Differencing the Risk of Reiterative Spatial Incidence of COVID-19 Using Space–Time 3D Bins of Geocoded Daily Cases
Abstract
:1. Introduction
2. Materials and Methods
2.1. Overview of the Study Area
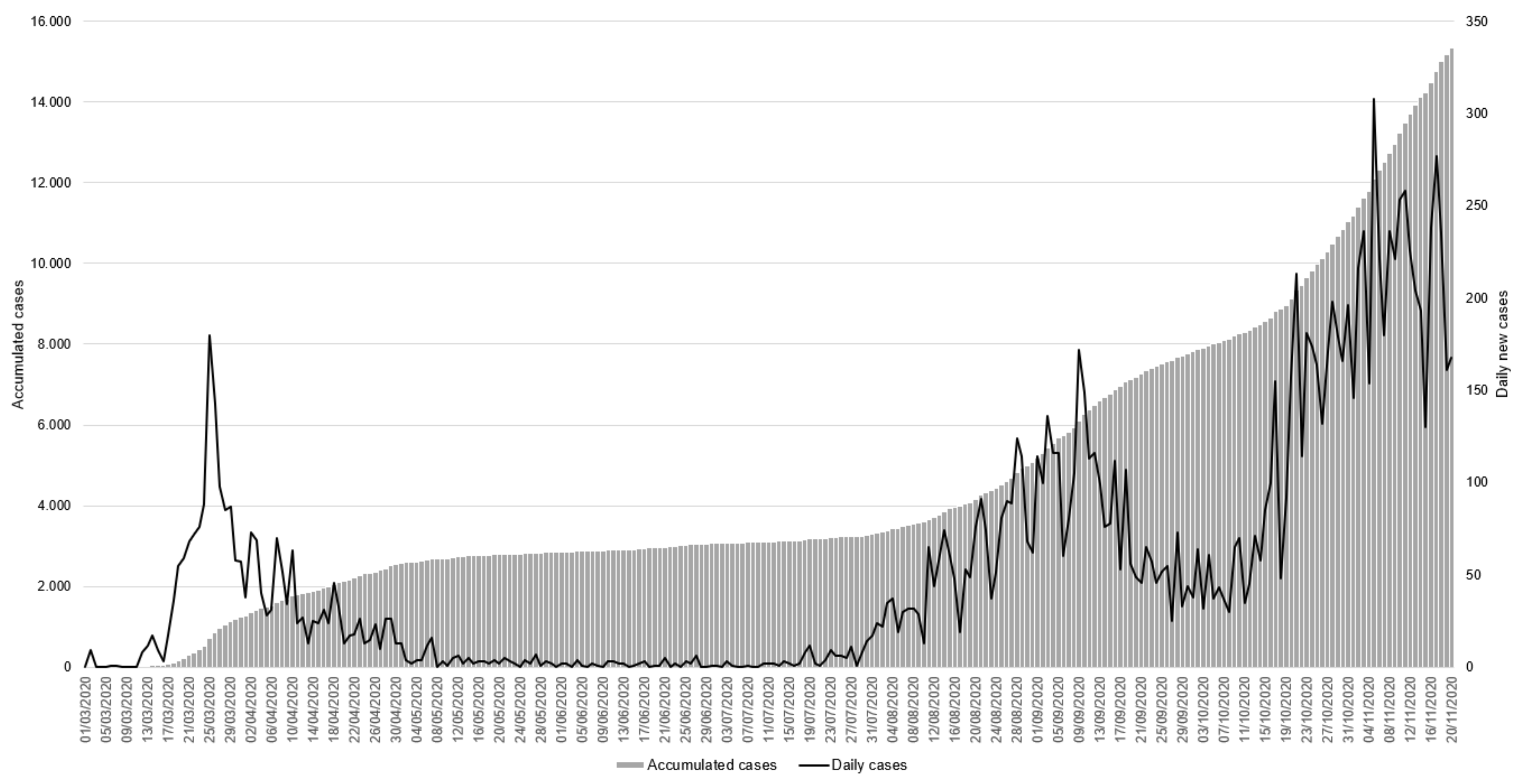
2.2. The Main Source: Daily Microdata on Reported Cases
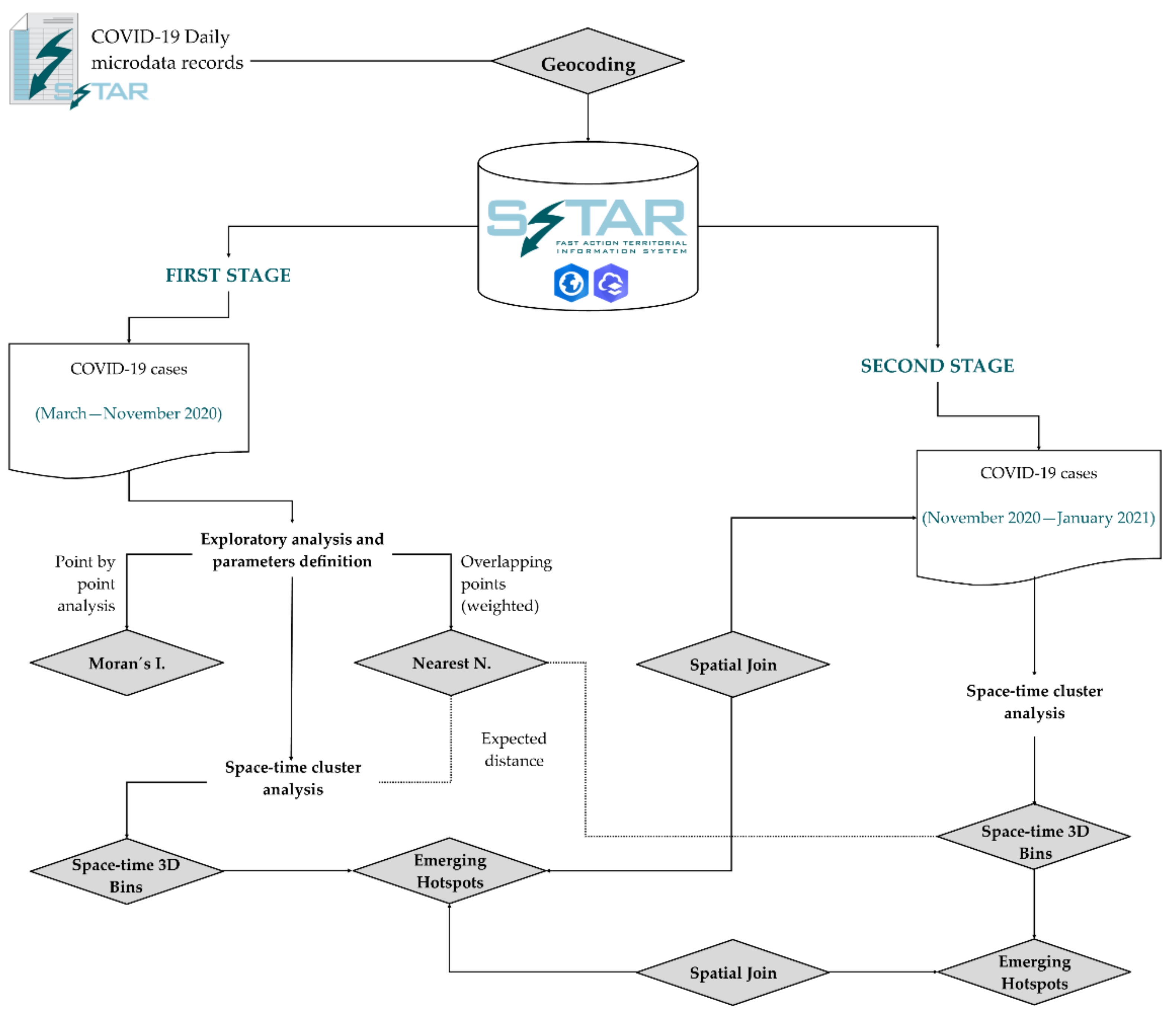
2.3. Research Methods
2.3.1. First Stage, from March to November 2020: 3D Bins and Emerging Hotspots Analysis
- The organisation of time intervals starts from three basic principles. The first are the temporal references used by health authorities, which frequently handle terms close to two weeks (for example, for confining close contacts and for estimating accumulated incidents of 14 or, where appropriate, 7 days to follow the trend). Secondly, we must consider the methodological restriction of the 3D bins creation tool that requires a minimum of ten time intervals and, finally, an adjusted number of intervals close to 10 as the excess of intervals in the tests performed created too many cube moments with no cases or very low counts. The effect of excessive time compartmentalisation tends to disconnect emerging countries from the general pattern of the pandemic wave. As a result of the three criteria described, the 4-week interval is suitable since it includes 2 periods of the usual reference time for the study of accumulated incidents (i.e., 14 days) and meets the condition for the method to be applied that establishes a minimum of 10 moments in time for the development of bins. As we mentioned before, exploratory analysis of slides based on 1 period (2 weeks instead of 4) produced too many bins (many of them with 0 cases) and the results were not adequate.
- The bin size is a metric based on expected distance (nearest neighbour analysis) from layer of points with COVID-19 cases and a weight field of location counts equivalent to number of positive cases in the same geographical coordinate. It is an objective size that improves the results for observed distance (bins too small and with 0 cases or only 1 case) and gives an objective parameter for reproducing this method at other times and elsewhere, with the nearest neighbour analysis needing to be calculated first. The size of the cubes based on the expected distance avoids distortions derived from cube sizes that are too small (which do not provide a spatial pattern with respect to the geocoded point layer) and also the possible cancellation or concealment of spatial differences if the cubes that are generated were too big. The tests carried out at both the municipal and regional levels support the value of the expected distance as a distance parameter of the bins of COVID-19.
2.3.2. Second Stage, from November 2020 to January 2021: Checking the Predictive Potential of Emerging Hotspots
- New cases outside the emerging hotspots modelled.
- New cases inside previous emerging hotspots, where there are two possible combinations: one corresponding to “no pattern detected” because statistical significance is not reached, and the other consisting of emerging hotspots with statistical significance.
3. Results
3.1. First Stage: 3D Bins and Emerging Hotspots in Cantabria from March to November 2020
- Two hundred and fifty-one bins (41.69%) are new hotspots, i.e., these are hotspots with statistical significance only in the final part of the period analysed (October–November). These bins contain 2190 cases (15.75% of the series analysed, 20.96% of cases in hotspots with statistical significance).
- One hundred and seventy-eight bins (29.57%) are oscillating hotspots. These are hotspots which are significant towards the end of the period (October–November) but show a previous trend in which they were significant cold spots. Less than 90% of the time intervals in these cases have shown significant hotspots. In Cantabria, this type is ranked second not only in number of bins but also in number of cases, with 3163 reported positive cases of COVID-19 (22.74% of the series analysed, 30.28% of cases in hotspots with statistical significance). This type is also where the lowest mean age of cases is detected (41.7 years).
- Ninety-eight bins (16.28%) are sporadic hotspots. This type corresponds to locations that switch from hotspot to non-hotspot status several times in the period considered. They show up as significant hotspots in less than 90% of the time intervals and never as significant cold spots. This type is striking in that in 16.28% of statistically significant cubes, it is the one with the most COVID-19 cases, with a total of 3988 cases (28.68% of the series analysed, 38.18% of cases in hotspots with statistical significance). Furthermore, possibly related to the large number of cases, it is in this hotspot pattern where most deaths are found when those in residential homes are excluded (42 deaths, equivalent to 31%).
- Seventy-five bins (12.46%) are consecutive hotspots. These are areas with significant hotspots in a single run without interruption in the final time intervals considered. These cubes were not significant hotspots before the last run. These hotspots are also the least common in terms of number of cases, accounting for 1105 cases (7.95% of the series analysed, 10.58% of cases in hotspots with statistical significance) in all.
3.2. Testing the Predictive Potential of the Emerging Initial Model: The Spread of the Pandemic Two Months Later
3.2.1. The Predictive Power of Analytics of Emerging Hotspots Is Confirmed
3.2.2. Emerging Patterns Detected Two Months Later
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- De Cos, O.; Castillo, V.; Cantarero, D. Facing a Second Wave from a Regional View: Spatial Patterns of COVID-19 as a Key Determinant for Public Health and Geoprevention Plans. Int. J. Environ. Res. Public Health 2020, 17, 8468. [Google Scholar]
- Cromley, E.K. Using GIS to address epidemiologic research questions. Curr. Epidemiol. Rep. 2019, 6, 162–173. [Google Scholar] [CrossRef]
- Gerber, T.D.; Ping, D.; Armstrong-Brown, J.; McNutt, L.A.; Cole, F.B. Charting a path to location intelligence for STD control. Public Health Rep. 2009, 124, 49–57. [Google Scholar] [CrossRef] [Green Version]
- Jindal, C.; Kumar, S.; Sharma, S.; Choi, Y.M.; Efird, J.T. The Prevention and Management of COVID-19: Seeking a Practical and Timely Solution. Int. J. Environ. Res. Public Health 2020, 17, 3986. [Google Scholar] [CrossRef] [PubMed]
- Kleinman, K.P.; Abrams, A.M.; Kulldorff, M.; Platt, R. A model-adjusted space-time scan statistic with an application to syndromic surveillance. Epidemio. Infect. 2005, 133, 409–419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kulldorff, M.; Heffernan, R.; Hartman, J.; Assuncao, R.; Mostashari, F. A space-time permutation scan statistic for disease outbreak detection. PLoS Med. 2005, 2, e59. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, F.; Cheng, S.; He, G.; Huang, F.; Zhang, H.; Xu, B.; Murimwa, T.C.; Cheng, J.; Hu, D.; Wang, L. Space-Time Clustering Characteristics of Tuberculosis in China, 2005–2011. PLoS ONE 2013, 8, e83605. [Google Scholar] [CrossRef] [PubMed]
- Pangilinan, M.A.; Gonzalez, D.P.; Leong, R.N.; Co, F.F. Spatial analysis of the distribution of report dengue incidence in the National Capital Region, Philippines. Acta Med. Philipp. 2017, 51, 126–132. [Google Scholar] [CrossRef]
- Tzai-Hung, W. Analyzing the patterns of space-time distances of tracking the diffusion of an epidemic. In Book Space-Time Integration in Geography and GIScience; Mei-Po, K., Douglas, R., Donggen, W., Chenghu, Z., Eds.; Springer: Dordrecht, The Netherlands, 2015; pp. 269–282. [Google Scholar] [CrossRef]
- Al-Ahmadi, K.; Alahmadi, S.; Al-Zahrani, A. Spatiotemporal Clustering of Middle East Resparatory Syndorme Coronavirus (MERS-CoV) Incidence in Saudi Arabia, 2012–2019. Int. J. Environ. Res. Public Health 2019, 16, 2520. [Google Scholar] [CrossRef] [Green Version]
- Mala, S.; Jat, M.K. Geographic information system based spatio-temporal dengue fever cluster analysis and mapping. Egypt. J. Remote Sens. Space Sci. 2019, 22, 297–304. [Google Scholar] [CrossRef]
- Hughey, S.M.; Kaczynski, A.T.; Porter, D.E.; Hibbert, J.; Turner-McGrievy, G.; Liu, J. Spatial clustering patterns of child weight status in a southeastern US county. Appl. Geogr. 2018, 99, 12–21. [Google Scholar] [CrossRef]
- Greenough, P.G.; Nelson, E.L. Beyond mapping: A case for geospatial analytics in humanitarian health. Conflict Health 2019, 13, 50. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.; Yue, Y.; Zhou, X.; Zhang, X. How urban factors affect to spatiotemporal distribution of infectious diseases in addition to intercity population movement in China. ISPRS Int. J. Geo Inf. 2020, 9, 615. [Google Scholar] [CrossRef]
- Whittle, R.S.; Díaz-Artiles, A. An ecological study of socioeconomic predictors of detection of COVID-19 cases across neighborhoods in New York City. BMC Med. 2020, 18, 271. [Google Scholar] [CrossRef] [PubMed]
- Dhaval, D.D. Urban Densities and the COVID-19 Pandemic: Upending the Sustainability Myth of Global Megacities. ORF Occas. Pap. 2020, 244, 1–42. [Google Scholar]
- Hamidi, S.; Sabouri, S.; Ewing, R. Does density aggravate the COVID-19 Pandemic? Early findings and lessons for planners. J. Am. Plan. Assoc. 2020. [Google Scholar] [CrossRef]
- Bamweyana, I.; Okello, D.A.; Ssengendo, R. Socio-Economic Vulnerability to COVID-19: The spatial case of Greater Kampala Metropolitan Area (GKMA). J. GIS 2020, 12, 302–318. [Google Scholar] [CrossRef]
- Borjas, G.J. Demographic Determinants of Testing Incidence and COVID-19 Infections in New York City Neighborhoods; Working Paper: 26952; National Bureau of Economic Research: Cambridge, MA, USA, 2020. [Google Scholar]
- Liu, Y.; He, Y.; Xia, Z. Space-time variation and spatial differentiation of COVID-19 confirmed cases in Hubei province based on extended GWR. ISPRS Int. J. Geo Inf. 2020, 9, 536. [Google Scholar] [CrossRef]
- Hashim, H.; Wan-Mohd, W.M.N.; Sadek, E.S.S.M.; Dimyati, K.M. Modeling urban crime patterns using spatial space time and regression analysis. Int. Arch. Photogramm. Remote Sens. Spatial Inf. Sci. 2019, XLII-4/W16, 247–254. [Google Scholar] [CrossRef] [Green Version]
- Kang, Y.; Cho, N.; Son, S. Spatiotemporal characteristics of elderly population´s traffic accidents in Seoul using space-time cube and space-time kernel density estimation. PLoS ONE 2018, 13, e0196845. [Google Scholar] [CrossRef] [Green Version]
- Sayed, M.H.; Masoud, P.; Masoud, B.; Manoochehr, K.; Meysam, O. Pattern mining analysis of pulmonary TB cases in Hamadan province: Using space-time cube. Int. J. Epidemil. Res. 2017, 4, 111–117. [Google Scholar]
- Chunbao, M.; Dechan, T.; Tingyu, M.; Chunhua, B.; Jian, Q.; Weiyi, P.; Zhiyong, Z. An analysis of spatiotemporal pattern for COVID-19 in China based on space-time cube. J. Med. Virol. 2020, 92, 1587–1595. [Google Scholar]
- Thakar, V. Unfolding events in space and time: Geospatial insights into COVID-19 diffusion in Washington State during the initial stage of the outbreak. ISPRS Int. J. Geo Inf. 2020, 9, 382. [Google Scholar] [CrossRef]
- Franch-Pardo, I.; Napoletano, B.M.; Rosete-Verges, F.; Billa, L. Spatial analysis and GIS in the study of COVID-19. A review. Sci. Total Environ. 2020, 739, 140033. [Google Scholar] [CrossRef] [PubMed]
- Hemant, B.; Anjum, S.; Kumar, S.; Gautam, S.; Gupta, A.; Kumbhare, H.; Anshul, A.; Kumar, R. Understanding COVID-19 transmission through Bayesian probabilistic modeling and GIS-based Voronoi approach a policy perspective. Environ. Dev. Sustain. 2020. [Google Scholar] [CrossRef]
- Batista, F.; Poelman, H. Mapping Population Density in Functional Urban Areas. A Method to Downscale Population Statistics to Urban Atlas Polygons; JRC Technical Reports; European Commission: Luxembourg, 2016; Available online: https://tinyurl.com/y9r2mbkt (accessed on 10 May 2020).
- De Cos, O.; Castillo, V.; Cantarero, D. Scalable Analysis of COVID-19 Spatiotemporal Patterns Based on Data Mining Tools: Using 3D Bins to Predict Short-time Focus Locations. Res. Sq. Prepr. 2020. [Google Scholar] [CrossRef]
- ESRI COVID-19 GIS Hub. Available online: https://coronavirus-resources.esri.com/ (accessed on 15 June 2020).
- The Geoenrichment Service. ArcGIS REST API. Available online: https://developers.arcgis.com/rest/geoenrichment/api-reference/geoenrichment-service-overview.htm (accessed on 10 August 2020).
- Mollalo, A.; Vahedi, B.; Rivera, K.M. GIS-based spatial modeling of COVID-19 incidence rate in the continental United States. Sci. Total Environ. 2020, 728, 138884. [Google Scholar] [CrossRef]
- Abdrakhmanov, S.K.; Mukhanbetkaliyev, Y.Y.; Korennoy, F.I.; Karatayev, B.S.; Mukhanbetkaliyeva, A.A.; Abdrakhmanova, A.S. Spatio-temporal analysis and visualization of the anthrax epidemic situation in livestock in Kazakhstan over the period 1933–2016. Geospat. Health 2017, 12, 316–329. [Google Scholar]
- Zhao, Y.; Ge, L.; Liu, J.; Liu, H.; Yu, L.; Wang, N.; Zhou, Y.; Ding, X. Anaylzing hemorrhagic fever with renal syndrome in Hubei Province, China: A space-time cube-based approach. J. Int. Med. Res. 2019, 47, 3371–3388. [Google Scholar] [CrossRef] [Green Version]
- Kulldorff, M. Prospective time periodic geographical disease surveillance using scan statistic. J. R. Statist. Soc. A 2001, 164 Pt 1, 61–72. [Google Scholar] [CrossRef]
- Ogneva-Himmelberger, Y. Spatial analysis of drug poisoning deaths and Access to substance-use disorder treatment in the United States. In Proceedings of the 5th International Conference on Geographical Information Systems Theory, Applications and Management (GISTAM), Crete, Greece, 3–5 May 2019; pp. 315–321. [Google Scholar] [CrossRef]
- Kamel, M.N.; Geraghty, E.M. Geographical tracking and mapping of coronavirus disease COVID-19/severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) epidemic and associated events around the world: How 21st century GIS technologies are supporting the global fight against outbreaks and epidemics. Int. J. Health Geogr. 2020, 19, 8. [Google Scholar] [CrossRef] [Green Version]
- Zhou, C.; Su, F.; Pei, T.; Zhang, A.; Du, Y.; Luo, B.; Cao, Z.; Wang, J.; Yuan, W.; Zhu, Y.; et al. COVID-19: Challenges to GIS with Big Data. Geogr. Sustain. 2020, 1, 77–87. [Google Scholar] [CrossRef]
- Hernando, F. La seguridad en las ciudades: El nuevo enfoque de la geoprevención. Scr. Nova 2008, 12, 270. [Google Scholar]
- Sarwar, S.; Waheed, R.; Sarwar, S.; Khan, A. COVID-19 challenges to Pakistan: Is GIS analysis useful to draw solutions? Sci. Total Environ. 2020, 730, 139089. [Google Scholar] [CrossRef] [PubMed]



| Cases (COVID-19 Records) | First Stage (March 2020 to November 2020) | Second Stage (November 2020 to January 2021) |
|---|---|---|
| Cases initially reported | 15,168 | 6530 |
| Total geocoded cases | 14,938 | 6200 |
| Geocoded cases studied | 13,907 | 5766 |
| Nearest Neighbour Results | Layer of Cases (Points) | Layer of Location Counts |
|---|---|---|
| Observed distance | 38.7 m | 126.5 m |
| Expected distance | 408.2 m | 538.5 m |
| Nearest neighbour ratio | 0.095 | 0.217 |
| Z Score | −201.58 | −122.07 |
| p value | <0.01 | <0.01 |
| Emerging Pattern | Number of Bins | % Bins | Number of Cases | % Cases |
|---|---|---|---|---|
| No pattern detected | 812 | 57.43 | 3,461 | 24.89 |
| Pattern detected: | ||||
| New hotspots | 251 | 17.75 * 41.69 ** | 2190 | 15.75 * 20.96 ** |
| Oscillating hotspots | 178 | 12.59 * 29.57 ** | 3163 | 22.74 * 30.28 ** |
| Sporadic hotspots | 98 | 6.93 * 16.28 ** | 3988 | 28.68 * 38.18 ** |
| Consecutive hotspots | 75 | 5.30 * 12.46 ** | 1105 | 7.95 * 10.58 ** |
| Total pattern detected | 602 | 42,57 | 10,446 | 75.11 |
| Total emerging | 1414 | 100.00 | 13,907 | 100.00 |
| Emerging Pattern | Previous Pattern (Stage 1) | New Cases (Stage 2) | ||
|---|---|---|---|---|
| Area | % Area | COVID-19 Cases | % Cases | |
| Outside previous emerging hotspot | − | − | 425 | 7.37 |
| Total inside previous emerging hotspot | 410.79 * 158.61 ** | 100.00 | 5341 | 92.63 |
| No pattern detected | 235.90 * 91.08 ** | 57.43 | 907 | 16.98 |
| Pattern detected: | 174.89 * 67.53 ** | 42.57 | 4,434 | 83.02 |
| New hotspots | 72.92 * 28.15 ** | 17.75 | 987 | 18.48 |
| Oscillating hotspots | 51.71 * 19.97 ** | 12.59 | 1,355 | 25.37 |
| Sporadic hotspots | 28.47 * 10.99 ** | 6.93 | 1,608 | 30.11 |
| Consecutive hotspots | 21.79 * 8.41 ** | 5.30 | 484 | 9.06 |
| Cross Emerging Patterns: Stage One–Stage Two * | Number of Bins | Average % Time Hot | Average % Time Cold | Number of Cases |
|---|---|---|---|---|
| Sporadic–Intensifying ** | 57 | 98.92 | 0.00 | 1433 |
| No Pattern–No Pattern | 331 | 0.49 | 5.32 | 1013 |
| New–No Pattern | 133 | 5.44 | 2.37 | 662 |
| Oscillating–Sporadic | 58 | 51.19 | 0.00 | 632 |
| Consecutive–Sporadic | 40 | 35.00 | 0.00 | 354 |
| Oscillating–Intensifying ** | 20 | 92.69 | 0.00 | 326 |
| Oscillating–Consecutive ** | 46 | 29.93 | 0.00 | 325 |
| New–Consecutive ** | 31 | 23.82 | 0.00 | 298 |
| Sporadic–Consecutive ** | 24 | 30.77 | 0.00 | 224 |
| Not previous–No pattern | 96 | 0.32 | 2.32 | 168 |
| Consecutive–Consecutive | 15 | 29.23 | 0.00 | 141 |
| Oscillating–New | 10 | 7.69 | 2.31 | 36 |
| No Pattern–Consecutive ** | 2 | 38.46 | 0.00 | 30 |
| New–Sporadic ** | 2 | 42.31 | 0.00 | 21 |
| Oscillating–Oscillating | 2 | 26.92 | 7.69 | 19 |
| No Pattern–New ** | 3 | 7.69 | 0.00 | 13 |
| No Pattern–Oscillating ** | 4 | 15.38 | 7.69 | 13 |
| Oscillating–No Pattern | 6 | 0.00 | 20.51 | 10 |
| New–Oscillating ** | 3 | 15.38 | 7.69 | 9 |
| Not previous–Consecutive ** | 4 | 28.85 | 0.00 | 7 |
| New–New ** | 2 | 7.69 | 7.69 | 6 |
| Sporadic–Sporadic | 2 | 80.77 | 0.00 | 6 |
| Consecutive–No Pattern | 1 | 15.38 | 0.00 | 5 |
| Not previous–Oscillating ** | 2 | 15.38 | 7.69 | 4 |
| Consecutive–New | 3 | 7.69 | 0.00 | 4 |
| Consecutive–Oscillating | 1 | 23.08 | 7.69 | 4 |
| Not previous–Sporadic | 2 | 30.77 | 0.00 | 3 |
| Total | 900 | − | − | 5766 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Cos, O.; Castillo, V.; Cantarero, D. Differencing the Risk of Reiterative Spatial Incidence of COVID-19 Using Space–Time 3D Bins of Geocoded Daily Cases. ISPRS Int. J. Geo-Inf. 2021, 10, 261. https://0-doi-org.brum.beds.ac.uk/10.3390/ijgi10040261
De Cos O, Castillo V, Cantarero D. Differencing the Risk of Reiterative Spatial Incidence of COVID-19 Using Space–Time 3D Bins of Geocoded Daily Cases. ISPRS International Journal of Geo-Information. 2021; 10(4):261. https://0-doi-org.brum.beds.ac.uk/10.3390/ijgi10040261
Chicago/Turabian StyleDe Cos, Olga, Valentín Castillo, and David Cantarero. 2021. "Differencing the Risk of Reiterative Spatial Incidence of COVID-19 Using Space–Time 3D Bins of Geocoded Daily Cases" ISPRS International Journal of Geo-Information 10, no. 4: 261. https://0-doi-org.brum.beds.ac.uk/10.3390/ijgi10040261






