Pan-Drug Resistant Acinetobacter baumannii, but Not Other Strains, Are Resistant to the Bee Venom Peptide Melittin
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Peptide
2.3. Determination of the Minimum Inhibitory Concentration (MIC)
2.4. Inhibition of Biofilm Formation
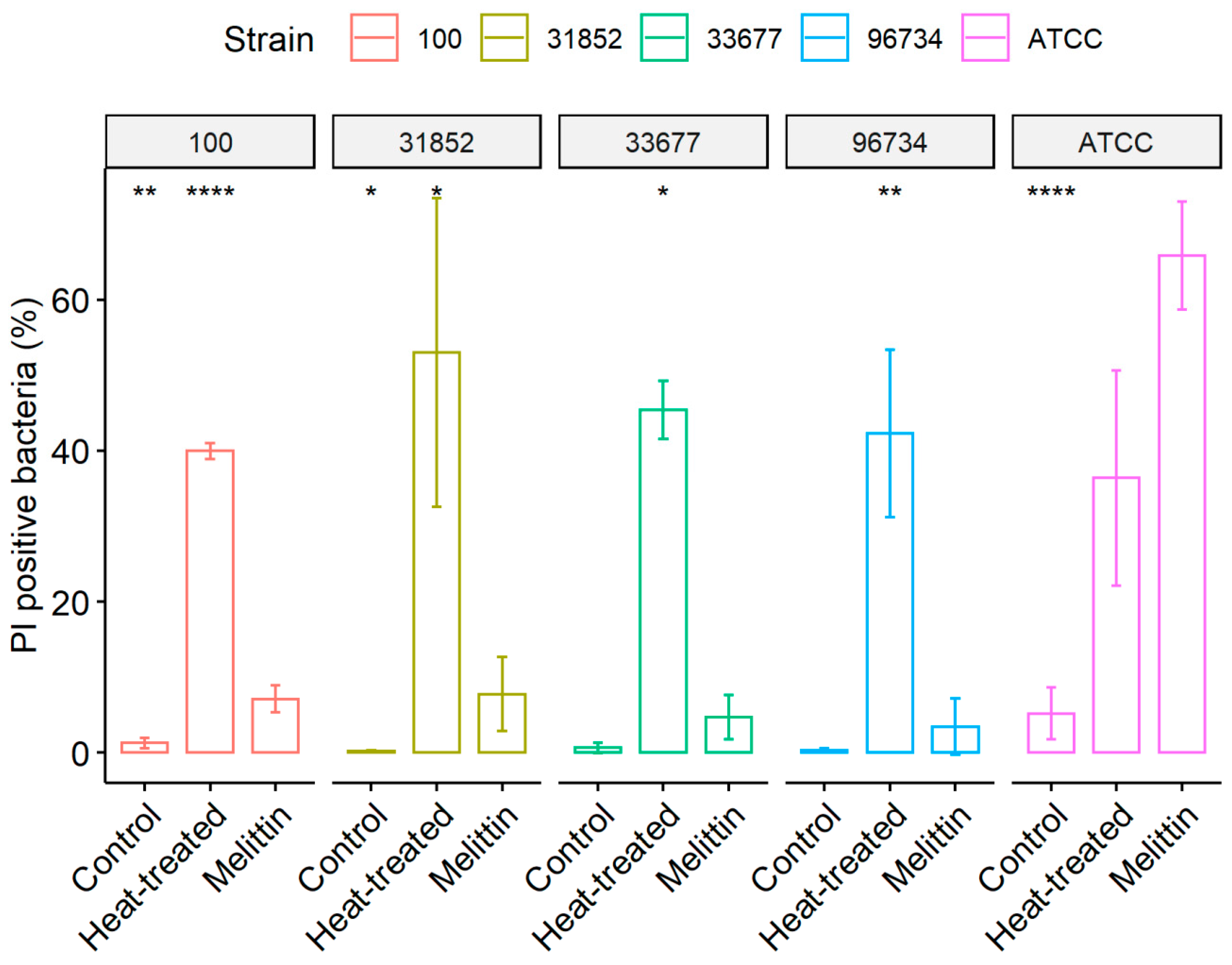
2.5. Membrane Permeability Assay
2.6. Analysis of Bacterial Proliferation
2.7. Statistical Analysis
3. Results
3.1. Determination of MIC for Melittin, Polymyxin and Imipenem in A. baumannii Strains
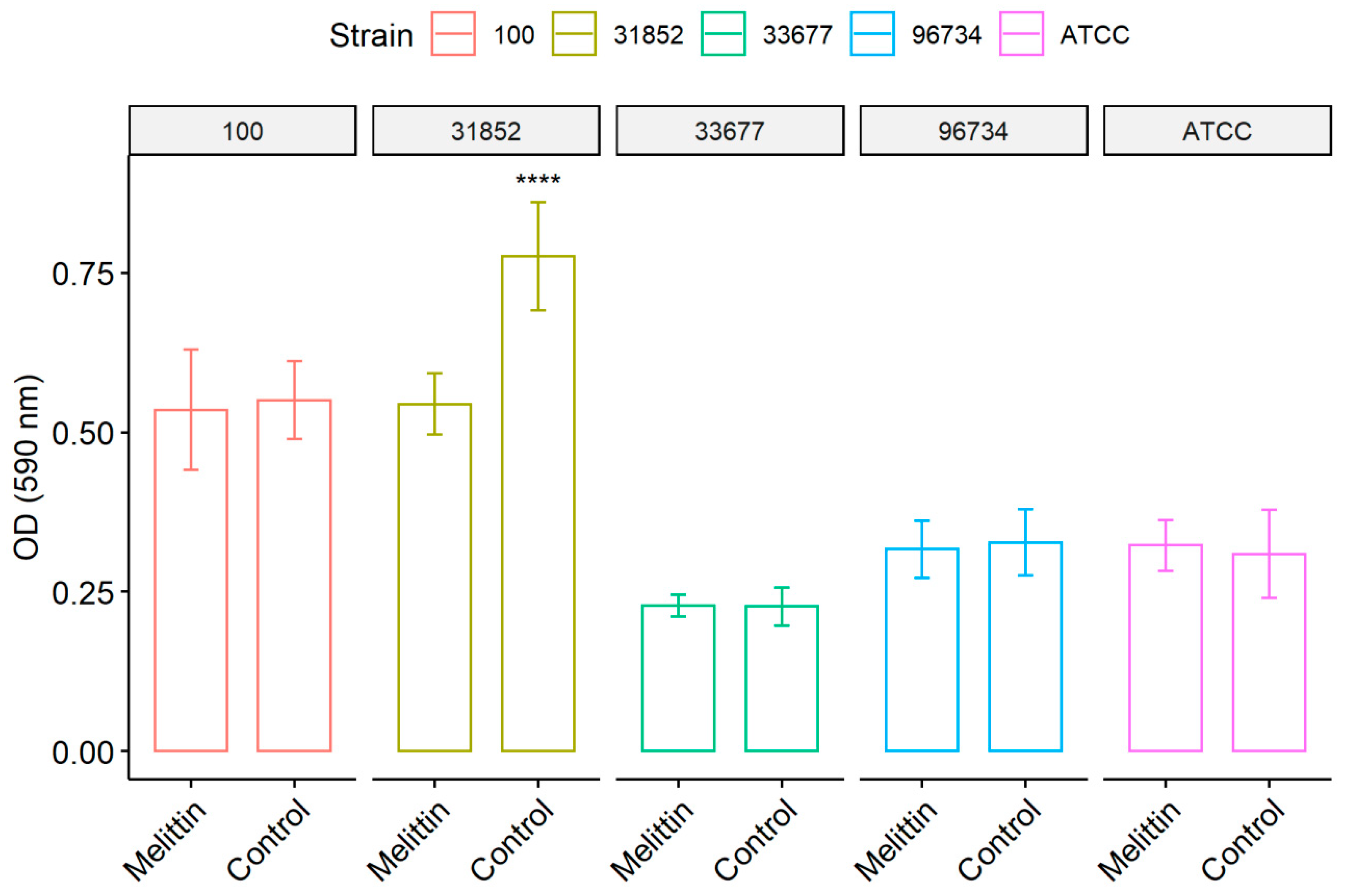
3.2. Biofilm Formation Test
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Qi, L.; Li, H.; Zhang, C.; Liang, B.; Li, J.; Wang, L.; Du, X.; Liu, X.; Qiu, S.; Song, H. Relationship between antibiotic resistance, biofilm formation, and biofilm-specific resistance in Acinetobacter baumannii. Front. Microbiol. 2016, 7, 483. [Google Scholar] [CrossRef] [Green Version]
- Safari, M.; Mozaffari Nejad, A.S.; Bahador, A.; Jafari, R.; Alikhani, M.Y. Prevalence of ESBL and MBL encoding genes in Acinetobacter baumannii strains isolated from patients of intensive care units (ICU). Saudi J. Biol. Sci. 2015, 22, 424–429. [Google Scholar] [CrossRef] [Green Version]
- Wong, D.; Nielsen, T.B.; Bonomo, R.A.; Pantapalangkoor, P.; Luna, B.; Spellberg, B. Clinical and pathophysiological overview of Acinetobacter infections: A century of challenges. Clin. Microbiol. Rev. 2017, 30, 409–447. [Google Scholar] [CrossRef] [Green Version]
- Harding, C.M.; Hennon, S.W.; Feldman, M.F. Uncovering the mechanisms of Acinetobacter baumannii virulence. Nat. Rev. Microbiol. 2018, 16, 91–102. [Google Scholar] [CrossRef]
- Center for Disease Control and Prevention (CDC). Antibiotic Resistance Threats in the United States, USA; CDC: Atlanta, GA, USA, 2013.
- Paterson, D.L.; Bonomo, R.A. Multidrug-resistant gram-negative pathogens: The urgent need for ‘old’ polymyxins. Adv. Exp. Med. Biol. 2019, 1145, 9–13. [Google Scholar] [PubMed]
- Romanin, P.; Palermo, R.L.; Cavalini, J.F.; Fávaro, L.D.S.; De Paula-Petroli, S.B.; Fernandes, E.V.; Dos Anjos Szczerepa, M.M.; Tognim, M.C.B.; Yamada-Ogatta, S.F.; Carrara-Marroni, F.E.; et al. Multidrug- and extensively drug-resistant Acinetobacter baumannii in a tertiary hospital from Brazil: The importance of carbapenemase encoding genes and epidemic clonal complexes in a 10-year study. Microb. Drug Resist. 2019, 25, 1365–1373. [Google Scholar] [CrossRef] [PubMed]
- Mao, T.; Zhai, H.; Duan, G.; Yang, H. Patterns of drug-resistant bacteria in a general hospital, China, 2011–2016. Pol. J. Microbiol. 2019, 68, 225–232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- World Health Organization (WHO). WHO Publishes List of Bacteria for Which New Antibiotics Are Urgently Needed; WHO: Geneva, Switzerland, 2017. [Google Scholar]
- Bardbari, A.M.; Arabestani, M.R.; Karami, M.; Keramat, F.; Alikhani, M.Y.; Bagheri, K.P. Correlation between ability of biofilm formation with their responsible genes and MDR patterns in clinical and environmental Acinetobacter baumannii isolates. Microb. Pathog. 2017, 108, 122–128. [Google Scholar] [CrossRef]
- Howard, A.; O’Donoghue, M.; Feeney, A.; Sleator, R.D. Acinetobacter baumannii: An emerging opportunistic pathogen. Virulence 2012, 3, 243–250. [Google Scholar] [CrossRef]
- Dahdouh, E.; Orgaz, B.; Gómez-Gil, R.; Mingorance, J.; Daoud, Z.; Suarez, M.; San Jose, C. Patterns of biofilm structure and formation kinetics among Acinetobacter baumannii clinical isolates with different antibiotic resistance profiles. Med. Chem. Commun. 2016, 7, 157–163. [Google Scholar] [CrossRef]
- Batoni, G.; Maisetta, G.; Esin, S. Antimicrobial peptides and their interaction with biofilms of medically relevant bacteria. Biochim. Biophys. Acta 2016, 1858, 1044–1060. [Google Scholar] [CrossRef] [PubMed]
- Gunn, J.S.; Bakaletz, L.O.; Wozniak, D.J. What’s on the outside matters: The role of the extracellular polymeric substance of Gram-negative biofilms in evading host immunity and as a target for therapeutic intervention. J. Biol. Chem. 2016, 291, 12538–12546. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramos-Gallardo, G. Chronic wounds in burn injury: A case report on importance of biofilms. World J. Plast. Surg. 2016, 5, 175–180. [Google Scholar] [PubMed]
- Vila-Farres, X.; Garcia de la Maria, C.; López-Rojas, R.; Pachón, J.; Giralt, E.; Vila, J. In Vitro activity of several antimicrobial peptides against colistin-susceptible and colistin-resistant Acinetobacter baumannii. Clin. Microbiol. Infect. 2012, 18, 383–387. [Google Scholar] [CrossRef] [Green Version]
- Sun, F.; Qu, F.; Ling, Y.; Mao, P.; Xia, P.; Chen, H.; Zhou, D. Biofilm-associated infections: Antibiotic resistance and novel therapeutic strategies. Future Microbiol. 2013, 8, 877–886. [Google Scholar] [CrossRef]
- Lashinsky, J.N.; Henig, O.; Pogue, J.M.; Kaye, K.S. Minocycline for the treatment of multidrug and extensively drug-resistant A. baumannii: A review. Infect. Dis. Ther. 2017, 6, 199–211. [Google Scholar] [CrossRef] [Green Version]
- Greene, C.; Wu, J.; Rickard, A.H.; Xi, C. Evaluation of the ability of Acinetobacter baumannii to form biofilms on six different biomedical relevant surfaces. Lett. Appl. Microbiol. 2016, 63, 233–239. [Google Scholar] [CrossRef] [Green Version]
- Sierra, J.M.; Fusté, E.; Rabanal, F.; Vinuesa, T.; Viñas, M. An overview of antimicrobial peptides and the latest advances in their development. Expert Opin. Biol. Ther. 2017, 17, 663–676. [Google Scholar] [CrossRef]
- Nuti, R.; Goud, N.S.; Saraswati, A.P.; Alvala, R.; Alvala, M. Antimicrobial peptides: A promising therapeutic strategy in tackling antimicrobial resistance. Curr. Med. Chem 2017, 24, 4303–4314. [Google Scholar] [CrossRef]
- Wimley, W.C. Describing the mechanism of antimicrobial peptide action with the interfacial activity model. ACS Chem. Biol. 2010, 5, 905–917. [Google Scholar] [CrossRef] [Green Version]
- Zasloff, M. Antimicrobial peptides of multicellular organisms. Nature 2002, 415, 389–395. [Google Scholar] [CrossRef]
- Koehbach, J.; Craik, D.J. The vast structural diversity of antimicrobial peptides. Trends Pharmacol. Sci. 2019, 40, 517–528. [Google Scholar] [CrossRef]
- Fox, J.L. Antimicrobial peptides stage a comeback. Nat. Biotechnol. 2013, 31, 379–382. [Google Scholar] [CrossRef]
- Dosler, S.; Karaaslan, E. Inhibition and destruction of Pseudomonas aeruginosa biofilms by antibiotics and antimicrobial peptides. Peptides 2014, 62, 32–37. [Google Scholar] [CrossRef]
- Rios, A.C.; Moutinho, C.G.; Pinto, F.C.; Del Fiol, F.S.; Jozala, A.; Chaud, M.V.; Vila, M.M.; Teixeira, J.A.; Balcão, V.M. Alternatives to overcoming bacterial resistances: State-of-the-art. Microbiol. Res. 2016, 191, 51–80. [Google Scholar] [CrossRef]
- Memariani, H.; Memariani, M.; Shahidi-Dadras, M.; Nasiri, S.; Akhavan, M.M.; Moravvej, H. Melittin: From honeybees to superbugs. Appl. Microbiol. Biotechnol. 2019, 103, 3265–3276. [Google Scholar] [CrossRef]
- Dorman, L.C.; Markley, L.D. Solid phase synthesis and antibacterial activity of N-terminal sequences of melittin. J. Med. Chem. 1971, 14, 5–9. [Google Scholar] [CrossRef]
- Steiner, H.; Hultmark, D.; Engström, Å.; Bennich, H.; Boman, H.G. Sequence and specificity of two antibacterial proteins involved in insect immunity. Nature 1981, 292, 246–248. [Google Scholar] [CrossRef]
- Dosler, S.; Karaaslan, E.; Alev Gerceker, A. Antibacterial and anti-biofilm activities of melittin and colistin, alone and in combination with antibiotics against Gram-negative bacteria. J. Chemother. 2016, 28, 95–103. [Google Scholar] [CrossRef]
- Pashaei, F.; Bevalian, P.; Akbari, R.; Pooshang Bagheri, K. Single dose eradication of extensively drug resistant Acinetobacter spp. In a mouse model of burn infection by melittin antimicrobial peptide. Microb. Pathog. 2019, 127, 60–69. [Google Scholar] [CrossRef]
- Sugden, R.; Kelly, R.; Davies, S. Combatting antimicrobial resistance globally. Nat. Microbiol. 2016, 1, 16187. [Google Scholar] [CrossRef]
- Yang, L.; Harroun, T.A.; Weiss, T.M.; Ding, L.; Huang, H.W. Barrel-stave model or toroidal model? A case study on melittin pores. Biophys. J. 2001, 81, 1475–1485. [Google Scholar] [CrossRef] [Green Version]
- Rady, I.; Siddiqui, I.A.; Rady, M.; Mukhtar, H. Melittin, a major peptide component of bee venom, and its conjugates in cancer therapy. Cancer Lett. 2017, 402, 16–31. [Google Scholar] [CrossRef] [Green Version]
- Andersson, M.; Ulmschneider, J.P.; Ulmschneider, M.B.; White, S.H. Conformational states of melittin at a bilayer interface. Biophys. J. 2013, 104, L12–L14. [Google Scholar] [CrossRef] [Green Version]
- Lee, M.-T.; Sun, T.-L.; Hung, W.-C.; Huang, H.W. Process of inducing pores in membranes by melittin. Proc. Natl. Acad. Sci. USA 2013, 110, 14243–14248. [Google Scholar] [CrossRef] [Green Version]
- Wu, Y.; Ma, L.; Cheley, S.; Bayley, H.; Cui, Q.; Chapman, E.R. Permeation of styryl dyes through nanometer-scale pores in membranes. Biochemistry 2011, 50, 7493–7502. [Google Scholar] [CrossRef] [Green Version]
- Paull, B.R.; Yunginger, J.W.; Gleich, G.J. Melittin: An allergen of honeybee venom. J. Allergy Clin. Immunol. 1977, 59, 334–338. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standands Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing; CLSI approved standard M100-S25; CLSI: Wayne, PA, USA, 2015. [Google Scholar]
- Lall, N.; Henley-Smith, C.J.; De Canha, M.N.; Oosthuizen, C.B.; Berrington, D. Viability reagent, prestoblue, in comparison with other available reagents, utilized in cytotoxicity and antimicrobial assays. Int. J. Microbiol. 2013, 2013, 420601. [Google Scholar] [CrossRef] [Green Version]
- Moskowitz, S.M.; Foster, J.M.; Emerson, J.; Burns, J.L. Clinically feasible biofilm susceptibility assay for isolates of Pseudomonas aeruginosa from patients with cystic fibrosis. J. Clin. Microbiol. 2004, 42, 1915–1922. [Google Scholar] [CrossRef] [Green Version]
- Eales, M.G.; Ferrari, E.; Goddard, A.D.; Lancaster, L.; Sanderson, P.; Miller, C. Mechanistic and phenotypic studies of bicarinalin, BP100 and colistin action on Acinetobacter baumannii. Res. Microbiol. 2018, 169, 296–302. [Google Scholar] [CrossRef] [Green Version]
- Tyagi, P.; Singh, M.; Kumari, H.; Kumari, A.; Mukhopadhyay, K. Bactericidal activity of curcumin I is associated with damaging of bacterial membrane. PLoS ONE 2015, 10, e0121313. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Travnickova, E.; Mikula, P.; Oprsal, J.; Bohacova, M.; Kubac, L.; Kimmer, D.; Soukupova, J.; Bittner, M. Resazurin assay for assessment of antimicrobial properties of electrospun nanofiber filtration membranes. AMB Express 2019, 9, 183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deslouches, B.; Steckbeck, J.D.; Craigo, J.K.; Doi, Y.; Burns, J.L.; Montelaro, R.C. Engineered cationic antimicrobial peptides to overcome multidrug resistance by ESKAPE pathogens. Antimicrob. Agents Chemother. 2015, 59, 1329–1333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mohamed, M.F.; Brezden, A.; Mohammad, H.; Chmielewski, J.; Seleem, M.N. Targeting biofilms and persisters of ESKAPE pathogens with P14KanS, a kanamycin peptide conjugate. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 848–859. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.; Wang, J.; Pan, H.; Hu, W.; Li, Y. Multidrug resistance and biofilm formation contribute to the nosocomial infections caused by Acinetobacter baumannii. In The Battle against Microbial Pathogens: Basic Science, Technological Advances and Educational Programs; Méndez-Vilas, A., Ed.; Formatex Research Center: Badajoz, Spain, 2015; Volume 1, pp. 452–461. [Google Scholar]
- Bjarnsholt, T.; Ciofu, O.; Molin, S.; Givskov, M.; Høiby, N. Applying insights from biofilm biology to drug development—Can a new approach be developed? Nat. Rev. Drug. Discov. 2013, 12, 791–808. [Google Scholar] [CrossRef]
- Hale, J.D.; Hancock, R.E. Alternative mechanisms of action of cationic antimicrobial peptides on bacteria. Expert Rev. Anti-Infect. Ther. 2007, 5, 951–959. [Google Scholar] [CrossRef]
- Memariani, H.; Shahbazzadeh, D.; Ranjbar, R.; Behdani, M.; Memariani, M.; Pooshang Bagheri, K. Design and characterization of short hybrid antimicrobial peptides from pEM-2, mastoparan-VT1, and mastoparan-B. Chem. Biol. Drug. Des. 2017, 89, 327–338. [Google Scholar] [CrossRef]
- Almeida, J.R.; Palacios, A.L.V.; Patiño, R.S.P.; Mendes, B.; Teixeira, C.A.S.; Gomes, P.; da Silva, S.L. Harnessing snake venom phospholipases A2 to novel approaches for overcoming antibiotic resistance. Drug Dev. Res. 2019, 80, 68–85. [Google Scholar] [CrossRef] [Green Version]
- Picoli, T.; Peter, C.M.; Zani, J.L.; Waller, S.B.; Lopes, M.G.; Boesche, K.N.; Vargas, G.D.Á.S.; Hübner, O.; Fischer, G. Melittin and its potential in the destruction and inhibition of the biofilm formation by Staphylococcus aureus, Escherichia coli and Pseudomonas aeruginosa isolated from bovine milk. Microb. Pathog. 2017, 112, 57–62. [Google Scholar] [CrossRef]
- Shi, W.; Li, C.; Li, M.; Zong, X.; Han, D.; Chen, Y.C. Antimicrobial peptide melittin against Xanthomonas oryzae pv. oryzae, the bacterial leaf blight pathogen in rice. Appl. Microbiol. Biotechnol. 2016, 100, 5059–5067. [Google Scholar] [CrossRef] [Green Version]
- Dosler, S.; Gerceker, A.A. In Vitro activities of antimicrobial cationic peptides; melittin and nisin, alone or in combination with antibiotics against Gram-positive bacteria. J. Chemother. 2012, 24, 137–143. [Google Scholar] [CrossRef]
- Gopal, R.; Lee, J.H.; Kim, Y.G.; Kim, M.S.; Seo, C.H.; Park, Y. Antimicrobial, anti-biofilm activities and cell selectivity of the NRC-16 peptide derived from witch flounder, Glyptocephalus cynoglossus. Mar. Drugs 2013, 11, 1836–1852. [Google Scholar] [CrossRef] [Green Version]
- Bardbari, A.M.; Arabestani, M.R.; Karami, M.; Keramat, F.; Aghazadeh, H.; Alikhani, M.Y.; Bagheri, K.P. Highly synergistic activity of melittin with imipenem and colistin in biofilm inhibition against multidrug-resistant strong biofilm producer strains of Acinetobacter baumannii. Eur. J. Clin. Microbiol. Infect. Dis. 2018, 37, 443–454. [Google Scholar] [CrossRef]
- Ying, C.; Li, Y.; Wang, Y.; Zheng, B.; Yang, C. Investigation of the molecular epidemiology of Acinetobacter baumannii isolated from patients and environmental contamination. J. Antibiot. (Tokyo) 2015, 68, 562–567. [Google Scholar] [CrossRef]
- Giacometti, A.; Cirioni, O.; Kamysz, W.; D’Amato, G.; Silvestri, C.; Del Prete, M.S.; Lukasiak, J.; Scalise, G. Comparative activities of cecropin A, melittin, and cecropin A-melittin peptide CA(1–7)M(2–9)NH2 against multidrug-resistant nosocomial isolates of Acinetobacter baumannii. Peptides 2003, 24, 1315–1318. [Google Scholar] [CrossRef]
- Socarras, K.M.; Theophilus, P.A.S.; Torres, J.P.; Gupta, K.; Sapi, E. Antimicrobial activity of bee venom and melittin against Borrelia burgdorferi. Antibiotics 2017, 6, 31. [Google Scholar] [CrossRef] [Green Version]
- Khozani, R.S.; Shahbazzadeh, D.; Harzandi, N.; Feizabadi, M.M.; Bagheri, K.P. Kinetics study of antimicrobial peptide, melittin, in simultaneous biofilm degradation and eradication of potent biofilm producing MDR Pseudomonas aeruginosa isolates. Int. J. Pept. Res. Ther. 2018, 25, 329–338. [Google Scholar] [CrossRef]
- Kumar, P.; Kizhakkedathu, J.N.; Straus, S.K. Antimicrobial peptides: Diversity, mechanism of action and strategies to improve the activity and biocompatibility in vivo. Biomolecules 2018, 8, 4. [Google Scholar] [CrossRef] [Green Version]
- Park, S.C.; Kim, J.Y.; Shin, S.O.; Jeong, C.Y.; Kim, M.H.; Shin, S.Y.; Cheong, G.W.; Park, Y.; Hahm, K.S. Investigation of toroidal pore and oligomerization by melittin using transmission electron microscopy. Biochem. Biophys. Res. Commun. 2006, 343, 222–228. [Google Scholar] [CrossRef]
- Sengupta, D.; Leontiadou, H.; Mark, A.E.; Marrink, S.J. Toroidal pores formed by antimicrobial peptides show significant disorder. Biochim. Biophys. Acta 2008, 1778, 2308–2317. [Google Scholar] [CrossRef] [Green Version]
- Bechinger, B.; Gorr, S.-U. Antimicrobial peptides: Mechanisms of action and resistance. J. Dent. Res. 2017, 96, 254–260. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 2003, 67, 593–656. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hurdle, J.G.; O’Neill, A.J.; Chopra, I.; Lee, R.E. Targeting bacterial membrane function: An underexploited mechanism for treating persistent infections. Nat. Rev. Microbiol. 2011, 9, 62–75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moerman, L.; Bosteels, S.; Noppe, W.; Willems, J.; Clynen, E.; Schoofs, L.; Thevissen, K.; Tytgat, J.; Van Eldere, J.; Van Der Walt, J.; et al. Antibacterial and antifungal properties of alpha-helical, cationic peptides in the venom of scorpions from southern Africa. Eur. J. Biochem. 2002, 269, 4799–4810. [Google Scholar] [CrossRef]
- Al-Ani, I.; Zimmermann, S.; Reichling, J.; Wink, M. Pharmacological synergism of bee venom and melittin with antibiotics and plant secondary metabolites against multi-drug resistant microbial pathogens. Phytomedicine 2015, 22, 245–255. [Google Scholar] [CrossRef]

| Strain/Resistance Profile | Melittin (µg/mL) | Polymyxin c (µg/mL) | Imipenem d (µg/mL) |
|---|---|---|---|
| A. baumannii ATCC 19606 | 17 | 0.25 | 0.25 |
| A. baumannii 31852 (S) | 20 | 0.25 | ≤0.125 |
| A. baumannii 33677 (XDR) | 31 | 0.25 | 16 |
| A. baumannii 96734 (XDR) | 45.5 | 0.25 | 16 |
| A. baumannii 100 (PDR b) | >284 | 8 | 32 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rangel, K.; Curty Lechuga, G.; Almeida Souza, A.L.; Rangel da Silva Carvalho, J.P.; Simões Villas Bôas, M.H.; De Simone, S.G. Pan-Drug Resistant Acinetobacter baumannii, but Not Other Strains, Are Resistant to the Bee Venom Peptide Melittin. Antibiotics 2020, 9, 178. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics9040178
Rangel K, Curty Lechuga G, Almeida Souza AL, Rangel da Silva Carvalho JP, Simões Villas Bôas MH, De Simone SG. Pan-Drug Resistant Acinetobacter baumannii, but Not Other Strains, Are Resistant to the Bee Venom Peptide Melittin. Antibiotics. 2020; 9(4):178. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics9040178
Chicago/Turabian StyleRangel, Karyne, Guilherme Curty Lechuga, André Luis Almeida Souza, João Pedro Rangel da Silva Carvalho, Maria Helena Simões Villas Bôas, and Salvatore Giovanni De Simone. 2020. "Pan-Drug Resistant Acinetobacter baumannii, but Not Other Strains, Are Resistant to the Bee Venom Peptide Melittin" Antibiotics 9, no. 4: 178. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics9040178





