Targeted Chemometrics Investigations of Source-, Age- and Gender-Dependencies of Oral Cavity Malodorous Volatile Sulphur Compounds
Abstract
:1. Summary
2. Data Description
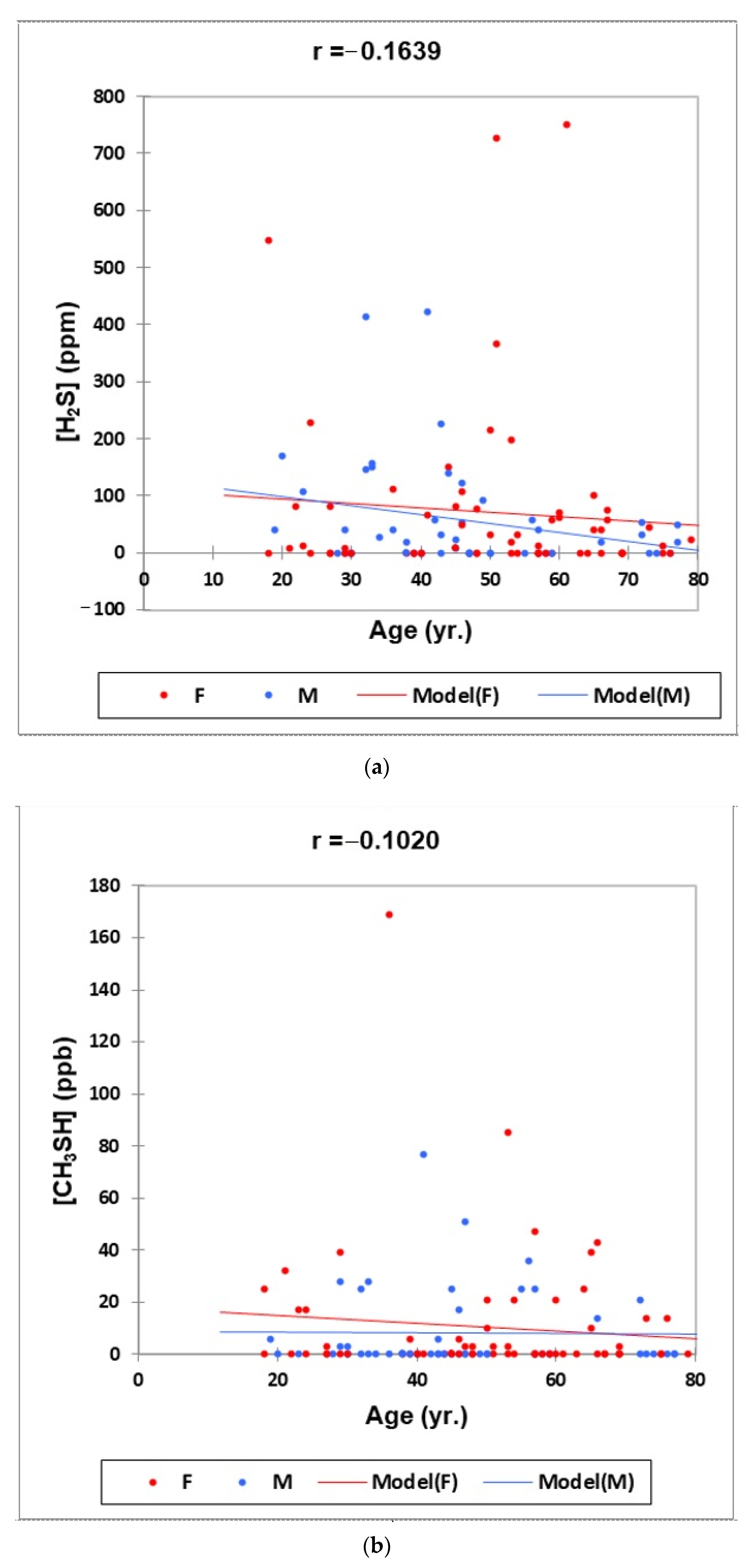
- Figure 1: Plots of (a) H2S, (b) CH3SH and (c) (CH3)2S oral cavity air concentrations versus participant age for females (red) and males (blue) for the outlier-free dataset (n = 114). Correlation coefficient (r) values for these relationships are also provided.
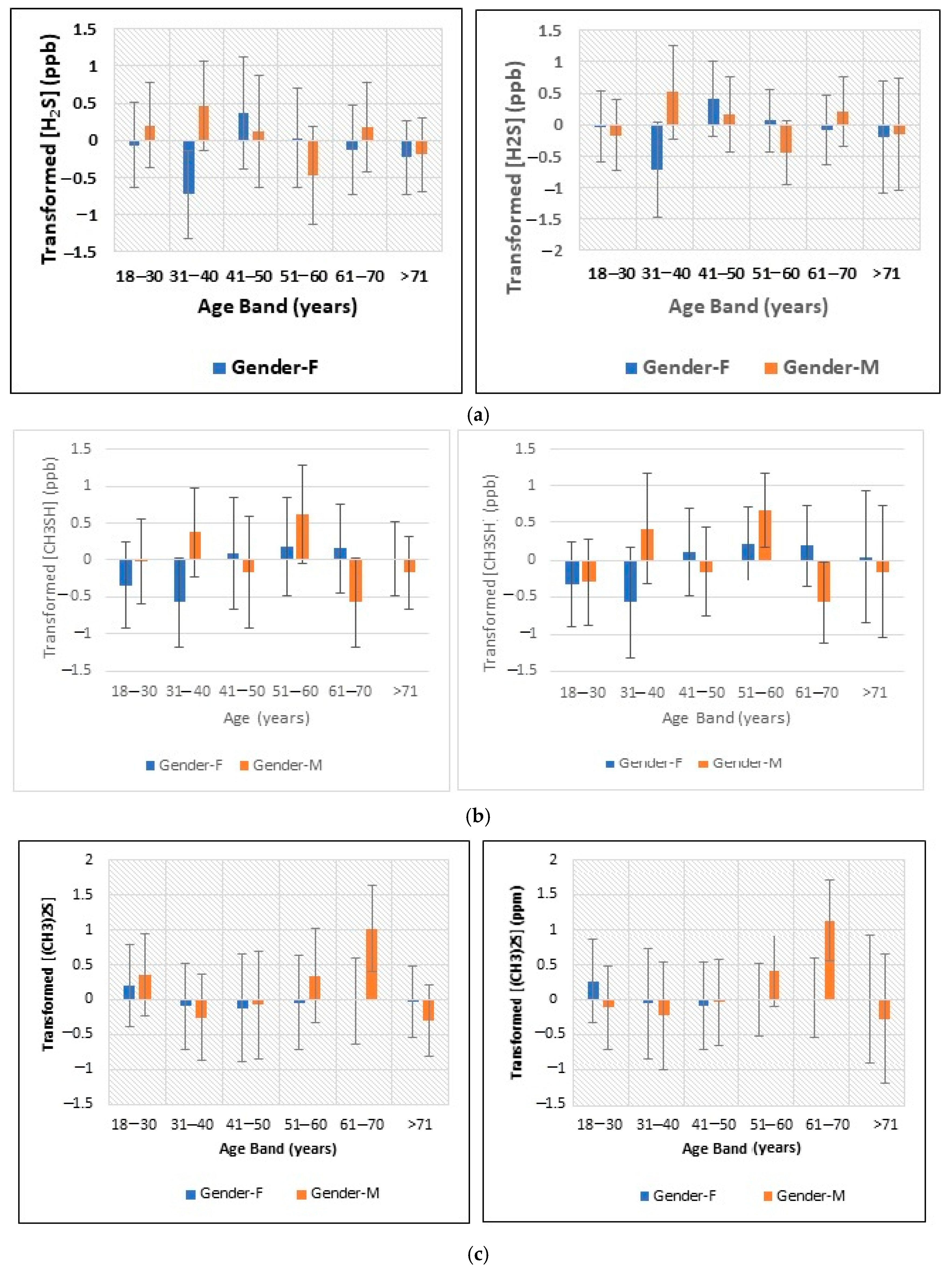
- Figure 2: Plots of least squares mean ± 95% confidence intervals (CIs) for glog-transformed and autoscaled oral cavity concentrations of (a) H2S, (b) CH3SH and (c) (CH3)2S for each age band and gender featured in the analysis-of-variance-based experimental design applied (results obtained both before and after the removal of n = 2 possible outlier participant samples are shown).
- Figure 3: A monoclustering heatmap diagram based on Student’s t-tests displaying ‘between-participant’ variations in oral cavity VSC concentrations for each age band investigated (VSC levels were glog-transformed and autoscaled prior to analysis).
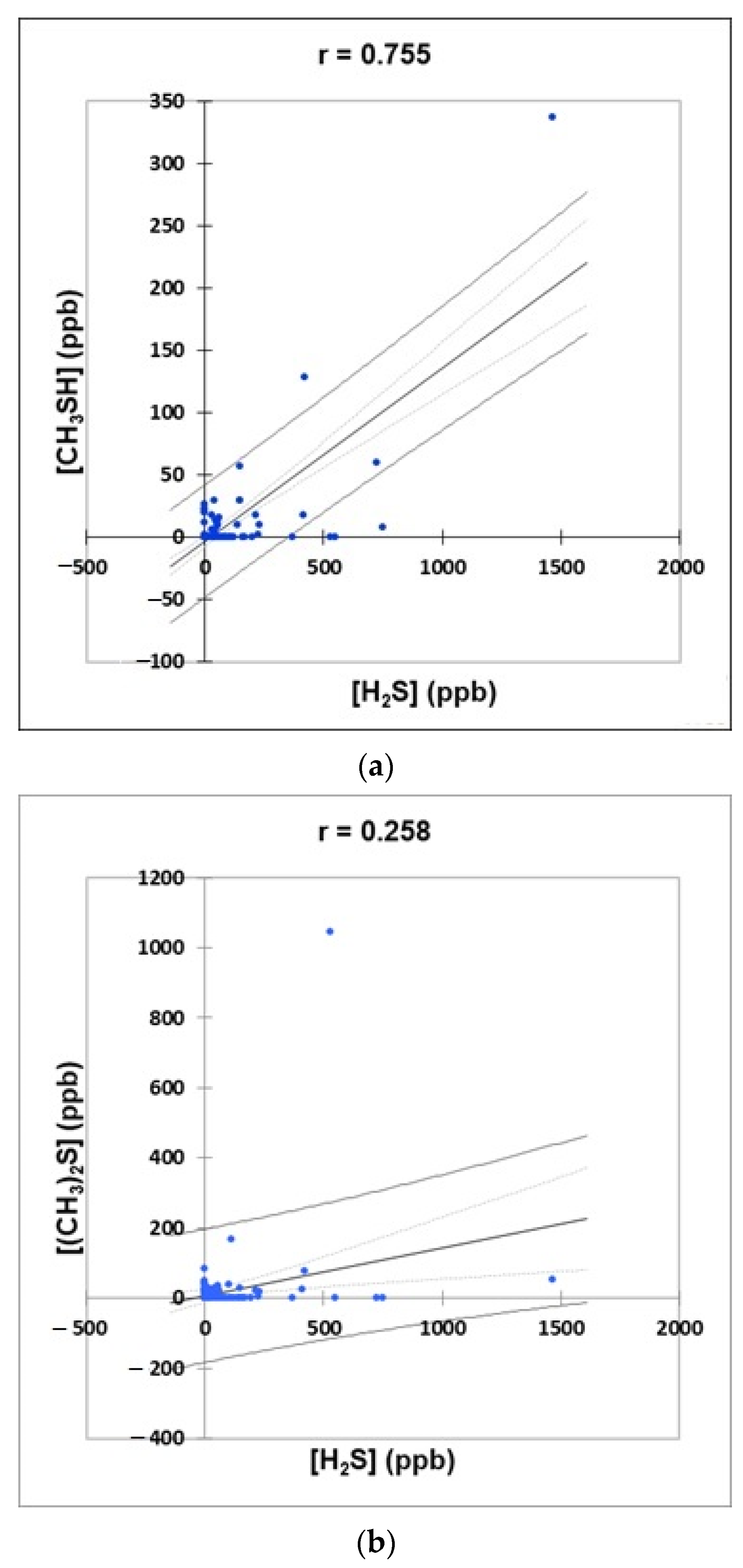
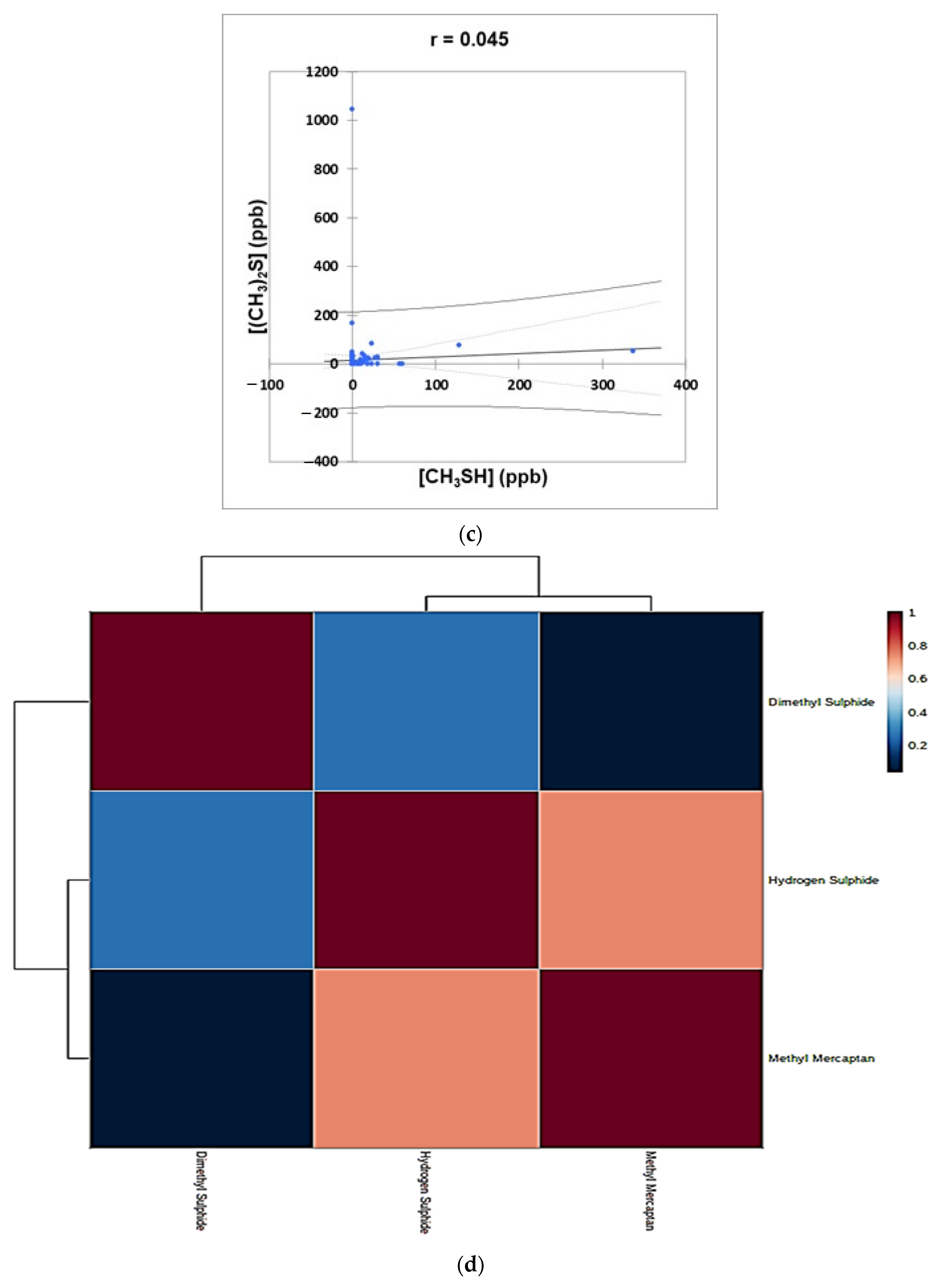
- Figure 4: Plots of (a) CH3SH versus H2S, (b) (CH3)2S versus H2S, and (c) (CH3)2S versus CH3SH concentrations for the complete oral cavity VSC dataset. 95% confidence intervals (CIs) for mean values and observations are denoted. A clustering correlation heatmap is shown in (d).
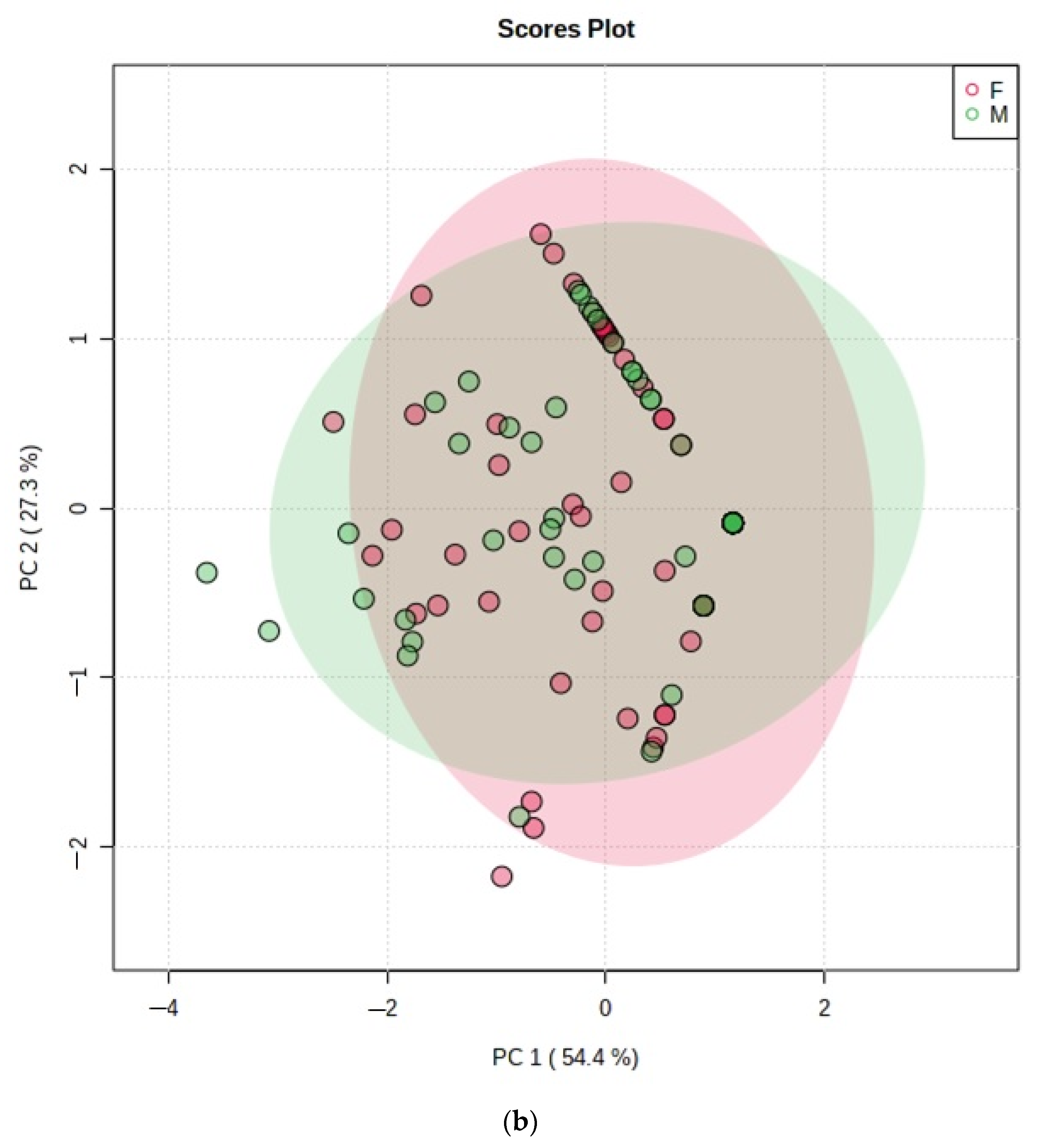
- Figure 5: PCA scores plot with a maximum of two components demonstrating no distinctive clusterings between (a) age bands and (b) genders arising from the VSC dataset. Estimated variance contributions of PC1 and PC2 for PCA models are indicated,
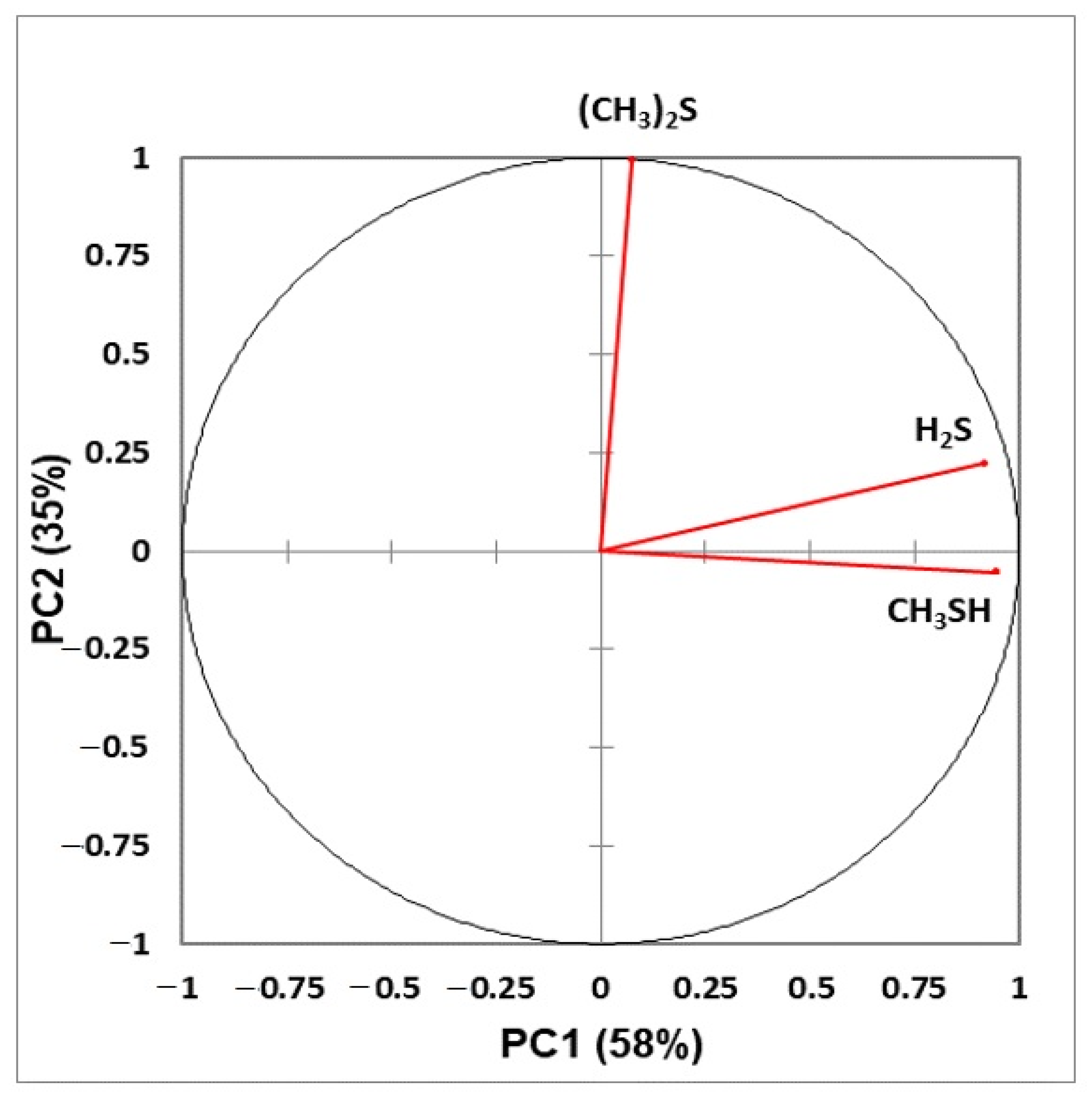
- Figure 6: Correlation circle diagram displaying correlations between all explanatory variables considered, and principal components 1 and 2 in a principal component analysis model applied to the complete VSC dataset.
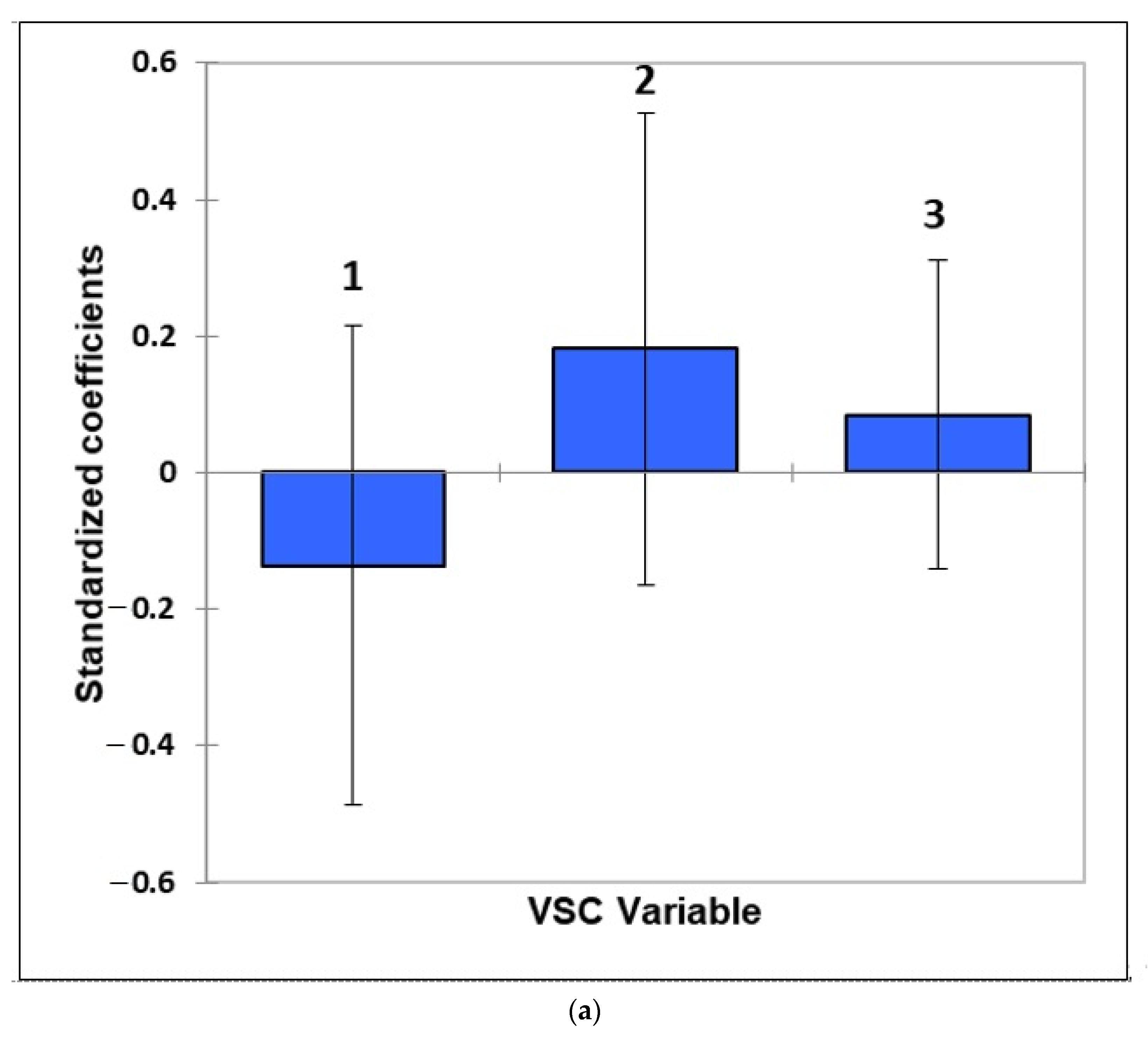
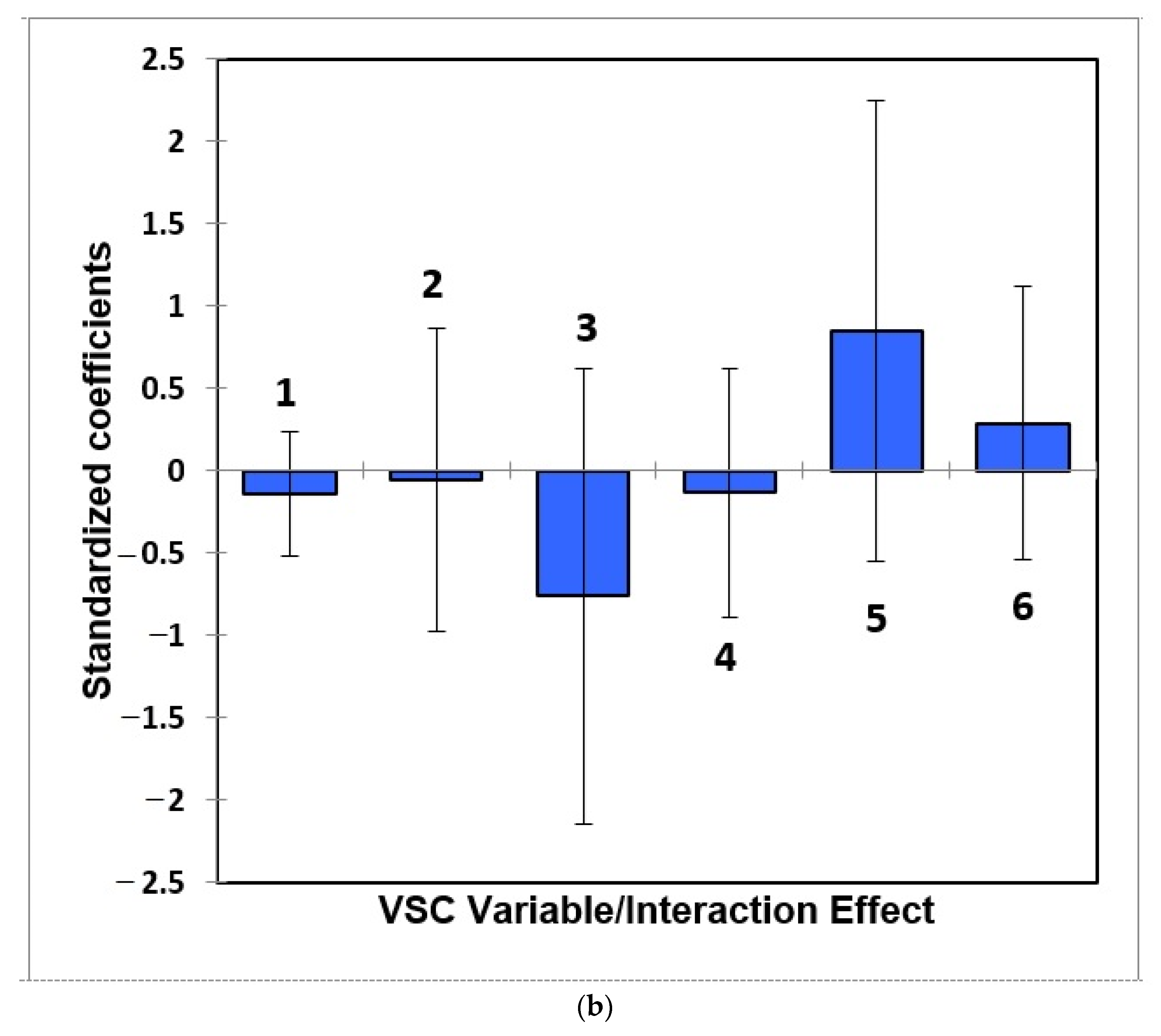
- Figure 7: Estimated standardised coefficients ± 95% CIs for logistic regression analysis models constructed for prediction of the binary participant gender score variable (0 for females, +1 for males) from a model featuring (a) 3 VSC levels alone, and (b), as (a), but with their 3 first-order interaction effects also considered.
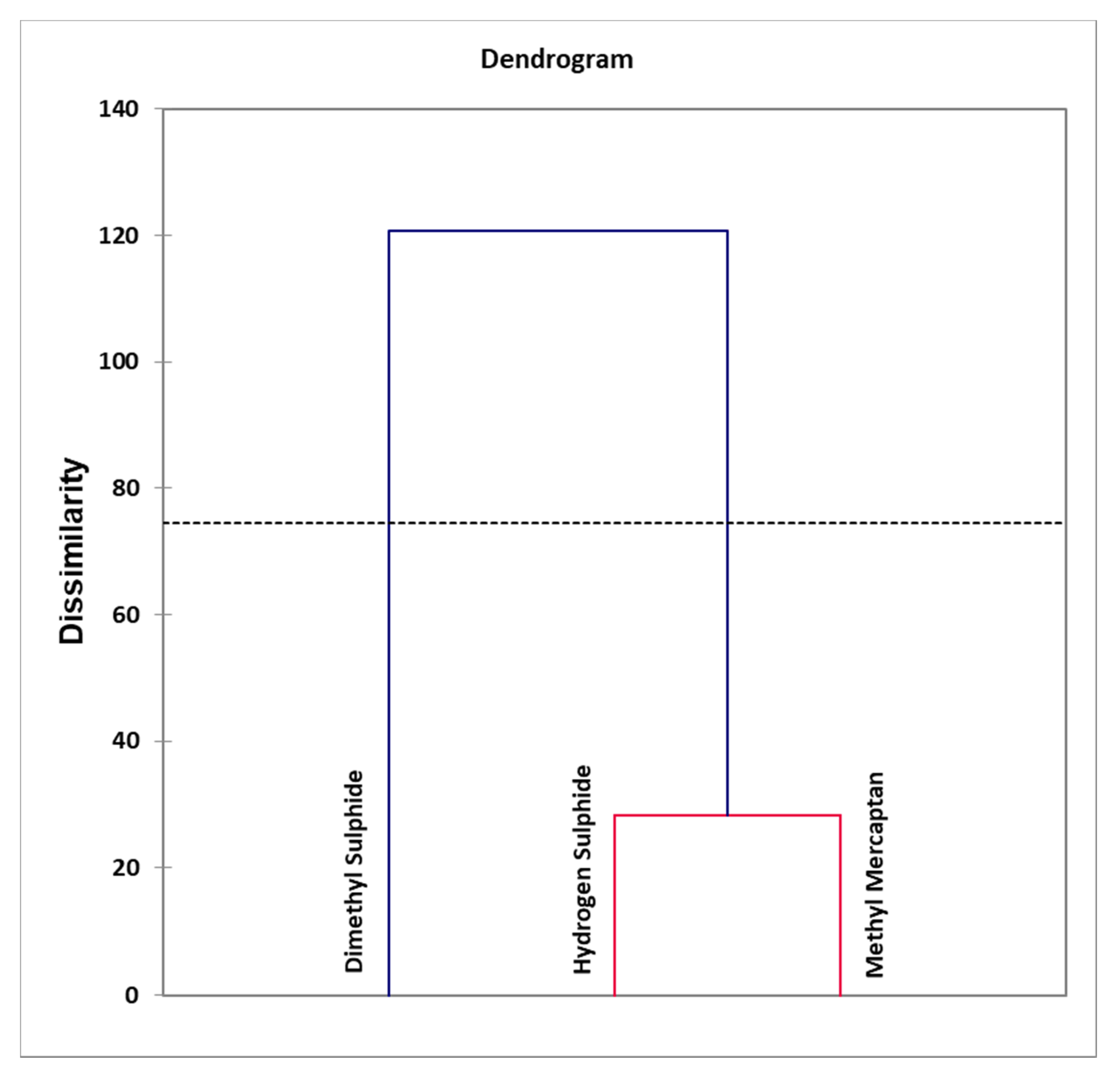
- Figure 8: Agglomerative hierarchal clustering (AHC) dendogram diagram of VSC variables, revealing clear distinctions between clusters arising from an orally-sourced combination of correlated oral cavity H2S and CH3SH levels, and that of uncorrelated (orthogonal) extra-oral (CH3)2S. An automatic entropy-derived threshold limit is also shown.
- Table 1: Frequency listing of numbers of study participants in each age band with oral cavity VSC levels greater than their threshold concentrations of malodorous objectionability (TCMO) limits.
- Table 2: Pearson (n − 1) correlation matrix (correlation coefficient r values) for the raw oral cavity VSC concentration dataset (partial correlation coefficients are also provided). VSCs are indicated in the column and row headings.
- Table 3: Principal component loadings vectors of VSC predictor variables (columns) following Varimax rotation and Kaiser normalisation featured in a principal component analysis (PCA) model of the complete dataset.
- Appendix A: Brief Historical Review of the Development and Applications of Methods for Evaluating Oral Malodour in Humans
- Appendix B: Short Outlines of the Principles and Applications of Multivariate (MV) Analysis Techniques Employed in Chemometrics and Metabolomics Investigations
- Supplement: Complete VSC dataset with gaseous oral cavity H2S, CH3SH and (CH3)2S concentrations (ppb) listed in columns 4, 5 and 6 respectively. Corresponding samples codes, age bands and genders are provided in columns 1, 2 and 3 respectively.
2.1. Study Experimental Design and Criteria
2.2. Statistical Analysis of Oral Cavity Volatile Sulphur Compound (VSC) Dataset
2.3. Results
2.3.1. Exploration of Dependencies of Frequencies of Participants with VSC Levels above Their Threshold Concentrations of Malodorous Objectionability (TCMO) Limits on Age Bands and Gender: Contingency Table Analysis
2.3.2. Preliminary Statistical Investigations: Correlations with Age and Gender
2.3.3. Analysis of Variance (ANOVA), Analysis of Covariance (ANCOVA) and Multivariate Analysis of Variance (MANOVA) of the VSC Dataset
2.3.4. Correlations between Oral Cavity VSC Levels
2.3.5. Application of Multivariate Analysis Techniques: Principal Component Analysis (PCA), Partial Least Squares-Discrimination Analysis (PLS-DA), Partial Least Squares-Discrimination Regression (PLS-R), Orthogonal Partial Least Squares-Discrimination Analysis (OPLS-DA), Principal Component Regression (PCR) and ANOVA-Simultaneous Component Analysis (ASCA)
2.3.6. Logistic Regression Analysis Models
2.3.7. Agglomerative Hierarchal Clustering (AHC) Analysis Model Conducted with Truncation Threshold Limit Setting
2.4. Discussion of Results Obtained
2.5. Conclusions
2.6. Potential Limitations and Strengths of the Study
3. Methods
3.1. Exclusion Criteria
3.2. Details of VSC Measurements
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
Brief Historical Review of the Development and Applications of Methods for Evaluating Oral Malodour in Humans
Appendix B
Appendix B.1. Short Outlines of the Principles and Applications of Multivariate (MV) Analysis Techniques
Appendix B.1.1. Multivariate Analysis-of-Variance (MANOVA)
Appendix B.1.2. Principal Component Analysis (PCA)
Appendix B.1.3. Principal Component Regression (PCR), Partial Least-Squares-Regression and -Discriminant Analysis (PLS-R and PLS-DA respectively), and Orthogonal Projections to Latent Structures-Discriminant Analysis (OPLS-DA)
Appendix B.1.4. ANOVA Simultaneous Component Analysis (ASCA)
Appendix B.1.5. Agglomerative Hierarchical Clustering (AHC)
Appendix B.1.6. Logistic Regression Analysis (logRA)
References
- Tonzetich, J. Production and origin of oral malodor: A review of mechanisms and methods of analysis. J. Periodontol. 1977, 48, 13–20. [Google Scholar] [CrossRef]
- Miyazaki, H.; Sakao, S.; Katoh, Y.; Takehara, T. Correlation between volatile sulphur compounds and certain oral health measurements in the general population. J. Periodontol. 1995, 66, 679–684. [Google Scholar] [CrossRef]
- Du, M.; Li, L.; Jiang, H.; Zheng, Y.; Zhang, J. Prevalence and relevant factors of halitosis in Chinese subjects: A clinical research. BMC Oral Health 2019, 19, 45. [Google Scholar] [CrossRef] [PubMed]
- Hammad, M.M.; Darwazeh, A.M.G.; Al-Waeli, H.; Tarakji, B.; Alhadithy, T.T. Prevalence and awareness of halitosis in a sample of Jordanian population. J. Int. Soc. Prevent. Commun. Dent. 2014, 4, S178–S186. [Google Scholar]
- McNamara, T.F.; Alexander, J.F.; Lee, M. The role of microorganisms in the production of oral malodour. Oral Surg. Oral Med. Oral Pathol. 1972, 34, 41–48. [Google Scholar] [CrossRef]
- Tonzetich, J.; McBride, B.C. Characterization of volatile sulphur production by pathogenic and non-pathogenic strains of oral Bacteroides. Arch. Oral Biol. 1981, 26, 963–969. [Google Scholar] [CrossRef]
- Kleinberg, I.; Westbay, G. Oral Malodor. Crit. Rev. Oral Biol. Med. 1990, 1, 247–260. [Google Scholar] [CrossRef]
- Kostelc, J.G.; Preti, G.; Zelson, P.R.; Brauner, L.; Bachni, P. Oral odors in early experimental gingivitis. J. Periodontal. Res. 1984, 19, 303–312. [Google Scholar] [CrossRef] [PubMed]
- Morris, P.P.; Read, R.R. Halitosis: Variations in mouth and total breath odour intensity resulting from prophylaxis and antisepsis. J. Dent. Res. 1949, 28, 324–333. [Google Scholar] [CrossRef]
- Tonzetich, J. Oral malodour: An indicator of health status and oral cleanliness. Int. J. Dent. Res. 1978, 28, 309–319. [Google Scholar]
- Spouge, J.D. Halitosis. A review of its causes and treatment. Dent. Pract. Dent. Rec. 1964, 14, 307–317. [Google Scholar]
- Prinz, H. Offensive breath, its causes and prevention. Dent. Cosmos 1930, 72, 700–707. [Google Scholar]
- Tachibana, Y. The relation between pyorrhea alveolaris and H2S producing bacteria in human mouth. Kokubyo Gakkai Zasshi 1957, 24, 219–229. [Google Scholar] [CrossRef]
- Yaegaki, K.; Sanada, K. Biochemical and clinical factors influencing oral malodour in periodontal patients. J. Periodontol. 1992, 63, 786–792. [Google Scholar] [CrossRef] [PubMed]
- Tangerman, A.; Winkel, E.G. Intra- and extra-oral halitosis: Finding of a new form of extra-oral blood-borne halitosis caused by dimethyl sulphide. J. Clin. Periodontol. 2007, 34, 748–755. [Google Scholar] [CrossRef]
- Tangerman, A.; Winkel, E.G. Extra-oral halitosis: An overview. J. Breath Res. 2010, 4, 017003. [Google Scholar] [CrossRef] [PubMed]
- Aydin, M.; Harvey-Woodworth, C. Halitosis: A new definition and classification. Br. Dent. J. 2014, 217, E1. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, U.; Sharma, G.; Juneja, M.; Nagpal, A. Halitosis: Current concepts on etiology, diagnosis and management. Eur. J. Dent. 2016, 10, 292–300. [Google Scholar] [CrossRef]
- Madhushankari, G.S.; Yamunadevi, A.; Selvamani, M.; Mohan Kumar, K.P.; Basandi, P.S. Halitosis—An overview: Part-I—Classification, etiology, and pathophysiology of halitosis. J. Pharm. Bioallied. Sci. 2015, 7, S339–S343. [Google Scholar] [CrossRef]
- Snel, J.; Burgering, M.; Smit, B.; Noordman, W.; Tangerman, A.; Winkel, E.G.; Kleerebezem, M. Volatile sulphur compounds in morning breath of human volunteers. Arch. Oral Biol. 2011, 56, 29–34. [Google Scholar] [CrossRef] [Green Version]
- Askin, R. Multicollinearity in regression: Review and examples. J. Forecasting 1982, 1, 281–292. [Google Scholar] [CrossRef]
- Loesche, W.J.; Kazor, C. Microbiology and treatment of halitosis. Periodontol. 2000 2002, 28, 256–279. [Google Scholar] [CrossRef] [PubMed]
- Porter, S.R. Diet and halitosis. Curr. Opin. Clin. Nutr. Metab. Care 2011, 14, 463–468. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Fujimoto, A.; Yoneda, M.; Watanabe, T.; Hirofuji, T.; Hanioka, T. Resting salivary flow independently associated with oral malodor. BMC Oral Health 2016, 17, 23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szabó, A.; Tarnai, Z.; Berkovits, C.; Novák, P.; Mohácsi, A.; Braunitzer, G.; Rakonczay, Z.; Turzó, K.; Nagy, K.; Szabó, G. Volatile sulphur compound measurement with OralChromaTM: A methodological improvement. J. Breath Res. 2015, 9, 016001. [Google Scholar] [CrossRef] [PubMed]
- Hanada, M.; Koda, H.; Onaga, K.; Tanaka, K.; Okabayashi, T.; Itoh, T.; Miyazaki, H. Portable oral malodor analyzer using highly sensitive In2O3 gas sensor combined with a simple gas chromatography system. Anal. Chim. Acta 2003, 475, 27–35. [Google Scholar] [CrossRef]
- Greenman, J.; Duffield, J.; Spencer, P.; Rosenberg, M.; Corry, D.; Saad, S.; Lenton, P.; Majerus, G.; Nachnani, S.S.; El-Maaytah, M. Study on the organoleptic intensity scale for measuring oral malodor. J. Dent. Res. 2004, 83, 81–85. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.-J.; Lee, J.-Y.; Kho, H.-S.; Chung, J.-W.; Park, H.-K.; Kim, Y.-K. A new organoleptic testing method for evaluating halitosis. J. Periodontol. 2009, 80, 93–97. [Google Scholar] [CrossRef]
- Solis-Gaffar, M.C.; Niles, H.P.; Rainieri, W.; Kestenbaum, R.C. Instrumental evaluation of mouth odor in a human clinical study. J. Dentl. Health 1975, 54, 351–357. [Google Scholar]
- Schmidt, N.F.; Missan, S.R.; Tarbet, W.J.; Couper, A.D. The correlation between organoleptic mouth-odor ratings and levels of volatile sulphur compounds. Oral Surg. 1978, 45, 560–566. [Google Scholar] [CrossRef]
- Brening, R.H.; Sulser, G.F.; Fosdick, L. The determination of halitosis by use of the osmoscope and the cryoscopic method. J. Dent. Res. 1939, 18, 127–132. [Google Scholar] [CrossRef]
- Rosenberg, M.; McCulloch, C.A.G. Measurement of oral malodor: Current methods and future prospects. J. Periodont. 1992, 63, 776–782. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, M.; Kulkarni, G.V.; Bosy, A.; McCulloch, C.A.G. Reproducibility and sensitivity of oral malodor measurements with a portable sulphide monitor. J. Dent. Res. 1991, 70, 1436–1440. [Google Scholar] [CrossRef]
- Rosenberg, M.; Septon, I.; Eli, I.; Bar-Ness, R.G.; Gelernter, I.; Brenner, S.; Gabbay, J. Halitosis measurements by an industrial sulphide monitor. J. Periodont. 1991, 62, 487–489. [Google Scholar] [CrossRef]
- Rosenberg, M.; Gelernter, I.; Barki, M.; Bar-Ness, R. Day-long reduction of oral malodor by a two phase oil:water mouthrinse as compared to chlorhexidine gluconate and placebo rinses. J. Periodont. 1992, 63, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Silwood, C.J.L.; Lynch, E.; Grootveld, M. A multifactorial investigation of the ability of oral health care products (OHCPs) to alleviate oral malodour. J. Clin. Periodont. 2001, 28, 634–641. [Google Scholar] [CrossRef]
- Awano, S.; Takata, Y.; Soh, I.; Yoshida, A.; Hamasaki, T.; Sonoki, K.; Ohsumi, T. Correlations between health status and OralChromaTM-determined volatile sulfide levels in mouth air of the elderly. J. Breath Res. 2011, 5, 046007. [Google Scholar] [CrossRef] [PubMed]
- Grootveld, K.L.; Lynch, E.; Grootveld, M. Twelve-hour longevity of the oral malodour-neutralising capacity of an oral rinse product containing the chlorine dioxide precursor sodium chlorite. J. Oral Health Dent. 2018, 1, 003. [Google Scholar]
- Grootveld, M. Metabolic Profiling: Disease and Xenobiotics; Issues in Toxicology Series; Royal Society of Chemistry: Cambridge, UK, 2014; ISBN 1849731632. [Google Scholar]
- López-Rubio, E.; Elizondo, D.A.; Grootveld, M.; Jerez, J.M.; Luque-Baena, R.M. Computational Intelligence Techniques in Medicine; Special issue of Computational and Mathematical Methods in Medicine; Hindawi Publishing Corporation: London, UK, 2014. [Google Scholar]



| Age Band | VSC | Total | ||
|---|---|---|---|---|
| H2S | CH3SH | (CH3)2S | ||
| 18–30 | 5 | 2 | 4 | 23 |
| 31–40 | 5 | 2 | 3 | 16 |
| 41–50 | 7 | 4 | 3 | 27 |
| 51–60 | 3 | 8 | 5 | 22 |
| 61–70 | 2 | 2 | 3 | 14 |
| >71 | 0 | 1 | 0 | 14 |
| Total | 22 | 19 | 18 | 116 |
| VSC Variables | H2S | CH3SH | (CH3)2S |
|---|---|---|---|
| H2S | 1 | 0.755 (0.770) ** | 0.258 (0.342) * |
| CH3SH | 0.755 (0.770) ** | 1 | 0.045 (−0.236) |
| (CH3)2S | 0.258 (0.342) * | 0.045 (−0.236) | 1 |
| PC Loadings after Varimax Rotation | ||
|---|---|---|
| VSC | PC1 | PC2 |
| H2S | 0.92 (0.84) | 0.22 (0.050) |
| CH3SH | 0.95 (0.90) | −0.05 (0.003) |
| (CH3)2S | 0.07 (0.005) | 0.99 (0.99) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grootveld, K.L.; Ruiz-Rodado, V.; Grootveld, M. Targeted Chemometrics Investigations of Source-, Age- and Gender-Dependencies of Oral Cavity Malodorous Volatile Sulphur Compounds. Data 2021, 6, 36. https://0-doi-org.brum.beds.ac.uk/10.3390/data6040036
Grootveld KL, Ruiz-Rodado V, Grootveld M. Targeted Chemometrics Investigations of Source-, Age- and Gender-Dependencies of Oral Cavity Malodorous Volatile Sulphur Compounds. Data. 2021; 6(4):36. https://0-doi-org.brum.beds.ac.uk/10.3390/data6040036
Chicago/Turabian StyleGrootveld, Kerry L., Victor Ruiz-Rodado, and Martin Grootveld. 2021. "Targeted Chemometrics Investigations of Source-, Age- and Gender-Dependencies of Oral Cavity Malodorous Volatile Sulphur Compounds" Data 6, no. 4: 36. https://0-doi-org.brum.beds.ac.uk/10.3390/data6040036






