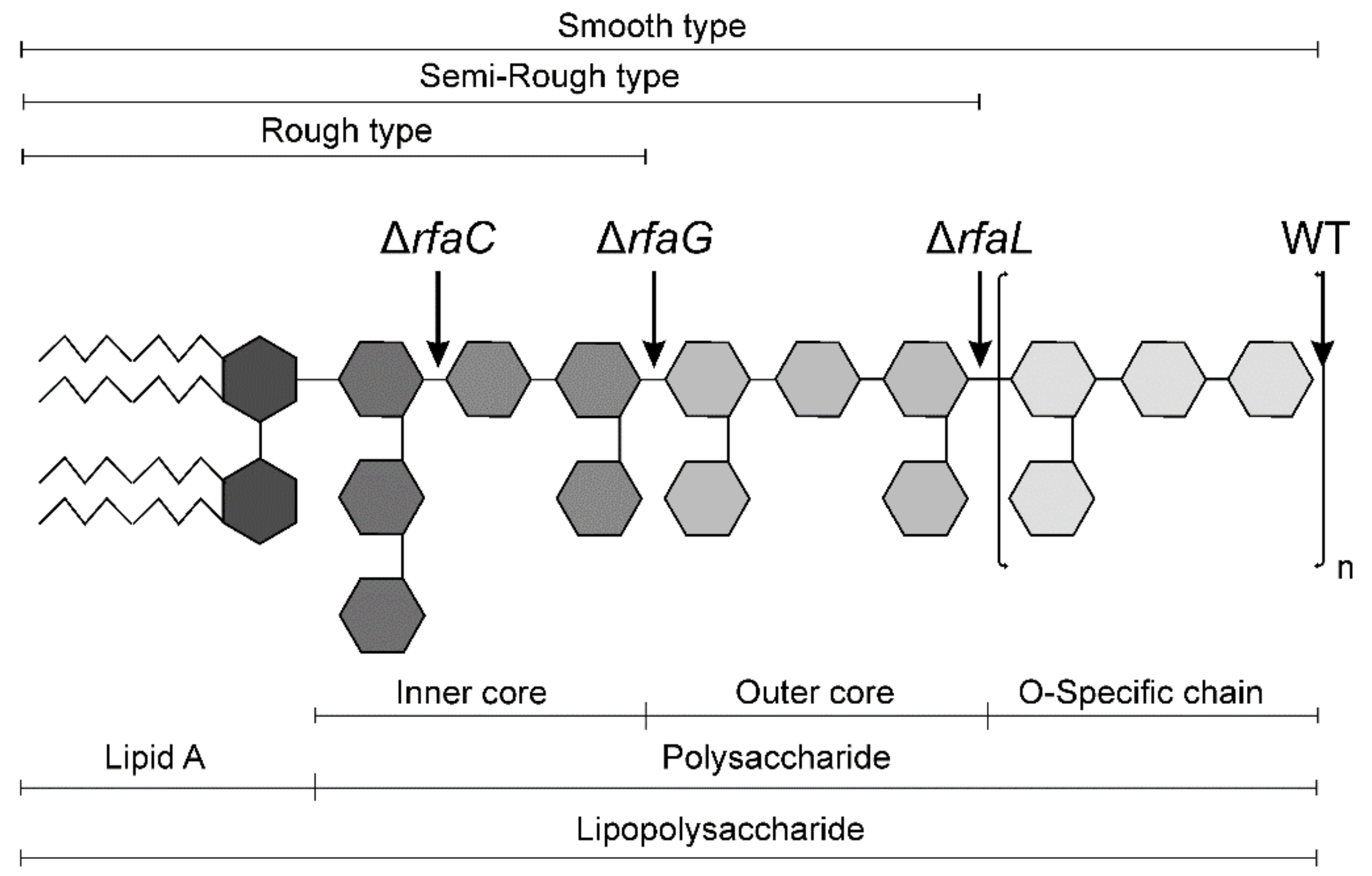
5.2. Salmonella Typhimurium LT2 Strain and Its Isogenic Δrfa Mutants
5.3. Gnotobiotic Piglets
5.4. Colonization/Infection of the Germ-Free Piglets
5.5. Clinical Signs
5.6. Bacterial Colonization and Translocation
5.7. Blood Plasma and Intestinal Lavages
5.8. Histologic Assessment
5.9. Isolation of Total RNA and Reverse Transcription
5.10. Real-Time PCR
5.11. ELISA
5.12. Statistical Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
References
- Hurley, D.; McCusker, M.P.; Fanning, S.; Martins, M. Salmonella-host interactions—Modulation of the host innate immune system. Front. Immunol. 2014, 5, 481. [Google Scholar] [CrossRef] [PubMed]
- Haraga, A.; Ohlson, M.B.; Miller, S.I. Salmonellae interplay with host cells. Nat. Rev. Microbiol. 2008, 6, 53–66. [Google Scholar] [CrossRef] [PubMed]
- Robarge, M.E.; Ramos-Vara, J.A. Pathology in practice. Enteric salmonellosis. J. Am. Vet. Med. Assoc. 2014, 245, 643–645. [Google Scholar] [CrossRef] [PubMed]
- Keestra-Gounder, A.M.; Tsolis, R.M.; Bäumler, A.J. Now you see me, now you don’t: The interaction of Salmonella with innate immune receptors. Nat. Rev. Microbiol. 2015, 13, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Santos, R.L.; Tsolis, R.M.; Bäumler, A.J.; Adams, L.G. Pathogenesis of Salmonella-induced enteritis. Braz. J. Med. Biol. Res. 2003, 36, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Nikaido, H. Prevention of drug access to bacterial targets: Permeability barriers and active efflux. Science 1994, 264, 382–388. [Google Scholar] [CrossRef] [PubMed]
- Raetz, C.R.; Whitfield, C. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 2002, 71, 635–700. [Google Scholar] [CrossRef]
- Beveridge, T.J. Structures of gram-negative cell walls and their derived membrane vesicles. J. Bacteriol. 1999, 181, 4725–4733. [Google Scholar]
- Mikolajczyk, A.; Zlotkowska, D. Subclinical lipopolysaccharide from Salmonella Enteritidis induces dysregulation of bioactive substances from selected brain sections and glands of neuroendocrine axes. Toxins 2019, 11, 91. [Google Scholar] [CrossRef]
- Wang, H.; Ma, S. The cytokine storm and factors determining the sequence and severity of organ dysfunction in multiple organ dysfunction syndrome. Am. J. Emerg. Med. 2008, 26, 711–715. [Google Scholar] [CrossRef]
- Caroff, M.; Karibian, D. Structure of bacterial lipopolysaccharides. Carbohydr. Res. 2003, 338, 2431–2447. [Google Scholar] [CrossRef] [PubMed]
- Kong, Q.; Yang, J.; Liu, Q.; Alamuri, P.; Roland, K.L.; Curtiss, R., III. Effect of deletion of genes involved in lipopolysaccharide core and O-antigen synthesis on virulence and immunogenicity of Salmonella enterica serovar Typhimurium. Infect. Immun. 2011, 79, 4227–4239. [Google Scholar] [CrossRef] [PubMed]
- Leyman, B.; Boyen, F.; Van, P.A.; Verbrugghe, E.; Haesebrouck, F.; Pasmans, F. Salmonella Typhimurium LPS mutations for use in vaccines allowing differentiation of infected and vaccinated pigs. Vaccine 2011, 29, 3679–3685. [Google Scholar] [CrossRef] [PubMed]
- Foster, N.; Lovell, M.A.; Marston, K.L.; Hulme, S.D.; Frost, A.J.; Bland, P.; Barrow, P.A. Rapid protection of gnotobiotic pigs against experimental salmonellosis following induction of polymorphonuclear leukocytes by avirulent Salmonella enterica. Infect. Immun. 2003, 71, 2182–2191. [Google Scholar] [CrossRef] [PubMed]
- Splichal, I.; Trebichavsky, I.; Splichalova, A.; Barrow, P.A. Protection of gnotobiotic pigs against Salmonella enterica serotype Typhimurium by rough mutant of the same serotype is accompanied by the change of local and systemic cytokine response. Vet. Immunol. Immunopathol. 2005, 103, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Crhanova, M.; Malcova, M.; Mazgajova, M.; Karasova, D.; Sebkova, A.; Fucikova, A.; Bortlicek, Z.; Pilousova, L.; Kyrova, K.; Dekanova, M.; et al. LPS structure influences protein secretion in Salmonella enterica. Vet. Microbiol. 2011, 152, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Karasova, D.; Sebkova, A.; Vrbas, V.; Havlickova, H.; Sisak, F.; Rychlik, I. Comparative analysis of Salmonella enterica serovar Enteritidis mutants with a vaccine potential. Vaccine 2009, 27, 5265–5270. [Google Scholar] [CrossRef]
- Matiasovic, J.; Stepanova, H.; Volf, J.; Kubala, L.; Ovesna, P.; Rychlik, I.; Faldyna, M. Influence of the lipopolysaccharide structure of Salmonella enterica serovar Enteritidis on interactions with pig neutrophils. Vet. Microbiol. 2011, 150, 167–172. [Google Scholar] [CrossRef]
- Stepanova, H.; Volf, J.; Malcova, M.; Matiasovic, J.; Faldyna, M.; Rychlik, I. Association of attenuated mutants of Salmonella enterica serovar Enteritidis with porcine peripheral blood leukocytes. FEMS Microbiol. Lett. 2011, 321, 37–42. [Google Scholar] [CrossRef]
- Lunney, J.K. Advances in swine biomedical model genomics. Int. J. Biol. Sci. 2007, 3, 179–184. [Google Scholar] [CrossRef]
- Meurens, F.; Summerfield, A.; Nauwynck, H.; Saif, L.; Gerdts, V. The pig: A model for human infectious diseases. Trends Microbiol. 2012, 20, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Goldfarb, R.D.; Dellinger, R.P.; Parrillo, J.E. Porcine models of severe sepsis: Emphasis on porcine peritonitis. Shock 2005, 24 (Suppl. S1), 75–81. [Google Scholar] [CrossRef] [PubMed]
- Michie, H.R. The value of animal models in the development of new drugs for the treatment of the sepsis syndrome. J. Antimicrob. Chemother. 1998, 41 (Suppl. S1), 47–49. [Google Scholar] [CrossRef] [PubMed]
- Stecher, B.; Hardt, W.D. Mechanisms controlling pathogen colonization of the gut. Curr. Opin. Microbiol. 2011, 14, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Tremaroli, V.; Backhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 2012, 489, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Mooser, C.; de Agüero, M.G.; Ganal-Vonarburg, S.C. Standardization in host-microbiota interaction studies: Challenges, gnotobiology as a tool, and perspective. Curr. Opin. Microbiol. 2018, 44, 50–60. [Google Scholar] [CrossRef]
- Salmon, H.; Berri, M.; Gerdts, V.; Meurens, F. Humoral and cellular factors of maternal immunity in swine. Dev. Comp. Immunol. 2009, 33, 384–393. [Google Scholar] [CrossRef]
- Splichalova, A.; Slavikova, V.; Splichalova, Z.; Splichal, I. Preterm life in sterile conditions: A study on preterm, germ-free piglets. Front. Immunol. 2018, 9, 220. [Google Scholar] [CrossRef]
- McClelland, M.; Sanderson, K.E.; Spieth, J.; Clifton, S.W.; Latreille, P.; Courtney, L.; Porwollik, S.; Ali, J.; Dante, M.; Du, F.; et al. Complete genome sequence of Salmonella enterica serovar Typhimurium LT2. Nature 2001, 413, 852–856. [Google Scholar] [CrossRef]
- Clarke, R.C.; Gyles, C.L. Virulence of wild and mutant strains of Salmonella typhimurium in ligated intestinal segments of calves, pigs, and rabbits. Am. J. Vet. Res. 1987, 48, 504–510. [Google Scholar]
- Trebichavsky, I.; Splichalova, A.; Rychlik, I.; Hojna, H.; Muneta, Y.; Mori, Y.; Splichal, I. Attenuated aroA Salmonella enterica serovar Typhimurium does not induce inflammatory response and early protection of gnotobiotic pigs against parental virulent LT2 strain. Vaccine 2006, 24, 4285–4289. [Google Scholar] [CrossRef] [PubMed]
- Barthel, M.; Hapfelmeier, S.; Quintanilla-Martinez, L.; Kremer, M.; Rohde, M.; Hogardt, M.; Pfeffer, K.; Russmann, H.; Hardt, W.D. Pretreatment of mice with streptomycin provides a Salmonella enterica serovar Typhimurium colitis model that allows analysis of both pathogen and host. Infect. Immun. 2003, 71, 2839–2858. [Google Scholar] [CrossRef] [PubMed]
- Dlabac, V.; Trebichavsky, I.; Rehakova, Z.; Hofmanova, B.; Splichal, I.; Cukrowska, B. Pathogenicity and protective effect of rough mutants of Salmonella species in germ-free piglets. Infect. Immun. 1997, 65, 5238–5243. [Google Scholar] [PubMed]
- Galen, J.E.; Buskirk, A.D.; Tennant, S.M.; Pasetti, M.F. Live Attenuated Human Salmonella Vaccine Candidates: Tracking the Pathogen in Natural Infection and Stimulation of Host Immunity. EcoSal Plus 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Griffiths, G.; Repnik, U.; Hornef, M. Seeing is understanding: Salmonella’s way to penetrate the intestinal epithelium. Int. J. Med. Microbiol. 2018, 308, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Gunzel, D.; Fromm, M. Claudins and other tight junction proteins. Compr. Physiol. 2012, 2, 1819–1852. [Google Scholar] [CrossRef]
- Liu, X.; Xia, B.; He, T.; Li, D.; Su, J.H.; Guo, L.; Wang, J.F.; Zhu, Y.H. Oral administration of a select mixture of Lactobacillus and Bacillus alleviates inflammation and maintains mucosal barrier integrity in the ileum of pigs challenged with Salmonella Infantis. Microorganisms 2019, 7, 135. [Google Scholar] [CrossRef]
- Edelblum, K.L.; Shen, L.; Weber, C.R.; Marchiando, A.M.; Clay, B.S.; Wang, Y.; Prinz, I.; Malissen, B.; Sperling, A.I.; Turner, J.R. Dynamic migration of gammadelta intraepithelial lymphocytes requires occludin. Proc. Natl. Acad. Sci. USA 2012, 109, 7097–7102. [Google Scholar] [CrossRef]
- Kohler, H.; Sakaguchi, T.; Hurley, B.P.; Kase, B.A.; Reinecker, H.C.; McCormick, B.A. Salmonella enterica serovar Typhimurium regulates intercellular junction proteins and facilitates transepithelial neutrophil and bacterial passage. Am. J. Physiol. Gastrointest. Liver Physiol. 2007, 293, G178–G187. [Google Scholar] [CrossRef]
- Loetscher, Y.; Wieser, A.; Lengefeld, J.; Kaiser, P.; Schubert, S.; Heikenwalder, M.; Hardt, W.D.; Stecher, B. Salmonella transiently reside in luminal neutrophils in the inflamed gut. PLoS ONE 2012, 7, e34812. [Google Scholar] [CrossRef]
- Splichalova, A.; Jenistova, V.; Splichalova, Z.; Splichal, I. Colonization of preterm gnotobiotic piglets with probiotic Lactobacillus rhamnosus GG and its interference with Salmonella Typhimurium. Clin. Exp. Immunol. 2019, 195, 381–394. [Google Scholar] [CrossRef] [PubMed]
- Eichner, M.; Protze, J.; Piontek, A.; Krause, G.; Piontek, J. Targeting and alteration of tight junctions by bacteria and their virulence factors such as Clostridium perfringens enterotoxin. Pflugers Arch. 2017, 469, 77–90. [Google Scholar] [CrossRef] [PubMed]
- Splichal, I.; Splichalova, A. Experimental enteric bacterial infections in pigs. J. Infect. Dis. 2018, 218, 504–505. [Google Scholar] [CrossRef] [PubMed]
- Baggiolini, M.; Walz, A.; Kunkel, S.L. Neutrophil-activating peptide-1/interleukin 8, a novel cytokine that activates neutrophils. J. Clin. Investig. 1989, 84, 1045–1049. [Google Scholar] [CrossRef] [PubMed]
- Sedger, L.M.; McDermott, M.F. TNF and TNF-receptors: From mediators of cell death and inflammation to therapeutic giants—past, present and future. Cytokine Growth Factor Rev. 2014, 25, 453–472. [Google Scholar] [CrossRef] [PubMed]
- Penaloza, H.F.; Schultz, B.M.; Nieto, P.A.; Salazar, G.A.; Suazo, I.; Gonzalez, P.A.; Riedel, C.A.; Alvarez-Lobos, M.M.; Kalergis, A.M.; Bueno, S.M. Opposing roles of IL-10 in acute bacterial infection. Cytokine Growth Factor Rev. 2016, 32, 17–30. [Google Scholar] [CrossRef]
- Jeong, K.I.; Zhang, Q.; Nunnari, J.; Tzipori, S. A piglet model of acute gastroenteritis induced by Shigella dysenteriae Type 1. J. Infect. Dis. 2010, 201, 903–911. [Google Scholar] [CrossRef] [PubMed]
- Splichalova, A.; Splichal, I. Local and systemic occurrences of HMGB1 in gnotobiotic piglets infected with E. coli O55 are related to bacterial translocation and inflammatory cytokines. Cytokine 2012, 60, 597–600. [Google Scholar] [CrossRef]
- Collado-Romero, M.; Arce, C.; Ramirez-Boo, M.; Carvajal, A.; Garrido, J.J. Quantitative analysis of the immune response upon Salmonella typhimurium infection along the porcine intestinal gut. Vet. Res. 2010, 41, 23. [Google Scholar] [CrossRef]
- Knetter, S.M.; Bearson, S.M.; Huang, T.H.; Kurkiewicz, D.; Schroyen, M.; Nettleton, D.; Berman, D.; Cohen, V.; Lunney, J.K.; Ramer-Tait, A.E.; et al. Salmonella enterica serovar Typhimurium-infected pigs with different shedding levels exhibit distinct clinical, peripheral cytokine and transcriptomic immune response phenotypes. Innate Immun. 2015, 21, 227–241. [Google Scholar] [CrossRef]
- Splichalova, A.; Splichal, I.; Chmelarova, P.; Trebichavsky, I. Alarmin HMGB1 is released in the small intestine of gnotobiotic piglets infected with enteric pathogens and its level in plasma reflects severity of sepsis. J. Clin. Immunol. 2011, 31, 488–497. [Google Scholar] [CrossRef] [PubMed]
- Ng, P.C.; Li, K.; Wong, R.P.; Chui, K.; Wong, E.; Li, G.; Fok, T.F. Proinflammatory and anti-inflammatory cytokine responses in preterm infants with systemic infections. Arch. Dis. Child. Fetal Neonatal Ed. 2003, 88, F209–F213. [Google Scholar] [CrossRef] [PubMed]
- Gogos, C.A.; Drosou, E.; Bassaris, H.P.; Skoutelis, A. Pro- versus anti-inflammatory cytokine profile in patients with severe sepsis: A marker for prognosis and future therapeutic options. J. Infect. Dis. 2000, 181, 176–180. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Splichal, I.; Muneta, Y.; Mori, Y.; Takahashi, E. Development and application of a pig IL-8 ELISA detection system. J. Immunoass. Immunochem. 2003, 24, 219–232. [Google Scholar] [CrossRef] [PubMed]
- Datsenko, K.A.; Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 2000, 97, 6640–6645. [Google Scholar] [CrossRef] [PubMed]


| Gene | 5′-Reverse Primer-3′ | 5′-Forward Primer-3′ | #LNA Probe |
|---|---|---|---|
| BACT 1 | TCCCTGGAGAAGAGCTACGA | AAGAGCGCCTCTGGACAC | 9 |
| CYPA 2 | CCTGAAGCATACGGGTCCT | AAAGACCACATGTTTGCCATC | 48 |
| CLD-1 3 | CACCACTTTGCAAGCAACC | TGGCCACAAAGATGGCTATT | 3 |
| CLD-2 4 | CTCGCGCCAAAGACAGAG | ATGAAGATTCCACGCAACG | 77 |
| OCLN 5 | AAAGAGCTCTCTCGACTGGATAAA | AGCAGCAGCCATGTACTCTTC | 42 |
| IL-8 | TTCTTCTTTATCCCCAAACTGG | CCACATGTCCTCAAGGTAGGA | 41 |
| TNF-α | TCAGGGATTCAGGGATGTGT | GAAGCCCCAGTTCCAATTC | 77 |
| IL-10 | TGCCTCCCACTTTCTCTTGT | TTCCTATGAGTGTAAGCGACTTTG | 23 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Splichalova, A.; Splichalova, Z.; Karasova, D.; Rychlik, I.; Trevisi, P.; Sinkora, M.; Splichal, I. Impact of the Lipopolysaccharide Chemotype of Salmonella Enterica Serovar Typhimurium on Virulence in Gnotobiotic Piglets. Toxins 2019, 11, 534. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11090534
Splichalova A, Splichalova Z, Karasova D, Rychlik I, Trevisi P, Sinkora M, Splichal I. Impact of the Lipopolysaccharide Chemotype of Salmonella Enterica Serovar Typhimurium on Virulence in Gnotobiotic Piglets. Toxins. 2019; 11(9):534. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11090534
Chicago/Turabian StyleSplichalova, Alla, Zdislava Splichalova, Daniela Karasova, Ivan Rychlik, Paolo Trevisi, Marek Sinkora, and Igor Splichal. 2019. "Impact of the Lipopolysaccharide Chemotype of Salmonella Enterica Serovar Typhimurium on Virulence in Gnotobiotic Piglets" Toxins 11, no. 9: 534. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11090534





