Modern Scientific Visualizations on the Web
Abstract
:1. Introduction
2. Survey Scope and Structure
Scientific Visualization (SciVis) Future Readiness Score
3. Representation Types and Techniques in Web-Based Visualization
3.1. Surface and Volume Rendering
3.2. Rendering in the Browser
3.3. Virtual and Augmented Reality in the Web
4. Data and File Formats
5. Scientific Applications of Web-Based Visualization
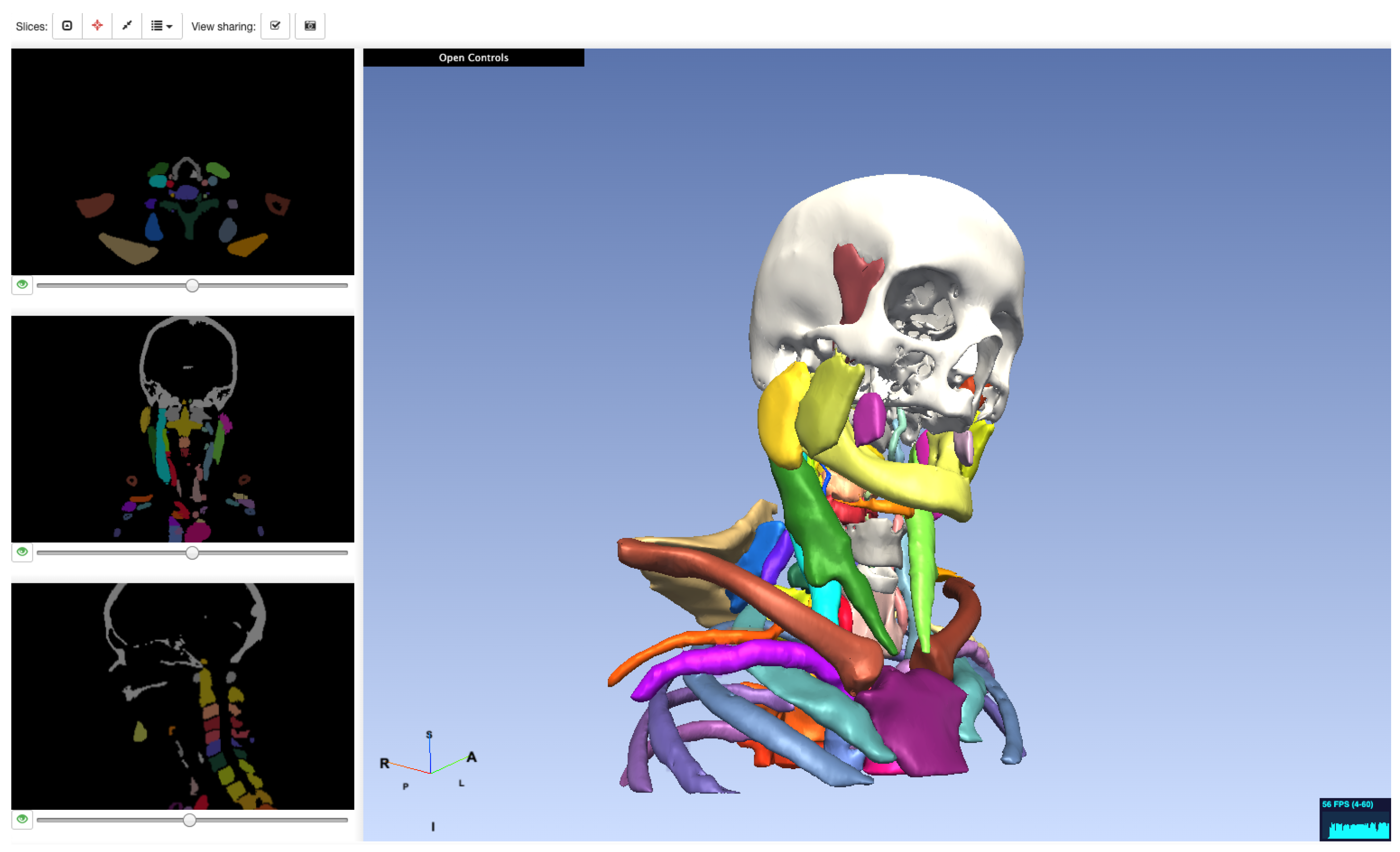
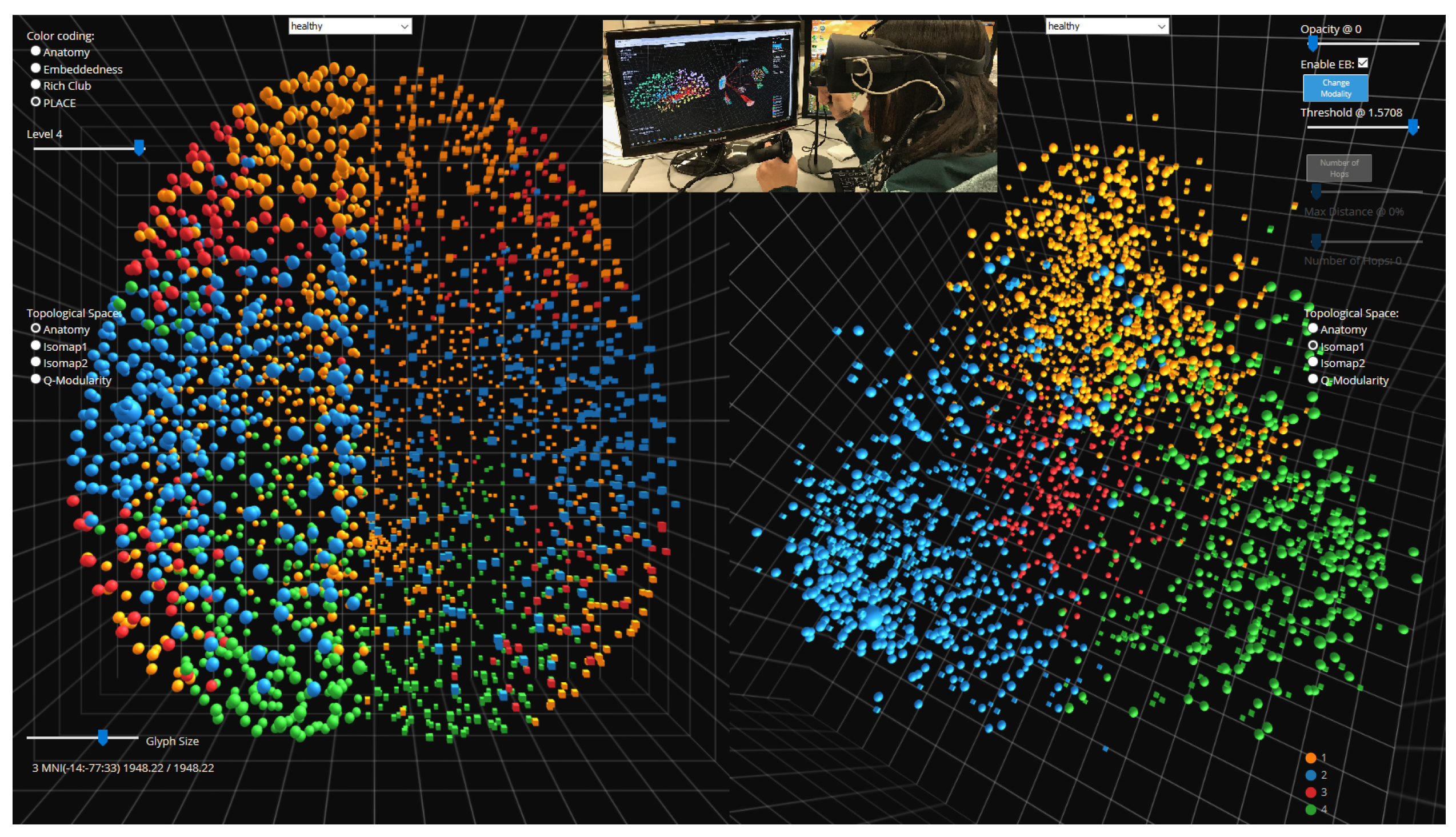
5.1. Medical Applications
5.2. Biology, Chemistry, and Molecular Visualizations
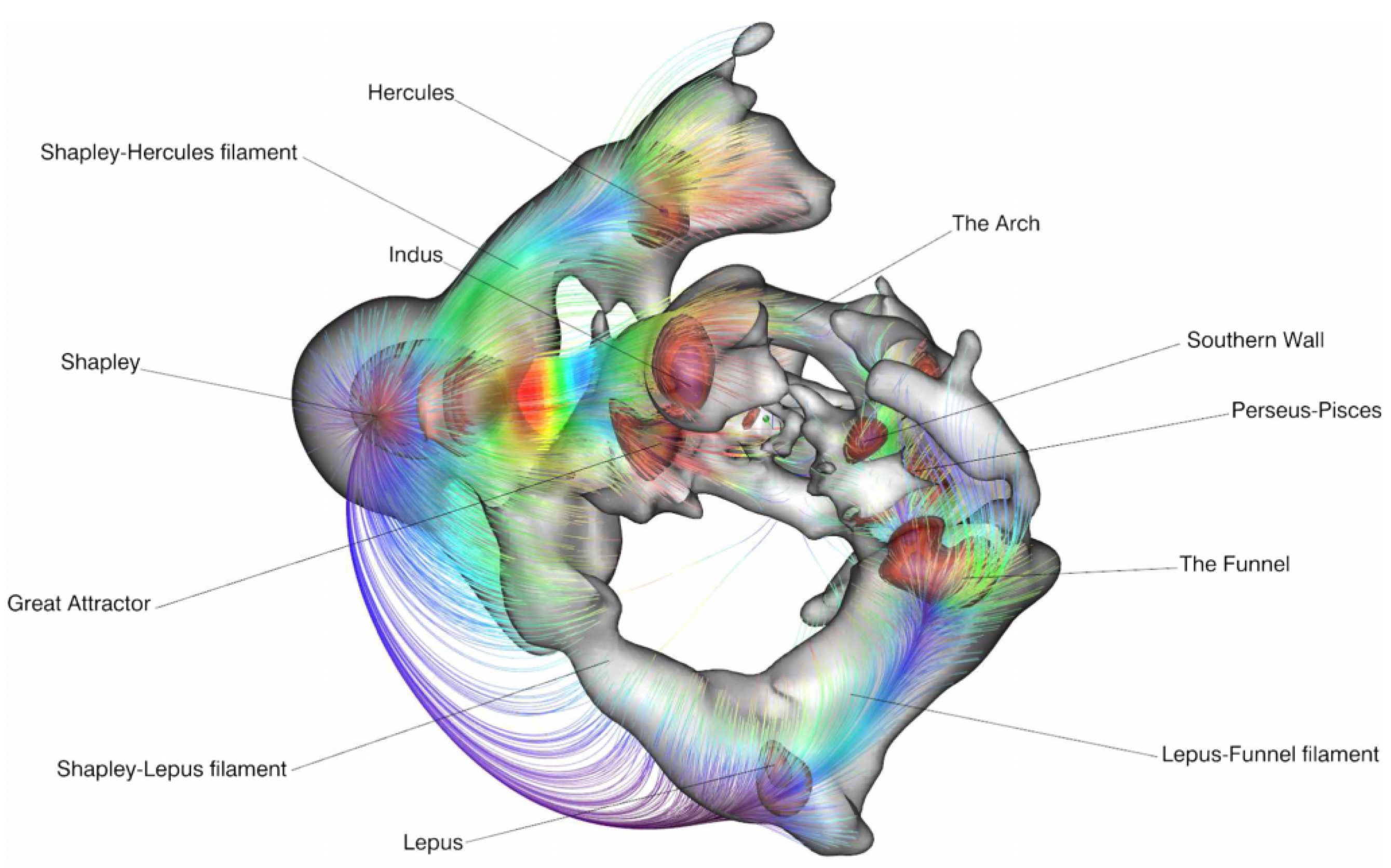
5.3. Physics
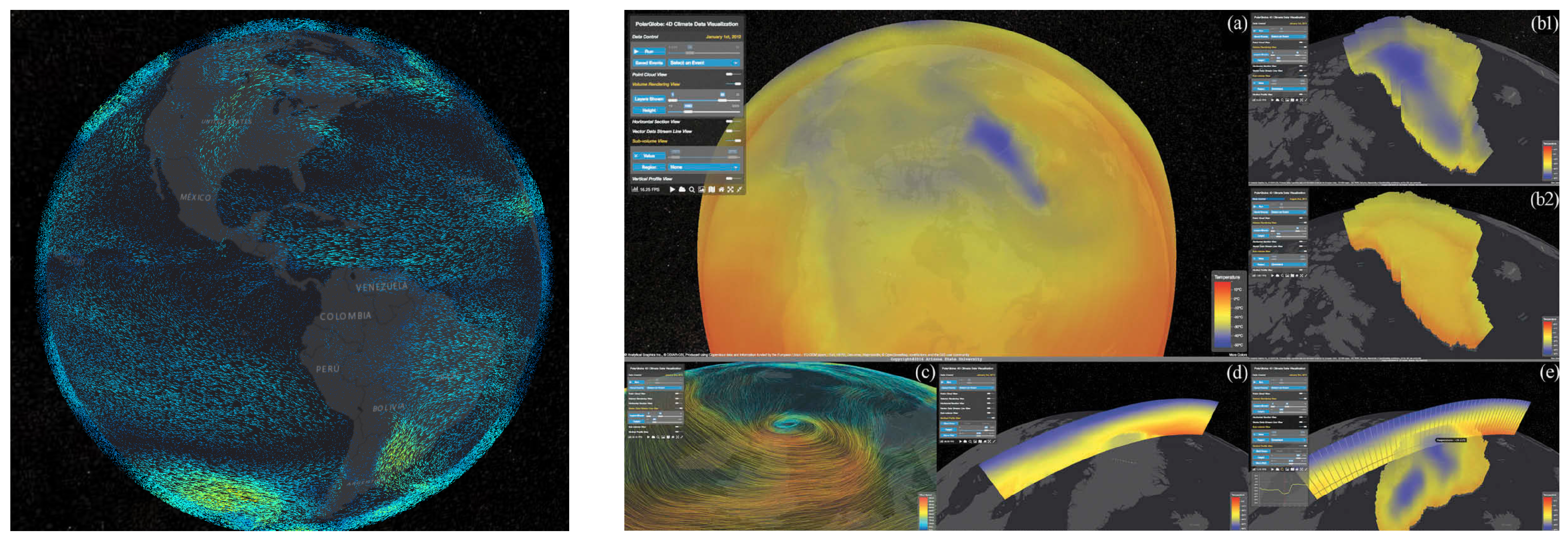
5.4. Geographical Visualizations
5.5. Cross Scientific Toolkits Supporting Scientific Visualization
6. Discussion and Directions for Future Work
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| HTML5 | Hyper Text Markup Language 5 |
| WebGL | Web Graphics Library |
| WebXR | Web Cross Reality (augmented and virtual reality) |
| WebRTC | Web Real-Time Protocol |
References
- Beyer, J.; Hadwiger, M.; Pfister, H. State-of-the-art in GPU-based large-scale volume visualization. In Computer Graphics Forum; The Eurographs Association & John Wiley & Sons, Ltd.: Chichester, UK, 2015; Volume 34, pp. 13–37. [Google Scholar]
- Jourdain, S.; Ayachit, U.; Geveci, B. Paraviewweb, a web framework for 3d visualization and data processing. In Proceedings of the IADIS International Conference on Web Virtual Reality And Three-Dimensional Worlds, Freiburg, Germany, 27–29 July 2010; Volume 7, p. 1. [Google Scholar]
- Pascucci, V.; Scorzelli, G.; Summa, B.; Bremer, P.T.; Gyulassy, A.; Christensen, C.; Philip, S.; Kumar, S. The ViSUS visualization framework. In High Performance Visualization: Enabling Extreme-Scale Scientific Insight; Chapman and Hall/CRC: Boca Raton, FL, USA, 2012; pp. 401–414. [Google Scholar]
- Sons, K.; Klein, F.; Rubinstein, D.; Byelozyorov, S.; Slusallek, P. XML3D: Interactive 3D graphics for the web. In Proceedings of the 15th International Conference on Web 3D Technology, Los Angeles, CA, USA, 24–25 July 2010; pp. 175–184. [Google Scholar]
- Congote, J.; Segura, A.; Kabongo, L.; Moreno, A.; Posada, J.; Ruiz, O. Interactive Visualization of Volumetric Data With Webgl in Real-Time. In Proceedings of the 16th International Conference on 3D Web Technology, Paris, France, 20–22 June 2011; pp. 137–146. [Google Scholar]
- Marrin, C. Webgl Specification; Khronos WebGL Working Group: Beaverton, OR, USA, 2011; Volume 3. [Google Scholar]
- Evans, A.; Romeo, M.; Bahrehmand, A.; Agenjo, J.; Blat, J. 3D graphics on the web: A survey. Comput. Graph. 2014, 41, 43–61. [Google Scholar]
- Potenziani, M.; Callieri, M.; Dellepiane, M.; Scopigno, R. Publishing and Consuming 3D Content on the Web: A Survey. Found. Trends Comput. Graph. Vis. 2018, 10, 244–333. [Google Scholar] [CrossRef]
- Friendly, M.; Denis, D.J. Milestones in the History of Thematic Cartography, Statistical Graphics, and Data Visualization. Citeseer 2001, 32, 13. [Google Scholar]
- Cabello, R. Three.js. 2010. Available online: Https://threejs.org/ (accessed on 8 June 2020).
- Haber, R.B.; McNabb, D.A. Visualization idioms: A conceptual model for scientific visualization systems. Vis. Sci. Comput. 1990, 74, 93. [Google Scholar]
- Greenberg, D.P.; Cohen, M.F.; Torrance, K.E. Radiosity: A method for computing global illumination. Vis. Comput. 1986, 2, 291–297. [Google Scholar]
- Whitted, T. An improved illumination model for shaded display. In ACM Siggraph 2005 Courses; Association for Computing Machinery: New York, NY, USA, 2005; p. 4-es. [Google Scholar]
- Appel, A. Some techniques for shading machine renderings of solids. In Proceedings of the Spring Joint Computer Conference, Association for Computing Machinery, New York, NY, USA, 30 April–2 May 1968; pp. 37–45. [Google Scholar]
- Drebin, R.A.; Carpenter, L.; Hanrahan, P. Volume rendering. ACM Siggraph Comput. Graph. 1988, 22, 65–74. [Google Scholar]
- Haehn, D.; Rannou, N.; Ahtam, B.; Grant, E.; Pienaar, R. Neuroimaging in the browser using the x toolkit. Front. Neuroinform. 2014, 101. [Google Scholar]
- Mwalongo, F.; Krone, M.; Becher, M.; Reina, G.; Ertl, T. GPU-based remote visualization of dynamic molecular data on the web. Graph. Model. 2016, 88, 57–65. [Google Scholar]
- Shi, S.; Hsu, C.H. A survey of interactive remote rendering systems. ACM Comput. Surv. CSUR 2015, 47, 1–29. [Google Scholar] [CrossRef]
- Discher, S.; Richter, R.; Döllner, J. Concepts and techniques for web-based visualization and processing of massive 3D point clouds with semantics. Graph. Model. 2019, 104, 101036. [Google Scholar] [CrossRef]
- Raji, M.; Hota, A.; Hobson, T.; Huang, J. Scientific visualization as a microservice. IEEE Trans. Vis. Comput. Graph. 2018, 26, 1760–1774. [Google Scholar] [CrossRef] [PubMed]
- Kitware. Vtk.js. 2017. Available online: https://kitware.github.io/vtk-js/ (accessed on 5 June 2020).
- Kitware. ParaView ArcticViewer, The Ultimate Data Viewer. 2019. Available online: https://kitware.github.io/arctic-viewer/ (accessed on 22 July 2020).
- Daly, L.; Brutzman, D. X3D: Extensible 3D graphics standard [standards in a nutshell]. IEEE Signal Process. Mag. 2007, 24, 130–135. [Google Scholar]
- Behr, J.; Eschler, P.; Jung, Y.; Zöllner, M. X3DOM: A DOM-based HTML5/X3D integration model. In Proceedings of the 14th International Conference on 3D Web Technology, Darmstadt, Germany, 16–17 June 2009; Association for Computing Machinery: New York, NY, USA, 2009; pp. 127–135. [Google Scholar]
- Tamm, G.; Slusallek, P. Web-enabled Server-based and Distributed Real-time Ray-Tracing. In EGPGV; Eurographics Association: Goslar, DEU, 2016; pp. 55–67. [Google Scholar]
- Vullo, R.P.; Catalfamo, M.A. Dynamically Generating Virtual Reality Scenes Using Molly and A-Frame. In Proceedings of the International Conference on Internet Computing and Internet of Things (ICOMP’17), Las Vegas, NV, USA, 17–20 July 2017; pp. 21–24. Available online: https://csce.ucmss.com/cr/books/2017/LFS/CSREA2017/ICM3266.pdf (accessed on 20 July 2020).
- Liu, Z.; Jiang, B.; Heer, J. imMens: Real-time visual querying of big data. In Computer Graphics Forum; Wiley Online Library: Hoboken, NJ, USA, 2013; Volume 32, pp. 421–430. [Google Scholar]
- Falk, M.; Grottel, S.; Krone, M.; Reina, G. Interactive gpu-based visualization of large dynamic particle data. Synth. Lect. Vis. 2016, 4, 1–121. [Google Scholar] [CrossRef]
- Lee, G.h.; Nam, J.h.; Han, H.s.; Kwon, S.c. A Study on 3D File Format for Web-based Scientific Visualization. Int. J. Adv. Cult. Technol. 2019, 7, 243–247. [Google Scholar]
- Mobeen, M.M.; Feng, L. Mobile visualization of biomedical volume datasets. J. Internet Technol. Secur. Trans. 2012, 1, 52–60. [Google Scholar] [CrossRef]
- Mani, G.; Li, W. 3D web based surgical training through comparative analysis. In Proceedings of the 18th International Conference on 3D Web Technology, Sebastian, Spain, 20–22 June 2013; pp. 83–86. [Google Scholar]
- Haehn, D. Slice: Drop: Collaborative medical imaging in the browser. In ACM SIGGRAPH 2013 Computer Animation Festival; Association for Computing Machinery: New York, NY, USA, 2013; p. 1. [Google Scholar]
- Virag, I.; Stoicu-Tivadar, L.; Amăricăi, E. Browser-based medical visualization system. In Proceedings of the 2014 IEEE 9th IEEE International Symposium on Applied Computational Intelligence and Informatics (SACI), Timisoara, Romania, 15–17 May 2014; IEEE: Piscataway, NJ, USA, 2014; pp. 355–359. [Google Scholar]
- Sherif, T.; Kassis, N.; Rousseau, M.É.; Adalat, R.; Evans, A.C. BrainBrowser: Distributed, web-based neurological data visualization. Front. Neuroinform. 2015, 8, 89. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Halle, M.; Demeusy, V.; Kikinis, R. The open anatomy browser: A collaborative web-based viewer for interoperable anatomy atlases. Front. Neuroinform. 2017, 11, 22. [Google Scholar] [CrossRef] [Green Version]
- Haehn, D.; Hoffer, J.; Matejek, B.; Suissa-Peleg, A.; Al-Awami, A.K.; Kamentsky, L.; Gonda, F.; Meng, E.; Zhang, W.; Schalek, R.; et al. Scalable interactive visualization for connectomics. In Informatics; Multidisciplinary Digital Publishing Institute: Basel, Switzerland, 2017; Volume 4, p. 29. [Google Scholar]
- Qiao, L.; Chen, X.; Zhang, Y.; Zhang, J.; Wu, Y.; Li, Y.; Mo, X.; Chen, W.; Xie, B.; Qiu, M. An html5-based pure website solution for rapidly viewing and processing large-scale 3d medical volume reconstruction on mobile internet. Int. J. Telemed. Appl. 2017, 2017, 4074137. [Google Scholar] [CrossRef]
- Bernal-Rusiel, J.L.; Rannou, N.; Gollub, R.L.; Pieper, S.; Murphy, S.; Robertson, R.; Grant, P.E.; Pienaar, R. Reusable client-side javascript modules for immersive web-based real-time collaborative neuroimage visualization. Front. Neuroinform. 2017, 11, 32. [Google Scholar] [CrossRef]
- Ledoux, L.P.; Morency, F.C.; Cousineau, M.; Houde, J.C.; Whittingstall, K.; Descoteaux, M. Fiberweb: Diffusion visualization and processing in the browser. Front. Neuroinform. 2017, 11, 54. [Google Scholar] [CrossRef] [Green Version]
- Keiriz, J.J.; Zhan, L.; Ajilore, O.; Leow, A.D.; Forbes, A.G. NeuroCave: A web-based immersive visualization platform for exploring connectome datasets. Netw. Neurosci. 2018, 2, 344–361. [Google Scholar] [CrossRef] [PubMed]
- Min, Q.; Liu, N.; Chen, Y. A Web-based Medical Image Viewer for 2D and 3D visualization. In Proceedings of the 2018 2nd International Conference on Management Engineering, Software Engineering and Service Sciences, Wuhan, China, 13–15 January 2018; 2018; pp. 261–264. [Google Scholar]
- Lavrič, P.; Bohak, C.; Marolt, M. Collaborative view-aligned annotations in web-based 3D medical data visualization. In Proceedings of the 2017 40th International Convention on Information and Communication Technology, Electronics and Microelectronics (MIPRO), Opatija, Croatia, 22–26 May 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 259–263. [Google Scholar]
- Kokelj, Ž.; Bohak, C.; Marolt, M. A web-based virtual reality environment for medical visualization. In Proceedings of the 2018 41st International Convention on Information and Communication Technology, Electronics and Microelectronics (MIPRO), Opatija, Croatia, 21–25 May 2018; IEEE: Piscataway, NJ, USA, 2018; pp. 299–302. [Google Scholar]
- Ganglberger, F.; Swoboda, N.; Frauenstein, L.; Kaczanowska, J.; Haubensak, W.; Bühler, K. BrainTrawler: A visual analytics framework for iterative exploration of heterogeneous big brain data. Comput. Graph. 2019, 82, 304–320. [Google Scholar] [CrossRef]
- Mullie, L.; Afilalo, J. CoreSlicer: A web toolkit for analytic morphomics. BMC Med. Imaging 2019, 19, 15. [Google Scholar] [CrossRef] [Green Version]
- Moraes, T.; Amorim, P.; Silva, J.; Pedrini, H. Web-Based Interactive Visualization of Medical Images in a Distributed System. In Proceedings of the 14th International Conference on Computer Graphics Theory and Applications, Prague, Czech Republic, 25–27 February 2019; pp. 346–353. [Google Scholar]
- Zhang, Q. Web-based medical data visualization and information sharing towards application in distributed diagnosis. Inform. Med. Unlocked 2019, 14, 69–81. [Google Scholar] [CrossRef]
- Zhang, Q. Medical data visual synchronization and information interaction using Internet-based graphics rendering and message-oriented streaming. Inform. Med. Unlocked 2019, 17, 100253. [Google Scholar] [CrossRef]
- Franke, L.; Weidele, D.K.I.; Zhang, F.; Cetin-Karayumak, S.; Pieper, S.; O’Donnell, L.J.; Rathi, Y.; Haehn, D. FiberStars: Visual Comparison of Diffusion Tractography Data between Multiple Subjects. arXiv 2020, arXiv:2005.08090. [Google Scholar]
- PV. PV—WebGL Protein Viewer. 2012. Available online: http://github.com/biasmv/pv (accessed on 17 July 2020).
- Hanson, R.M.; Prilusky, J.; Renjian, Z.; Nakane, T.; Sussman, J.L. JSmol and the next-generation web-based representation of 3D molecular structure as applied to proteopedia. Isr. J. Chem. 2013, 53, 207–216. [Google Scholar] [CrossRef]
- Pettit, J.B.; Marioni, J.C. bioWeb3D: An online webGL 3D data visualisation tool. BMC Bioinform. 2013, 14, 185. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Leung, K.S.; Nakane, T.; Wong, M.H. Iview: An interactive WebGL visualizer for protein-ligand complex. BMC Bioinform. 2014, 15, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Mohebifar, M.; Sajadi, F. Chemozart: A web-based 3D molecular structure editor and visualizer platform. J. Chemin. 2015, 7, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Burger, M.C. ChemDoodle Web Components: HTML5 toolkit for chemical graphics, interfaces, and informatics. J. Chemin. 2015, 7, 1–7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rego, N.; Koes, D. 3Dmol. js: Molecular visualization with WebGL. Bioinformatics 2015, 31, 1322–1324. [Google Scholar] [CrossRef] [Green Version]
- Grebner, C.; Norrby, M.; Enström, J.; Nilsson, I.; Hogner, A.; Henriksson, J.; Westin, J.; Faramarzi, F.; Werner, P.; Boström, J. 3D-Lab: A collaborative web-based platform for molecular modeling. Future Med. Chem. 2016, 8, 1739–1752. [Google Scholar] [CrossRef] [PubMed]
- Norrby, M.; Grebner, C.; Eriksson, J.; Bostrom, J. Molecular rift: Virtual reality for drug designers. J. Chem. Inf. Model. 2015, 55, 2475–2484. [Google Scholar] [CrossRef]
- Skjærven, L.; Jariwala, S.; Yao, X.Q.; Grant, B.J. Online interactive analysis of protein structure ensembles with Bio3D-web. Bioinformatics 2016, 32, 3510–3512. [Google Scholar] [CrossRef] [Green Version]
- Bekker, G.J.; Nakamura, H.; Kinjo, A.R. Molmil: A molecular viewer for the PDB and beyond. J. Chemin. 2016, 8, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Kovanci, G.; Ghaffar, M.; Sommer, B. Web-based hybrid-dimensional visualization and exploration of cytological localization scenarios. J. Integr. Bioinform. 2016, 13, 47–58. [Google Scholar] [CrossRef]
- Djekidel, M.N.; Wang, M.; Zhang, M.Q.; Gao, J. HiC-3DViewer: A new tool to visualize Hi-C data in 3D space. Quant. Biol. 2017, 5, 183–190. [Google Scholar] [CrossRef] [Green Version]
- Sehnal, D.; Deshpande, M.; Vařeková, R.S.; Mir, S.; Berka, K.; Midlik, A.; Pravda, L.; Velankar, S.; Koča, J. LiteMol suite: Interactive web-based visualization of large-scale macromolecular structure data. Nat. Methods 2017, 14, 1121–1122. [Google Scholar] [CrossRef]
- Shi, M.; Gao, J.; Zhang, M.Q. Web3DMol: Interactive protein structure visualization based on WebGL. Nucleic Acids Res. 2017, 45, W523–W527. [Google Scholar] [CrossRef] [Green Version]
- Zhou, G.; Xia, J. OmicsNet: A web-based tool for creation and visual analysis of biological networks in 3D space. Nucleic Acids Res. 2018, 46, W514–W522. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rose, A.S.; Bradley, A.R.; Valasatava, Y.; Duarte, J.M.; Prlić, A.; Rose, P.W. NGL viewer: Web-based molecular graphics for large complexes. Bioinformatics 2018, 34, 3755–3758. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rose, A.S.; Hildebrand, P.W. NGL Viewer: A web application for molecular visualization. Nucleic Acids Res. 2015, 43, W576–W579. [Google Scholar] [CrossRef] [PubMed]
- Carrillo-Tripp, M.; Alvarez-Rivera, L.; Lara-Ramírez, O.I.; Becerra-Toledo, F.J.; Vega-Ramírez, A.; Quijas-Valades, E.; González-Zavala, E.; González-Vázquez, J.C.; García-Vieyra, J.; Santoyo-Rivera, N.B.; et al. HTMoL: Full-stack solution for remote access, visualization, and analysis of molecular dynamics trajectory data. J. Comput. Aided Mol. Des. 2018, 32, 869–876. [Google Scholar] [CrossRef] [Green Version]
- Gralka, P.; Becher, M.; Braun, M.; Frieß, F.; Müller, C.; Rau, T.; Schatz, K.; Schulz, C.; Krone, M.; Reina, G.; et al. MegaMol–a comprehensive prototyping framework for visualizations. Eur. Phys. J. Spec. Top. 2019, 227, 1817–1829. [Google Scholar] [CrossRef]
- Wang, J.; Youkharibache, P.; Zhang, D.; Lanczycki, C.J.; Geer, R.C.; Madej, T.; Phan, L.; Ward, M.; Lu, S.; Marchler, G.H.; et al. iCn3D, a web-based 3D viewer for sharing 1D/2D/3D representations of biomolecular structures. Bioinformatics 2020, 36, 131–135. [Google Scholar] [CrossRef] [Green Version]
- Cassidy, K.C.; Šefčík, J.; Raghav, Y.; Chang, A.; Durrant, J.D. ProteinVR: Web-based molecular visualization in virtual reality. PLoS Comput. Biol. 2020, 16, e1007747. [Google Scholar] [CrossRef]
- Cignoni, P.; Callieri, M.; Corsini, M.; Dellepiane, M.; Ganovelli, F.; Ranzuglia, G. MeshLab: An Open-Source Mesh Processing Tool. In Eurographics Italian Chapter Conference; Scarano, V., Chiara, R.D., Erra, U., Eds.; The Eurographics Association: Allaville, Switzerland, 2008. [Google Scholar] [CrossRef]
- Chandler, J.; Obermaier, H.; Joy, K.I. WebGL-Enabled Remote Visualization of Smoothed Particle Hydrodynamics Simulations; EuroVis: Geneve, Switzerland, 2015. [Google Scholar]
- McCauley, T. A browser-based event display for the CMS Experiment at the LHC using WebGL. J. Phys. Conf. Ser. 2017, 898, 072030. [Google Scholar] [CrossRef]
- Yeh, A. Programming driven 3D modeling on the web. In Proceedings of the 22nd International Conference on 3D Web Technology, Brisbane, Australia, 5–8 June 2017; pp. 1–9. [Google Scholar]
- Diblen, F.; Attema, J.; Bakhshi, R.; Caron, S.; Hendriks, L.; Stienen, B. SPOT: Open Source framework for scientific data repository and interactive visualization. SoftwareX 2019, 9, 328–331. [Google Scholar] [CrossRef]
- Lupinetti, K.; Cabiddu, D.; Giannini, F.; Monti, M. CAD3A: A web-based application to visualize and semantically enhance CAD assembly models. In Proceedings of the 2019 15th International Conference on Signal-Image Technology & Internet-Based Systems (SITIS), Sorrento, Italy, 26–29 November 2019; IEEE: Piscataway, NJ, USA, 2019; pp. 462–469. [Google Scholar]
- Bracci, M.; Tarini, M.; Pietroni, N.; Livesu, M.; Cignoni, P. HexaLab. net: An online viewer for hexahedral meshes. Comput. Aided Des. 2019, 110, 24–36. [Google Scholar] [CrossRef] [Green Version]
- Kaboudian, A.; Cherry, E.M.; Fenton, F.H. Large-scale interactive numerical experiments of chaos, solitons and fractals in real time via GPU in a web browser. Chaos Solitons Fractals 2019, 121, 6–29. [Google Scholar]
- Figueiras, E.; Olivieri, D.N.; Paredes, A.; Michinel, H. QMwebJS—An Open Source Software Tool to Visualize and Share Time-Evolving Three-Dimensional Wavefunctions. Mathematics 2020, 8, 430. [Google Scholar]
- Potenziani, M.; Callieri, M.; Dellepiane, M.; Corsini, M.; Ponchio, F.; Scopigno, R. 3DHOP: 3D heritage online presenter. Comput. Graph. 2015, 52, 129–141. [Google Scholar]
- Müller, R.D.; Qin, X.; Sandwell, D.T.; Dutkiewicz, A.; Williams, S.E.; Flament, N.; Maus, S.; Seton, M. The GPlates portal: Cloud-based interactive 3D visualization of global geophysical and geological data in a web browser. PLoS ONE 2016, 11, e0150883. [Google Scholar]
- Koeva, M.; Luleva, M.; Maldjanski, P. Integrating spherical panoramas and maps for visualization of cultural heritage objects using virtual reality technology. Sensors 2017, 17, 829. [Google Scholar]
- Koeva, M. 3D modelling and interactive web-based visualization of cultural heritage objects. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2016, 41, 297–303. [Google Scholar]
- Li, W.; Wang, S. PolarGlobe: A web-wide virtual globe system for visualizing multidimensional, time-varying, big climate data. Int. J. Geogr. Inf. Sci. 2017, 31, 1562–1582. [Google Scholar]
- Evangelidis, K.; Papadopoulos, T.; Papatheodorou, K.; Mastorokostas, P.; Hilas, C. 3D geospatial visualizations: Animation and motion effects on spatial objects. Comput. Geosci. 2018, 111, 200–212. [Google Scholar]
- Discher, S.; Richter, R.; Döllner, J. A scalable webGL-based approach for visualizing massive 3D point clouds using semantics-dependent rendering techniques. In Proceedings of the 23rd International ACM Conference on 3D Web Technology, Poznań, Poland, 20–22 June 2018; pp. 1–9. [Google Scholar]
- Liu, D.; Peng, J.; Wang, Y.; Huang, M.; He, Q.; Yan, Y.; Ma, B.; Yue, C.; Xie, Y. Implementation of interactive three-dimensional visualization of air pollutants using WebGL. Environ. Model. Softw. 2019, 114, 188–194. [Google Scholar]
- Liu, L.; Silver, D.; Bemis, K. Visualizing three-dimensional ocean eddies in web browsers. IEEE Access 2019, 7, 44734–44747. [Google Scholar]
- Boutsi, A.M.; Ioannidis, C.; Soile, S. An Integrated Approach to 3D Web Visualization of Cultural Heritage Heterogeneous Datasets. Remote Sens. 2019, 11, 2508. [Google Scholar] [CrossRef] [Green Version]
- Desprat, C.; Jessel, J.P.; Luga, H. A 3D collaborative editor using WebGL and WebRTC. In Proceedings of the 20th International Conference on 3D Web Technology, Heraklion, Greece, 18–21 June 2015; pp. 157–158. [Google Scholar]
- Hadjar, H.; Meziane, A.; Gherbi, R.; Setitra, I.; Aouaa, N. WebVR based interactive visualization of open health data. In Proceedings of the 2nd International Conference on Web Studies, Samara, Russia, 26–27 November 2018; pp. 56–63. [Google Scholar]
- Yang, W.; Tao, Y.; Lin, H. Voxer—A platform for creating, customizing, and sharing scientific visualizations. J. Vis. 2019, 22, 1161–1176. [Google Scholar] [CrossRef]
- Matelsky, J.K.; Downs, J.; Cowley, H.P.; Wester, B.; Gray-Roncal, W. A substrate for modular, extensible data-visualization. Big Data Anal. 2020, 5, 1. [Google Scholar] [CrossRef] [Green Version]
- Qualter, J.; Sculli, F.; Oliker, A.; Napier, Z.; Lee, S.; Garcia, J.; Frenkel, S.; Harnik, V.; Triola, M. The biodigital human: A web-based 3D platform for medical visualization and education. Stud. Health Technol. Inform. 2012, 173, 359–361. [Google Scholar]
- Violante, M.G.; Vezzetti, E. Design and implementation of 3D Web-based interactive medical devices for educational purposes. Int. J. Interact. Des. Manuf. IJIDeM 2017, 11, 31–44. [Google Scholar] [CrossRef] [Green Version]
- Smit, N.; Hofstede, C.W.; Kraima, A.; Jansma, D.; deRuiter, M.; Eisemann, E.; Vilanova, A. The online anatomical human: Web-based anatomy education. Proc. Eurogr. Educ. Pap. 2016, 2016, 37–40. [Google Scholar]
- Petersson, H.; Sinkvist, D.; Wang, C.; Smedby, Ö. Web-based interactive 3D visualization as a tool for improved anatomy learning. Anat. Sci. Educ. 2009, 2, 61–68. [Google Scholar]
- Birr, S.; Mönch, J.; Sommerfeld, D.; Preim, U.; Preim, B. The LiverAnatomyExplorer: A WebGL-based surgical teaching tool. IEEE Comput. Graph. Appl. 2013, 33, 48–58. [Google Scholar]
- Preim, B.; Saalfeld, P. A survey of virtual human anatomy education systems. Comput. Graph. 2018, 71, 132–153. [Google Scholar]
- John, N.W. The impact of Web3D technologies on medical education and training. Comput. Educ. 2007, 49, 19–31. [Google Scholar] [CrossRef]
- Jacinto, H.; Kéchichian, R.; Desvignes, M.; Prost, R.; Valette, S. A web interface for 3D visualization and interactive segmentation of medical images. In Proceedings of the 17th International Conference on 3D Web Technology, Los Angeles, CA, USA, 4–5 August 2012; pp. 51–58. [Google Scholar]
- Jiménez, J.; López, A.; Cruz, J.; Esteban, F.J.; Navas, J.; Villoslada, P.; de Miras, J.R. A Web platform for the interactive visualization and analysis of the 3D fractal dimension of MRI data. J. Biomed. Inform. 2014, 51, 176–190. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marion, C.; Jomier, J. Real-time collaborative scientific WebGL visualization with WebSocket. In Proceedings of the 17th International Conference on 3D Web Technology, Los Angeles, CA, USA, 4–5 August 2012; pp. 47–50. [Google Scholar]
- Noguera, J.M.; Jiménez, J.R. Visualization of Very Large 3D Volumes on Mobile Devices and WebGL. Available online: http://wscg.zcu.cz/WSCG2012/short/B71-full.pdf (accessed on 8 August 2020).
- Arbelaiz, A.; Moreno, A.; Kabongo, L.; García-Alonso, A. Volume visualization tools for medical applications in ubiquitous platforms. In eHealth 360; Springer: Berlin/Heidelberg, Germany, 2017; pp. 443–450. [Google Scholar]
- Hou, X.; Sun, J.; Zhang, J. A web-based solution for 3D medical image visualization. In Medical Imaging 2015: PACS and Imaging Informatics: Next Generation and Innovations; SPIE: Orlando, FL, USA, 2015; Volume 9418, p. 941810. [Google Scholar]
- Lorensen, W.E.; Cline, H.E. Marching cubes: A high resolution 3D surface construction algorithm. ACM Siggraph Comput. Graph. 1987, 21, 163–169. [Google Scholar] [CrossRef]
- Ashwini, A.; Kwon, J. Image processing pipeline for web-based real-time 3d visualization of teravoxel volumes. In Proceedings of the International Conference on Data Mining and Big Data, Shanghai, China, June 17–22, 2018; Springer: Berlin/Heidelberg, Germany, 2018; pp. 203–212. [Google Scholar]
- Haehn, D.; Franke, L.; Zhang, F.; Karayumak, S.C.; Pieper, S.; O’Donnell, L.; Rathi, Y. TRAKO: Efficient Transmission of Tractography Data for Visualization. arXiv 2020, arXiv:2004.13630. [Google Scholar]
- Mwalongo, F.; Krone, M.; Reina, G.; Ertl, T. State-of-the-Art Report in Web-based Visualization. In Computer Graphics Forum, Wiley Online Library; Eurographics Association: Geneve, Switzerland, 2016; Volume 35, pp. 553–575. [Google Scholar]
- Callieri, M.; Andrei, R.M.; Di Benedetto, M.; Zoppè, M.; Scopigno, R. Visualization methods for molecular studies on the web platform. In Proceedings of the 15th International Conference on Web 3D Technology, Los Angeles, CA, USA, 24–25 July 2010; pp. 117–126. [Google Scholar]
- Rose, A.S.; Bradley, A.R.; Valasatava, Y.; Duarte, J.M.; Prlić, A.; Rose, P.W. Web-based molecular graphics for large complexes. In Proceedings of the 21st International Conference on Web3D Technology, Anaheim, CA, USA, 22–24 July 2016; pp. 185–186. [Google Scholar]
- Jiang, C.; Jin, X.; Dong, Y.; Chen, M. Kekule. js: An open source javascript chemoinformatics toolkit. J. Chem. Inf. Model. 2016, 56, 1132–1138. [Google Scholar] [CrossRef] [PubMed]
- Tiemann, J.K.; Guixa-Gonzalez, R.; Hildebrand, P.W.; Rose, A.S. MDsrv: Viewing and sharing molecular dynamics simulations on the web. Nat. Methods 2017, 14, 1123–1124. [Google Scholar] [CrossRef]
- Hildebrand, P.W.; Rose, A.S.; Tiemann, J.K. Bringing molecular dynamics simulation data into view. Trends Biochem. Sci. 2019, 44, 902–913. [Google Scholar] [CrossRef] [Green Version]
- Abriata, L.A. Web apps come of age for molecular sciences. In Informatics; Multidisciplinary Digital Publishing Institute: Basel, Switzerland, 2017; Volume 4, p. 28. [Google Scholar]
- Martinez, X.; Krone, M.; Alharbi, N.; Rose, A.S.; Laramee, R.S.; O’Donoghue, S.; Baaden, M.; Chavent, M. Molecular graphics: Bridging structural biologists and computer scientists. Structure 2019, 27, 1617–1623. [Google Scholar] [CrossRef]
- Yuan, S.; Chan, H.S.; Hu, Z. Implementing WebGL and HTML5 in macromolecular visualization and modern computer-aided drug design. Trends Biotechnol. 2017, 35, 559–571. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The protein data bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [Green Version]
- Burley, S.K.; Berman, H.M.; Christie, C.; Duarte, J.M.; Feng, Z.; Westbrook, J.; Young, J.; Zardecki, C. RCSB Protein Data Bank: Sustaining a living digital data resource that enables breakthroughs in scientific research and biomedical education. Protein Sci. 2018, 27, 316–330. [Google Scholar] [CrossRef] [Green Version]
- Berman, H.M.; Olson, W.K.; Beveridge, D.L.; Westbrook, J.; Gelbin, A.; Demeny, T.; Hsieh, S.H.; Srinivasan, A.; Schneider, B. The nucleic acid database. A comprehensive relational database of three-dimensional structures of nucleic acids. Biophys. J. 1992, 63, 751. [Google Scholar] [CrossRef] [Green Version]
- Li, S.; Olson, W.K.; Lu, X.J. Web 3DNA 2.0 for the analysis, visualization, and modeling of 3D nucleic acid structures. Nucleic Acids Res. 2019, 47, W26–W34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sedova, M.; Iyer, M.; Li, Z.; Jaroszewski, L.; Post, K.W.; Hrabe, T.; Porta-Pardo, E.; Godzik, A. Cancer3D 2.0: Interactive analysis of 3D patterns of cancer mutations in cancer subsets. Nucleic Acids Res. 2019, 47, D895–D899. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sedova, M.; Jaroszewski, L.; Godzik, A. Protael: Protein data visualization library for the web. Bioinformatics 2016, 32, 602–604. [Google Scholar] [CrossRef] [Green Version]
- Figueiras, E.; Olivieri, D.; Paredes, A.; Michinel, H. QMBlender: Particle-based visualization of 3D quantum wave function dynamics. J. Comput. Sci. 2019, 35, 44–56. [Google Scholar] [CrossRef]
- Community, B.O. Blender—A 3D Modelling and Rendering Package; Blender Foundation; Stichting Blender Foundation: Amsterdam, The Netherlands, 2018. [Google Scholar]
- Kent, B.R. 3D Scientific Visualization With Blender; Morgan & Claypool Publishers: San Rafael, CA, USA, 2015. [Google Scholar]
- Ghaffar, M.; Biere, N.; Jäger, D.; Klein, K.; Schreiber, F.; Kruse, O.; Sommer, B. 3D modelling and visualisation of heterogeneous cell membranes in Blender. In Proceedings of the 11th International Symposium on Visual Information Communication and Interaction, Växjö, Sweden, 13–15 August 2018; pp. 64–71. [Google Scholar]
- Ihmsen, M.; Akinci, N.; Akinci, G.; Teschner, M. Unified spray, foam and air bubbles for particle-based fluids. Vis. Comput. 2012, 28, 669–677. [Google Scholar] [CrossRef]
- Naiman, J. AstroBlend: An astrophysical visualization package for Blender. Astron. Comput. 2016, 15, 50–60. [Google Scholar] [CrossRef] [Green Version]
- Rajendiran, N.; Durrant, J.D. Pyrite: A blender plugin for visualizing molecular dynamics simulations using industry-standard rendering techniques. J. Comput. Chem. 2018, 39, 748–755. [Google Scholar] [CrossRef]
- Rene, K.M.; Jeff Gay, M.M. OpenJSCAD. 2019. Available online: https://openjscad.org/ (accessed on 3 July 2020).
- Dykes, T.; Hassan, A.; Gheller, C.; Croton, D.; Krokos, M. Interactive 3D visualization for theoretical virtual observatories. Mon. Not. R. Astron. Soc. 2018, 477, 1495–1507. [Google Scholar] [CrossRef]
- Dolag, K.; Reinecke, M.; Gheller, C.; Imboden, S. Splotch: Visualizing cosmological simulations. New J. Phys. 2008, 10, 125006. [Google Scholar] [CrossRef]
- Bertin, E.; Pillay, R.; Marmo, C. Web-based visualization of very large scientific astronomy imagery. Astron. Comput. 2015, 10, 43–53. [Google Scholar]
- Rosenfield, P.; Fay, J.; Gilchrist, R.K.; Cui, C.; Weigel, A.D.; Robitaille, T.; Otor, O.J.; Goodman, A. AAS WorldWide telescope: A seamless, cross-platform data visualization engine for astronomy research, education, and democratizing data. Astrophys. J. Suppl. Ser. 2018, 236, 22. [Google Scholar]
- Pomarède, D.; Courtois, H.M.; Hoffman, Y.; Tully, R.B. Cosmography and data visualization. Publ. Astron. Soc. Pac. 2017, 129, 058002. [Google Scholar]
- Pomarede, D.; Hoffman, Y.; Courtois, H.M.; Tully, R.B. The Cosmic V-Web. Astrophys. J. 2017, 845, 55. [Google Scholar]
- Feng, L.; Wang, C.; Li, C.; Li, Z. A research for 3D WebGIS based on WebGL. In Proceedings of 2011 International Conference on Computer Science and Network Technology, Harbin, China, 24–26 December 2011; IEEE: Piscataway, NJ, USA, 2011; Volume 1, pp. 348–351. [Google Scholar]
- Loesch, B.; Christen, M.; Nebiker, S. OpenWebGlobe–an open source SDK for creating large-scale virtual globes on a WebGL basis. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2012, 39, B4. [Google Scholar] [CrossRef] [Green Version]
- Graphics, A. Cesium–WebGL Virtual Globe and Map Engine. 2013. Available online: https://cesium.com/cesiumjs/ (accessed on 17 June 2020).
- Krämer, M.; Gutbell, R. A case study on 3D geospatial applications in the web using state-of-the-art WebGL frameworks. In Proceedings of the 20th International Conference on 3D Web Technology, Heraklion, Greece, 18–21 June 2015; pp. 189–197. [Google Scholar]
- Miao, R.; Song, J.; Zhu, Y. 3D geographic scenes visualization based on WebGL. In Proceedings of the 2017 6th International Conference on Agro-Geoinformatics, Fairfax, VA, USA, 7–10 August 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 1–6. [Google Scholar]
- Resch, B.; Wohlfahrt, R.; Wosniok, C. Web-based 4D visualization of marine geo-data using WebGL. Cartogr. Geogr. Inf. Sci. 2014, 41, 235–247. [Google Scholar]
- Galeazzi, F.; Callieri, M.; Dellepiane, M.; Charno, M.; Richards, J.; Scopigno, R. Web-based visualization for 3D data in archaeology: The ADS 3D viewer. J. Archaeol. Sci. Rep. 2016, 9, 1–11. [Google Scholar]
- Raji, M.; Hota, A.; Huang, J. Scalable web-embedded volume rendering. In Proceedings of the 2017 IEEE 7th Symposium on Large Data Analysis and Visualization (LDAV), Phoenix, AZ, USA, 2 October 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 45–54. [Google Scholar]
- Wald, I.; Johnson, G.P.; Amstutz, J.; Brownlee, C.; Knoll, A.; Jeffers, J.; Günther, J.; Navrátil, P. Ospray-a cpu ray tracing framework for scientific visualization. IEEE Trans. Vis. Comput. Graph. 2016, 23, 931–940. [Google Scholar]
- Lavoué, G.; Chevalier, L.; Dupont, F. Streaming compressed 3D data on the web using JavaScript and WebGL. In Proceedings of the 18th international Conference on 3D Web Technology, San Sebastian, Spain, 20–22 June 2013; pp. 19–27. [Google Scholar]
- Tamm, G.; Slusallek, P. Plugin free remote visualization in the browser. In Visualization and Data Analysis 2015; SPIE/IS&T Electronic Imaging: San Francisco, CA, USA, 2015; Volume 9397, p. 939705. [Google Scholar]








| Name, Authors, Reference & URL 1 | Codebase | Infra-Structure | Responsive Design | VR/ AR 2 | License | SciVis FRS 3 | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| Code Available | Updates | Documentation | Support | WebGL | ||||||
| Medicine | ||||||||||
| Movania and Feng [30] | ✗ | ✗ | ✗ | ✗ | 1.0 | client | ✓ | ✗ | n/a | 0.3125 |
| Mani and Li [31] | ✗ | ✗ | ✗ | ✗ | 1.0 | cloud-based | ✗ | ✗ | n/a | 0.1875 |
| SliceDrop [32] https://slicedrop.com/ | ✓ | ✓ | ✓ | ✓ | 1.0 | client | ✓ | ✗ | MIT | 0.8125 |
| XTK [16] https://github.com/xtk/X | ✓ | ✓ | ✓ | ✓ | 1.0 | client | ✓ | ✗ | MIT | 0.8125 |
| Virag et al. [33] | ✗ | ✗ | ✗ | ✗ | 1.0 | client-server | ✓ | ✓ | n/a | 0.375 |
| BrainBrowser [34] https://brainbrowser.cbrain.mcgill.ca/ | ✓ | ✓ | ✓ | ✓ | 1.0 | client-server | ✓ | ✗ | GPL | 0.75 |
| OpenAnatomyBrowser [35] https://github.com/mhalle/oabrowser/tree/master | ✓ | ✗ | ✓ | ✗ | 1.0 | client-server | ✓ | ✗ | n/a | 0.5 |
| BUTTERFLY [36] https://github.com/Rhoana/butterfly | ✓ | ✓ | ✓ | ✓ | 1.0 | client-server | ✓ | ✗ | MIT | 0.75 |
| Qiao et al. [37] | ✗ | ✗ | ✗ | ✗ | 1.0 | client-server | ✓ | ✓/✗ | n/a | 0.3125 |
| MedView [38] https://github.com/FNNDSC/medview | ✓ | ✓ | ✓ | ✓ | 1.0 | client-server | ✗ | ✗ | MIT | 0.625 |
| FiberWeb [39] www.imeka.ca/fiberweb | ✗ | ✗ | ✗ | ✗ | 1.0 | client | ✓ | ✗ | closed source | 0.3125 |
| NeuroCave [40] https://creativecodinglab.github.io/NeuroCave/ | ✓ | ✗ | ✓ | ✓ | 2.0 | client(-server) | ✓ | ✓ | n/a | 0.8125 |
| Min et al. [41] | ✗ | ✗ | ✗ | ✗ | ✗ | client-server | ✗ | ✗ | n/a | 0.07 |
| Med3D [42] https://github.com/UL-FRI-LGM/Med3D | ✓ | ✗ | ✓ | ✓ | 2.0 | client-server | ✓ | ✗ | BSD | 0.6875 |
| Kokelj et al. [43] | ✗ | ✗ | ✗ | ✗ | 2.0 | client-server | ✓ | ✓ | n/a | 0.4375 |
| BrainTrawler [44] | upon request | ✗ | ✗ | ✗ | ✗ | client-server | ✓ | ✗ | n/a | 0.2857 |
| CoreSlicer [45] https://github.com/louismullie/coreslicer | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✗ | ✗ | MIT | 0.6875 |
| Moraes et al. [46] (integrated in https://github.com/tfmoraes/invesalius3) | integrated in other platform | ✗ | ✗ | ✗ | 2.0 | client-server | ✓ | ✗ | GPL | 0.4375 |
| Zhang [47,48] | ✗ | ✓ | ✗ | ✗ | 2.0 | client-server | ✓ | ✗ | n/a | 0.4375 |
| FiberStars [49] https://lorifranke.github.io/FiberStars/ | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✓ | ✗ | MIT | 0.8125 |
| Biology, Chemistry & Molecular Science | ||||||||||
| PV [50] https://biasmv.github.io/pv | ✓ | ✗ | ✓ | ✓ | 2.0 | client-server | ✗ | ✗ | MIT | 0.5625 |
| JSmol [51] https://sourceforge.net/projects/jsmol | ✓ | ✓ | ✓ | ✓ | 1.0/✗ | client-server | ✓ | ✗ | LGPL | 0.75 |
| bioWeb3D [52] https://github.com/jbogp/bioWeb3D | ✓ | ✓ | ✓ | ✓ | 1.0 | client-server | ✓ | ✗ | AFL | 0.75 |
| iView [53] https://github.com/HongjianLi/iview | ✓ | ✗ | ✗ | ✗ | 1.0 | client | ✓ | ✗ | Apache/MIT | 0.4375 |
| Chemozart [54] https://github.com/mohebifar/chemozart | ✓ | ✗ | ✗ | ✗ | 1.0 | client-server | ✓ | ✗ | Apache | 0.375 |
| ChemDoodle [55] https://web.chemdoodle.com | ✓ | ✓ | ✓ | ✓ | 1.0 | client | ✓ | ✗ | GPL | 0.8125 |
| 3Dmol.js [56] https://github.com/3dmol/3Dmol.js/http://3Dmol.csb.pitt.edu | ✓ | ✓ | ✓ | ✓ | 1.0 | client | ✓ | ✗ | BSD | 0.8125 |
| 3D-Lab / Molecular Rift [57,58] https://github.com/Magnusnorrby/MolecularRift | ✓ | ✓ | ✓ | ✓ | ✗ | client-server | ✓ | ✓ | GPL | 0.9286 |
| Bio3D-web [59] http://thegrantlab.org/bio3d/Webapps | ✓ | ✓ | ✓ | ✓ | ✗ | client-server | ✗ | ✗ | GPL2 | 0.6429 |
| Mwalongo et al. [17] | ✗ | ✗ | ✗ | ✗ | 2.0 | client-server/ cloud-based | ✓ | ✗ | n/a | 0.375 |
| MolMil [60] http://gjbekker.github.io/molmil//https://github.com/gjbekker/molmil | ✓ | ✗ | ✓ | ✓ | 1.0 | client | ✓ | ✗ | LGPL | 0.6875 |
| CmPIweb [61] http://CmPIweb.CELLmicrocosmos.org. | ✓ | ✗ | ✗ | ✗ | 1.0 | client-server | futurework | ✗ | n/a | 0.3125 |
| HiC-3D-Viewer [62] https://github.com/mohamed-amine-guerras/HiC3DViewer | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✗ | ✗ | GPL | 0.6875 |
| LiteMol [63] https://www.litemol.org/ | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✓ | ✓/✗ | Apache | 0.875 |
| Web3DMol [64] https://web3dmol.net/ | downloadable on website | ✗ | ✓ | ✓ | 1.0 | client | ✓ | ✗ | n/a | 0.6875 |
| OmicsNet [65] http://www.omicsnet.ca. | ✗ | ✓ | ✓ | ✓ | 2.0 | client-server | ✗ | futurework | n/a | 0.625 |
| NGL [66,67] http://arose.github.io/ngl / https://github.com/arose/ngl | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✓ | ✗ | MIT | 0.8125 |
| HTMol [68] http://htmol.tripplab.com/https://github.com/tripplab/HTMoL | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | futurework | ✗ | MIT | 0.75 |
| MegaMol [69] https://megamol.org//https://github.com/UniStuttgart-VISUS/megamol | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✓ | ✗ | BSD | 0.8125 |
| iCn3d [70] https://github.com/ncbi/icn3d | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✗ | ✗ | public domain | 0.6875 |
| ProteinVR [71] https://durrantlab.pitt.edu/protein-vr/ | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✓ | ✓ | BSD | 0.9375 |
| Physics | ||||||||||
| MeshLabJS [72] https://www.meshlabjs.net// https://github.com/cnr-isti-vclab/meshlabjs | ✓ | ✗ | ✓ | ✓ | 1.0 | client-server | ✗ | ✗ | AGPL | 0.5 |
| Chandler et al. [73] | ✗ | ✗ | ✗ | ✗ | 1.0 | client-server | futurework | ✗ | n/a | 0.1875 |
| OpenJSCAD https://openjscad.org// https://github.com/jscad/OpenJSCAD.org | ✓ | ✓ | ✓ | ✓ | ✗ | client or client-server | ✗ | ✗ | MIT | 0.714 |
| iSpy WebGL [74] http://cern.ch/ispy-webgl/ https://github.com/cms-outreach/ispy-webgl | ✓ | ✓ | ✓ | ✓ | 2.0 | client | ✓ | futurework | MIT | 0.9375 |
| VRMath2 [75] https://vrmath2.net/ https://vrmath2.net/VRM2/ | ✓ | ✓ | ✓ | ✗ | 1.0 | client-server | ✓ | ✓ | copyright | 0.75 |
| SPOT [76] https://github.com/ElsevierSoftwareX/SOFTX_2018_178 | ✓ | ✗ | ✓ | ✓ | ✗ | client-server | ✗ | ✗ | Apache | 0.5 |
| CAD3A [77] http://cad3a.ge.imati.cnr.it/webapp/ / https://github.com/KKaty/CAD_PatternComputation | ✗ | ✓ | ✗ | ✓ | 2.0 | client-server | ✓ | ✗ | MIT | 0.5625 |
| HexaLab [78] https://github.com/cnr-isti-vclab/HexaLab / https://www.hexalab.net/ | ✓ | ✓ | ✓ | ✓ | 2.0 | client | ✗ | ✗ | MIT | 0.75 |
| Abubu.js [79] https://github.com/kaboudian/abubujs / https://chaos.gatech.edu/NGL_CSF/ | ✓ | ✓ | ✓ | ✓ | 2.0 | client | ✓ | ✗ | MIT | 0.875 |
| QMWebJS [80] http://www.parvis3d.org.es/qmweb/ / https://github.com/EdgarFigueiras/QM_Particles_WebGL | ✓ | ✓ | ✗ | ✓ | 2.0 | cloud-based | ✗ | beta | n/a | 0.625 |
| WWT http://worldwidetelescope.org/webclient/ / https://github.com/WorldWideTelescope | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✓ | ✗ | MIT | 0.8125 |
| Geography, Meteorology & Archaeology | ||||||||||
| 3DHOP [81] https://github.com/cnr-isti-vclab/3DHOP | ✓ | ✓ | ✓ | ✓ | 2.0 | client or client-server | ✓ | futurework | GPL | 0.9375 |
| GPlates portal [82] http://portal.gplates.org/ | ✗ | ✓ | ✓ | ✓ | 2.0 | cloud-based | ✓ | ✗ | GPL | 0.75 |
| WebGL Globe https://experiments.withgoogle.com/chrome/globe | ✓ | ✓ | ✓ | ✓ | 2.0 | client | ✓ | ✗ | Apache | 0.875 |
| Koeva et al. [83,84] no url provided | ✓ | |||||||||
| PolarGlobe [85] http://cici.lab.asu.edu/polarglobe/ | ✗ | ✗ | ✓ | ✓ | 2.0 | client-server | ✓ | ✗ | copyright | 0.5625 |
| 3DAV [86] https://github.com/Prieston/3dav | ✓ | ✗ | ✓ | ✓ | 2.0 | client | ✗ | ✗ | n/a | 0.625 |
| Discher et al. [19,87] | ✗ | ✗ | ✗ | ✗ | 2.0 | client-server | ✓ | ✓ | n/a | 0.4375 |
| Liu et al. [88] https://github.com/liusir2000/visAirPollutant | ✓ | ✗ | ✓ | ✗ | 2.0 | client-server | ✗ | ✗ | n/a | 0.4375 |
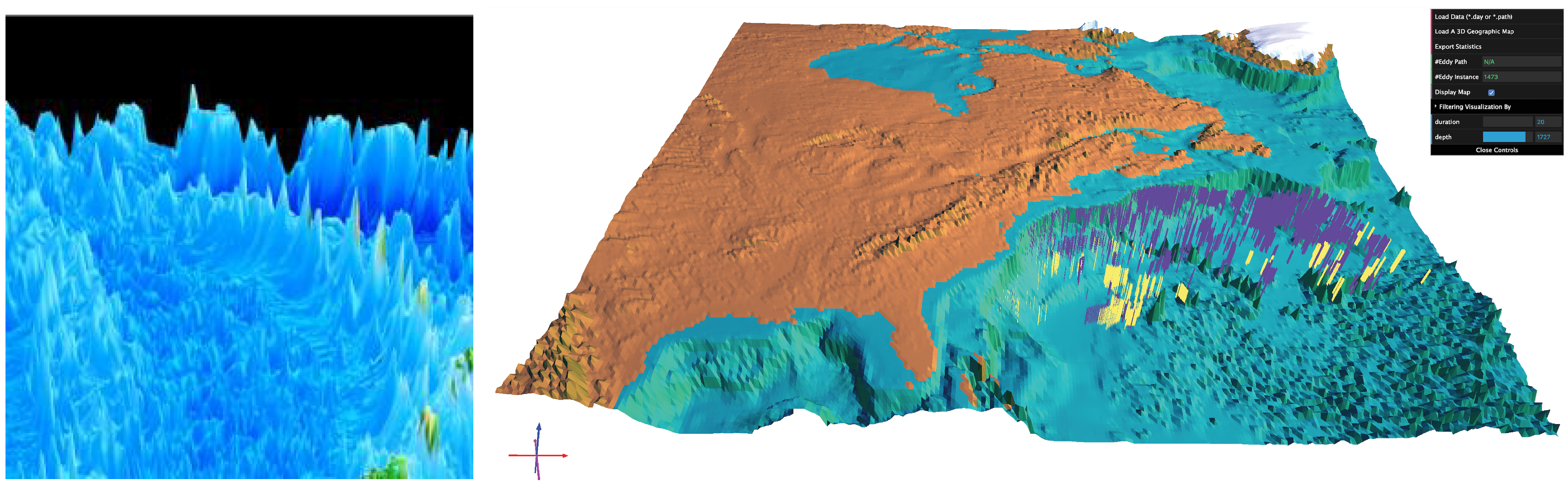
| EddyViz [89] https://vizlab.rutgers.edu/eddyviz.html | ✓ | ✗ | ✓ | ✓ | 2.0 | client-server | ✗ | ✗ | n/a | 0.5625 |
| Boutsi et al. [90] | ✗ | ✓ | ✗ | ✗ | 2.0 | client-server | futurework | ✓ | n/a | 0.5 |
| Cross-Scientific Applications | ||||||||||
| ParaViewWeb [2] https://www.paraview.org/web | ✓ | ✓ | ✓ | ✓ | ✗ | client-server | ✓ | ✗ | BSD | 0.6875 |
| Desprat et al. [91] https://github.com/caro3801/3DP2P | ✓ | ✗ | ✗ | ✗ | 1.0 | client-server | ✗ | ✗ | n/a | 0.25 |
| Tapestry [20] https://github.com/seelabutk/tapestry | ✓ | ✗ | ✓ | ✗ | ✗ | cloud-based | ✓ | ✗ | MIT | 0.5714 |
| Hadjar et al. [92] http://193.194.91.152/test/ / https://datavizcerist.shinyapps.io/dataviz/ | ✗ | ✗ | ✗ | ✗ | 2.0 | client-server | ✓ | ✓ | n/a | 0.4375 |
| Voxer [93] https://github.com/cad420/voxer | ✓ | ✓ | ✓ | ✓ | ✗ | client-server | ✓ | ✗ | n/a | 0.7857 |
| Substrate [94] https://github.com/aplbrain/substrate | ✓ | ✓ | ✓ | ✓ | 2.0 | client-server | ✗ | ✗ | Apache | 0.6875 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Franke, L.; Haehn, D. Modern Scientific Visualizations on the Web. Informatics 2020, 7, 37. https://0-doi-org.brum.beds.ac.uk/10.3390/informatics7040037
Franke L, Haehn D. Modern Scientific Visualizations on the Web. Informatics. 2020; 7(4):37. https://0-doi-org.brum.beds.ac.uk/10.3390/informatics7040037
Chicago/Turabian StyleFranke, Loraine, and Daniel Haehn. 2020. "Modern Scientific Visualizations on the Web" Informatics 7, no. 4: 37. https://0-doi-org.brum.beds.ac.uk/10.3390/informatics7040037






