Nuclear Envelope Regulation of Oncogenic Processes: Roles in Pancreatic Cancer
Abstract
:1. Introduction
2. Chromatin Organization and Dynamics
3. The Nuclear Envelope
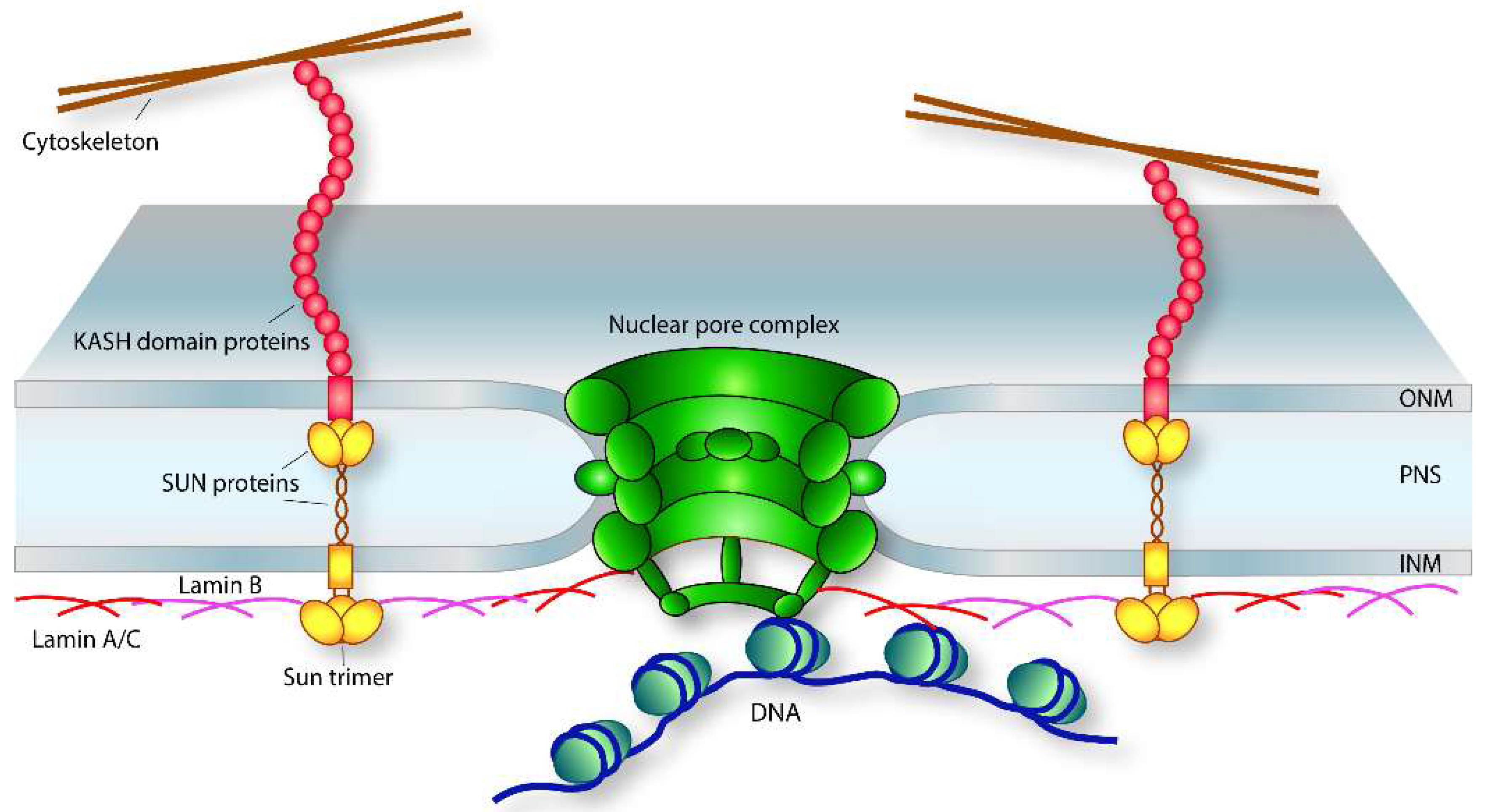
3.1. LINC Complexes
3.2. Nuclear Lamina
3.3. Nuclear Pore Complex
3.4. Potential Regulation of Epigenomic Dynamics by the Nuclear Envelope in the Setting of Pancreatic Cancer
4. Summary and Future Directions
Funding
Conflicts of Interest
References
- Torre, L.A.; Bray, F.; Siegel, R.L.; Ferlay, J.; Lortet-Tieulent, J.; Jemal, A. Global cancer statistics, 2012. CA Cancer J. Clin. 2015, 65, 87–108. [Google Scholar] [CrossRef] [PubMed]
- McGuire, S. World Cancer Report 2014. Geneva, Switzerland: World Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Adv. Nutr. 2016, 7, 418–419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cronin, K.A.; Lake, A.J.; Scott, S.; Sherman, R.L.; Noone, A.M.; Howlader, N.; Henley, S.J.; Anderson, R.N.; Firth, A.U.; Ma, J.; et al. Annual Report to the Nation on the Status of Cancer, part I: National cancer statistics. Cancer 2018, 124, 2785–2800. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics, 2017. CA Cancer J. Clin. 2017, 67, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.; Sharma, P.; Zakalik, D. Comparison of Demographics, Tumor Characteristics, and Survival Between Pancreatic Adenocarcinomas and Pancreatic Neuroendocrine Tumors: A Population-based Study. Am. J. Clin. Oncol. 2018, 41, 485–491. [Google Scholar] [CrossRef] [PubMed]
- Hruban, R.H.; Klimstra, D.S. Adenocarcinoma of the pancreas. Semin. Diagn. Pathol. 2014, 31, 443–451. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martinez-Bosch, N.; Vinaixa, J.; Navarro, P. Immune Evasion in Pancreatic Cancer: From Mechanisms to Therapy. Cancers 2018, 10, 6. [Google Scholar] [CrossRef] [PubMed]
- Bahrami, A.; Khazaei, M.; Bagherieh, F.; Ghayour-Mobarhan, M.; Maftouh, M.; Hassanian, S.M.; Avan, A. Targeting stroma in pancreatic cancer: Promises and failures of targeted therapies. J. Cell. Physiol. 2017, 232, 2931–2937. [Google Scholar] [CrossRef] [PubMed]
- Bakkevold, K.E.; Arnesjo, B.; Kambestad, B. Carcinoma of the pancreas and papilla of Vater: Presenting symptoms, signs, and diagnosis related to stage and tumour site. A prospective multicentre trial in 472 patients. Norwegian Pancreatic Cancer Trial. Scand. J. Gastroenterol. 1992, 27, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Porta, M.; Fabregat, X.; Malats, N.; Guarner, L.; Carrato, A.; de Miguel, A.; Ruiz, L.; Jariod, M.; Costafreda, S.; Coll, S.; et al. Exocrine pancreatic cancer: Symptoms at presentation and their relation to tumour site and stage. Clin. Transl. Oncol. 2005, 7, 189–197. [Google Scholar] [CrossRef] [PubMed]
- El Guesmi, S.; Ben Nasr, S.; Afrit, M.; Labidi, S.; Boussen, H. Exocrine Pancreatic carcinoma in Tunisia: A retrospective study about 158 cases. Tunis. Med. 2015, 93, 73–75. [Google Scholar] [PubMed]
- Lowenfels, A.B.; Maisonneuve, P. Epidemiology and risk factors for pancreatic cancer. Best Pract. Res. Clin. Gastroenterol. 2006, 20, 197–209. [Google Scholar] [CrossRef] [PubMed]
- Jones, S.; Zhang, X.; Parsons, D.W.; Lin, J.C.; Leary, R.J.; Angenendt, P.; Mankoo, P.; Carter, H.; Kamiyama, H.; Jimeno, A.; et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 2008, 321, 1801–1806. [Google Scholar] [CrossRef] [PubMed]
- Biankin, A.V.; Waddell, N.; Kassahn, K.S.; Gingras, M.C.; Muthuswamy, L.B.; Johns, A.L.; Miller, D.K.; Wilson, P.J.; Patch, A.M.; Wu, J.; et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 2012, 491, 399–405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Klein, A.P.; Wolpin, B.M.; Risch, H.A.; Stolzenberg-Solomon, R.Z.; Mocci, E.; Zhang, M.; Canzian, F.; Childs, E.J.; Hoskins, J.W.; Jermusyk, A.; et al. Genome-wide meta-analysis identifies five new susceptibility loci for pancreatic cancer. Nat. Commun. 2018, 9, 556. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.Z.; Spektor, A.; Cornils, H.; Francis, J.M.; Jackson, E.K.; Liu, S.; Meyerson, M.; Pellman, D. Chromothripsis from DNA damage in micronuclei. Nature 2015, 522, 179–184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morris, L.G.; Chan, T.A. Therapeutic targeting of tumor suppressor genes. Cancer 2015, 121, 1357–1368. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.Y.; Muller, W.J. Oncogenes and tumor suppressor genes. Cold Spring Harb. Perspect. Biol. 2010, 2, a003236. [Google Scholar] [CrossRef] [PubMed]
- Inokawa, H.; Katayama, N.; Nakao, M. Evaluation of multidrug cancer chronotherapy based on cell cycle model under influences of circadian clock. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2016, 2016, 1439–1442. [Google Scholar] [PubMed]
- Sotak, M.; Sumova, A.; Pacha, J. Cross-talk between the circadian clock and the cell cycle in cancer. Ann. Med. 2014, 46, 221–232. [Google Scholar] [CrossRef] [PubMed]
- Lomberk, G.; Blum, Y.; Nicolle, R.; Nair, A.; Gaonkar, K.S.; Marisa, L.; Mathison, A.; Sun, Z.; Yan, H.; Elarouci, N.; et al. Distinct epigenetic landscapes underlie the pathobiology of pancreatic cancer subtypes. Nat. Commun. 2018, 9, 1978. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Micco, R.; Sulli, G.; Dobreva, M.; Liontos, M.; Botrugno, O.A.; Gargiulo, G.; dal Zuffo, R.; Matti, V.; d’Ario, G.; Montani, E.; et al. Interplay between oncogene-induced DNA damage response and heterochromatin in senescence and cancer. Nat. Cell Biol. 2011, 13, 292–302. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lawrimore, J.; Barry, T.M.; Barry, R.M.; York, A.C.; Friedman, B.; Cook, D.M.; Akialis, K.; Tyler, J.; Vasquez, P.; Yeh, E.; et al. Microtubule dynamics drive enhanced chromatin motion and mobilize telomeres in response to DNA damage. Mol. Biol. Cell 2017, 28, 1701–1711. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amendola, M.; van Steensel, B. Mechanisms and dynamics of nuclear lamina-genome interactions. Curr. Opin. Cell Biol. 2014, 28, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, I.A.; Dundr, M. Chromatin loops and causality loops: The influence of RNA upon spatial nuclear architecture. Chromosoma 2017, 126, 541–557. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, I.A.; Dundr, M. Nuclear bodies: Built to boost. J. Cell Biol. 2016, 213, 509–511. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Serizay, J.; Ahringer, J. Genome organization at different scales: Nature, formation and function. Curr. Opin. Cell Biol. 2018, 52, 145–153. [Google Scholar] [CrossRef] [PubMed]
- Rada-Iglesias, A.; Grosveld, F.G.; Papantonis, A. Forces driving the three-dimensional folding of eukaryotic genomes. Mol. Syst. Biol. 2018, 14, e8214. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.D.; Tanizawa, H.; Iwasaki, O.; Noma, K. Transcription factors mediate condensin recruitment and global chromosomal organization in fission yeast. Nat. Genet. 2016, 48, 1242–1252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iwasaki, O.; Corcoran, C.J.; Noma, K. Involvement of condensin-directed gene associations in the organization and regulation of chromosome territories during the cell cycle. Nucleic Acids Res. 2016, 44, 3618–3628. [Google Scholar] [CrossRef] [PubMed]
- Miloshev, G.; Staneva, D.; Uzunova, K.; Vasileva, B.; Draganova-Filipova, M.; Zagorchev, P.; Georgieva, M. Linker histones and chromatin remodelling complexes maintain genome stability and control cellular ageing. Mech. Ageing Dev. 2018. [Google Scholar] [CrossRef] [PubMed]
- Fatakia, S.N.; Kulashreshtha, M.; Mehta, I.S.; Rao, B.J. Chromosome territory relocation paradigm during DNA damage response: Some insights from molecular biology to physics. Nucleus 2017, 8, 449–460. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mehta, I.S.; Kulashreshtha, M.; Chakraborty, S.; Kolthur-Seetharam, U.; Rao, B.J. Chromosome territories reposition during DNA damage-repair response. Genome Biol. 2013, 14, R135. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.Q.; Bosco, G. Gene Positioning Effects on Expression in Eukaryotes. Annu. Rev. Genet. 2015, 49, 627–646. [Google Scholar] [CrossRef] [PubMed]
- Kinney, N.A.; Sharakhov, I.V.; Onufriev, A.V. Chromosome-nuclear envelope attachments affect interphase chromosome territories and entanglement. Epigenet. Chromatin 2018, 11, 3. [Google Scholar] [CrossRef] [PubMed]
- Cremer, T.; Cremer, M.; Cremer, C. The 4D Nucleome: Genome Compartmentalization in an Evolutionary Context. Biochemistry 2018, 83, 313–325. [Google Scholar] [CrossRef] [PubMed]
- Cremer, T.; Cremer, M.; Hubner, B.; Strickfaden, H.; Smeets, D.; Popken, J.; Sterr, M.; Markaki, Y.; Rippe, K.; Cremer, C. The 4D nucleome: Evidence for a dynamic nuclear landscape based on co-aligned active and inactive nuclear compartments. FEBS Lett. 2015, 589, 2931–2943. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pederson, T. Dynamics and genome-centricity of interchromatin domains in the nucleus. Nat. Cell Biol. 2002, 4, E287–E291. [Google Scholar] [CrossRef] [PubMed]
- Pascarella, A.; Ferrandino, G.; Credendino, S.C.; Moccia, C.; D’Angelo, F.; Miranda, B.; D’Ambrosio, C.; Bielli, P.; Spadaro, O.; Ceccarelli, M.; et al. DNAJC17 is localized in nuclear speckles and interacts with splicing machinery components. Sci. Rep. 2018, 8, 7794. [Google Scholar] [CrossRef] [PubMed]
- Rieder, D.; Ploner, C.; Krogsdam, A.M.; Stocker, G.; Fischer, M.; Scheideler, M.; Dani, C.; Amri, E.Z.; Muller, W.G.; McNally, J.G.; et al. Co-expressed genes prepositioned in spatial neighborhoods stochastically associate with SC35 speckles and RNA polymerase II factories. Cell. Mol. Life Sci. 2014, 71, 1741–1759. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.C.; Chang, P.Y.; Chao, C.C. CITED2 silencing sensitizes cancer cells to cisplatin by inhibiting p53 trans-activation and chromatin relaxation on the ERCC1 DNA repair gene. Nucleic Acids Res. 2015, 43, 10760–10781. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heinz, K.S.; Casas-Delucchi, C.S.; Torok, T.; Cmarko, D.; Rapp, A.; Raska, I.; Cardoso, M.C. Peripheral re-localization of constitutive heterochromatin advances its replication timing and impairs maintenance of silencing marks. Nucleic Acids Res. 2018, 46, 6112–6128. [Google Scholar] [CrossRef] [PubMed]
- Peixoto, P.; Castronovo, V.; Matheus, N.; Polese, C.; Peulen, O.; Gonzalez, A.; Boxus, M.; Verdin, E.; Thiry, M.; Dequiedt, F.; et al. HDAC5 is required for maintenance of pericentric heterochromatin, and controls cell-cycle progression and survival of human cancer cells. Cell Death Differ. 2012, 19, 1239–1252. [Google Scholar] [CrossRef] [PubMed]
- Sabari, B.R.; Dall’Agnese, A.; Boija, A.; Klein, I.A.; Coffey, E.L.; Shrinivas, K.; Abraham, B.J.; Hannett, N.M.; Zamudio, A.V.; Manteiga, J.C.; et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 2018, 361. [Google Scholar] [CrossRef] [PubMed]
- Hnisz, D.; Young, R.A. New Insights into Genome Structure: Genes of a Feather Stick Together. Mol. Cell 2017, 67, 730–731. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ji, X.; Dadon, D.B.; Powell, B.E.; Fan, Z.P.; Borges-Rivera, D.; Shachar, S.; Weintraub, A.S.; Hnisz, D.; Pegoraro, G.; Lee, T.I.; et al. 3D Chromosome Regulatory Landscape of Human Pluripotent Cells. Cell Stem Cell 2016, 18, 262–275. [Google Scholar] [CrossRef] [PubMed]
- Andricovich, J.; Perkail, S.; Kai, Y.; Casasanta, N.; Peng, W.; Tzatsos, A. Loss of KDM6A Activates Super-Enhancers to Induce Gender-Specific Squamous-like Pancreatic Cancer and Confers Sensitivity to BET Inhibitors. Cancer Cell 2018, 33, 512–526.e8. [Google Scholar] [CrossRef] [PubMed]
- Hodges, H.C.; Stanton, B.Z.; Cermakova, K.; Chang, C.Y.; Miller, E.L.; Kirkland, J.G.; Ku, W.L.; Veverka, V.; Zhao, K.; Crabtree, G.R. Dominant-negative SMARCA4 mutants alter the accessibility landscape of tissue-unrestricted enhancers. Nat. Struct. Mol. Biol. 2018, 25, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Sharakhov, I.V.; Bondarenko, S.M.; Artemov, G.N.; Onufriev, A.V. The Role of Chromosome-Nuclear Envelope Attachments in 3D Genome Organization. Biochemistry 2018, 83, 350–358. [Google Scholar] [CrossRef] [PubMed]
- De Magistris, P.; Antonin, W. The Dynamic Nature of the Nuclear Envelope. Curr. Biol. 2018, 28, R487–R497. [Google Scholar] [CrossRef] [PubMed]
- de Las Heras, J.I.; Schirmer, E.C. The nuclear envelope and cancer: A diagnostic perspective and historical overview. Adv. Exp. Med. Biol. 2014, 773, 5–26. [Google Scholar] [PubMed]
- Chow, K.H.; Factor, R.E.; Ullman, K.S. The nuclear envelope environment and its cancer connections. Nat. Rev. Cancer 2012, 12, 196–209. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- English, A.R.; Voeltz, G.K. Endoplasmic reticulum structure and interconnections with other organelles. Cold Spring Harb. Perspect. Biol. 2013, 5, a013227. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.L.; Burke, B. LINC complexes and nuclear positioning. Semin. Cell Dev. Biol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Meinke, P.; Schirmer, E.C. LINC’ing form and function at the nuclear envelope. FEBS Lett. 2015, 589, 2514–2521. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Bethmann, C.; Worth, N.F.; Davies, J.D.; Wasner, C.; Feuer, A.; Ragnauth, C.D.; Yi, Q.; Mellad, J.A.; Warren, D.T.; et al. Nesprin-1 and -2 are involved in the pathogenesis of Emery Dreifuss muscular dystrophy and are critical for nuclear envelope integrity. Hum. Mol. Genet. 2007, 16, 2816–2833. [Google Scholar] [CrossRef] [PubMed]
- Patel, J.T.; Bottrill, A.; Prosser, S.L.; Jayaraman, S.; Straatman, K.; Fry, A.M.; Shackleton, S. Mitotic phosphorylation of SUN1 loosens its connection with the nuclear lamina while the LINC complex remains intact. Nucleus 2014, 5, 462–473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Camozzi, D.; Capanni, C.; Cenni, V.; Mattioli, E.; Columbaro, M.; Squarzoni, S.; Lattanzi, G. Diverse lamin-dependent mechanisms interact to control chromatin dynamics. Focus on laminopathies. Nucleus 2014, 5, 427–440. [Google Scholar] [CrossRef] [PubMed]
- Athirasala, A.; Hirsch, N.; Buxboim, A. Nuclear mechanotransduction: Sensing the force from within. Curr. Opin. Cell Biol. 2017, 46, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Jahed, Z.; Soheilypour, M.; Peyro, M.; Mofrad, M.R. The LINC and NPC relationship—It’s complicated! J. Cell Sci. 2016, 129, 3219–3229. [Google Scholar] [CrossRef] [PubMed]
- Nikolova-Krstevski, V.; Leimena, C.; Xiao, X.H.; Kesteven, S.; Tan, J.C.; Yeo, L.S.; Yu, Z.Y.; Zhang, Q.; Carlton, A.; Head, S.; et al. Nesprin-1 and actin contribute to nuclear and cytoskeletal defects in lamin A/C-deficient cardiomyopathy. J. Mol. Cell. Cardiol. 2011, 50, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Gimpel, P.; Lee, Y.L.; Sobota, R.M.; Calvi, A.; Koullourou, V.; Patel, R.; Mamchaoui, K.; Nedelec, F.; Shackleton, S.; Schmoranzer, J.; et al. Nesprin-1alpha-Dependent Microtubule Nucleation from the Nuclear Envelope via Akap450 Is Necessary for Nuclear Positioning in Muscle Cells. Curr. Biol. 2017, 27, 2999–3009.e9. [Google Scholar] [CrossRef] [PubMed]
- Ketema, M.; Wilhelmsen, K.; Kuikman, I.; Janssen, H.; Hodzic, D.; Sonnenberg, A. Requirements for the localization of nesprin-3 at the nuclear envelope and its interaction with plectin. J. Cell Sci. 2007, 120, 3384–3394. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, W.; Schneider, M.; Neumann, S.; Jaeger, V.M.; Taranum, S.; Munck, M.; Cartwright, S.; Richardson, C.; Carthew, J.; Noh, K.; et al. Nesprin interchain associations control nuclear size. Cell. Mol. Life Sci. 2012, 69, 3493–3509. [Google Scholar] [CrossRef] [PubMed]
- Jahed, Z.; Vu, U.T.; Fadavi, D.; Ke, H.; Rathish, A.; Kim, S.C.J.; Feng, W.; Mofrad, M.R.K. A Molecular Model for LINC complex regulation: Activation of SUN2 for KASH binding. Mol. Biol. Cell 2018. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Stumpf, M.; Muller, R.; Eichinger, L.; Glockner, G.; Noegel, A.A. The function of the inner nuclear envelope protein SUN1 in mRNA export is regulated by phosphorylation. Sci. Rep. 2017, 7, 9157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kelkar, P.; Walter, A.; Papadopoulos, S.; Mross, C.; Munck, M.; Peche, V.S.; Noegel, A.A. Nesprin-2 mediated nuclear trafficking and its clinical implications. Nucleus 2015, 6, 479–489. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, P.; Noegel, A.A. Inner nuclear envelope protein SUN1 plays a prominent role in mammalian mRNA export. Nucleic Acids Res. 2015, 43, 9874–9888. [Google Scholar] [CrossRef] [PubMed]
- Imaizumi, H.; Sato, K.; Nishihara, A.; Minami, K.; Koizumi, M.; Matsuura, N.; Hieda, M. X-ray-enhanced cancer cell migration requires the linker of nucleoskeleton and cytoskeleton complex. Cancer Sci 2018, 109, 1158–1165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Infante, E.; Castagnino, A.; Ferrari, R.; Monteiro, P.; Aguera-Gonzalez, S.; Paul-Gilloteaux, P.; Domingues, M.J.; Maiuri, P.; Raab, M.; Shanahan, C.M.; et al. LINC complex-Lis1 interplay controls MT1-MMP matrix digest-on-demand response for confined tumor cell migration. Nat. Commun. 2018, 9, 2443. [Google Scholar] [CrossRef] [PubMed]
- Isermann, P.; Lammerding, J. Nuclear mechanics and mechanotransduction in health and disease. Curr. Biol. 2013, 23, R1113–R1121. [Google Scholar] [CrossRef] [PubMed]
- Clever, M.; Mimura, Y.; Funakoshi, T.; Imamoto, N. Regulation and coordination of nuclear envelope and nuclear pore complex assembly. Nucleus 2013, 4, 105–114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, Y.; Kim, Y.; Shimi, T.; Goldman, R.D.; Zheng, Y. Concentration-dependent lamin assembly and its roles in the localization of other nuclear proteins. Mol. Biol. Cell 2014, 25, 1287–1297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Osmanagic-Myers, S.; Dechat, T.; Foisner, R. Lamins at the crossroads of mechanosignaling. Genes Dev. 2015, 29, 225–237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harr, J.C.; Luperchio, T.R.; Wong, X.; Cohen, E.; Wheelan, S.J.; Reddy, K.L. Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J. Cell Biol. 2015, 208, 33–52. [Google Scholar] [CrossRef] [PubMed]
- Zullo, J.M.; Demarco, I.A.; Pique-Regi, R.; Gaffney, D.J.; Epstein, C.B.; Spooner, C.J.; Luperchio, T.R.; Bernstein, B.E.; Pritchard, J.K.; Reddy, K.L.; et al. DNA sequence-dependent compartmentalization and silencing of chromatin at the nuclear lamina. Cell 2012, 149, 1474–1487. [Google Scholar] [CrossRef] [PubMed]
- Pope, B.D.; Ryba, T.; Dileep, V.; Yue, F.; Wu, W.; Denas, O.; Vera, D.L.; Wang, Y.; Hansen, R.S.; Canfield, T.K.; et al. Topologically associating domains are stable units of replication-timing regulation. Nature 2014, 515, 402–405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turgay, Y.; Eibauer, M.; Goldman, A.E.; Shimi, T.; Khayat, M.; Ben-Harush, K.; Dubrovsky-Gaupp, A.; Sapra, K.T.; Goldman, R.D.; Medalia, O. The molecular architecture of lamins in somatic cells. Nature 2017, 543, 261–264. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimi, T.; Kittisopikul, M.; Tran, J.; Goldman, A.E.; Adam, S.A.; Zheng, Y.; Jaqaman, K.; Goldman, R.D. Structural organization of nuclear lamins A, C, B1, and B2 revealed by superresolution microscopy. Mol. Biol. Cell 2015, 26, 4075–4086. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ungricht, R.; Kutay, U. Establishment of NE asymmetry-targeting of membrane proteins to the inner nuclear membrane. Curr. Opin. Cell Biol. 2015, 34, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Lammerding, J.; Fong, L.G.; Ji, J.Y.; Reue, K.; Stewart, C.L.; Young, S.G.; Lee, R.T. Lamins A and C but not lamin B1 regulate nuclear mechanics. J. Biol. Chem. 2006, 281, 25768–25780. [Google Scholar] [CrossRef] [PubMed]
- Harada, T.; Swift, J.; Irianto, J.; Shin, J.W.; Spinler, K.R.; Athirasala, A.; Diegmiller, R.; Dingal, P.C.; Ivanovska, I.L.; Discher, D.E. Nuclear lamin stiffness is a barrier to 3D migration, but softness can limit survival. J. Cell Biol. 2014, 204, 669–682. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stephens, A.D.; Banigan, E.J.; Adam, S.A.; Goldman, R.D.; Marko, J.F. Chromatin and lamin A determine two different mechanical response regimes of the cell nucleus. Mol. Biol. Cell 2017, 28, 1984–1996. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guilluy, C.; Osborne, L.D.; Van Landeghem, L.; Sharek, L.; Superfine, R.; Garcia-Mata, R.; Burridge, K. Isolated nuclei adapt to force and reveal a mechanotransduction pathway in the nucleus. Nat. Cell Biol. 2014, 16, 376–381. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pagliara, S.; Franze, K.; McClain, C.R.; Wylde, G.; Fisher, C.L.; Franklin, R.J.M.; Kabla, A.J.; Keyser, U.F.; Chalut, K.J. Auxetic nuclei in embryonic stem cells exiting pluripotency. Nat. Mater. 2014, 13, 638–644. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thiam, H.R.; Vargas, P.; Carpi, N.; Crespo, C.L.; Raab, M.; Terriac, E.; King, M.C.; Jacobelli, J.; Alberts, A.S.; Stradal, T.; et al. Perinuclear Arp2/3-driven actin polymerization enables nuclear deformation to facilitate cell migration through complex environments. Nat. Commun. 2016, 7, 10997. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shah, P.; Wolf, K.; Lammerding, J. Bursting the Bubble—Nuclear Envelope Rupture as a Path to Genomic Instability? Trends Cell Biol. 2017, 27, 546–555. [Google Scholar] [CrossRef] [PubMed]
- Hatch, E.M.; Fischer, A.H.; Deerinck, T.J.; Hetzer, M.W. Catastrophic nuclear envelope collapse in cancer cell micronuclei. Cell 2013, 154, 47–60. [Google Scholar] [CrossRef] [PubMed]
- Capo-Chichi, C.D.; Yeasky, T.M.; Smith, E.R.; Xu, X.X. Nuclear envelope structural defect underlies the main cause of aneuploidy in ovarian carcinogenesis. BMC Cell Biol. 2016, 17, 37. [Google Scholar] [CrossRef] [PubMed]
- Ly, P.; Cleveland, D.W. Rebuilding Chromosomes After Catastrophe: Emerging Mechanisms of Chromothripsis. Trends Cell Biol. 2017, 27, 917–930. [Google Scholar] [CrossRef] [PubMed]
- Utani, K.; Kawamoto, J.K.; Shimizu, N. Micronuclei bearing acentric extrachromosomal chromatin are transcriptionally competent and may perturb the cancer cell phenotype. Mol. Cancer Res. 2007, 5, 695–704. [Google Scholar] [CrossRef] [PubMed]
- Fischer, A.H. The diagnostic pathology of the nuclear envelope in human cancers. Adv. Exp. Med. Biol. 2014, 773, 49–75. [Google Scholar] [PubMed]
- Chang, P.; Li, Y.; Li, D. Micronuclei levels in peripheral blood lymphocytes as a potential biomarker for pancreatic cancer risk. Carcinogenesis 2011, 32, 210–215. [Google Scholar] [CrossRef] [PubMed]
- Edens, L.J.; White, K.H.; Jevtic, P.; Li, X.; Levy, D.L. Nuclear size regulation: From single cells to development and disease. Trends Cell Biol. 2013, 23, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Jevtic, P.; Edens, L.J.; Li, X.; Nguyen, T.; Chen, P.; Levy, D.L. Concentration-dependent Effects of Nuclear Lamins on Nuclear Size in Xenopus and Mammalian Cells. J. Biol. Chem. 2015, 290, 27557–27571. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, R.N.; Chen, P.; Levy, D.L. Recent advances in understanding nuclear size and shape. Nucleus 2016, 7, 167–186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Preston, C.C.; Wyles, S.P.; Reyes, S.; Storm, E.C.; Eckloff, B.W.; Faustino, R.S. NUP155 insufficiency recalibrates a pluripotent transcriptome with network remodeling of a cardiogenic signaling module. BMC Syst. Biol. 2018, 12, 62. [Google Scholar] [CrossRef] [PubMed]
- von Appen, A.; Beck, M. Structure Determination of the Nuclear Pore Complex with Three-Dimensional Cryo electron Microscopy. J. Mol. Biol. 2016, 428, 2001–2010. [Google Scholar] [CrossRef] [PubMed]
- von Appen, A.; Kosinski, J.; Sparks, L.; Ori, A.; DiGuilio, A.L.; Vollmer, B.; Mackmull, M.T.; Banterle, N.; Parca, L.; Kastritis, P.; et al. In situ structural analysis of the human nuclear pore complex. Nature 2015, 526, 140–143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Friedman, A.K.; Baker, L.A. Synthetic hydrogel mimics of the nuclear pore complex display selectivity dependent on FG-repeat concentration and electrostatics. Soft Matter 2016, 12, 9477–9484. [Google Scholar] [CrossRef] [PubMed]
- Frey, S.; Richter, R.P.; Gorlich, D. FG-rich repeats of nuclear pore proteins form a three-dimensional meshwork with hydrogel-like properties. Science 2006, 314, 815–817. [Google Scholar] [CrossRef] [PubMed]
- Yoneda, Y.; Hieda, M.; Nagoshi, E.; Miyamoto, Y. Nucleocytoplasmic protein transport and recycling of Ran. Cell Struct. Funct. 1999, 24, 425–433. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.A. Interplay between nuclear transport and ubiquitin/SUMO modifications in the regulation of cancer-related proteins. Semin. Cancer Biol. 2014, 27, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Mahipal, A.; Malafa, M. Importins and exportins as therapeutic targets in cancer. Pharmacol. Ther. 2016, 164, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Faustino, R.S.; Nelson, T.J.; Terzic, A.; Perez-Terzic, C. Nuclear transport: Target for therapy. Clin. Pharmacol. Ther. 2007, 81, 880–886. [Google Scholar] [CrossRef] [PubMed]
- Yan, W.Q.; Du, J.; Jiang, H.; Hou, J. Effect of nuclear receptor inhibitor importazole on the proliferation and apoptosis of multiple myeloma cells. Zhonghua Xue Ye Xue Za Zhi 2013, 34, 323–326. [Google Scholar] [PubMed]
- Soderholm, J.F.; Bird, S.L.; Kalab, P.; Sampathkumar, Y.; Hasegawa, K.; Uehara-Bingen, M.; Weis, K.; Heald, R. Importazole, a small molecule inhibitor of the transport receptor importin-beta. ACS Chem. Biol. 2011, 6, 700–708. [Google Scholar] [CrossRef] [PubMed]
- Muqbil, I.; Azmi, A.S.; Mohammad, R.M. Nuclear Export Inhibition for Pancreatic Cancer Therapy. Cancers 2018, 10, 138. [Google Scholar] [CrossRef] [PubMed]
- Azmi, A.S.; Li, Y.; Muqbil, I.; Aboukameel, A.; Senapedis, W.; Baloglu, E.; Landesman, Y.; Shacham, S.; Kauffman, M.G.; Philip, P.A.; et al. Exportin 1 (XPO1) inhibition leads to restoration of tumor suppressor miR-145 and consequent suppression of pancreatic cancer cell proliferation and migration. Oncotarget 2017, 8, 82144–82155. [Google Scholar] [CrossRef] [PubMed]
- Kashyap, T.; Argueta, C.; Aboukameel, A.; Unger, T.J.; Klebanov, B.; Mohammad, R.M.; Muqbil, I.; Azmi, A.S.; Drolen, C.; Senapedis, W.; et al. Selinexor, a Selective Inhibitor of Nuclear Export (SINE) compound, acts through NF-kappaB deactivation and combines with proteasome inhibitors to synergistically induce tumor cell death. Oncotarget 2016, 7, 78883–78895. [Google Scholar] [CrossRef] [PubMed]
- Kazim, S.; Malafa, M.P.; Coppola, D.; Husain, K.; Zibadi, S.; Kashyap, T.; Crochiere, M.; Landesman, Y.; Rashal, T.; Sullivan, D.M.; et al. Selective Nuclear Export Inhibitor KPT-330 Enhances the Antitumor Activity of Gemcitabine in Human Pancreatic Cancer. Mol. Cancer Ther. 2015, 14, 1570–1581. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laurila, E.; Vuorinen, E.; Savinainen, K.; Rauhala, H.; Kallioniemi, A. KPNA7, a nuclear transport receptor, promotes malignant properties of pancreatic cancer cells in vitro. Exp. Cell Res. 2014, 322, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Vuorinen, E.M.; Rajala, N.K.; Rauhala, H.E.; Nurminen, A.T.; Hytonen, V.P.; Kallioniemi, A. Search for KPNA7 cargo proteins in human cells reveals MVP and ZNF414 as novel regulators of cancer cell growth. Biochim. Biophys. Acta 2017, 1863, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Vuorinen, E.M.; Rajala, N.K.; Ihalainen, T.O.; Kallioniemi, A. Depletion of nuclear import protein karyopherin alpha 7 (KPNA7) induces mitotic defects and deformation of nuclei in cancer cells. BMC Cancer 2018, 18, 325. [Google Scholar] [CrossRef] [PubMed]
- Fornerod, M.; Boer, J.; van Baal, S.; Jaegle, M.; von Lindern, M.; Murti, K.G.; Davis, D.; Bonten, J.; Buijs, A.; Grosveld, G. Relocation of the carboxyterminal part of CAN from the nuclear envelope to the nucleus as a result of leukemia-specific chromosome rearrangements. Oncogene 1995, 10, 1739–1748. [Google Scholar] [PubMed]
- Nakamura, T.; Largaespada, D.A.; Lee, M.P.; Johnson, L.A.; Ohyashiki, K.; Toyama, K.; Chen, S.J.; Willman, C.L.; Chen, I.M.; Feinberg, A.P.; et al. Fusion of the nucleoporin gene NUP98 to HOXA9 by the chromosome translocation t(7;11)(p15;p15) in human myeloid leukaemia. Nat. Genet. 1996, 12, 154–158. [Google Scholar] [CrossRef] [PubMed]
- Takeda, A.; Yaseen, N.R. Nucleoporins and nucleocytoplasmic transport in hematologic malignancies. Semin. Cancer Biol. 2014, 27, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Makise, M.; Nakamura, H.; Kuniyasu, A. The role of vimentin in the tumor marker Nup88-dependent multinucleated phenotype. BMC Cancer 2018, 18, 519. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhao, J.; Li, Y. Multiple biological processes may be associated with tumorigenesis under NUP88-overexpressed condition. Genes Chromosomes Cancer 2017, 56, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Naylor, R.M.; Jeganathan, K.B.; Cao, X.; van Deursen, J.M. Nuclear pore protein NUP88 activates anaphase-promoting complex to promote aneuploidy. J. Clin. Investig. 2016, 126, 543–559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simon, D.N.; Rout, M.P. Cancer and the nuclear pore complex. Adv. Exp. Med. Biol. 2014, 773, 285–307. [Google Scholar] [PubMed]
- Shimozono, N.; Jinnin, M.; Masuzawa, M.; Masuzawa, M.; Wang, Z.; Hirano, A.; Tomizawa, Y.; Etoh-Kira, T.; Kajihara, I.; Harada, M.; et al. NUP160-SLC43A3 is a novel recurrent fusion oncogene in angiosarcoma. Cancer Res. 2015, 75, 4458–4465. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Liu, Y.; Feng, W.; Lei, L.; Du, Y.; Wu, J.; Wang, S. NUP37, a positive regulator of YAP/TEAD signaling, promotes the progression of hepatocellular carcinoma. Oncotarget 2017, 8, 98004–98013. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.R.; Zhang, L.J.; Wang, Y.Y.; Li, F.; Wang, M.W.; Sun, X.F. Increased serum level of Nup88 protein is associated with the development of colorectal cancer. Med. Oncol. 2012, 29, 1789–1795. [Google Scholar] [CrossRef] [PubMed]
- Mendez, M.G.; Kojima, S.; Goldman, R.D. Vimentin induces changes in cell shape, motility, and adhesion during the epithelial to mesenchymal transition. FASEB J. 2010, 24, 1838–1851. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, X.; Carstens, J.L.; Kim, J.; Scheible, M.; Kaye, J.; Sugimoto, H.; Wu, C.C.; LeBleu, V.S.; Kalluri, R. Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature 2015, 527, 525–530. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hayashi, A.; Tanoshima, R.; Tsujimoto, S.I.; Yanagimachi, M.; Takeuchi, M.; Sasaki, K.; Ikeda, J.; Kajiwara, R.; Ito, S.; Takahashi, H. Crizotinib treatment for refractory pediatric acute myeloid leukemia with RAN-binding protein 2-anaplastic lymphoma kinase fusion gene. Blood Cancer J. 2016, 6, e456. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, Y.; Wang, B.; Lan, H.; Liu, Y.; Chen, F.; Zhang, J.; Luo, J. Sumoylation in p27kip1 via RanBP2 promotes cancer cell growth in cholangiocarcinoma cell line QBC939. BMC Mol. Biol. 2017, 18, 23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ashikari, D.; Takayama, K.; Tanaka, T.; Suzuki, Y.; Obinata, D.; Fujimura, T.; Urano, T.; Takahashi, S.; Inoue, S. Androgen induces G3BP2 and SUMO-mediated p53 nuclear export in prostate cancer. Oncogene 2017, 36, 6272–6281. [Google Scholar] [CrossRef] [PubMed]
- Vecchione, L.; Gambino, V.; Raaijmakers, J.; Schlicker, A.; Fumagalli, A.; Russo, M.; Villanueva, A.; Beerling, E.; Bartolini, A.; Mollevi, D.G.; et al. A Vulnerability of a Subset of Colon Cancers with Potential Clinical Utility. Cell 2016, 165, 317–330. [Google Scholar] [CrossRef] [PubMed]
- Snow, C.J.; Paschal, B.M. Roles of the nucleoporin Tpr in cancer and aging. Adv. Exp. Med. Biol. 2014, 773, 309–322. [Google Scholar] [PubMed]
- Neureiter, D.; Jager, T.; Ocker, M.; Kiesslich, T. Epigenetics and pancreatic cancer: Pathophysiology and novel treatment aspects. World J. Gastroenterol. 2014, 20, 7830–7848. [Google Scholar] [CrossRef] [PubMed]
- Kind, J.; Pagie, L.; Ortabozkoyun, H.; Boyle, S.; de Vries, S.S.; Janssen, H.; Amendola, M.; Nolen, L.D.; Bickmore, W.A.; van Steensel, B. Single-cell dynamics of genome-nuclear lamina interactions. Cell 2013, 153, 178–192. [Google Scholar] [CrossRef] [PubMed]
- Kind, J.; Pagie, L.; de Vries, S.S.; Nahidiazar, L.; Dey, S.S.; Bienko, M.; Zhan, Y.; Lajoie, B.; de Graaf, C.A.; Amendola, M.; et al. Genome-wide maps of nuclear lamina interactions in single human cells. Cell 2015, 163, 134–147. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Du, Y.; Kong, X.; Li, Z.; Jia, Z.; Cui, J.; Gao, J.; Wang, G.; Xie, K. Lamin B1 is a novel therapeutic target of betulinic acid in pancreatic cancer. Clin. Cancer Res. 2013, 19, 4651–4661. [Google Scholar] [CrossRef] [PubMed]
- Camps, J.; Wangsa, D.; Falke, M.; Brown, M.; Case, C.M.; Erdos, M.R.; Ried, T. Loss of lamin B1 results in prolongation of S phase and decondensation of chromosome territories. FASEB J. 2014, 28, 3423–3434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Somech, R.; Shaklai, S.; Geller, O.; Amariglio, N.; Simon, A.J.; Rechavi, G.; Gal-Yam, E.N. The nuclear-envelope protein and transcriptional repressor LAP2beta interacts with HDAC3 at the nuclear periphery, and induces histone H4 deacetylation. J. Cell Sci. 2005, 118, 4017–4025. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.G.; Song, J.; Wang, Z.; Dormann, H.L.; Casadio, F.; Li, H.; Luo, J.L.; Patel, D.J.; Allis, C.D. Haematopoietic malignancies caused by dysregulation of a chromatin-binding PHD finger. Nature 2009, 459, 847–851. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franks, T.M.; McCloskey, A.; Shokirev, M.N.; Benner, C.; Rathore, A.; Hetzer, M.W. Nup98 recruits the Wdr82-Set1A/COMPASS complex to promoters to regulate H3K4 trimethylation in hematopoietic progenitor cells. Genes Dev. 2017, 31, 2222–2234. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Yang, D.; Sabbatini, M.E.; Colby, A.H.; Grinstaff, M.W.; Oberlies, N.H.; Pearce, C.; Liu, K. Contrasting roles of H3K4me3 and H3K9me3 in regulation of apoptosis and gemcitabine resistance in human pancreatic cancer cells. BMC Cancer 2018, 18, 149. [Google Scholar] [CrossRef] [PubMed]
- Lomberk, G.A.; Urrutia, R. The Triple-Code Model for Pancreatic Cancer: Cross Talk Among Genetics, Epigenetics, and Nuclear Structure. Surg. Clin. N. Am. 2015, 95, 935–952. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A.; Hieda, M.; Yokoyama, Y.; Nishioka, Y.; Yoshidome, K.; Tsujimoto, M.; Matsuura, N. Global loss of a nuclear lamina component, lamin A/C, and LINC complex components SUN1, SUN2, and nesprin-2 in breast cancer. Cancer Med. 2015, 4, 1547–1557. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yajun, C.; Chen, Y.; Xiaosa, L.; Xiao, W.; Jia, C.; Zhong, W.; Bin, X. Loss of Sun2 promotes the progression of prostate cancer by regulating fatty acid oxidation. Oncotarget 2017, 8, 89620–89630. [Google Scholar] [CrossRef] [PubMed]
- Aymard, F.; Aguirrebengoa, M.; Guillou, E.; Javierre, B.M.; Bugler, B.; Arnould, C.; Rocher, V.; Iacovoni, J.S.; Biernacka, A.; Skrzypczak, M.; et al. Genome-wide mapping of long-range contacts unveils clustering of DNA double-strand breaks at damaged active genes. Nat. Struct. Mol. Biol. 2017, 24, 353–361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Connor, A.A.; Denroche, R.E.; Jang, G.H.; Timms, L.; Kalimuthu, S.N.; Selander, I.; McPherson, T.; Wilson, G.W.; Chan-Seng-Yue, M.A.; Borozan, I.; et al. Association of Distinct Mutational Signatures With Correlates of Increased Immune Activity in Pancreatic Ductal Adenocarcinoma. JAMA Oncol. 2017, 3, 774–783. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Nakano, K.; Obchoei, S.; Setoguchi, K.; Matsumoto, M.; Yamamoto, T.; Obika, S.; Shimada, K.; Hiraoka, N. Small Nucleolar Noncoding RNA SNORA23, Up-Regulated in Human Pancreatic Ductal Adenocarcinoma, Regulates Expression of Spectrin Repeat-Containing Nuclear Envelope 2 to Promote Growth and Metastasis of Xenograft Tumors in Mice. Gastroenterology 2017, 153, 292–306.e2. [Google Scholar] [CrossRef] [PubMed]
- Janin, A.; Bauer, D.; Ratti, F.; Millat, G.; Mejat, A. Nuclear envelopathies: A complex LINC between nuclear envelope and pathology. Orphanet J. Rare Dis. 2017, 12, 147. [Google Scholar] [CrossRef] [PubMed]
- Cappello, P.; Curcio, C.; Mandili, G.; Roux, C.; Bulfamante, S.; Novelli, F. Next Generation Immunotherapy for Pancreatic Cancer: DNA Vaccination is Seeking New Combo Partners. Cancers 2018, 10, 51. [Google Scholar] [CrossRef] [PubMed]
- Hilmi, M.; Bartholin, L.; Neuzillet, C. Immune therapies in pancreatic ductal adenocarcinoma: Where are we now? World J. Gastroenterol. 2018, 24, 2137–2151. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Saif, M.W. T cell optimization for the treatment of pancreatic cancer. Expert Opin. Biol. Ther. 2017, 17, 1493–1501. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.W.; Kochenderfer, J.N.; Stetler-Stevenson, M.; Cui, Y.K.; Delbrook, C.; Feldman, S.A.; Fry, T.J.; Orentas, R.; Sabatino, M.; Shah, N.N.; et al. T cells expressing CD19 chimeric antigen receptors for acute lymphoblastic leukaemia in children and young adults: A phase 1 dose-escalation trial. Lancet 2015, 385, 517–528. [Google Scholar] [CrossRef]
- Kochenderfer, J.N.; Dudley, M.E.; Kassim, S.H.; Somerville, R.P.; Carpenter, R.O.; Stetler-Stevenson, M.; Yang, J.C.; Phan, G.Q.; Hughes, M.S.; Sherry, R.M.; et al. Chemotherapy-refractory diffuse large B-cell lymphoma and indolent B-cell malignancies can be effectively treated with autologous T cells expressing an anti-CD19 chimeric antigen receptor. J. Clin. Oncol. 2015, 33, 540–549. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, A.; Mahalingam, D. Immunotherapy in pancreatic adenocarcinoma-overcoming barriers to response. J. Gastrointest. Oncol. 2018, 9, 143–159. [Google Scholar] [CrossRef] [PubMed]
| Types of Mutation | Gene Symbol | Gene Description | Entrez Gene ID | Ensembl Gene ID |
|---|---|---|---|---|
| Somatic | AKT2 | AKT serine/threonine kinase 2 | 208 | ENSG00000105221 |
| ARID1A | AT-rich interaction domain 1A | 8289 | ENSG00000117713 | |
| BRAF | B-Raf proto-oncogene, serine/threonine kinase | 673 | ENSG00000157764 | |
| CCSER1/FAM190A | Coiled-coil serine rich protein 1 | 401,145 | ENSG00000184305 | |
| CDKN2A | Cyclin dependent kinase inhibitor 2A | 1029 | ENSG00000147889 | |
| EP300 | E1A binding protein p300 | 2033 | ENSG00000100393 | |
| KMT2C/MLL3 | Lysine methyltransferase 2C | 58,508 | ENSG00000055609 | |
| KRAS | KRAS proto-oncogene, GTPase | 3845 | ENSG00000133703 | |
| MYB | MYB proto-oncogene, transcription factor | 4602 | ENSG00000118513 | |
| NCOA3/AIB1 | Nuclear receptor coactivator 3 | 8202 | ENSG00000124151 | |
| SMAD4 | SMAD family member 4 | 4089 | ENSG00000141646 | |
| TGFBR2 | Transforming growth factor beta receptor 2 | 7048 | ENSG00000163513 | |
| TP53 | Tumor protein p53 | 7157 | ENSG00000141510 | |
| USP9X | Ubiquitin specific peptidase 9, X-linked | 8239 | ENSG00000124486 | |
| Germline | ATM | ATM serine/threonine kinase | 472 | ENSG00000149311 |
| BRCA2 | BRCA2, DNA repair associated | 675 | ENSG00000139618 | |
| CDKN2A | Cyclin dependent kinase inhibitor 2A | 1029 | ENSG00000147889 | |
| PALB2 | Partner and localizer of BRCA2 | 79,728 | ENSG00000083093 | |
| PRSS1 | Protease, serine 1 | 5644 | ENSG00000204983 | |
| STK11 | Serine/threonine kinase 11 | 6794 | ENSG00000118046 |
| Cluster No. (E Score) | Database | Functional Term | Fold Enrichment | p Value | Benjamini |
|---|---|---|---|---|---|
| Cluster 1 (3.12) | UP_KEYWORDS | Acetylation | 4.723 | 0.0000028 | 0.0002505 |
| KEGG_PATHWAY | Cell cycle | 20.264 | 0.0006173 | 0.0033905 | |
| UP_KEYWORDS | Transcription | 4.904 | 0.0002617 | 0.0046472 | |
| UP_KEYWORDS | Transcription regulation | 5.043 | 0.0002192 | 0.0048656 | |
| GOTERM_BP_DIRECT | Positive regulation of transcription, DNA-templated | 15.049 | 0.0000176 | 0.0066071 | |
| BIOCARTA | First Multivalent Nuclear Factor | 28.889 | 0.0001714 | 0.0068348 | |
| UP_KEYWORDS | Nucleus | 2.803 | 0.0011438 | 0.0101343 | |
| UP_KEYWORDS | Phosphoprotein | 2.139 | 0.0013327 | 0.0107323 | |
| GOTERM_CC_DIRECT | Nuclear chromatin | 29.054 | 0.0002398 | 0.0112081 | |
| GOTERM_CC_DIRECT | Nucleoplasm | 4.028 | 0.0007497 | 0.0174706 | |
| GOTERM_CC_DIRECT | Nucleus | 2.589 | 0.0015626 | 0.0242026 | |
| GOTERM_MF_DIRECT | DNA binding | 5.430 | 0.0005138 | 0.0263695 | |
| Cluster 2 (2.95) | KEGG_PATHWAY | FoxO signaling pathway | 28.128 | 0.0000006 | 0.0000098 |
| KEGG_PATHWAY | Adherens junction | 26.543 | 0.0044414 | 0.0168064 | |
| KEGG_PATHWAY | TGF-beta signaling pathway | 22.435 | 0.0061682 | 0.0224314 | |
| Cluster 3 (2.86) | KEGG_PATHWAY | Chronic myeloid leukemia | 61.073 | 0.0000000 | 0.0000000 |
| KEGG_PATHWAY | Pancreatic cancer | 67.650 | 0.0000000 | 0.0000000 | |
| KEGG_PATHWAY | Colorectal cancer | 60.792 | 0.0000000 | 0.0000003 | |
| KEGG_PATHWAY | HTLV-I infection | 19.631 | 0.0000000 | 0.0000003 | |
| KEGG_PATHWAY | Pathways in cancer | 12.787 | 0.0000002 | 0.0000037 | |
| KEGG_PATHWAY | FoxO signaling pathway | 28.128 | 0.0000006 | 0.0000098 | |
| KEGG_PATHWAY | Non-small cell lung cancer | 56.088 | 0.0000008 | 0.0000111 | |
| KEGG_PATHWAY | Glioma | 48.322 | 0.0000014 | 0.0000178 | |
| KEGG_PATHWAY | Melanoma | 44.238 | 0.0000021 | 0.0000226 | |
| KEGG_PATHWAY | Prostate cancer | 35.692 | 0.0000049 | 0.0000482 | |
| KEGG_PATHWAY | Thyroid hormone signaling pathway | 27.552 | 0.0000137 | 0.0001230 | |
| KEGG_PATHWAY | Bladder cancer | 61.286 | 0.0000226 | 0.0001865 | |
| KEGG_PATHWAY | Hepatitis B | 21.661 | 0.0000354 | 0.0002697 | |
| KEGG_PATHWAY | Endometrial cancer | 48.322 | 0.0000465 | 0.0003286 | |
| KEGG_PATHWAY | Renal cell carcinoma | 38.657 | 0.0000910 | 0.0006002 | |
| UP_KEYWORDS | Disease mutation | 5.188 | 0.0000396 | 0.0011754 | |
| KEGG_PATHWAY | MAPK signaling pathway | 12.317 | 0.0003191 | 0.0019727 | |
| KEGG_PATHWAY | Neurotrophin signaling pathway | 20.939 | 0.0005607 | 0.0032611 | |
| KEGG_PATHWAY | Hepatitis C | 18.893 | 0.0007577 | 0.0037452 | |
| KEGG_PATHWAY | Thyroid cancer | 64.984 | 0.0007496 | 0.0038995 | |
| UP_KEYWORDS | Proto-oncogene | 24.916 | 0.0003911 | 0.0043423 | |
| KEGG_PATHWAY | Proteoglycans in cancer | 12.564 | 0.0024661 | 0.0115730 | |
| KEGG_PATHWAY | Viral carcinogenesis | 12.257 | 0.0026465 | 0.0118544 | |
| KEGG_PATHWAY | Acute myeloid leukemia | 33.653 | 0.0027845 | 0.0119306 | |
| KEGG_PATHWAY | Central carbon metabolism in cancer | 29.446 | 0.0036227 | 0.0148594 | |
| KEGG_PATHWAY | Long-term potentiation | 28.554 | 0.0038486 | 0.0151538 | |
| UP_SEQ_FEATURE | Mutagenesis site | 5.233 | 0.0001733 | 0.0202413 | |
| KEGG_PATHWAY | MicroRNAs in cancer | 8.817 | 0.0067131 | 0.0227320 | |
| KEGG_PATHWAY | ErbB signaling pathway | 21.661 | 0.0066041 | 0.0231553 | |
| KEGG_PATHWAY | Progesterone-mediated oocyte maturation | 21.661 | 0.0066041 | 0.0231553 | |
| GOTERM_CC_DIRECT | Cytosol | 3.383 | 0.0021984 | 0.0255285 | |
| UP_KEYWORDS | Isopeptide bond | 6.493 | 0.0043625 | 0.0274111 | |
| UP_KEYWORDS | Apoptosis | 10.971 | 0.0041342 | 0.0279639 | |
| KEGG_PATHWAY | PI3K-Akt signaling pathway | 7.283 | 0.0113878 | 0.0370899 | |
| KEGG_PATHWAY | Sphingolipid signaling pathway | 15.705 | 0.0122869 | 0.0387127 | |
| UP_KEYWORDS | Kinase | 8.000 | 0.0099232 | 0.0434085 | |
| KEGG_PATHWAY | Signaling pathways regulating pluripotency of stem cells | 13.461 | 0.0164870 | 0.0486500 | |
| KEGG_PATHWAY | Insulin signaling pathway | 13.656 | 0.0160424 | 0.0488026 | |
| Cluster 4 (2.60) | KEGG_PATHWAY | Cell cycle | 20.264 | 0.0006173 | 0.0033905 |
| KEGG_PATHWAY | Viral carcinogenesis | 12.257 | 0.0026465 | 0.0118544 | |
| KEGG_PATHWAY | MicroRNAs in cancer | 8.817 | 0.0067131 | 0.0227320 | |
| GOTERM_MF_DIRECT | p53 binding | 58.144 | 0.0009983 | 0.0340310 | |
| GOTERM_BP_DIRECT | Apoptotic process | 11.391 | 0.0005128 | 0.0470735 | |
| Cluster 5 (2.58) | KEGG_PATHWAY | Hepatitis B | 21.661 | 0.0000354 | 0.0002697 |
| KEGG_PATHWAY | Cell cycle | 20.264 | 0.0006173 | 0.0033905 | |
| UP_KEYWORDS | Ubl conjugation | 6.035 | 0.0003300 | 0.0048835 | |
| GOTERM_BP_DIRECT | Negative regulation of transcription from RNA polymerase II promoter | 10.764 | 0.0000880 | 0.0164117 | |
| UP_KEYWORDS | Isopeptide bond | 6.493 | 0.0043625 | 0.0274111 | |
| GOTERM_BP_DIRECT | Positive regulation of transcription from RNA polymerase II promoter | 7.900 | 0.0003776 | 0.0462357 | |
| KEGG_PATHWAY | Wnt signaling pathway | 13.656 | 0.0160424 | 0.0488026 | |
| Cluster 6 (1.94) | UP_KEYWORDS | Acyltransferase | 25.791 | 0.0050407 | 0.0295385 |
| Cluster No. (E Score) | Database | Functional Term | Fold Enrichment | p Value | Benjamini |
|---|---|---|---|---|---|
| Cluster 1 (4.25) | GOTERM_BP_DIRECT | Strand displacement | 322.923 | 0.0000230 | 0.0038316 |
| GOTERM_BP_DIRECT | DNA synthesis involved in DNA repair | 239.886 | 0.0000420 | 0.0035042 | |
| GOTERM_BP_DIRECT | Double-strand break repair via homologous recombination | 113.459 | 0.0001900 | 0.0105193 | |
| Cluster 2 (2.74) | UP_KEYWORDS | Tumor suppressor | 96.897 | 0.0000000 | 0.0000015 |
| GOTERM_CC_DIRECT | Nucleoplasm | 5.455 | 0.0023864 | 0.0557288 | |
| Cluster 3 (1.76) | GOTERM_BP_DIRECT | Cell cycle arrest | 59.546 | 0.0006886 | 0.0283490 |
| Cluster 4 (1.72) | UP_KEYWORDS | Cell cycle | 21.109 | 0.0002990 | 0.0056659 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Preston, C.C.; Faustino, R.S. Nuclear Envelope Regulation of Oncogenic Processes: Roles in Pancreatic Cancer. Epigenomes 2018, 2, 15. https://0-doi-org.brum.beds.ac.uk/10.3390/epigenomes2030015
Preston CC, Faustino RS. Nuclear Envelope Regulation of Oncogenic Processes: Roles in Pancreatic Cancer. Epigenomes. 2018; 2(3):15. https://0-doi-org.brum.beds.ac.uk/10.3390/epigenomes2030015
Chicago/Turabian StylePreston, Claudia C., and Randolph S. Faustino. 2018. "Nuclear Envelope Regulation of Oncogenic Processes: Roles in Pancreatic Cancer" Epigenomes 2, no. 3: 15. https://0-doi-org.brum.beds.ac.uk/10.3390/epigenomes2030015





