Prevalence and Antibiotic Resistance of ESKAPE Pathogens Isolated in the Emergency Department of a Tertiary Care Teaching Hospital in Hungary: A 5-Year Retrospective Survey
Abstract
:1. Introduction
2. Results
2.1. Distribution of Bacterial Isolates
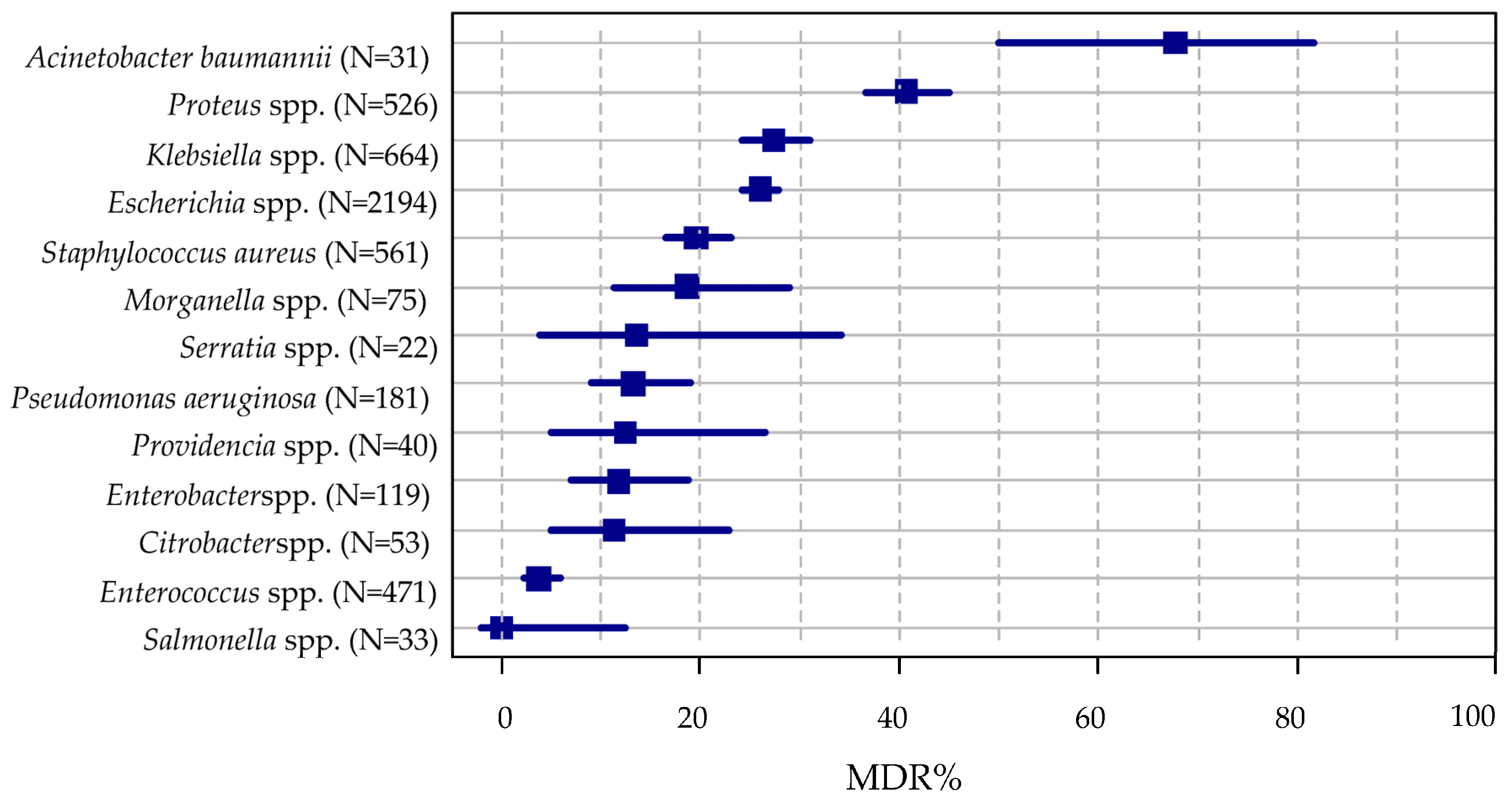
2.2. Bacterial Resistance According to Specific Indicators
2.3. Bacterial Resistance to Specific Antibacterials
3. Discussion
Strengths and Limitations
4. Materials and Methods
4.1. Study Site, Data Collection and Management, Inclusion Criteria
4.2. Laboratory Procedures, Microbial Identification
4.3. Antimicrobial Susceptibility Testing
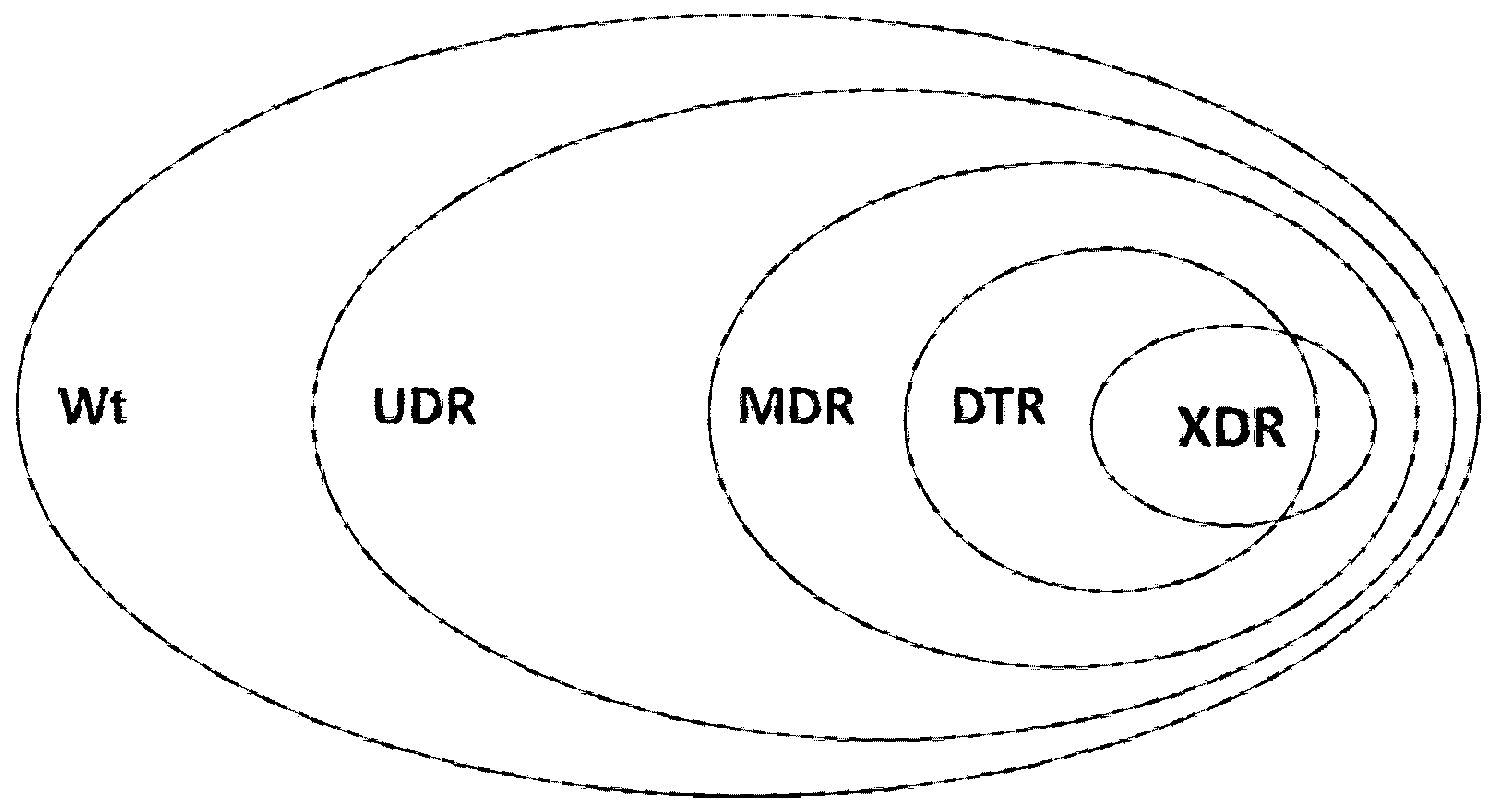
4.4. Definitions for the Classification of Resistance Groups
4.5. Statistical Analyses
4.6. Ethical Considerations
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Data Availability
References
- Cosgrove, S.E. The relationship between antimicrobial resistance and patient outcomes: Mortality, length of hospital stay, and health care costs. Clin. Infect. Dis. 2006, 42 (Suppl. 2), S82–S89. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naylor, N.R.; Atun, R.; Zhu, N.; Kulasabanathan, K.; Silva, S.; Chatterjee, A.; Knight, G.M.; Robotham, J.V. Estimating the burden of antimicrobial resistance: A systematic literature review. Antimicrob. Resist. Infect. Control. 2018, 7, 58. [Google Scholar] [CrossRef] [PubMed]
- Cassini, A.; Högberg, L.D.; Plachouras, D.; Quattrocchi, A.; Hoxha, A.; Simonsen, G.S.; Colomb-Cotinat, M.; Kretzschmar, M.E.; Devleesschauwer, B.; Cecchini, M.; et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the European Union and the European Economic Area in 2015: A population-level health estimate. Lancet Infect. Dis. 2019, 19, 56–66. [Google Scholar] [CrossRef] [Green Version]
- Tacconelli, E.; Pezzani, M.D. Public health burden of antimicrobial resistance in Europe. Lancet Infect. Dis. 2019, 19, 4–6. [Google Scholar] [CrossRef] [Green Version]
- Lim, L.M.; Ly, N.; Anderson, D.; Yang, J.C.; Macander, L.; Jarkowski, A., 3rd; Forrest, A.; Bulitta, J.B.; Tsuji, B.T. Resurgence of colistin: A review of resistance, toxicity, pharmacodynamics, and dosing. Pharmacotherapy 2010, 30, 1279–1291. [Google Scholar] [CrossRef]
- Bassetti, M.; Righi, E. Safety profiles of old and new antimicrobials for the treatment of MRSA infections. Expert Opin. Drug Saf. 2016, 15, 467–481. [Google Scholar] [CrossRef] [PubMed]
- Sipahi, O.R. Economics of antibiotic resistance. Expert Rev. Anti-Infect. Ther. 2008, 6, 523–539. [Google Scholar] [CrossRef]
- Van Duin, D.; Paterson, D.L. Multidrug-resistant bacteria in the community: Trends and lessons learned. Infect. Dis. Clin. N. Am. 2016, 30, 377–390. [Google Scholar] [CrossRef] [Green Version]
- Zhen, X.; Lundborg, C.S.; Sun, X.; Hu, X.; Dong, H. Economic burden of antibiotic resistance in ESKAPE organisms: A systematic review. Antimicrob. Resist. Infect. Control. 2019, 8, 137. [Google Scholar] [CrossRef] [Green Version]
- Pendleton, J.N.; Gorman, S.P.; Gilmore, B.F. Clinical relevance of the ESKAPE pathogens. Expert Rev. Anti-Infect. Ther. 2013, 11, 297–308. [Google Scholar] [CrossRef]
- Rice, L.B. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No ESKAPE. J. Infect. Dis. 2008, 197, 1079–1081. [Google Scholar] [CrossRef] [PubMed]
- Santajit, S.; Indrawattana, N. Mechanisms of antimicrobial resistance in ESKAPE pathogens. Biomed Res. Int. 2016, 2016, 2475067. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mulani, M.S.; Kamble, E.E.; Kumkar, S.N.; Tawre, M.S.; Pardesi, K.R. Emerging strategies to combat ESKAPE pathogens in the era of antimicrobial resistance: A review. Front. Microbiol. 2019, 10, 539. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Prioritization of Pathogens to Guide Discovery, Research and Development of New Antibiotics for Drug Resistant Bacterial Infections, Including Tuberculosis; World Health Organization: Geneva, Switzerland, 2017. [Google Scholar]
- Yang, S.; Xu, H.; Sun, J.; Sun, S. Shifting trends and age distribution of ESKAPEEc resistance in bloodstream infection, Southwest China, 2012–2017. Antimicrob. Resist. Infect. Control. 2019, 8, 61. [Google Scholar] [CrossRef] [Green Version]
- Llaca-Díaz, J.M.; Mendoza-Olazarán, S.; Camacho-Ortiz, A.; Flores, S.; Garza-González, E. One-year surveillance of ESKAPE pathogens in an intensive care unit of Monterrey, Mexico. Chemotherapy 2012, 58, 475–481. [Google Scholar] [CrossRef]
- An, S.-Q.; Berg, G. Stenotrophomonas maltophilia. Trends Microbiol. 2018, 26, 637–638. [Google Scholar] [CrossRef] [Green Version]
- Ward, M.M.; Diekema, D.J.; Yankey, J.W.; Vaughn, T.E.; BootsMiller, B.J.; Pendergast, J.F.; Doebbeling, B.N. Implementation of strategies to prevent and control the emergence and spread of antimicrobial-resistant microorganisms in U.S. hospitals. Infect. Control. Hosp. Epidemiol. 2005, 26, 21–30. [Google Scholar] [CrossRef]
- Costelloe, C.; Metcalfe, C.; Lovering, A.; Mant, D.; Hay, A.D. Effect of antibiotic prescribing in primary care on antimicrobial resistance in individual patients: Systematic review and meta-analysis. BMJ 2010, 340, c2096. [Google Scholar] [CrossRef] [Green Version]
- Byrne, M.K.; Miellet, S.; McGlinn, A.; Fish, J.; Meedya, S.; Reynolds, N.; van Oijen, A.M. The drivers of antibiotic use and misuse: The development and investigation of a theory driven community measure. BMC Public Health 2019, 19, 1425. [Google Scholar] [CrossRef] [Green Version]
- May, L.; Cosgrove, S.; L’Archeveque, M.; Talan, D.A.; Payne, P.; Jordan, J.; Rothman, R.E. A call to action for antimicrobial stewardship in the emergency department: Approaches and strategies. Ann. Emerg. Med. 2012, 62, 69–77. [Google Scholar] [CrossRef] [Green Version]
- Forero, R.; Nahidi, S.; de Costa, J.; Fatovich, D.; FitzGerald, G.; Toloo, S.; McCarthy, S.; Mountain, D.; Gibson, N.; Mohsin, M.; et al. Perceptions and experiences of emergency department staff during the implementation of the four-hour rule/national emergency access target policy in Australia: A qualitative social dynamic perspective. BMC Health Serv. Res. 2019, 19, 82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goulopoulos, A.; Rofe, O.; Kong, D.; Maclean, A.; O’Reilly, M. Attitudes and beliefs of Australian emergency department clinicians on antimicrobial stewardship in the emergency department: A qualitative study. Emerg. Med. Australas 2019, 31, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Colliers, A.; Adriaenssens, N.; Anthierens, S.; Bartholomeeusen, S.; Philips, H.; Remmen, R.; Coenen, S. Antibiotic prescribing quality in out-of-hours primary care and critical appraisal of disease-specific quality indicators. Antibiotics 2019, 8, 79. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adriaenssens, N.; Bartholomeeusen, S.; Ryckebosch, P.; Coenen, S. Quality of antibiotic prescription during office hours and out-of-hours in Flemish primary care, using European quality indicators. Eur. J. Gen. Pract. 2014, 20, 114–120. [Google Scholar] [CrossRef]
- Colliers, A.; Coenen, S.; Remmen, R.; Philips, H.; Anthierens, S. How do general practitioners and pharmacists experience antibiotic use in out-of-hours primary care? An exploratory qualitative interview study to inform a participatory action research project. BMJ Open 2018, 8, e023154. [Google Scholar] [CrossRef] [PubMed]
- Yunquera-Romero, L.; Márquez-Gómez, I.; Henares-López, A.; Morales-Lara, M.J.; Gallego Fernández, C.; Asensi-Díez, R. Appropriateness of antimicrobial prescriptions in the emergency department of a tertiary hospital. Rev. Esp. Quimioter. 2018, 31, 209–216. [Google Scholar]
- Denny, K.J.; Gartside, J.G.; Alcorn, K.; Cross, J.W.; Maloney, S.; Keijzers, G. Appropriateness of antibiotic prescribing in the emergency department. J. Antimicrob. Chemother. 2019, 74, 515–520. [Google Scholar] [CrossRef]
- Labricciosa, F.M.; Sartelli, M.; Correia, S.; Abbo, L.M.; Severo, M.; Ansaloni, L.; Coccolini, F.; Alves, C.; Melo, R.B.; Baiocchi, G.L.; et al. Emergency surgeons’ perceptions and attitudes towards antibiotic prescribing and resistance: A worldwide cross-sectional survey. World J. Emerg. Surg. 2018, 13, 27. [Google Scholar] [CrossRef]
- Draper, H.M.; Farland, J.B.; Heidel, R.E.; May, L.S.; Suda, K.J. Comparison of bacteria isolated from emergency department patients versus hospitalized patients. Am. J. Health Syst. Pharm. 2013, 70, 2124–2128. [Google Scholar] [CrossRef]
- Rothe, K.; Wantia, N.; Spinner, C.D.; Schneider, J.; Lahmer, T.; Waschulzik, B.; Schmid, R.M.; Busch, D.H.; Katchanov, J. Antimicrobial resistance of bacteraemia in the emergency department of a German university hospital (2013–2018): Potential carbapenem-sparing empiric treatment options in light of the new EUCAST recommendations. BMC Infect. Dis. 2019, 19, 1091. [Google Scholar] [CrossRef] [Green Version]
- Kao, C.-H.; Kuo, Y.-C.; Chen, C.-C.; Chang, Y.-T.; Chen, Y.-S.; Wann, S.-R.; Liu, Y.-C. Isolated pathogens and clinical outcomes of adult bacteremia in the emergency department: A retrospective study in a tertiary referral center. J. Microbiol. Immunol. Infect. 2011, 44, 215–221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Merritt, C.; Haran, J.P.; Mintzer, J.; Stricker, J.; Merchant, R.C. All purulence is local—Epidemiology and management of skin and soft tissue infections in three urban emergency departments. BMC Emerg. Med. 2013, 13, 26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zatorski, C.; Jordan, J.A.; Cosgrove, S.E.; Zocchi, M.; May, L. Comparison of antibiotic susceptibility of escherichia coli in urinary isolates from an emergency department with other institutional susceptibility data. Am. J. Health Syst. Pharm. 2015, 72, 2176–2180. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.-C.; Hung, Y.-P.; Lin, H.-J.; Liu, H.-C.; Lee, J.-C.; Wu, Y.-H.; Li, C.-W.; Li, M.-C.; Ko, W.-C. Antimicrobial susceptibility of clinical enterobacteriaceae isolates at the emergency department in a regional hospital: A threat of extended spectrum beta-lactamase-producers among nursing home residents. J. Microbiol. Immunol. Infect. 2016, 49, 584–590. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jorgensen, S.; Zurayk, M.; Yeung, S.; Terry, J.; Dunn, M.; Nieberg, P.; Wong-Beringer, A. Emergency department urinary antibiograms differ by specific patient group. J. Clin. Microbiol. 2017, 55, 2629–2636. [Google Scholar] [CrossRef] [Green Version]
- Frazee, B.W.; Trivedi, T.; Montgomery, M.; Petrovic, D.-F.; Yamaji, R.; Riley, L. Emergency department urinary tract infections caused by extended-spectrum β-lactamase-producing enterobacteriaceae: Many patients have no identifiable risk factor and discordant empiric therapy is common. Ann. Emerg. Med. 2018, 72, 449–456. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control. Surveillance of Antimicrobial Resistance in Europe. Annual Report of the European Antimicrobial Resistance Surveillance Network (EARS-Net) 2018; ECDC: Stockholm, Sweden, 2019. [Google Scholar]
- Poon, E.; Self, L.; McLeod, S.L.; Caine, S.; Borgundvaag, B. Uncomplicated urinary tract infections in the emergency department: A review of local practice patterns. CJEM 2018, 20, 572–577. [Google Scholar] [CrossRef] [Green Version]
- De Angelis, G.; Fiori, B.; Menchinelli, G.; D’Inzeo, T.; Liotti, F.M.; Morandotti, G.A.; Sanguinetti, M.; Posteraro, B.; Spanu, T. Incidence and antimicrobial resistance trends in bloodstream infections caused by ESKAPE and escherichia coli at a large teaching hospital in Rome, a 9-year analysis (2007–2015). Eur. J. Clin. Microbiol. Infect. Dis. 2018, 37, 1627–1636. [Google Scholar] [CrossRef]
- Magiorakos, A.-P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [Green Version]
- Kadri, S.S.; Adjemian, J.; Lai, Y.L.; Spaulding, A.B.; Ricotta, E.; Prevots, D.R.; Palmore, T.N.; Rhee, C.; Klompas, M.; Dekker, J.P.; et al. Difficult-to-treat resistance in gram-negative bacteremia at 173 US hospitals: Retrospective cohort analysis of prevalence, predictors, and outcome of resistance to all first-line agents. Clin. Infect. Dis. 2018, 67, 1803–1814. [Google Scholar] [CrossRef] [Green Version]
- Kadri, S.S.; Lai, Y.L.E.; Ricotta, E.E.; Strich, J.R.; Babiker, A.; Rhee, C.; Klompas, M.; Dekker, J.P.; Powers, J.H., 3rd; Danner, R.L.; et al. External validation of difficult-to-treat resistance prevalence and mortality risk in gram-negative bloodstream infection using electronic health record data from 140 U.S. hospitals. Open Forum. Infect. Dis. 2019, 6, ofz110. [Google Scholar] [CrossRef] [PubMed]
- Huh, K.; Chung, D.R.; Ha, Y.E.; Ko, J.-H.; Kim, S.-H.; Kim, M.-J.; Huh, H.J.; Lee, N.Y.; Cho, S.Y.; Kang, C.-I.; et al. Impact of difficult-to-treat resistance in gram-negative bacteremia on mortality: Retrospective analysis of nationwide surveillance data. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Giannella, M.; Bussini, L.; Pascale, R.; Bartoletti, M.; Malagrinò, M.; Pancaldi, L.; Toschi, A.; Ferraro, G.; Marconi, L.; Ambretti, S.; et al. Prognostic utility of the new definition of difficult-to-treat resistance among patients with gram-negative bloodstream infections. Open Forum. Infect. Dis. 2019, 6, ofz505. [Google Scholar] [CrossRef] [PubMed]
- Muntean, D.; Horhat, F.-G.; Bădițoiu, L.; Dumitrașcu, V.; Bagiu, I.-C.; Horhat, D.-I.; Coșniță, D.A.; Krasta, A.; Dugăeşescu, D.; Licker, M. Multidrug-resistant gram-negative bacilli: A retrospective study of trends in a tertiary healthcare unit. Medicina 2018, 54, 92. [Google Scholar] [CrossRef] [Green Version]
- Giani, T.; Antonelli, A.; Caltagirone, M.; Mauri, C.; Nicchi, J.; Arena, F.; Nucleo, E.; Bracco, S.; Pantosti, A.; Luzzaro, F.; et al. Evolving beta-lactamase epidemiology in enterobacteriaceae from Italian nationwide surveillance, October 2013: KPC-carbapenemase spreading among outpatients. Eurosurveillance 2017, 22, 30583. [Google Scholar] [CrossRef]
- Touati, A.; Mairi, A. Epidemiology of carbapenemase-producing enterobacterales in the Middle East: A systematic review. Expert Rev. Anti-Infect. Ther. 2020, 18, 241–250. [Google Scholar] [CrossRef]
- Manenzhe, R.I.; Zar, H.J.; Nicol, M.P.; Kaba, M. The spread of carbapenemase-producing bacteria in Africa: A systematic review. J. Antimicrob. Chemother. 2015, 70, 23–40. [Google Scholar] [CrossRef] [Green Version]
- Tumbarello, M.; Sanguinetti, M.; Montuori, E.; Trecarichi, E.M.; Posteraro, B.; Fiori, B.; Citton, R.; D’Inzeo, T.; Fadda, G.; Cauda, R.; et al. Predictors of mortality in patients with bloodstream infections caused by extended-spectrum-beta-lactamase-producing enterobacteriaceae: Importance of inadequate initial antimicrobial treatment. Antimicrob. Agents Chemother. 2007, 51, 1987–1994. [Google Scholar] [CrossRef] [Green Version]
- Lee, C.-C.; Lee, C.-H.; Hong, M.-Y.; Hsieh, C.-C.; Tang, H.-J.; Ko, W.-C. Propensity-matched analysis of the impact of extended-spectrum β-lactamase production on adults with community-onset escherichia coli, klebsiella species, and proteus mirabilis bacteremia. J. Microbiol. Immunol. Infect. 2018, 51, 519–526. [Google Scholar] [CrossRef]
- Gajdács, M.; Bátori, Z.; Ábrók, M.; Lázár, A.; Burián, K. Characterization of resistance in gram-negative urinary isolates using existing and novel indicators of clinical relevance: A 10-year data analysis. Life 2020, 10, 16. [Google Scholar] [CrossRef] [Green Version]
- ECDC European Centre for Disease Prevention and Control (ECDC) Antimicrobial Consumption Database (ESAC-Net). Available online: https://ecdc.europa.eu/en/antimicrobial-consumption/surveillance-and-disease-data/database (accessed on 10 August 2020).
- Gajdács, M. Carbapenem-resistant but cephalosporin-susceptible pseudomonas aeruginosa in urinary tract infections: Opportunity for colistin sparing. Antibiotics 2020, 9, 153. [Google Scholar] [CrossRef] [Green Version]
- Leber, A.L. Clinical Microbiology Procedures Handbook, 4th ed.; ASM Press: Washington, DC, USA, 2016; ISBN 978-1-55581-880-7. [Google Scholar]
- Ábrók, M.; Lázár, A.; Szécsényi, M.; Deák, J.; Urbán, E. Combination of MALDI-TOF MS and PBP2′ latex agglutination assay for rapid MRSA detection. J. Microbiol. Methods 2018, 144, 122–124. [Google Scholar] [CrossRef] [PubMed]
- Sali, M.; De Maio, F.; Tarantino, M.; Garofolo, G.; Tittarelli, M.; Sacchini, L.; Zilli, K.; Pasquali, P.; Petrucci, P.; Marianelli, C.; et al. Rapid and safe one-step extraction method for the identification of brucella strains at genus and species level by MALDI-TOF mass spectrometry. PLoS ONE 2018, 13, e0197864. [Google Scholar] [CrossRef]
- Rawat, D.; Nair, D. Extended-spectrum β-lactamases in gram negative bacteria. J. Glob. Infect. Dis. 2010, 2, 263–274. [Google Scholar] [CrossRef] [PubMed]
- McDonnell, A.; Rex, J.H.; Goossens, H.; Bonten, M.; Fowler, V.G.J.; Dane, A. Efficient delivery of investigational antibacterial agents via sustainable clinical trial networks. Clin. Infect. Dis. 2016, 63 (Suppl. 2), S57–S59. [Google Scholar] [CrossRef] [PubMed]
- European Committee on Antimicrobial Susceptibility Testing EUCAST advice on intrinsic resistance and exceptional phenotypes v 3.2. 2020. Available online: https://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Expert_Rules/2020/Intrinsic_Resistance_and_Unusual_Phenotypes_Tables_v3.2_20200225.pdf (accessed on 2 March 2020).
| Gram Stain | Bacterial Family/Genus/Species | Frequency (%) |
|---|---|---|
| Gram-positive n = 1032 (20.75%) | Enterococcus spp. | 471 (9.47%) |
| Staphylococcus aureus | 561 (11.28%) | |
| Gram-negative n = 3942 (79.25%) | Stenotrophomonas maltophilia | 4 (0.08%) |
| Klebsiella spp. | 664 (13.35%) | |
| Acinetobacter baumannii | 31 (0.62%) | |
| Pseudomonas aeruginosa | 181 (3.64%) | |
| Enterobacterales: Escherichia spp. | 2194 (44.11%) | |
| Enterobacterales: Proteus spp. | 526 (10.57%) | |
| Enterobacterales: Enterobacter spp. | 119 (2.39%) | |
| Enterobacterales:Morganella spp. | 75 (1.51%) | |
| Enterobacterales: Citrobacter spp. | 53 (1.07%) | |
| Enterobacterales: Providencia spp. | 40 (0.8%) | |
| Enterobacterales: Salmonella spp. | 33 (0.66%) | |
| Enterobacterales: Serratia spp. | 22 (0.44%) | |
| Total | 4974 (100%) |
| Acro-nym | Bacterial Family/Genus/Species | Blood Culture (%) | Catheter-Specimen Urine (%) | Midstream Urine (%) | Wound or Abscess (%) | Others (%) | Total (%) |
|---|---|---|---|---|---|---|---|
| E | Enterococcus spp. | 93 (19.7%) | 191 (40.6%) | 76 (16.1%) | 48 (10.2%) | 63 (13.4%) | 471 (100%) |
| S | Staphylococcus aureus | 252 (64.5%) | 12 (3.1%) | 6 (1.5%) | 121 (30.9%) | 0 (0%) | 391 (100%) |
| Stenotrophomonas maltophilia | (0%) | 1 (0.6%) | (0%) | 3 (1.7%) | 170 (97.7%) | 174 (100%) | |
| K | Klebsiella spp. | 212 (31.9%) | 234 (35.2%) | 118 (17.8%) | 49 (7.4%) | 51 (7.7%) | 664 (100%) |
| A | Acinetobacter baumannii | 6 (19.4%) | 6 (19.4%) | (0%) | 5 (16.1%) | 14 (45.2%) | 31 (100%) |
| P | Pseudomonas aeruginosa | 47 (26%) | 53 (29.3%) | 16 (8.8%) | 25 (13.8%) | 40 (22.1%) | 181 (100%) |
| E | Enterobacterales: Citrobacter spp. | 13 (24.5%) | 17 (32.1%) | 7 (13.2%) | 9 (17%) | 7 (13.2%) | 53 (100%) |
| Enterobacterales: Enterobacter spp. | 27 (22.7%) | 30 (25.2%) | 9 (7.6%) | 24 (20.2%) | 29 (24.4%) | 119 (100%) | |
| Enterobacterales: Escherichia spp. | 676 (30.8%) | 823 (37.5%) | 531 (24.2%) | 75 (3.4%) | 89 (4.1%) | 2194 (100%) | |
| Enterobacterales: Morganella morganii | 25 (33.3%) | 11 (14.7%) | 2 (2.7%) | 21 (28%) | 16 (21.3%) | 75 (100%) | |
| Enterobacterales: Proteus spp. | 116 (22.1%) | 174 (33.1%) | 44 (8.4%) | 99 (18.8%) | 93 (17.7%) | 526 (100%) | |
| Enterobacterales: Providencia spp. | 6 (15%) | 15 (37.5%) | 2 (5%) | 8 (20%) | 9 (22.5%) | 40 (100%) | |
| Enterobacterales: Salmonella spp. | 29 (87.9%) | 3 (9.1%) | 1 (3%) | 0 (0%) | 0 (0%) | 33 (100%) | |
| Enterobacterales: Serratia spp. | 5 (22.7%) | 1 (4.5%) | 2 (9.1%) | 7 (31.8%) | 7 (31.8%) | 22 (100%) | |
| Total | 1507 (30.3%) | 1571 (31.6%) | 814 (16.4%) | 494 (9.9%) | 588 (11.8%) | 4974 (100%) | |
| Bacterial Family/Genus/Species | Isolates (100%) | Wt (%) | UDR (%) | MDR (%) | DTR (%) | XDR (%) |
|---|---|---|---|---|---|---|
| Enterococcus spp. | 471 | 18 (3.82%) | 0 (0%) | |||
| Staphylococcus aureus | 561 | 110 (19.61%) | 0 (0%) | |||
| Stenotrophomonas maltophilia | 4 | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | |
| Klebsiella spp. | 664 | 390 (58.73%) | 274 (41.27%) | 182 (27.41%) | 1 (0.15%) | 0 (0%) |
| Acinetobacter baumannii | 31 | 4 (12.90%) | 27 (87.10%) | 21 (67.74%) | 13 (41.94%) | 0 (0%) |
| Pseudomonas aeruginosa | 181 | 117 (64.64%) | 64 (35.36%) | 24 (13.26%) | 8 (4.42%) | 1 (0.55%) |
| Citrobacter spp. | 53 | 38 (71.70%) | 15 (28.30%) | 6 (11.32%) | 0 (0%) | 0 (0%) |
| Enterobacter spp. | 119 | 79 (66.39%) | 40 (33.61%) | 14 (11.76%) | 0 (0%) | 0 (0%) |
| Escherichia spp. | 2194 | 891 (40.61%) | 1303 (59.39%) | 572 (26.07%) | 0 (0%) | 0 (0%) |
| Morganellas spp. | 75 | 12 (16%) | 63 (84.00%) | 14 (18.67%) | 0 (0%) | 0 (0%) |
| Proteus spp. | 526 | 129 (24.52%) | 397 (75.48%) | 214 (40.68%) | 1 (0.19%) | 0 (0%) |
| Providencia spp. | 40 | 14 (35%) | 26 (65.00%) | 5 (12.50%) | 0 (0%) | 0 (0%) |
| Salmonella spp. | 33 | 28 (84.85%) | 5 (15.15%) | 0 (0%) | 0 (0%) | 0 (0%) |
| Serratia spp. | 22 | 13 (59.09%) | 9 (40.91%) | 3 (13.64%) | 0 (0%) | 0 (0%) |
| Total | 4974 | 2751 (55.31%) | 2223 (44.69%) | 1183 (23.78%) | 23 (0.46%) | 1 (0.02%) |
| OXA | AMP | CC | CIP * | ERY | GM | LNZ | TEC | VA | TGC | SXT | MDR | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Enterococcus faecalis | IR | 100.0 | IR | 56.8 | IR | IR | 99.1 | 100.0 | 100.0 | 99.3 | IR | 0.7 |
| Enterococcus faecium | IR | IR | IR | 7.1 | IR | IR | 100.0 | 78.8 | 66.7 | 100.0 | IR | 45.5 |
| Staphylococcus aureus | 83.1 | IR | 79.4 | 80.7 | 78.0 | 98.6 | 100.0 | 100.0 | 100.0 | 100.0 | 98.6 | 19.6 |
| MSSA | - | IR | 89.2 | 92.8 | 88.2 | 99.4 | 100.0 | 99.8 | 100.0 | 100.0 | 98.7 | 3.2 |
| MRSA | - | IR | 30.9 | 22.1 | 27.7 | 94.7 | 100.0 | 100.0 | 100.0 | 100.0 | 97.9 | 100.0 |
| AMP | AMC | CXM | CTX | CAX | CAZ | FEP | TZP | ETP | MEM | CIP | AK | GM | SXT | MDR | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Klebsiella aerogenes | 0.0 | IR | 66.7 | 75.0 | 80.6 | 80.6 | 85.7 | 80.0 | 100.0 | 100.0 | 87.1 | 100.0 | 96.8 | 93.5 | 22.6 |
| Klebsiella oxytoca | 0.0 | 91.1 | 96.8 | 98.0 | 97.5 | 97.5 | 98.0 | 98.0 | 100.0 | 100.0 | 100.0 | 98.0 | 100.0 | 98.7 | 2.53 |
| Klebsiella pneumoniae | 0.0 | 67.5 | 67.2 | 73.0 | 69.4 | 69.4 | 76.6 | 76.1 | 99.8 | 99.8 | 62.8 | 79.3 | 79.1 | 65.5 | 32.6 |
| Klebsiella variicola | 0.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 0.0 |
| Acinetobacter baumannii | NT | IR | NT | NT | IR | NT | NT | NT | IR | 55.2 | 13.8 | 37.9 | 40.7 | 21.4 | 65.5 |
| Pseudomonas aeruginosa | NT | IR | NT | NT | IR | 90.6 | 91.7 | 87.2 | IR | 85.6 | 77.9 | 96.7 | 90.1 | NT | 13.3 |
| Citrobacter freundii | 0.0 | IR | 20.0 | 82.4 | 71.4 | 71.4 | 82.4 | 82.4 | 100.0 | 100.0 | 90.5 | 100.0 | 100.0 | 100.0 | 19 |
| Citrobacter koseri | 0.0 | 81.5 | 77.3 | 100.0 | 100.0 | 100.0 | 100.0 | 85.7 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 3.7 |
| Enterobacter cloacae | 0.0 | IR | 38.8 | 84.4 | 83.0 | 83.0 | 84.4 | 84.1 | 100.0 | 100.0 | 93.3 | 98.4 | 98.9 | 93.2 | 11.2 |
| Escherichia coli | 48.5 | 81.8 | 85.4 | 85.8 | 86.1 | 86.2 | 86.7 | 85.0 | 100.0 | 100.0 | 67.9 | 89.2 | 90.3 | 72.7 | 26.1 |
| Morganella morganii | 0.0 | IR | 0.0 | 93.3 | 90.3 | 90.3 | 93.3 | 90.0 | 100.0 | 100.0 | 90.3 | 91.5 | 91.7 | 72.2 | 18.1 |
| Proteus mirabilis | 40.4 | 62.3 | 63.0 | 71.7 | 69.1 | 69.1 | 75.5 | 74.7 | 99.8 | 100.0 | 57.7 | 75.7 | 77.2 | 32.1 | 42.7 |
| Proteus vulgaris | 0.0 | 40.0 | 0.0 | 95.5 | 91.4 | 91.4 | 95.5 | 95.5 | 100.0 | 100.0 | 85.7 | 95.5 | 91.4 | 65.7 | 14.3 |
| Providencia stuartii | 0.0 | IR | 31.6 | 93.3 | 96.6 | 96.6 | 92.9 | 92.9 | 100.0 | 100.0 | 72.4 | 36.4 | 28.6 | 29.6 | 17.2 |
| Salmonella spp. | 90.6 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 96.0 | 100.0 | 100.0 | 96.9 | 0.0 |
| Enterobacterales | 34.6 | 70.4 | 76.0 | 83.1 | 82.3 | 82.3 | 84.9 | 83.5 | 99.9 | 100.0 | 69.4 | 86.1 | 87.5 | 68.5 | 27.1 |
| Enterobacterales non ESBL | 100.0 | 100.0 | 82.2 | 93.5 | 93.8 | 77.1 | 11.8 | ||||||||
| Enterobacterales ESBL | 99.7 | 99.8 | 9.6 | 46.8 | 58.6 | 27.9 | 98.5 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Benkő, R.; Gajdács, M.; Matuz, M.; Bodó, G.; Lázár, A.; Hajdú, E.; Papfalvi, E.; Hannauer, P.; Erdélyi, P.; Pető, Z. Prevalence and Antibiotic Resistance of ESKAPE Pathogens Isolated in the Emergency Department of a Tertiary Care Teaching Hospital in Hungary: A 5-Year Retrospective Survey. Antibiotics 2020, 9, 624. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics9090624
Benkő R, Gajdács M, Matuz M, Bodó G, Lázár A, Hajdú E, Papfalvi E, Hannauer P, Erdélyi P, Pető Z. Prevalence and Antibiotic Resistance of ESKAPE Pathogens Isolated in the Emergency Department of a Tertiary Care Teaching Hospital in Hungary: A 5-Year Retrospective Survey. Antibiotics. 2020; 9(9):624. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics9090624
Chicago/Turabian StyleBenkő, Ria, Márió Gajdács, Mária Matuz, Gabriella Bodó, Andrea Lázár, Edit Hajdú, Erika Papfalvi, Peter Hannauer, Péter Erdélyi, and Zoltán Pető. 2020. "Prevalence and Antibiotic Resistance of ESKAPE Pathogens Isolated in the Emergency Department of a Tertiary Care Teaching Hospital in Hungary: A 5-Year Retrospective Survey" Antibiotics 9, no. 9: 624. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics9090624






