Antimicrobial Resistance of Non-O157 Shiga Toxin-Producing Escherichia coli Isolated from Humans and Domestic Animals
Abstract
:1. Introduction
2. Results
2.1. The Distribution of O:H Serotypes and Stx Subtypes
2.2. Antimicrobial Susceptibility
2.3. Antimicrobial Resistance Genes
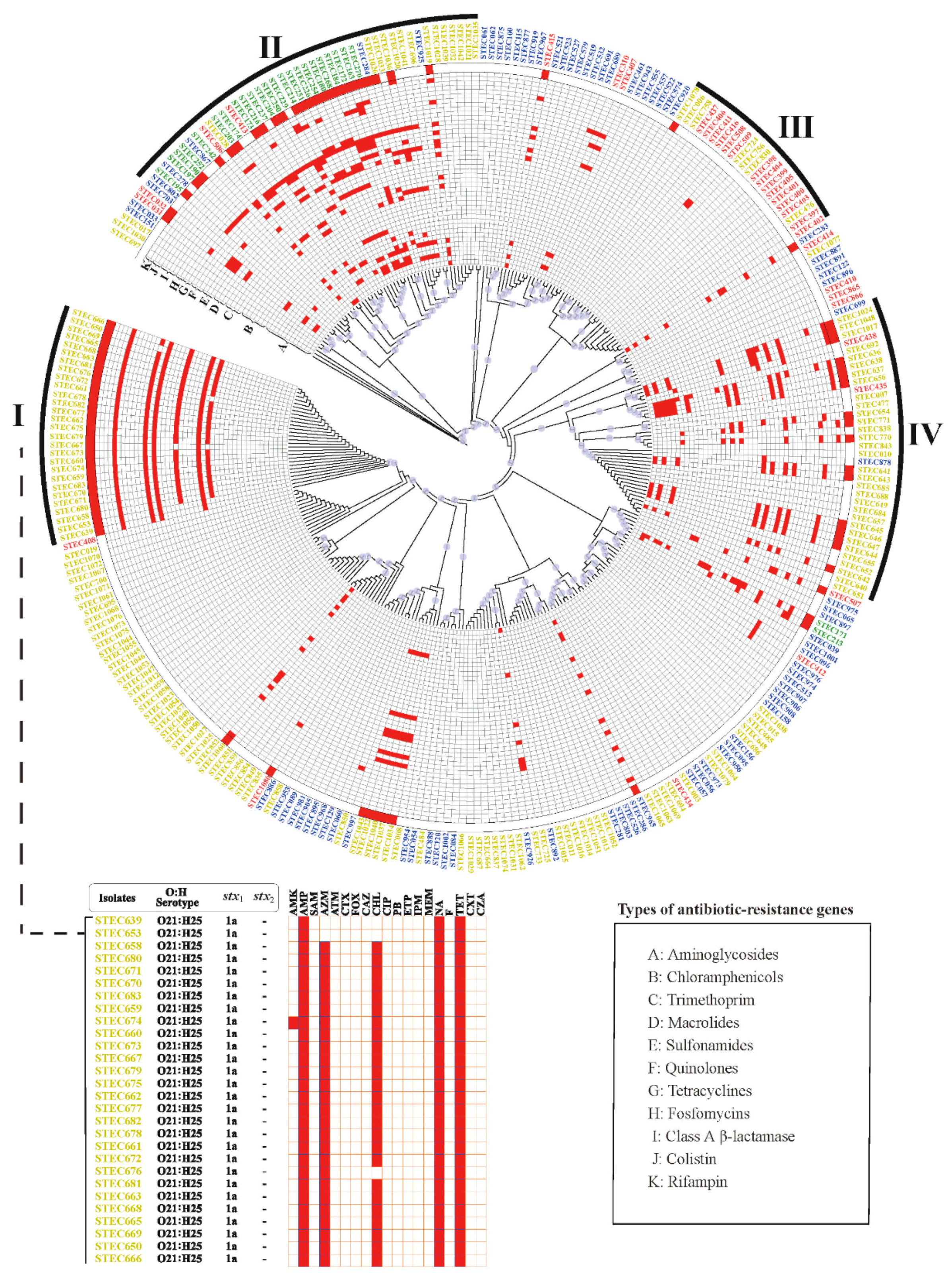
2.4. Phylogenetic Analysis
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains
4.2. Antimicrobial Susceptibility Testing
4.3. Whole Genome Sequencing (WGS)
4.4. In Silico O:H Serotyping, Stx Subtyping and AMR Gene Screening
4.5. SNP-Based Phylogenetic Analyses
4.6. Statistical Analysis
4.7. Data Availability
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Newell, D.G.; La Ragione, R.M. Enterohaemorrhagic and other Shiga toxin-producing Escherichia coli (STEC): Where are we now regarding diagnostics and control strategies? Transbound. Emerg. Dis. 2018, 65 (Suppl. 1), 49–71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mughini-Gras, L.; van Pelt, W.; van der Voort, M.; Heck, M.; Friesema, I.; Franz, E. Attribution of human infections with Shiga toxin-producing Escherichia coli (STEC) to livestock sources and identification of source-specific risk factors, The Netherlands (2010–2014). Zoonoses Public Health 2018, 65. [Google Scholar] [CrossRef] [PubMed]
- Heiman, K.E.; Mody, R.K.; Johnson, S.D.; Griffin, P.M.; Gould, L.H. Escherichia coli O157 outbreaks in the United States, 2003–2012. Emerg. Infect. Dis. 2015, 21, 1293–1301. [Google Scholar] [CrossRef] [PubMed]
- Valilis, E.; Ramsey, A.; Sidiq, S.; DuPont, H.L. Non-O157 Shiga toxin-producing Escherichia coli—A poorly appreciated enteric pathogen: Systematic review. Int. J. Infect. Dis. 2018, 76, 82–87. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joseph, A.; Cointe, A.; Kurkdjian, P.M.; Rafat, C.; Hertig, A. Shiga toxin-associated hemolytic uremic syndrome: A narrative review. Toxins 2020, 12, 67. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kinnula, S.; Hemminki, K.; Kotilainen, H.; Ruotsalainen, E.; Tarkka, E.; Salmenlinna, S.; Hallanvuo, S.; Leinonen, E.; Jukka, O.; Rimhanen-Finne, R. Outbreak of multiple strains of non-O157 Shiga toxin-producing and enteropathogenic associated with rocket salad, Finland, autumn 2016. Euro Surveill. 2018, 23. [Google Scholar] [CrossRef] [Green Version]
- Baker, K.S.; Dallman, T.J.; Thomson, N.R.; Jenkins, C. An outbreak of a rare Shiga-toxin-producing Escherichia coli serotype (O117:H7) among men who have sex with men. Microb. Genom. 2018, 4. [Google Scholar] [CrossRef]
- Crowe, S.J.; Bottichio, L.; Shade, L.N.; Whitney, B.M.; Corral, N.; Melius, B.; Arends, K.D.; Donovan, D.; Stone, J.; Allen, K.; et al. Shiga toxin-producing E. coli infections associated with flour. N. Engl. J. Med. 2017, 377, 2036–2043. [Google Scholar] [CrossRef]
- Kakoullis, L.; Papachristodoulou, E.; Chra, P.; Panos, G. Shiga toxin-induced haemolytic uraemic syndrome and the role of antibiotics: A global overview. J. Infect. 2019, 79, 75–94. [Google Scholar] [CrossRef]
- Melton-Celsa, A.R. Shiga toxin (Stx) classification, structure, and function. Microbiol. Spectr. 2014, 2, EHEC-0024-2013. [Google Scholar] [CrossRef] [Green Version]
- Muhlen, S.; Dersch, P. Treatment strategies for infections with Shiga toxin-producing Escherichia coli. Front. Cell. Infect. Microbiol. 2020, 10, 169. [Google Scholar] [CrossRef] [PubMed]
- Brigotti, M.; He, X.; Carnicelli, D.; Arfilli, V.; Porcellini, E.; Galassi, E.; Tazzari, P.L.; Ricci, F.; Patfield, S.A.; Testa, S.; et al. Particulate Shiga toxin 2 in blood is associated to the development of hemolytic uremic syndrome in children. Thromb. Haemost. 2020, 120, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Puño-Sarmiento, J.; Anderson, E.M.; Park, A.J.; Khursigara, C.M.; Foster, D.B. Potentiation of antibiotics by a novel antimicrobial peptide against Shiga toxin producing E. coli O157:H7. Sci. Rep. 2020, 10, 10029. [Google Scholar] [CrossRef] [PubMed]
- Berger, M.; Aijaz, I.; Berger, P.; Dobrindt, U.; Koudelka, G. Transcriptional and translational inhibitors block SOS response and Shiga toxin expression in enterohemorrhagic Escherichia coli. Sci. Rep. 2019, 9, 18777. [Google Scholar] [CrossRef] [PubMed]
- Oporto, B.; Ocejo, M.; Alkorta, M.; Marimón, J.M.; Montes, M.; Hurtado, A. Zoonotic approach to Shiga toxin-producing Escherichia coli: Integrated analysis of virulence and antimicrobial resistance in ruminants and humans. Epidemiol. Infect. 2019, 147, e164. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.S.; Lee, M.S.; Kim, J.H. Recent updates on outbreaks of Shiga toxin-producing Escherichia coli and its potential reservoirs. Front. Cell. Infect. Microbiol. 2020, 10, 273. [Google Scholar] [CrossRef]
- de Assis, D.C.S.; da Silva, T.M.L.; Brito, R.F.; da Silva, L.C.G.; Lima, W.G.; Brito, J.C.M. Shiga toxin-producing Escherichia coli (STEC) in bovine meat and meat products over the last 15 years in Brazil: A systematic review and meta-analysis. Meat Sci. 2020, 173, 108394. [Google Scholar] [CrossRef]
- Gonzalez, A.G.M.; Cerqueira, A.M.F. Shiga toxin-producing Escherichia coli in the animal reservoir and food in Brazil. J. Appl. Microbiol. 2020, 128, 1568–1582. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.-Y.; Wang, Y.; Walsh, T.R.; Yi, L.-X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Liu, J.-Y.; Liao, T.-L.; Huang, W.-C.; Liu, Y.-M.; Wu, K.-M.; Lauderdale, T.-L.; Tsai, S.-F.; Kuo, S.-C.; Kuo, H.-C. Increased mcr-1 in pathogenic Escherichia coli from diseased swine, Taiwan. J. Microbiol. Immunol. Infect. 2020, 53, 751–756. [Google Scholar] [CrossRef]
- Mariani, B.; Corbella, M.; Merla, C.; Tallarita, M.; Piralla, A.; Girello, A.; Castelli, M.; Bracchi, C.; Marone, P.; Cambieri, P. Bloodstream infections caused by Escherichia coli carrying mcr-1 gene in hospitalized patients in northern Italy from 2012 to 2018. Infection 2020, 48, 223–230. [Google Scholar] [CrossRef]
- Fan, R.; Shao, K.; Yang, X.; Bai, X.; Fu, S.; Sun, H.; Xu, Y.; Wang, H.; Li, Q.; Hu, B.; et al. High prevalence of non-O157 Shiga toxin-producing Escherichia coli in beef cattle detected by combining four selective agars. BMC Microbiol. 2019, 19, 213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, Q.; Bai, X.; Zhao, A.; Lan, R.; Du, H.; Wang, T.; Shi, C.; Yuan, X.; Bai, X.; Ji, S.; et al. Characterization of Shiga toxin-producing Escherichia coli isolated from healthy pigs in China. BMC Microbiol. 2014, 14, 5. [Google Scholar] [CrossRef] [Green Version]
- Mora, A.; Blanco, J.E.; Blanco, M.; Alonso, M.P.; Dhabi, G.; Echeita, A.; González, E.A.; Bernárdez, M.I.; Blanco, J. Antimicrobial resistance of Shiga toxin (verotoxin)-producing Escherichia coli O157:H7 and non-O157 strains isolated from humans, cattle, sheep and food in Spain. Res. Microbiol. 2005, 156, 793–806. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xu, C.; Zhang, R.; Chen, Y.; Shen, Y.; Hu, F.; Liu, D.; Lu, J.; Guo, Y.; Xia, X.; et al. Changes in colistin resistance and mcr-1 abundance in Escherichia coli of animal and human origins following the ban of colistin-positive additives in China: An epidemiological comparative study. Lancet Infect. Dis. 2020, 20, 1161–1171. [Google Scholar] [CrossRef]
- Rehman, M.A.; Yin, X.; Persaud-Lachhman, M.G.; Diarra, M.S. First detection of a fosfomycin resistance gene, fosA7, in Salmonella enterica serovar Heidelberg isolated from broiler chickens. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poirel, L.; Bonnin, R.A.; Nordmann, P. Genetic support and diversity of acquired extended-spectrum β-lactamases in Gram-negative rods. Infect. Genet. Evol. 2012, 12, 883–893. [Google Scholar] [CrossRef] [PubMed]
- Chong, Y.; Ito, Y.; Kamimura, T. Genetic evolution and clinical impact in extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae. Infect. Genet. Evol. 2011, 11, 1499–1504. [Google Scholar] [CrossRef]
- Hayashi, W.; Ohsaki, Y.; Taniguchi, Y.; Koide, S.; Kawamura, K.; Suzuki, M.; Kimura, K.; Wachino, J.-I.; Nagano, Y.; Arakawa, Y.; et al. High prevalence of bla among genetically diverse Escherichia coli recovered from retail raw chicken meat portions in Japan. Int. J. Food Microbiol. 2018, 284. [Google Scholar] [CrossRef]
- Bevan, E.R.; Jones, A.M.; Hawkey, P.M. Global epidemiology of CTX-M β-lactamases: Temporal and geographical shifts in genotype. J. Antimicrob. Chemother. 2017, 72, 2145–2155. [Google Scholar] [CrossRef] [Green Version]
- Gatica, J.; Jurkevitch, E.; Cytryn, E. Comparative metagenomics and network analyses provide novel insights into the scope and distribution of β-lactamase homologs in the environment. Front. Microbiol. 2019, 10, 146. [Google Scholar] [CrossRef] [Green Version]
- Bai, L.; Wang, L.; Yang, X.; Wang, J.; Gan, X.; Wang, W.; Xu, J.; Chen, Q.; Lan, R.; Fanning, S.; et al. Prevalence and molecular characteristics of extended-spectrum β-lactamase genes in isolated from diarrheic patients in China. Front. Microbiol. 2017, 8, 144. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.; Sun, H.; Bai, X.; Fu, S.; Fan, R.; Xiong, Y. Occurrence of multidrug-resistant and ESBL-producing atypical enteropathogenic in China. Gut Pathog. 2018, 10, 8. [Google Scholar] [CrossRef] [Green Version]
- Bielaszewska, M.; Idelevich, E.A.; Zhang, W.; Bauwens, A.; Schaumburg, F.; Mellmann, A.; Peters, G.; Karch, H. Effects of antibiotics on Shiga toxin 2 production and bacteriophage induction by epidemic Escherichia coli O104:H4 strain. Antimicrob. Agents Chemother. 2012, 56, 3277–3282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mir, R.A.; Kudva, I.T. Antibiotic-resistant Shiga toxin-producing Escherichia coli: An overview of prevalence and intervention strategies. Zoonoses Public Health 2019, 66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gibbons, J.F.; Boland, F.; Egan, J.; Fanning, S.; Markey, B.K.; Leonard, F.C. Antimicrobial resistance of faecal Escherichia coli isolates from pig farms with different durations of in-feed antimicrobial use. Zoonoses Public Health 2016, 63, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.; Wang, H.; Xin, Y.; Wei, R.; Tang, X.; Zhao, A.; Sun, H.; Zhang, W.; Wang, Y.; Xu, Y.; et al. Prevalence and characteristics of Shiga toxin-producing Escherichia coli isolated from retail raw meats in China. Int. J. Food Microbiol. 2015, 200, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [Green Version]
- Souvorov, A.; Agarwala, R.; Lipman, D.J. SKESA: Strategic k-mer extension for scrupulous assemblies. Genome Biol. 2018, 19, 153. [Google Scholar] [CrossRef] [Green Version]
- Shen, W.; Le, S.; Li, Y.; Hu, F. SeqKit: A cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 2016, 11, e0163962. [Google Scholar] [CrossRef]
- Chin, C.-S.; Alexander, D.H.; Marks, P.; Klammer, A.A.; Drake, J.; Heiner, C.; Clum, A.; Copeland, A.; Huddleston, J.; Eichler, E.E.; et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 2013, 10, 563–569. [Google Scholar] [CrossRef]
- Joensen, K.G.; Tetzschner, A.M.; Iguchi, A.; Aarestrup, F.M.; Scheutz, F. Rapid and easy in silico serotyping of Escherichia coli isolates by use of whole-genome sequencing data. J. Clin. Microbiol. 2015, 53, 2410–2426. [Google Scholar] [CrossRef] [Green Version]
- Feldgarden, M.; Brover, V.; Haft, D.H.; Prasad, A.B.; Slotta, D.J.; Tolstoy, I.; Tyson, G.H.; Zhao, S.; Hsu, C.-H.; McDermott, P.F.; et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob. Agents Chemother. 2019, 63. [Google Scholar] [CrossRef] [Green Version]
- Jia, B.; Raphenya, A.R.; Alcock, B.; Waglechner, N.; Guo, P.; Tsang, K.K.; Lago, B.A.; Dave, B.M.; Pereira, S.; Sharma, A.N.; et al. CARD 2017: Expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017, 45, D566–D573. [Google Scholar] [CrossRef]
- Gordon, N.C.; Price, J.R.; Cole, K.; Everitt, R.; Morgan, M.; Finney, J.; Kearns, A.M.; Pichon, B.; Young, B.; Wilson, D.J.; et al. Prediction of Staphylococcus aureus antimicrobial resistance by whole-genome sequencing. J. Clin. Microbiol. 2014, 52, 1182–1191. [Google Scholar] [CrossRef] [Green Version]
- Leaché, A.D.; Banbury, B.L.; Felsenstein, J.; de Oca, A.N.-M.; Stamatakis, A. Short tree, long tree, right tree, wrong tree: New acquisition bias corrections for inferring SNP phylogenies. Syst. Biol. 2015, 64, 1032–1047. [Google Scholar] [CrossRef] [Green Version]
- Croucher, N.J.; Page, A.J.; Connor, T.R.; Delaney, A.J.; Keane, J.A.; Bentley, S.D.; Parkhill, J.; Harris, S.R. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 2015, 43, e15. [Google Scholar] [CrossRef] [Green Version]
- Page, A.J.; Taylor, B.; Delaney, A.J.; Soares, J.; Seemann, T.; Keane, J.A.; Harris, S.R. Rapid efficient extraction of SNPs from multi-FASTA alignments. Microb. Genom. 2016, 2, e000056. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Zhou, X.; Shen, X.-X.; Hittinger, C.T.; Rokas, A. Evaluating fast Maximum Likelihood-based phylogenetic programs using empirical phylogenomic data sets. Mol. Biol. Evol. 2018, 35, 486–503. [Google Scholar] [CrossRef] [Green Version]

| No. of Antimicrobial Groups | No. of Resistant Isolates from Different Sources (%) | Total | |||
|---|---|---|---|---|---|
| Cattle | Sheep | Pig | Human | ||
| 0 | 75 (93.8) | 85 (53) | 0 (0.0) | 23 (69.7) | 183 (61.4) |
| 1 | 1 (1.3) | 14 (8.8) | 3 (12.0) | 2 (6.1) | 20 (6.7) |
| 2 | 1 (1.3) | 5 (3) | 1 (4.0) | 3 (9.1) | 10 (3.4) |
| ≥3 | 3 (3.8) a | 56 b (35.0) | 21 (84.0) c | 5 (15.2) a,b | 85 (28.5) |
| Antibiotic Class | No. of Isolates (%) | ||||
|---|---|---|---|---|---|
| (No. of Isolates) | Resistance Genes | Cattle | Sheep | Pig | Human |
| Chloramphenicols | cmlA5 | 1 (50.0) | - | - | - |
| 50 | cmlA6 | 1 (50.0) | - | 4 (66.6) | - |
| floR | - | 37 (90.2) | - | - | |
| cmlA5 + floR | - | 3 (7.3) | - | - | |
| cmlA6 + floR | - | - | 2 (33.3) | - | |
| catB3 + catA1 | - | - | - | 1 (100) | |
| Trimethoprim | dfrA12 | 1 (33.3) | 1 (2.3) | 6 (37.5) | - |
| 50 | dfrA14 | 1 (33.3) | 36 (83.7) | 1 (6.2) | - |
| dfrA17 | 1 (33.3) | 1 (2.3) | 6 (37.5) | 2 (66.6) | |
| dfrA15 | - | 5 (11.6) | 2 (12.5) | 1 (33.3) | |
| dfrA12 + dfrA14 | - | - | 1 (6.2) | - | |
| Macrolides | mphA | 2 (100) | 45 (100) | - | 3 (100) |
| 53 | mefB | - | - | 2 (66.6) | - |
| ermB + mphA | - | - | 1 (33.3) | - | |
| Quinolones | qnrS1 | 1 (100) | 48 (87.2) | - | 1 (100.0) |
| 66 | qnrB17 | - | 5 (9.0) | - | - |
| oqxAB | - | 1 (1.8) | 5 (55.5) | - | |
| qnrS1 + oqxAB | - | 1 (1.8) | 4 (44.4) | - | |
| Sulfonamides | sul1 | 1 (33.3) | 8 (28.5) | 4 (21.0) | 2 (28.5) |
| 57 | sul2 | 1 (33.3) | 11 (39.2) | 2 (10.5) | 3 (42.8) |
| sul3 | 1 (33.3) | 5 (17.89) | 5 (26.3) | - | |
| sul1 + sul2 | - | 3 (10.7) | 4 (21.0) | 2 (28.5) | |
| sul1 + sul3 | - | - | 3 (15.7) | - | |
| sul2 + sul3 | - | 1 (3.5) | - | - | |
| sul1 + sul2 + sul3 | - | - | 1(5.2) | - | |
| Tetracyclines | tetA | 3 (100.0) | 21 (84.0) | 21 (84.0) | 7 (77.7) |
| 90 | tetA + tetD | - | 4 (16.0) | 4 (16.0) | 2 (22.2) |
| Fosfomycins | fosA3 | 1 (25.0) | - | - | - |
| 8 | fosA7 | 3 (75.0) | 1 (100) | - | 3 (100) |
| class A β-lactamase | blaTEM-1 | 1 (50.0) | 39 (92.8) | 4 (66.6) | 1 (50.0) |
| 52 | blaCTX-M-55 | 1 (50.0) | 1 (2.3) | 1 (16.6) | - |
| blaCTX-M-65 | - | 2 (4.7) | - | - | |
| blaCTX-M-65 | - | - | 1 (16.6) | - | |
| +blaTEM-1 | |||||
| blaCTX-M-15 | - | - | - | 1 (50.0) | |
| +blaTEM-1 | |||||
| Colistin | mcr-1 | - | - | 7 (87.5) | - |
| 8 | mcr-3 | - | - | 1 (12.5) | - |
| Rifampin | arr-2 | 1 (100) | 7 (58.3) | - | - |
| 13 | arr-3 | - | 5 (41.6) | - | - |
| Aminoglycosides | ant(3″)-Ia | 1 (33.3) | 7 (23.3) | 4 (21.1) | 1 (16.7) |
| 58 | aac(3)-IIa + aph(3″)-Ib + aph(6)-Id | - | 10 (33.3) | - | 1 (16.7) |
| aph(3″)-Ib + aph(6)-Id | - | 4 (13.3) | - | 2 (33.3) | |
| aph(3″)-Ib + aph(3′)-Ia+ aph(6)-Id + ant(3″)-Ia | - | - | 6 (31.6) | - | |
| aph(3″)-Ib + aph(6)-Id + ant(3″)-Ia | - | 1 (3.3) | 1 (5.3) | 2 (33.3) | |
| aac(3)-IV + aph(3″)-Ib + aph(3′)-Ia + aph(4)-Ia + aph(6)-Id + ant(3″)-Ia | - | 4 (13.3) | - | - | |
| others | 2 (66.6) | 4 (13.3) | 8 (42.1) | - | |
| Antimicrobial Agent | No. of Phenotypic Resistant Isolates | No. of Phenotypic Susceptible Isolates | Sensitivity (%) | Specificity (%) | ||
|---|---|---|---|---|---|---|
| Susceptible by Genotype | Resistant by Genotype | Susceptible by Genotype | Resistant by Genotype | |||
| Chloramphenicol | 6 | 50 | 242 | 0 | 89 | 100 |
| Quinolone * | 37 | 38 | 195 | 28 | 51 | 87 |
| Aminoglycoside | 1 | 1 | 239 | 57 | 50 | 81 |
| Macrolide | 6 | 50 | 239 | 3 | 89 | 99 |
| Tetracycline | 8 | 89 | 200 | 1 | 92 | 100 |
| Trimethoprim-Sulfamethoxazole | 9 | 33 | 255 | 1 | 79 | 100 |
| Colistin | 2 | 6 | 289 | 1 | 75 | 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pan, Y.; Hu, B.; Bai, X.; Yang, X.; Cao, L.; Liu, Q.; Sun, H.; Li, J.; Zhang, J.; Jin, D.; et al. Antimicrobial Resistance of Non-O157 Shiga Toxin-Producing Escherichia coli Isolated from Humans and Domestic Animals. Antibiotics 2021, 10, 74. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics10010074
Pan Y, Hu B, Bai X, Yang X, Cao L, Liu Q, Sun H, Li J, Zhang J, Jin D, et al. Antimicrobial Resistance of Non-O157 Shiga Toxin-Producing Escherichia coli Isolated from Humans and Domestic Animals. Antibiotics. 2021; 10(1):74. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics10010074
Chicago/Turabian StylePan, Yanyu, Bin Hu, Xiangning Bai, Xi Yang, Lijiao Cao, Qian Liu, Hui Sun, Juan Li, Ji Zhang, Dong Jin, and et al. 2021. "Antimicrobial Resistance of Non-O157 Shiga Toxin-Producing Escherichia coli Isolated from Humans and Domestic Animals" Antibiotics 10, no. 1: 74. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics10010074








