The Effect of Blue Light on the Production of Citrinin in Monascus purpureus M9 by Regulating the mraox Gene through lncRNA AOANCR
Abstract
:1. Introduction
2. Results
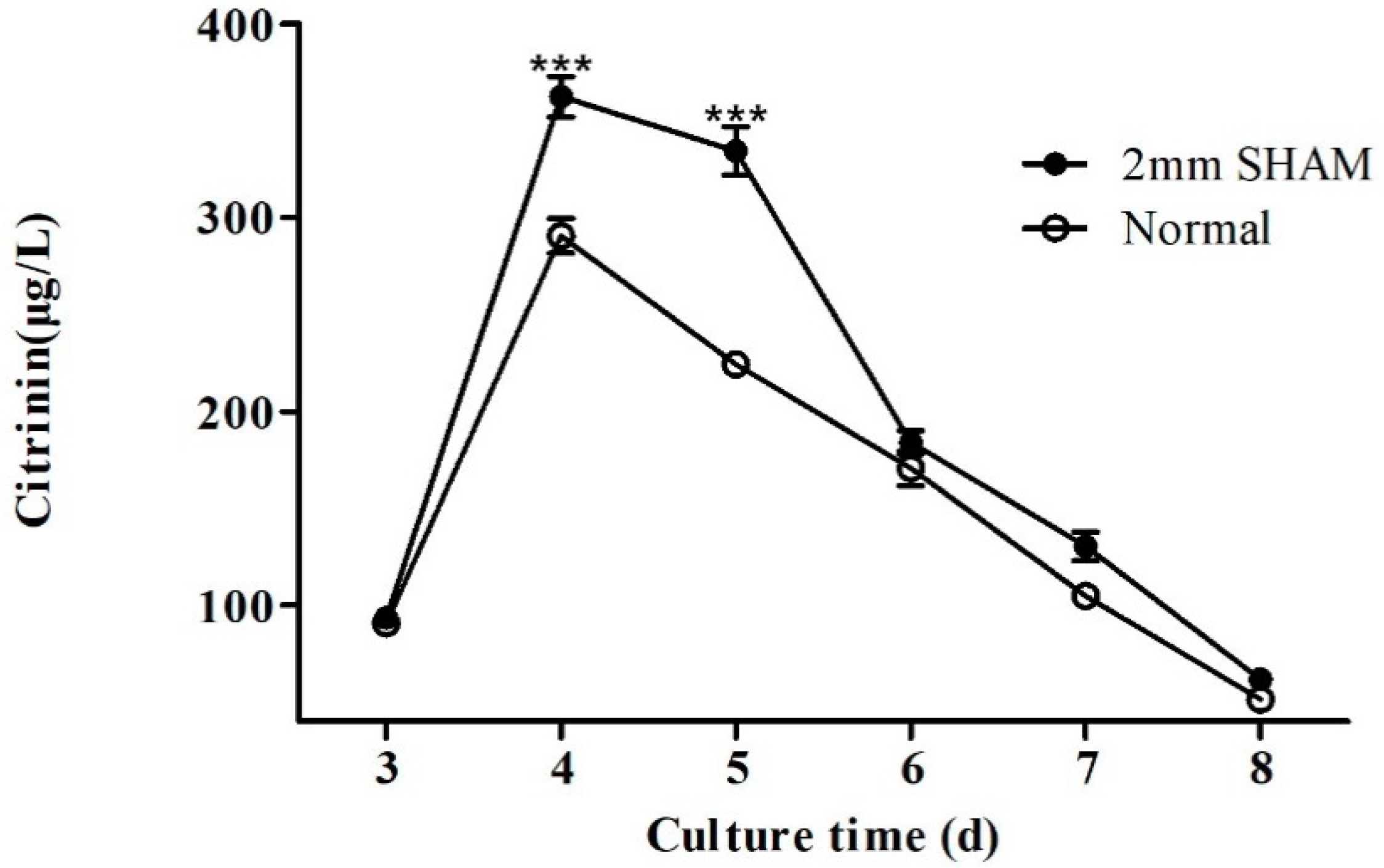
2.1. Effect of Blue Light on Citrinin Production in M9
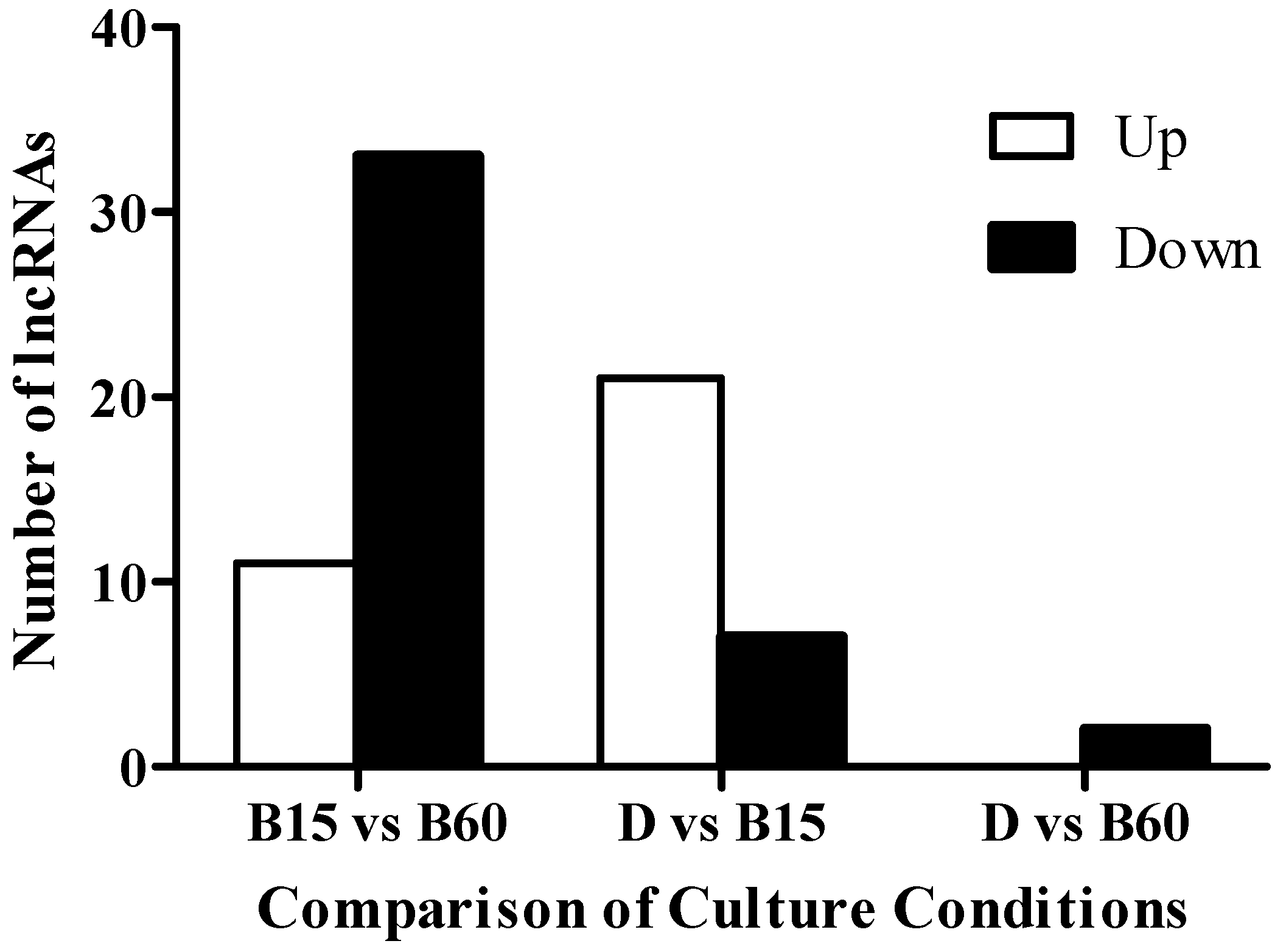
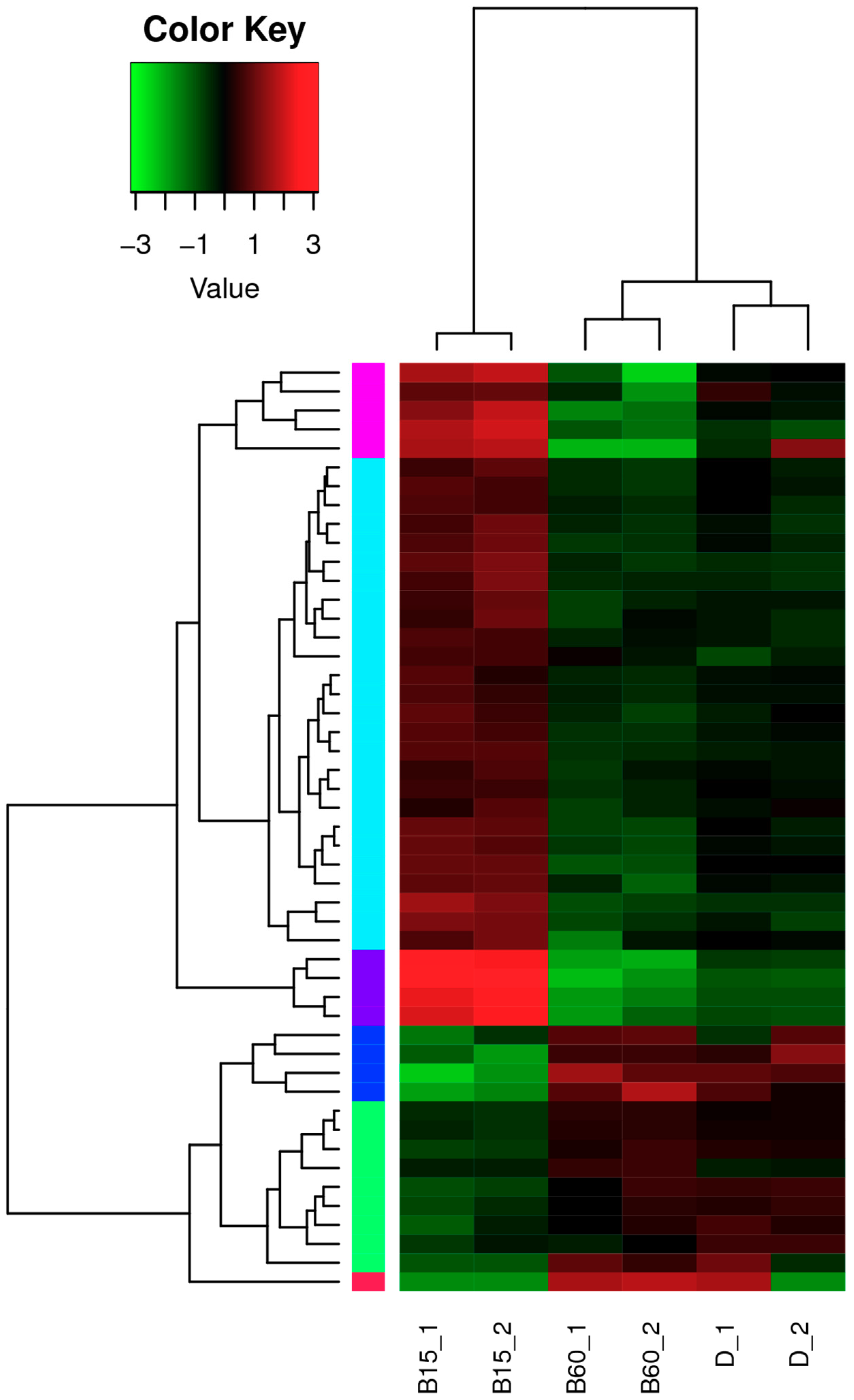
2.2. Analysis of lncRNA in the Transcriptome of Monascus M9
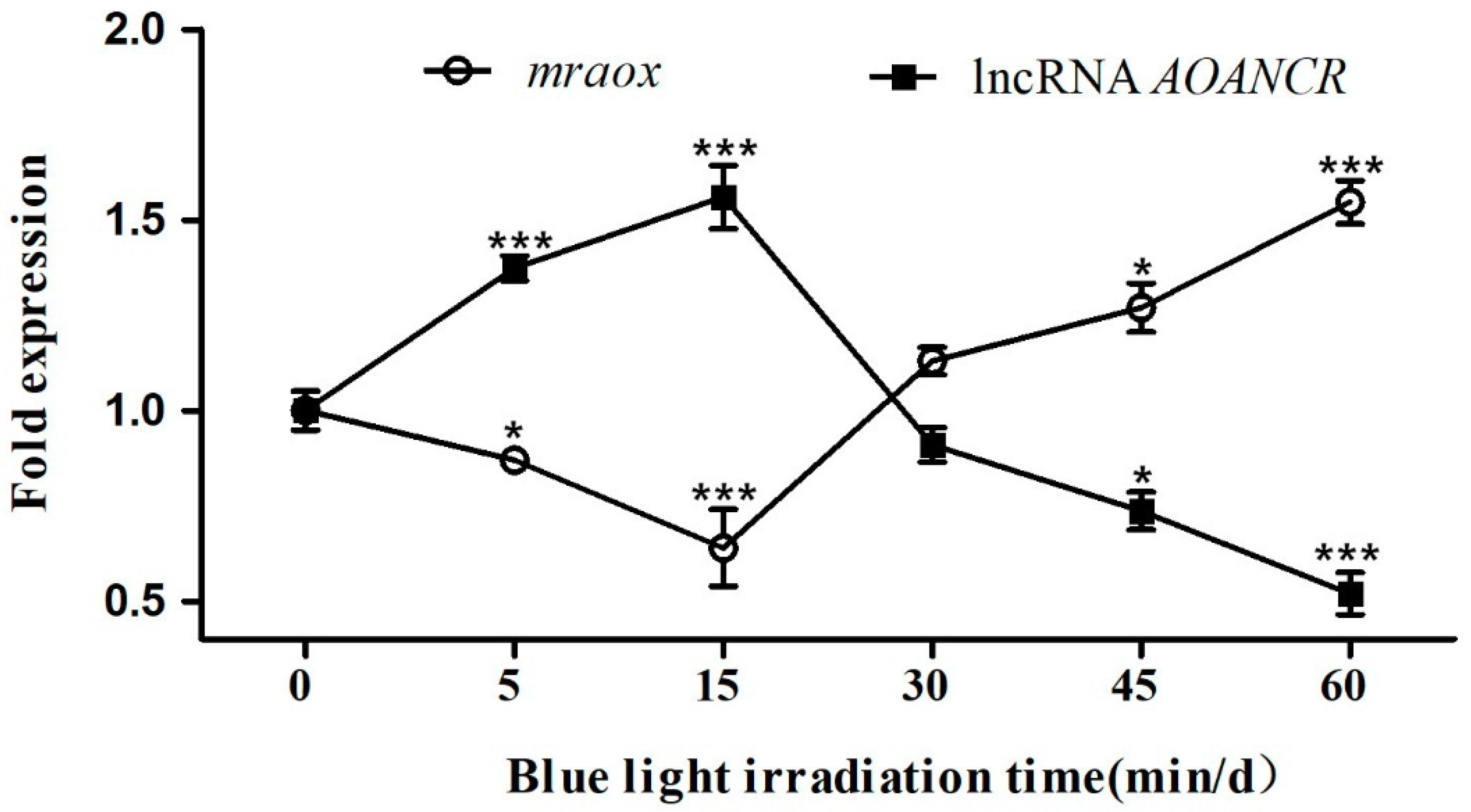
2.3. Expression of lncRNA AOANCR and Mraox Detection in Different Blue Light Conditions
2.4. Effects of Alternating Respiratory Pathway Mediated by Alternate Oxidase on Citrinin Synthesis in Monascus M9
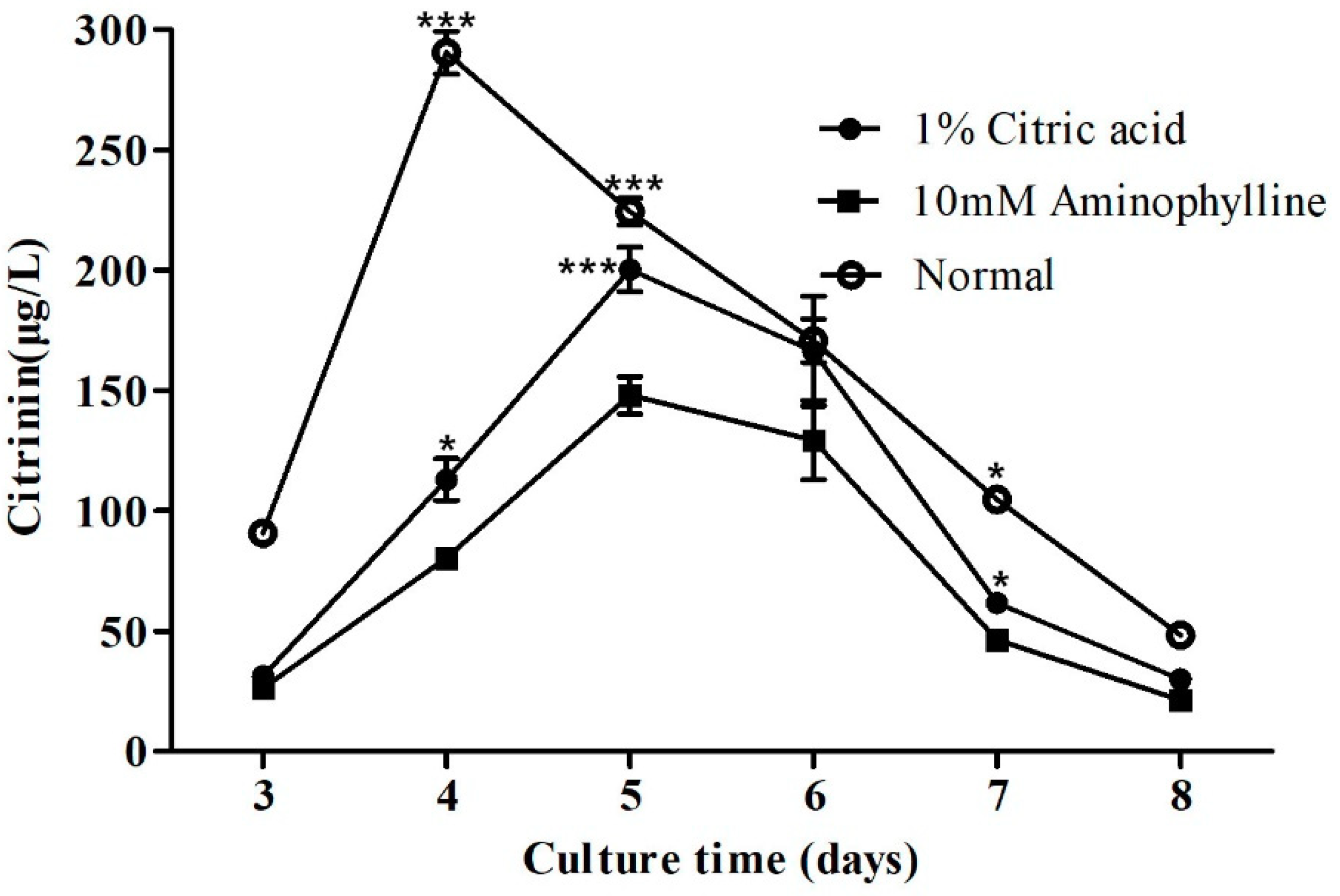
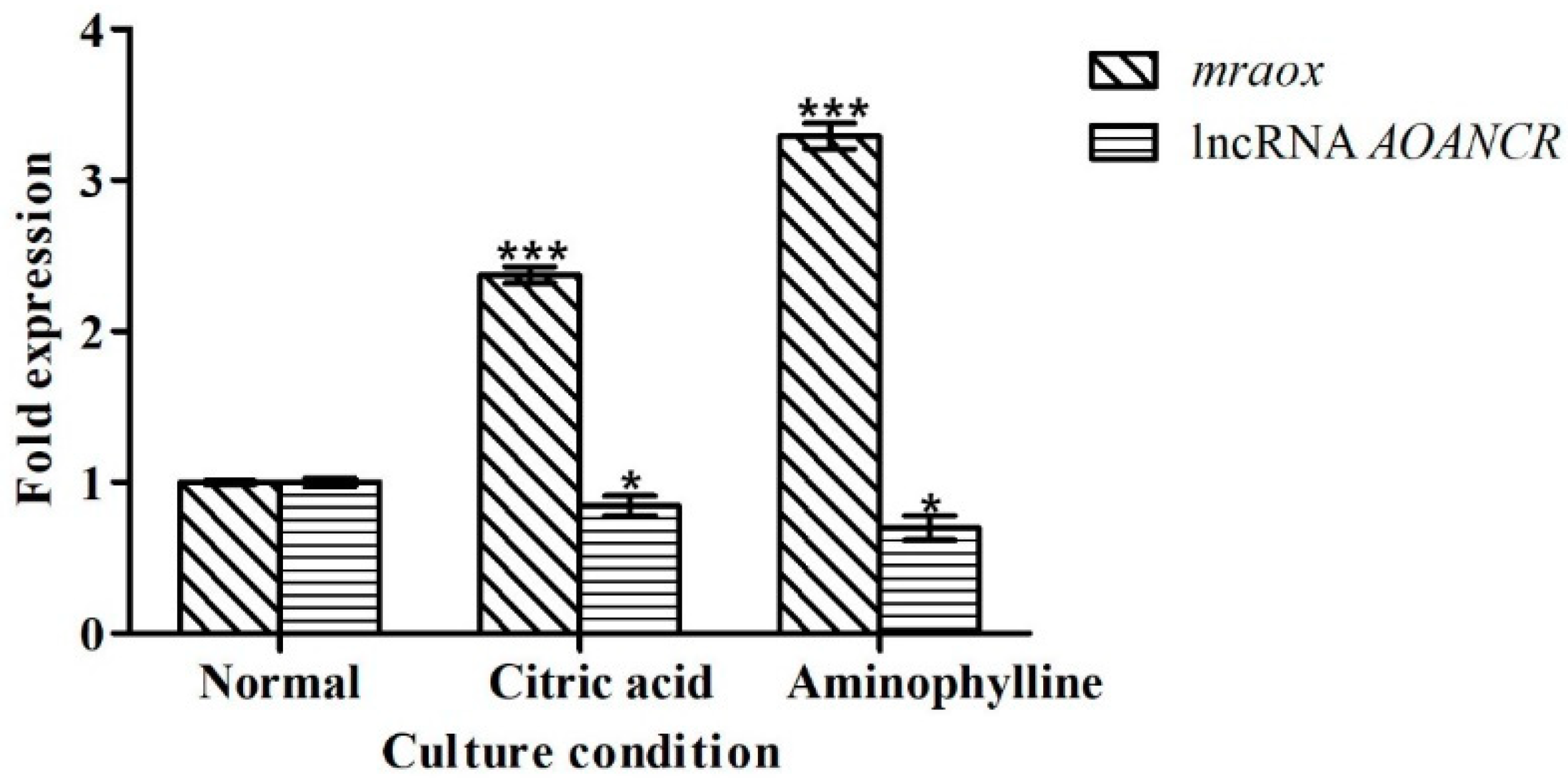
2.5. Citric Acid and Aminophylline Regulate Citrinin Production by Regulating Mraox
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Fungal Strain and Culture Conditions
5.2. Light Exposure Conditions
5.3. Transcriptome Data Acquisition
5.4. Bioinformatics Analysis
5.5. Analysis of Citrinin Production
5.6. QPCR Analysis
Author Contributions
Funding
Conflicts of Interest
References
- Istvan, E.S.; Deisenhofer, J. Structural mechanism for statin inhibition of HMG-CoA reductase. Science 2001, 292, 1160–1164. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Shao, Y.; Chen, F. Monascus pigments. Appl. Microbiol. Biotechnol. 2012, 96, 1421–1440. [Google Scholar] [CrossRef] [PubMed]
- Petera, P. Monascus secondary metabolites: Production and biological activity. J. Ind. Microbiol. Biotechnol. 2013, 40, 169–181. [Google Scholar] [CrossRef]
- Wang, C.; Chen, D.; Chen, M.; Wang, Y.; Li, Z.; Li, F. Stimulatory effects of blue light on the growth, monascin and ankaflavin production in monascus. Biotechnol. Lett. 2015, 37, 1043–1048. [Google Scholar] [CrossRef]
- Su, N.; Lin, Y.; Lee, M.; Ho, C. Ankaflavin from monascus-fermented red rice exhibits selective cytotoxic effect and induces cell death on Hep G2 cells. J. Agric. Food Chem. 2005, 53, 1949–1954. [Google Scholar] [CrossRef] [PubMed]
- Hsu, W.; Liao, T.; Lee, B.; Hsu, Y.; Pan, T. Tzuming, P. Ankaflavin regulates adipocyte function and attenuates hyperglycemia caused by high-fat diet via PPAR-gamma activation. J. Funct. Foods 2013, 5, 124–132. [Google Scholar] [CrossRef]
- Hsu, W.; Huang, Y.; Lee, B.; Hsu, Y.; Pan, T. The improvements of ankaflavin isolated from Monascus-fermented products on dyslipidemia in high-fat diet-induced hamster. J. Funct. Food 2013, 5, 434–443. [Google Scholar] [CrossRef]
- Hsu, W.; Chen, T.; Lee, B.; Hsu, Y.; Pan, T.; Tzuming, P. Monascin and ankaflavin act as natural AMPK activators with PPARα agonist activity to down-regulate nonalcoholic steatohepatitis in high-fat diet-fed C57BL/6 mice. Food Chem. Toxicol. 2014, 64, 94–103. [Google Scholar] [CrossRef]
- Liu, G.; Hou, T.; Yuan, Y.; Hang, P.; Zhao, J.; Sun, L.; Zhao, G.; Zhao, J.; Dong, J.; Wang, X.; et al. Fenofibrate inhibits atrial metabolic remodelling in atrial fibrillation through PPAR-α/sirtuin 1/PGC-1α pathway. Br. J. Pharmacol. 2016, 6, 1095–1109. [Google Scholar] [CrossRef]
- He, Y.; Cox, R.J. The molecular steps of citrinin biosynthesis in fungi. Chem. Sci. 2016, 7, 2119–2127. [Google Scholar] [CrossRef]
- Ostry, V.; Malir, F.; Ruprich, J. Producers and important dietary sources of ochratoxin A and citrinin. Toxins 2013, 5, 1574–1586. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Yu, Y.; Liu, B.; Yu, F. Development of a Sensitive Enzyme-Linked Immunosorbent Assay and Rapid Gold Nanoparticle Immunochromatographic Strip for Detecting Citrinin in Monascus Fermented Food. Toxins 2018, 10, 354. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Su, B.; Xu, Y.; Li, L.; Li, Y. Determination of two potential toxicity metabolites derived from the disruption of the pksCT gene in Monascus aurantiacus Li As3.4384. J. Sci. Food Agric. 2017, 97, 4190–4197. [Google Scholar] [CrossRef] [PubMed]
- Natalia, A.M.; Vicente, R.E.; Plácido, A.F.; Ana, M.G.; Laura, G. Occurrence of Mycotoxins in Swine Feeding from Spain. Toxins 2019, 11, 342. [Google Scholar] [CrossRef]
- Martins, C.; Vidal, A.; De, B.M.; De, S.S.; Nunes, C.; Torres, D.; Goios, A.; Lopes, C.; Assunção, R.; Alvito, P. Exposure assessment of Portuguese population to multiple mycotoxins: The human biomonitoring approach. Int. J. Hyg. Environ. Health 2019. [Google Scholar] [CrossRef] [PubMed]
- Gong, L.; Zhu, H.; Li, T.; Ming, G.; Duan, X.; Wang, J.; Jiang, Y. Molecular signatures of cytotoxic effects in human embryonic kidney 293 cells treated with single and mixture of ochratoxin A and citrinin. Food Chem. Toxicol. 2019, 123, 374–384. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Zhang, H.; Zhao, L.; Zhang, X.; Yang, Q.; Li, Q.; Sun, Y.; Cheng, Y. Progress in controlling citrinin in foods. Microbiol. China 2017, 44, 1451–1457. [Google Scholar] [CrossRef]
- Zhang, X.; Lin, Z.; Apaliya, M.T.; Gu, X.; Zheng, X.; Zhao, L.; Abdelhai, M.H.; Zhang, H.; Hu, W. The Possible Mechanisms Involved in Citrinin Elimination by Cryptococcus podzolicus Y3 and the Effects of Extrinsic Factors on the Degradation of Citrinin. J. Microbiol. Biotechnol. 2017, 27, 2119–2128. [Google Scholar] [CrossRef]
- Moghadam, H.D.; Shahidi, F.; Yazdi, F.T.; Jamab, M.S.; Eshaghi, Z. Biological detoxification of Monascus purpureus pigments by heat-treated Saccharomyces cerevisiae. J. Sci. Food Agric. 2019, 99, 4439–4444. [Google Scholar] [CrossRef]
- De Oliveira Filho, J.W.G.; Islam, M.T.; Ali, E.S.; Uddin, S.J.; Santos, J.V.O.; de Alencar, M.V.O.B.; Júnior, A.L.G.; Paz, M.F.C.J.; de Brito, M.D.R.M.; ESousa, J.M.C.; et al. A comprehensive review on biological properties of citrinin. Food Chem. Toxicol. 2017, 110, 130–141. [Google Scholar] [CrossRef]
- Tsuyoshi, M.; Akira, M.; Toshie, K.; Tadashi, O.; Yasuaki, U.; Fumihiro, S.; Hiroyuki, S.; Akira, W.; Masahiro, K. Light effects on cell development and secondary metabolism in Monascus. J. Ind. Microbiol. Biotechnol. 2005, 32, 103–108. [Google Scholar] [CrossRef]
- Markus, S.H.; Corinna, R.; Frank, R.; Anja, B.; Giancarlo, P.; Rolf, G. Influence of light on food relevant fungi with emphasis on ochratoxin producing species. Int. J. Food Microbiol. 2011, 145, 229–237. [Google Scholar] [CrossRef]
- Wang, C.; Yang, H.; Chen, M.; Wang, Y.; Li, F.; Luo, C.; Zhao, S.; He, D. Real-time quantitative analysis of the influence of blue light on citrinin biosynthetic gene cluster expression in Monascus. Biotechnol. Lett. 2012, 34, 1745–1748. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Xue, C.; Chen, M.; Wu, S.; Li, Z.; Wang, C. Effects of blue light on pigment biosynthesis of Monascus. J. Microbiol. 2016, 54, 305–310. [Google Scholar] [CrossRef] [PubMed]
- Spizzo, R.; Almeida, M.I.; Colombatti, A.; Calin, G.A. Long non-coding RNAs and cancer: A new frontier of translational research. Oncogene 2012, 31, 4577–4587. [Google Scholar] [CrossRef]
- Yamini, A.; Christian, H.; Sam, G.J.; Susan K, C. Natural Antisense Transcripts and Long Non-Coding RNA in Neurospora crassa. PLoS ONE 2014, 9, e91353. [Google Scholar] [CrossRef]
- Li, H.; Wang, Y.; Chen, M.; Xiao, P.; Hu, C.; Zeng, Z.; Wang, C.; Wang, J.; Hu, Z. Genome-wide long non-coding RNA screening, identification and characterization in a model microorganism Chlamydomonas reinhardtii. Sci. Rep. 2016, 6, 34109. [Google Scholar] [CrossRef]
- Chen, D.; Chen, M.; Wu, S.; Li, Z.; Yang, H.; Wang, C. The Molecular Mechanisms of Monascus Purpureus M9 Responses to Blue Light Based on the Transcriptome Analysis. Sci. Rep. 2017, 7, 5537. [Google Scholar] [CrossRef]
- Shao, Y.; Li, Q.; Zhou, Y.; Chen, F. Effects of an alternative oxidase gene on conidia viability under external stresses in Monascus ruber M7. J. Basic Microbiol. 2017, 9999, 1–6. [Google Scholar] [CrossRef]
- Linden, H.; Ballario, P.; Macino, G. Blue light regulation in Neurospora crassa. Fungal Genet. Biol. 1997, 22, 141–150. [Google Scholar] [CrossRef]
- Chen, C.H.; Ringelberg, C.S.; Gross, R.H.; Dunlap, J.C.; Loros, J.J. Genome-wide analysis of light-inducible responses reveals hierarchical light signalling in Neurospora. EMBO J. 2009, 28, 1029–1042. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Dunlap, J.C.; Loros, J.J. Neurospora illuminates fungal photoreception. Fungal Genet. Biol. 2010, 47, 922–929. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heintzen, C.; Loros, J.J.; Dunlap, J.C. The PAS Protein VIVID Defines a Clock-Associated Feedback Loop that Represses Light Input, Modulates Gating, and Regulates Clock Resetting. Cell 2001, 104, 453–464. [Google Scholar] [CrossRef]
- Hunt, S.M.; Thompson, S.; Elvin, M.; Heintzen, C. VIVID interacts with the WHITE COLLAR complex and FREQUENCY-interacting RNA helicase to alter light and clock responses in Neurospora. Proc. Natl. Acad. Sci. USA 2010, 107, 16709. [Google Scholar] [CrossRef] [PubMed]
- Malzahn, E.; Ciprianidis, S.; Káldi, K.; Schafmeier, T.; Brunner, M. Photoadaptation in Neurospora by competitive interaction of activating and inhibitory LOV domains. Cell 2010, 142, 762–772. [Google Scholar] [CrossRef] [PubMed]
- Schwerdtfeger, C.; Linden, H. Localization and light-dependent phosphorylation of white collar 1 and 2, the two central components of blue light signaling in Neurospora crassa. Eur. J. Biochem. 2000, 267, 414. [Google Scholar] [CrossRef]
- Schwerdtfeger, C.; Linden, H. VIVID is a flavoprotein and serves as a fungal blue light photoreceptor for photoadaptation. EMBO J. 2003, 22, 4846–4855. [Google Scholar] [CrossRef] [PubMed]
- Shrode, L.B.; Lewis, Z.A.; White, L.D.; Bell-Pedersen, D.; Ebbole, D.J. vvd Is Required for Light Adaptation of Conidiation-Specific Genes of Neurospora crassa, but Not Circadian Conidiation. Fungal Genet. Biol. 2001, 32, 169–181. [Google Scholar] [CrossRef]
- Terauchi, K.; Ohmori, M. Blue light stimulates cyanobacterial motility via a cAMP signal transduction system. Mol. Microbiol. 2004, 52, 303–309. [Google Scholar] [CrossRef]
- Miyake, T.; Zhang, M.; Kono, I.; Nozaki, N.; Sammoto, H. Repression of Secondary Metabolite Production by Exogenous cAMP in Monascus. Biosci. Biotechnol. Biochem. 2006, 70, 1521–1523. [Google Scholar] [CrossRef]
- Nemcovic, M.; Farkas, V. Stimulation of conidiation by derivatives of cAMP in Trichoderma viride. Folia Microbiol. 1998, 43, 399–402. [Google Scholar] [CrossRef]
- Li, Q. Cloning and Characterization of the Mraox1 Gene from Monascus Ruber. Master Dissertation, Huazhong Agricultural University, Wuhan, China, 2010. [Google Scholar] [CrossRef]
- Sarikaya-Bayram, Ö.; Palmer, J.M.; Keller, N.; Braus, G.H.; Bayram, Ö. One Juliet and four Romeos: VeA and its methyltransferases. Front. Microbiol. 2015, 6, 1. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Bai, Z.; O’Donnell, A.; Harvey, L.M.; Hoskisson, P.A.; McNeil, B. Oxidative stress in fungal fermentation processes: The roles of alternative respiration. Biotechnol. Lett. 2011, 33, 457–467. [Google Scholar] [CrossRef] [PubMed]
- Kakizaki, Y.; Ito, K. Engineering plant alternative oxidase function in mammalian cells: Substitution of the motif-likes sequence ENV for QDT diminishes catalytic activity of Arum concinnatum AOX1a expressed in HeLa Cells. Appl. Biochem. Biotechnol. 2013, 170, 1229–1240. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Liu, J. Enhanced fatty acid accumulation in Isochrysis galbana by inhibition of the mitochondrial alternative oxidase pathway under nitrogen deprivation. Bioresour. Technol. 2016, 211, 783–786. [Google Scholar] [CrossRef] [PubMed]
- Challabathula, D.; Abhaypratap, V.; Raghavendra, A.S.; Kollipara, P. Alternative oxidase pathway optimizes photosynthesis during osmotic and temperature stress by regulating cellular ROS, malate valve and antioxidative systems. Front. Plant Sci. 2016, 7, 68. [Google Scholar] [CrossRef]
- Hu, W.; Yan, X.; Yu, J. Importance of the mitochondrial alternative oxidase (AOX) pathway in alleviating photo inhibition in cucumber leaves under chilling injury and subsequent recovery when leaves are subjected to high light intensity. J. Hortic. Sci. Biotechnol. 2016, 1219239. [Google Scholar] [CrossRef]
- Garcia-Neto, W.; Cabrera-Orefice, A.; Uribe-Carvajal, S.; Kowaltowski, A.J.; Alberto Luévano-Martínez, L. High osmolarity environments activate the mitochondrial alternative oxidase in Debaryomyces Hansenii. PLoS ONE 2017, 12, e0169621. [Google Scholar] [CrossRef]
- Tardu, M.; Bulut, S.; Kavakli, I.H. MerR and ChrR mediate blue light induced photo-oxidative stress response at the transcriptional level in Vibrio cholerae. Sci. Rep. 2017, 7, 40817. [Google Scholar] [CrossRef]
- Martins, V.P.; Dinamarco, T.M.; Soriani, F.M.; Tudella, V.G.; Oliveira, S.C.; Goldman, G.H.; Curti, C.; Uyemura, S.A. Involvement of an alternative oxidase in oxidativestress and mycelium-to-yeast differentiation in Paracoccidioides brasiliensis. Eukaryot. Cell 2011, 10, 237–248. [Google Scholar] [CrossRef]
- Møller, I.M. Plant mitochondria and oxidativestress: Electron transport, NADPH turnover, andmetabolism of reactive oxygen species. Ann. Rev. Plant Physiol. Mol. Biol. 2001, 52, 561–591. [Google Scholar] [CrossRef] [PubMed]
- El-Khoury, R.; Dufour, E.; Rak, M.; Ramanantsoa, N.; Grandchamp, N.; Csaba, Z.; Duvillié, B.; Bénit, P.; Gallego, J.; Gressens, P.; et al. Alternative oxidase expression in the mouse enables by passing cytocherome c oxidise blockade and limits mitochondrial ROS overproduction. PLoS Genet. 2013, 9, e1003182. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Shi, K.; Fu, L.; Zhang, S.; Li, X.; Dong, D.; Jiang, Y.; Zhou, Y.; Xia, X.; Liang, W.; et al. The reduction of reactive oxygen species formation by mitochondrial alternative respiration in tomato basal defense against TMV infection. Planta 2012, 235, 225–238. [Google Scholar] [CrossRef] [PubMed]
- Qi, Z.; Smith, K.M.; Bredeweg, E.L.; Bosnjak, N.; Freitag, M.; Nargang, F.E. Alternative Oxidase Transcription Factors AOD2 and AOD5 of Neurospora crassa Control the Expression of Genes Involved in Energy Production and Metabolism. G3 (Bethesda) 2017, 9, 449–466. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Xu, J.; Wang, Y.; Cao, X. An interferon-independent lncRNA promotes viral replication by modulating cellular metabolism. Science 2017, 358, 1051–1055. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, J.; Yan, B.; Qu, Y.; Qin, F.; Yang, Y.; Hao, X.; Yu, J.; Zhao, Q.; Zhu, D.; Ao, G. Zm401 a short-open reading-frame mRNA or noncoding RNA, is essential for tapetum and microspore development and can regulate the floret formation in maize. J. Cell. Biochem. 2008, 105, 136–146. [Google Scholar] [CrossRef] [PubMed]
- Amor, B.B.; Wirth, S.; Merchan, F.; Philippe, L.; Yves, D.; Judith, H.; Alexis, M.; Allison, M.; Antoine, L.; Jean, M.; et al. Novel long non-proteincoding RNAs involved in Arabidopsis differentiation and stress responses. Genome Res. 2009, 19, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, A.; Shichino, Y.; Yamamoto, M. The long non-coding RNA world in yeasts. Biochim. Biophys. Acta 2016, 1859, 147–154. [Google Scholar] [CrossRef]
- Wang, C.; Fu, Z.; Chen, M.; Ban, Z.; Wang, Y.; Zhang, X. Blue Light Effects on Pigment and Citrinin Production in Monascus. In Proceedings of the 2009 3rd International Conference on Bioinformatics and Biomedical Engineering, Beijing, China, 11–13 June 2009. [Google Scholar]
- Fatima, A.; Aftab, U.; Shaaban, K.A.; Thorson, J.S.; Sajid, I. Spore forming Actinobacterial diversity of Cholistan Desert Pakistan: Polyphasic taxonomy, antimicrobial potential and chemical profiling. BMC Microbiol. 2019, 19, 49. [Google Scholar] [CrossRef]
- Tineke, L.L.; Antoine, C.; Carson, C.C.; Daniel, R.L. Single-molecule imaging reveals a switch between spurious and functional ncRNA transcription article single-molecule imaging reveals a switch between spurious and functional ncRNA transcription. Mol. Cell 2015, 60, 597–610. [Google Scholar] [CrossRef]
- Cloutier, S.C.; Wang, S.; Ma, W.K.; Al Husini, N.; Dhoondia, Z.; Ansari, A.; Pascuzzi, P.E.; Tran, E.J. Regulated Formation of lncRNA-DNA Hybrids Enables Faster Transcriptional Induction and Environmental Adaptation. Mol. Cell 2016, 62, 148. [Google Scholar] [CrossRef] [PubMed]
- Castelnuovo, M.; Rahman, S.; Guffanti, E.; Infantino, V.; Stutz, F.; Zenklusen, D. Bimodal expression of PHO84 is modulated by early termination of antisense transcription. Nat. Struct. Mol. Biol. 2013, 20, 851–858. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, Y.; Chen, H.; Teng, S. Intragenic transcription of a noncoding RNA modulates expression of ASP3 in budding yeast. RNA 2010, 16, 2085–2093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, T.; Xu, Z.; Clauder-Münster, S.; Steinmetz, L.M.; Buratowski, S. Set3 HDAC mediates effects of overlapping non-coding transcription on gene induction kinetics. Cell 2012, 150, 1158–1169. [Google Scholar] [CrossRef] [PubMed]
- Hainer, S.J.; Pruneski, J.A.; Mitchell, R.D.; Monteverde, R.M.; Martens, J.A. Intergenic transcription causes repression by directing nucleosome assembly. Genes Dev. 2011, 25, 29–40. [Google Scholar] [CrossRef]
- Davis, N.D.; Dalby, D.K.; Diener, U.L.; Snasnig, G.A. Medium-Scale production of Citrinin by Penicillium citrinum in a Semisynthetic Medium. Appl. Microbiol. 1975, 29, 118–120. [Google Scholar] [PubMed]
- Wang, L.; Dai, Y.; Chen, W.; Shao, Y.; Chen, F. Effects of light intensity and color on the biomass, extracellular red pigment and citrinin production of Monascus ruber. J. Agric. Food Chem. 2016, 64, 9506–9514. [Google Scholar] [CrossRef]
- Bühler, R.M.M.; Müller, B.L.; Moritz, D.E.; Vendruscolo, F.; de Oliveira, D.; Ninow, J.L. Influence of Light Intensity on Growth and Pigment Production by Monascus ruber in Submerged Fermentation. Appl. Biochem. Biotechnol. 2015, 176, 1277–1289. [Google Scholar] [CrossRef]
- Kang, B.; Zhang, X.; Wu, Z.; Wang, Z.; Park, S. Production of citrinin-free Monascus pigments by submerged culture at low pH. Enzyme Microb. Technol. 2014, 55, 50–57. [Google Scholar] [CrossRef]
- Zhen, Z.; Xiong, X.; Liu, Y.; Zhang, J.; Wang, S.; Li, L.; Gao, M. NaCl Inhibits Citrinin and Stimulates Monascus Pigments and Monacolin K Production. Toxins 2019, 11, 118. [Google Scholar] [CrossRef]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 2011, 39 (Suppl. 2), W29–W37. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Zhang, Y.; Ye, Z.; Liu, X.; Zhao, S.; Wei, L.; Gao, G. CPC: Assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007, 35 (Suppl. 2), W345–W349. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Terms | Value |
|---|---|
| Total sequences | 1455 |
| Total bases | 1056220 |
| Min sequence length | 202 |
| Max sequence length | 6927 |
| Average sequence length | 725.92 |
| Median sequence length | 310 |
| As | 25.29% |
| Ts | 24.90% |
| Gs | 24.78% |
| Cs | 25.03% |
| (A+T)s | 50.19% |
| (G+C)s | 49.81% |
| Ns | 0.00% |
| Primers | Sequence (5′–3′) |
|---|---|
| lncRNA AOANCR | |
| Forward primer | CGGGTGGCTAAGGGATTCAT |
| Reverse primer | GCCTGCTTGACTGCTGTTTC |
| mraox | |
| Forward primer | AAGAAACTGACCGACCGACC |
| Reverse primer | TGCAGTGTCCATTCCCCATC |
| actin | |
| Forward primer | GAGCGCGGCTACACCTTTAC |
| Reverse primer | GGGCAACGTAGCAGAGCTTCT |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, H.; Wang, X.; Li, Z.; Guo, Q.; Yang, M.; Chen, D.; Wang, C. The Effect of Blue Light on the Production of Citrinin in Monascus purpureus M9 by Regulating the mraox Gene through lncRNA AOANCR. Toxins 2019, 11, 536. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11090536
Yang H, Wang X, Li Z, Guo Q, Yang M, Chen D, Wang C. The Effect of Blue Light on the Production of Citrinin in Monascus purpureus M9 by Regulating the mraox Gene through lncRNA AOANCR. Toxins. 2019; 11(9):536. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11090536
Chicago/Turabian StyleYang, Hua, Xufeng Wang, Zhenjing Li, Qingbin Guo, Mingguan Yang, Di Chen, and Changlu Wang. 2019. "The Effect of Blue Light on the Production of Citrinin in Monascus purpureus M9 by Regulating the mraox Gene through lncRNA AOANCR" Toxins 11, no. 9: 536. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11090536




