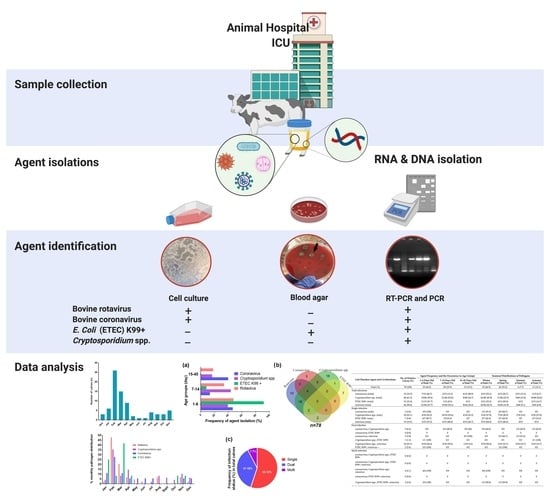
Seasonal and Age-Associated Pathogen Distribution in Newborn Calves with Diarrhea Admitted to ICU
Abstract
:1. Introduction
2. Material and Methods
2.1. Calves and Sampling
Ethics Statement
2.2. Isolation of Viral RNAs and Molecular Viral Screening from Samples
2.3. Genomic DNA Extraction from Stool Samples for the Screening of ETEC K99+ and Cryptosporidium spp.
2.4. Isolation of Viral Pathogens in Cell Cultures
2.5. Isolation of E. coli from Fecal Samples and Colony Screening
2.6. Statistical Analysis
3. Results
3.1. Study Population and Inclusion to Study
3.2. Identified Diarrheal Pathogens
3.3. Causative Agent Isolation Studies
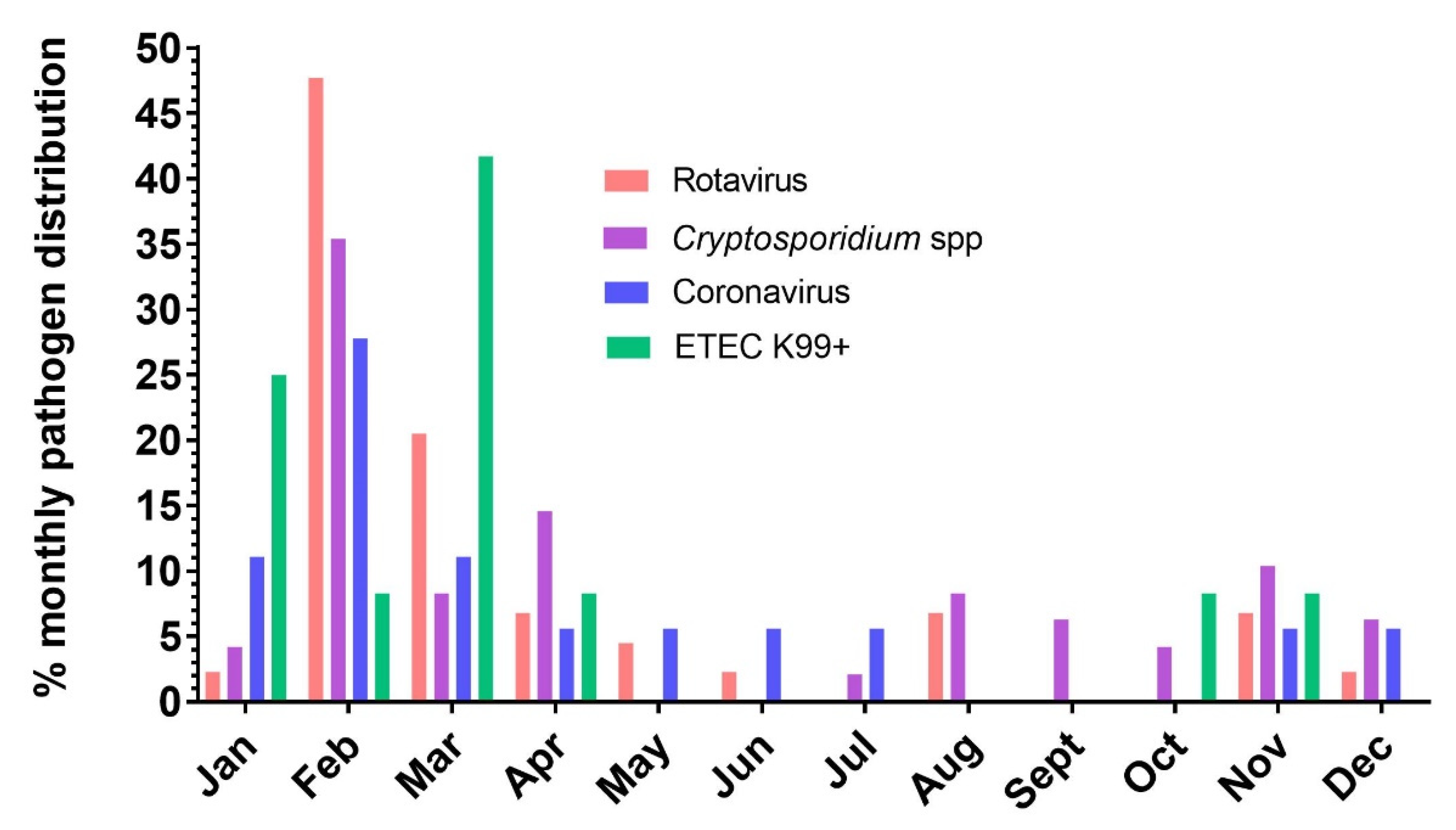
3.4. Risk Factor Analysis and Epidemiological Relevance
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Van Metre, D.C.; Tennant, B.C.; Whitlock, R.H. Infectious Diseases of the Gastrointestinal Tract. In Rebhun’s Diseases of Dairy Cattle, 2nd ed.; Thomas, J.D., Simon, F.P., Eds.; Saunders ELSEVIER: Saint Louis, MO, USA, 2008; pp. 200–294. [Google Scholar] [CrossRef]
- Cho, Y.-I.; Yoon, K.-J. An overview of calf diarrhea–infectious etiology, diagnosis, and intervention. J. Vet. Sci. 2014, 15, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Meganck, V.; Hoflack, G.; Opsomer, G. Advances in prevention and therapy of neonatal dairy calf diarrhoea: A systematical review with emphasis on colostrum management and fluid therapy. Acta Vet. Scand. 2014, 56, 75. [Google Scholar] [CrossRef] [Green Version]
- Perez, E.; Noordhuizen, J.P.T.M.; van Wuijkhuise, L.A.; Stassen, E.N. Management factors related to calf morbidity and mortality rates. Livest. Prod. Sci. 1990, 25, 79–93. [Google Scholar] [CrossRef]
- Bartels, C.J.M.; Holzhauer, M.; Jorritsma, R.; Swart, W.A.J.M.; Lam, T.J.G.M. Prevalence, prediction and risk factors of enteropathogens in normal and non-normal faeces of young Dutch dairy calves. Prev. Vet. Med. 2010, 93, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Nydam, D.V.; Mohammed, H.O. Quantitative risk assessment of Cryptosporidium species infection in dairy calves. J. Dairy Sci. 2005, 88, 3932–3943. [Google Scholar] [CrossRef]
- Timurkan, M.Ö.; Alkan, F. Identification of rotavirus A strains in small ruminants: First detection of G8P [1] genotypes in sheep in Turkey. Arch. Virol. 2020, 165, 425–431. [Google Scholar] [CrossRef]
- Gomez, D.E.; Arroyo, L.G.; Poljak, Z.; Viel, L.; Weese, J.S. Detection of bovine coronavirus in healthy and diarrheic dairy calves. J. Vet. Intern. Med. 2017, 31, 1884–1891. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Windeyer, M.C.; Leslie, K.E.; Godden, S.M.; Hodgins, D.C.; Lissemore, K.D.; LeBlanc, S.J. Factors associated with morbidity, mortality, and growth of dairy heifer calves up to 3 months of age. Prev. Vet. Med. 2014, 113, 231–240. [Google Scholar] [CrossRef]
- Naylor, J.M. Neonatal Calf Diarrhea. In Food Animal Practice, 5th ed.; Anderson, D.E., Rings, D.M., Eds.; W.B. Saunders ELSEVIER: Saint Louis, MO, USA, 2009; pp. 70–77. [Google Scholar] [CrossRef]
- Waltner-Toews, D.; Martin, S.W.; Meek, A.H. Dairy calf management, morbidity and mortality in Ontario Holstein herds. II. Age and seasonal patterns. Prev. Vet. Med. 1986, 4, 125–135. [Google Scholar] [CrossRef]
- Martin, S.W.; Schwabe, C.W.; Franti, C.E. Dairy calf mortality rate: Influence of meteorologic factors on calf mortality rate in Tulare County, California. Am. J. Vet. Res. 1975, 36, 1105–1109. [Google Scholar]
- Kumar, A.; Charpilienne, A.; Cohen, J. Nucleotide sequence of the gene encoding for the RNA binding protein (VP2) of RF bovine rotavirus. Nucleic Acids Res. 1989, 17, 2126. [Google Scholar] [CrossRef] [Green Version]
- Matthijnssens, J.; Ciarlet, M.; Heiman, E.; Arijs, I.; Delbeke, T.; McDonald, S.M.; Palombo, E.A.; Iturriza-Gómara, M.; Maes, P.; Patton, J.T.; et al. Full genome-based classification of rotaviruses reveals a common origin between human Wa-Like and porcine rotavirus strains and human DS-1-like and bovine rotavirus strains. J. Virol. 2008, 82, 3204–3219. [Google Scholar] [CrossRef] [Green Version]
- Dhama, K.; Chauhan, R.S.; Mahendran, M.; Malik, S.V.S. Rotavirus diarrhea in bovines and other domestic animals. Vet. Res. Commun. 2009, 33, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Boileau, M.J.; Kapil, S. Bovine coronavirus associated syndromes. Vet. Clin. N. Am. Food. Anim. Pract. 2010, 26, 123–146. [Google Scholar] [CrossRef] [PubMed]
- Oma, V.S.; Tråvén, M.; Alenius, S.; Myrmel, M.; Stokstad, M. Bovine coronavirus in naturally and experimentally exposed calves; viral shedding and the potential for transmission. Virol. J. 2016, 13, 100. [Google Scholar] [CrossRef] [Green Version]
- Mebus, C.A.; Stair, E.L.; Rhodes, M.B.; Twiehaus, M.J. Pathology of neonatal calf diarrhea induced by a coronavirus-like agent. Vet. Pathol. 1973, 10, 45–64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, C.J.; Hoet, A.E.; Sreevatsan, S.; Wittum, T.E.; Briggs, R.E.; Duff, G.C.; Saif, L.J. Transmission of bovine coronavirus and serologic responses in feedlot calves under field conditions. Am. J. Vet. Res. 2006, 67, 1412–1420. [Google Scholar] [CrossRef] [PubMed]
- Trotz-Williams, L.A.; Wayne Martin, S.; Leslie, K.E.; Duffield, T.; Nydam, D.V.; Peregrine, A.S. Calf-level risk factors for neonatal diarrhea and shedding of Cryptosporidium parvum in Ontario dairy calves. Prev. Vet. Med. 2007, 82, 12–28. [Google Scholar] [CrossRef] [PubMed]
- Feltus, D.C.; Giddings, C.W.; Schneck, B.L.; Monson, T.; Warshauer, D.; McEvoy, J.M. Evidence supporting zoonotic transmission of Cryptosporidium spp. in Wisconsin. J. Clin. Microbiol. 2006, 44, 4303–4308. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Graczyk, T.K.; Fayer, R.; Cranfield, M.R. Zoonotic transmission of Cryptosporidium parvum: Implications for water-borne cryptosporidiosis. Parasitol. Today 1997, 13, 348–351. [Google Scholar] [CrossRef]
- Razakandrainibe, R.; Diawara, E.H.I.; Costa, D.; Le Goff, L.; Lemeteil, D.; Ballet, J.J.; Gargala, G.; Favennec, L. Common occurrence of Cryptosporidium hominis in asymptomatic and symptomatic calves in France. PLoS Negl. Trop. Dis. 2018, 12, e0006355. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hunter, P.R.; Nichols, G. Epidemiology and clinical features of Cryptosporidium infection in immunocompromised patients. Clin. Microbiol. Rev. 2002, 15, 145–154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Semenza, J.C.; Nichols, G. Cryptosporidiosis surveillance and water-borne outbreaks in Europe. Eurosurveillance 2007, 12, 121–123. [Google Scholar] [CrossRef] [Green Version]
- Ryan, U.; Zahedi, A.; Paparini, A. Cryptosporidium in humans and animals—A one health approach to prophylaxis. Parasite Immunol. 2016, 38, 535–547. [Google Scholar] [CrossRef] [Green Version]
- Xiao, L.; Morgan, U.M.; Fayer, R.; Thompson, R.C.A.; Lal, A.A. Cryptosporidium systematics and implications for public health. Parasitol. Today 2000, 16, 287–292. [Google Scholar] [CrossRef]
- Santín, M.; Trout, J.M.; Xiao, L.; Zhou, L.; Greiner, E.; Fayer, R. Prevalence and age-related variation of Cryptosporidium species and genotypes in dairy calves. Vet. Parasitol. 2004, 122, 103–117. [Google Scholar] [CrossRef]
- Dubreuil, J.D.; Isaacson, R.E.; Schifferli, D.M. Animal enterotoxigenic Escherichia coli. EcoSal. Plus 2016, 7, 1–47. [Google Scholar] [CrossRef] [Green Version]
- Shams, Z.; Tahamtan, Y.; Pourbakhsh, A.; Hosseiny, M.H.; Kargar, M.; Hayati, M. Detection of enterotoxigenic K99 (F5) and F41 from fecal sample of calves by molecular and serological methods. Comp. Clin. Path. 2012, 21, 475–478. [Google Scholar] [CrossRef] [PubMed]
- Wattiaux, M.A. Neonatal diarrhoea. In Heifer Raising—Birth to Weaning; University of Wisconsin-Medison, Babcock Institute for International Dairy and Development: Madison, WI, USA, 2005; pp. 121–124. [Google Scholar]
- Chomczynski, P.; Sacchi, N. The single-step method of RNA isolation by acid guanidinium thiocyanate–phenol–chloroform extraction: Twenty-something years on. Nat. Protoc. 2006, 1, 581–585. [Google Scholar] [CrossRef] [PubMed]
- Iturriza Gómara, M.; Wong, C.; Blome, S.; Desselberger, U.; Gray, J. Molecular characterization of VP6 genes of human rotavirus isolates: Correlation of genogroups with subgroups and evidence of independent segregation. J. Virol. 2002, 76, 6596. [Google Scholar] [CrossRef] [Green Version]
- Cho, K.-O.; Hoet, A.E.; Loerch, S.C.; Wittum, T.E.; Saif, L.J. Evaluation of concurrent shedding of bovine coronavirus via the respiratory tract and enteric route in feedlot cattle. Am. J. Vet. Res. 2001, 62, 1436–1441. [Google Scholar] [CrossRef]
- Ryan, U.; Xiao, L.; Read, C.; Zhou, L.; Lal, A.A.; Pavlasek, I. Identification of novel Cryptosporidium genotypes from the Czech Republic. Appl. Environ. Microbiol. 2003, 69, 4302–4307. [Google Scholar] [CrossRef] [Green Version]
- Tang, J.-n.; Zeng, Z.-g.; Wang, H.-n.; Yang, T.; Zhang, P.-j.; Li, Y.-l.; Zhang, A.-y.; Fan, W.-q.; Zhang, Y.; Yang, X.; et al. An effective method for isolation of DNA from pig faeces and comparison of five different methods. J. Microbiol. Methods 2008, 75, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Arnold, M.; Patton, J.T.; McDonald, S.M. Culturing, storage, and quantification of rotaviruses. Curr. Protoc. Microbiol. 2009, 15, 15C–3. [Google Scholar] [CrossRef] [PubMed]
- Wyatt, R.; James, W.; Bohl, E.; Theil, K.; Saif, L.; Kalica, A.; Greenberg, H.; Kapikian, A.; Chanock, R. Human rotavirus type 2: Cultivation in vitro. Science 1980, 207, 189–191. [Google Scholar] [CrossRef] [PubMed]
- Saif, L.J.; Heckert, R.A.; Miller, K.L.; Tarek, M.M. Cell culture propagation of bovine coronavirus. J. Tissue Cult. Methods 1988, 11, 139–145. [Google Scholar] [CrossRef]
- Francis, D.H.; Remmers, G.A.; DeZeeuw, P.S. Production of K88, K99, and 987P antigens by Escherichia coli cultured on synthetic and complex media. J. Clin. Microbiol. 1982, 15, 181–183. [Google Scholar] [CrossRef] [Green Version]
- Güler, L.; Gündüz, K.; Ok, Ü. Virulence factors and antimicrobial susceptibility of Escherichia coli isolated from calves in Turkey. Zoonoses Public Health 2008, 55, 249–257. [Google Scholar] [CrossRef]
- Demir, P.A.; Aydin, E.; Ayvazoglu, C. Estimation of the economic losses related to calf mortalities Kars province in Turkey. Vet. Fak. Derg. 2019, 3, 283–290. [Google Scholar]
- Günlü, A. Buzağı Kayıpları ve Buzağı Hastalıklarının Ekonomik Değerlendirmesi. In Buzaği Sağliği ve Yetiştiriciliği, 1st ed.; Erdem, H., Çiftci, E., Işik, K., Yorgancilar, Ü., Cevdet, Y., Eds.; Medisan: Istanbul, Turkey, 2020; pp. 64–65. [Google Scholar]
- Renaud, D.L.; Rot, C.; Marshall, J.; Steele, M.A. The effect of Cryptosporidium parvum, rotavirus, and coronavirus infection on the health and performance of male dairy calves. J. Dairy Sci. 2021, 104, 2151–2163. [Google Scholar] [CrossRef]
- Benito, A.A.; Monteagudo, L.V.; Arnal, J.L.; Baselga, C.; Quílez, J. Occurrence and genetic diversity of rotavirus A in faeces of diarrheic calves submitted to a veterinary laboratory in Spain. Prev. Vet. Med. 2020, 185, 105196. [Google Scholar] [CrossRef] [PubMed]
- Bertoni, E.A.; Bok, M.; Vega, C.; Martinez, G.M.; Cimino, R.; Parreño, V. Influence of individual or group housing of newborn calves on rotavirus and coronavirus infection during the first 2 months of life. Trop. Anim. Health Prod. 2021, 53, 62. [Google Scholar] [CrossRef] [PubMed]
- Monney, J.D.; Adjogoua, E.V.; Karamoko, Y.; Akran, A. Incidences of calf diarrhea and the associated risk factors in ivory coast (2015–2017). IJAVMS 2020, 19, 454–461. [Google Scholar] [CrossRef]
- Uhde, F.L.; Kaufmann, T.; Sager, H.; Albini, S.; Zanoni, R.; Schelling, E.; Meylan, M. Prevalence of four enteropathogens in the faeces of young diarrhoeic dairy calves in Switzerland. Vet. Rec. 2008, 163, 362–366. [Google Scholar] [CrossRef] [PubMed]
- Çitil, M.; Gunes, V.; Karademir, B. 1996-2001 yılları arasında KAÜ Veteriner Fakültesi İç Hastalıkları Kliniğine getirilen ishalli buzağılar üzerine retrospektif bir çalışma. Vet. Fak. Derg. 2003, 9, 39–42. [Google Scholar]
- Yildirim, A.; Adanir, R.; Inci, A.; Yukari, B.A.; Duzlu, O.; Onder, Z.; Ciloglu, A.; Simsek, E. Prevalence and genotyping of bovine Cryptosporidium species in the Mediterranean and Central Anatolia region of Turkey. Comp. Immunol. Microbiol. Infect. Dis. 2020, 69, 101425. [Google Scholar] [CrossRef]
- Içen, H.; Arserim, N.B.; IŞIK, N.; Özkan, C.; Kaya, A. Prevalence of four enteropathogens with immunochromatographic rapid test in the feces of diarrheic calves in east and southeast of Turkey. Pak. Vet. J. 2013, 33, 496–499. [Google Scholar]
- Brenner, J.; Elad, D.; Markovics, A.; Grinberg, A.; Trainin, Z. Epidemiological study of neonatal calf diarrhoea in Israel–A one-year survey of faecal samples. Isr. J. Vet. Med. 1993, 48, 113. [Google Scholar]
- García, A.; Ruiz-Santa-Quiteria, J.A.; Orden, J.A.; Cid, D.; Sanz, R.; Gómez-Bautista, M.; de la Fuente, R. Rotavirus and concurrent infections with other enteropathogens in neonatal diarrheic dairy calves in Spain. Comp. Immunol. Microbiol. Infect. Dis. 2000, 23, 175–183. [Google Scholar] [CrossRef]
- Reynolds, D.J.; Morgan, J.H.; Chanter, N.; Jones, P.W.; Bridger, J.C.; Debney, T.G.; Bunch, K.J. Microbiology of calf diarrhoea in southern Britain. Vet. Rec. 1986, 119, 34–39. [Google Scholar] [CrossRef]
- Altuğ, N.; Yüksek, N.; Özkan, C.; Keleş, I.; Başbuğan, Y.; Agaoglu, Z.T.; Kaya, A.; Akgul, Y. Rapid etiological diagnosis of neonatal calf diarrhoea with immunochromatographi test kits. Van Vet. J. 2013, 24, 123–128. [Google Scholar]
- Chao, D.L.; Roose, A.; Roh, M.; Kotloff, K.L.; Proctor, J.L. The seasonality of diarrheal pathogens: A retrospective study of seven sites over three years. PLoS Negl. Trop. Dis. 2019, 13, e0007211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Paredes-Paredes, M.; Okhuysen, P.C.; Flores, J.; Mohamed, J.A.; Padda, R.S.; Gonzalez-Estrada, A.; Haley, C.A.; Carlin, L.G.; Nair, P.; DuPont, H.L. Seasonality of diarrheagenic Escherichia coli pathotypes in the US students acquiring diarrhea in Mexico. J. Travel Med. 2011, 18, 121–125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mattioli, M.C.; Boehm, A.B.; Davis, J.; Harris, A.R.; Mrisho, M.; Pickering, A.J. Enteric pathogens in stored drinking water and on caregiver’s hands in tanzanian households with and without reported cases of child diarrhea. PLoS ONE 2014, 9, e84939. [Google Scholar] [CrossRef]
- Cruvinel, L.B.; Ayres, H.; Zapa, D.M.B.; Nicaretta, J.E.; Couto, L.F.M.; Heller, L.M.; Bastos, T.S.A.; Cruz, B.C.; Soares, V.E.; Teixeira, W.F.; et al. Prevalence and risk factors for agents causing diarrhea (coronavirus, rotavirus, Cryptosporidium spp., Eimeria spp., and nematodes helminthes) according to age in dairy calves from Brazil. Trop. Anim. Health Prod. 2020, 52, 777–791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gumusova, S.O.; Yazici, Z.; Albayrak, H.; Meral, Y. Rotavirus and coronavirus prevalence in healthy calves and calves with diarrhea. Medycnya Weterinaria 2007, 63, 62–64. [Google Scholar]
- Torres-Medina, A.; Schlafer, D.H.; Mebus, C.A. Rotaviral and coronaviral diarrhea. Vet. Clin. N. Am. Food Anim. Pract. 1985, 1, 471–493. [Google Scholar] [CrossRef]


| Pathogen | Primer | Target-Loci | Sequence (5′-3′) | Size (bp) | Ref. |
|---|---|---|---|---|---|
| coronavirus | BCoVF | N | GCAATCCAGTAGTAGAGCGT | 730 | Cho et al. [34] |
| BCoVR | CTTAGTGGCATCCTTGCCAA | ||||
| Cryptosporidium spp. | 18SiCF2 | SSU rRNA | GACATATCATTCAAGTTTCTGACC | 763 | Ryan et al. [35] |
| 18SiCR2 | CTGAAGGAGTAAGGAACAACC | ||||
| 18SiCF1 | CCTATCAGCTTTAGACGGTAGG | 587 | |||
| 18SiCR1 | TCTAAGAATTTCACCTCTGACTG | ||||
| E. coli K99+ | K99F | F5 | TATTATCTTAGGTGGTATGG | 314 | Shams et al. [30] |
| K99R | GGTATCCTTTAGCAGCAGTATTTC | ||||
| rotavirus | 157-R | VP6 | GTTTTCCAAGAGTDATHAHYTCAGC | 405 | Iturriza et al. [33] |
| VP6-F | GACGGVGCRACTACATGGT |
| Calf Diarrhea Agent and Co-Infections | No. of Positive Calves (%) | Agent Frequency and the Occurrence in Age Groups | Seasonal Distributions of Pathogens | |||||
|---|---|---|---|---|---|---|---|---|
| 1–6 Days Old n/Total (%) | 7–14 Days Old n/Total (%) | 15–45 Days Old n/Total (%) | Winter n/Total (%) | Spring n/Total (%) | Summer n/Total (%) | Autumn n/Total (%) | ||
| Total (%) | 78 (100) | 35 (44.9) | 28 (35.9) | 15 (19.2) | 35 (44.9) | 26 (33.3) | 6 (7.7) | 11 (14.1) |
| Total infections | ||||||||
| coronavirus (total) | 15 (19.2) | 7/15 (46.7) | 2/15 (13.3) | 6/15 (40.0) | 8/15 (53.3) | 4/15 (26.7) | 2/15 (13.3) | 1/15 (6.7) |
| Cryptosporidium spp. (total) | 48 (61.5) | 19/48 (39.6) | 21/48 (43.8) | 8/48 (16.7) | 22/48 (45.8) | 11/48 (22.9) | 5/48 (10.4) | 10/48 (20.8) |
| ETEC K99+ (total) | 12 (15.4) | 11/12 (91.7) | 1/12 (8.3) | 0/12 | 4/12 (33.3) | 6/12 (50.0) | 0/12 | 2/12 (16.7) |
| rotavirus (total) | 44 (56.4) | 21/44 (47.7) | 15/44 (34.1) | 8/44 (18.2) | 23/44 (52.3) | 14/44 (31.8) | 4/44 (9.1) | 3/44 (6.8) |
| Single infection | ||||||||
| coronavirus (only) | 3 (3.8) | 3/3 (100) | 0/3 | 0/3 | 1/3 (33.3) | 2/3 (66.7) | 0/3 | 0/3 |
| Cryptosporidium spp. (only) | 18 (23.1) | 4/18 (22.2) | 10/18 (55.6) | 4/18 (22.2) | 5/18 (27.8) | 7/18 (38.9) | 1/18 (5.6) | 5/18 (27.8) |
| ETEC K99+ (only) | 7 (9.0) | 6/7 (85.7) | 1/7(14.3) | 0/7 | 3/7 (42.9) | 3/7 (42.9) | 0/7 | 1/7 (14.3) |
| rotavirus (only) | 15 (19.2) | 5/15 (33.3) | 6/15 (40.0) | 4/15 (26.7) | 9/15 (60.0) | 6/15 (40.0) | 0/15 | 0/15 |
| Dual infection | ||||||||
| coronavirus; Cryptosporidium spp. | 5 (6.4) | 0/5 | 2/5 (40.0) | 3/5 (60) | 3/5 (60.0) | 0/5 | 1/5 (20.0) | 1/5 (20.0) |
| coronavirus; ETEC K99+ | 0 (0.0) | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| coronavirus; rotavirus | 3 (3.8) | 0/3 | 0/3 | 3/3 (100) | 0/3 | 2/3 (66.7) | 1/3 (33.3) | 0/3 |
| Cryptosporidium spp.; ETEC K99+ | 1 (1.3) | 1/1 (100) | 0/1 | 0/1 | 0/1 | 0/1 | 0/1 | 1/1 (100) |
| Cryptosporidium spp.; rotavirus | 18 (23.1) | 8/18 (44.4) | 9/18 (50.0) | 1/18 (5.6) | 9/18 (50.0) | 3/18 (16.7) | 3/18 (16.7) | 3/18 (16.7) |
| ETEC K99+; rotavirus + | 2 (2.6) | 2/2 (100) | 0/2 | 0/2 | 0/2 | 2/2 (100) | 0/2 | 0/2 |
| Multi-infection | ||||||||
| coronavirus; Cryptosporidium spp.; ETEC K99+ | 0 (0.0) | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| coronavirus; Cryptosporidium spp.; ETEC K99+; rotavirus | 0 (0.0) | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| coronavirus; Cryptosporidium spp.; rotavirus | 4 (5.1) | 4/4 (100) | 0/4 | 0/4 | 4/4 (100) | 0/4 | 0/4 | 0/4 |
| coronavirus; ETEC K99+; rotavirus | 0 (0.0) | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Cryptosporidium spp.; ETEC K99+; rotavirus | 2 (2.6) | 2/2 (100) | 0/2 | 0/2 | 1/2 (50.0) | 1/2 (50.0) | 0/2 | 0/2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Berber, E.; Çanakoğlu, N.; Sözdutmaz, İ.; Simsek, E.; Sursal, N.; Ekinci, G.; Kökkaya, S.; Arıkan, E.; Ambarcıoğlu, P.; Göksu, A.G.; et al. Seasonal and Age-Associated Pathogen Distribution in Newborn Calves with Diarrhea Admitted to ICU. Vet. Sci. 2021, 8, 128. https://0-doi-org.brum.beds.ac.uk/10.3390/vetsci8070128
Berber E, Çanakoğlu N, Sözdutmaz İ, Simsek E, Sursal N, Ekinci G, Kökkaya S, Arıkan E, Ambarcıoğlu P, Göksu AG, et al. Seasonal and Age-Associated Pathogen Distribution in Newborn Calves with Diarrhea Admitted to ICU. Veterinary Sciences. 2021; 8(7):128. https://0-doi-org.brum.beds.ac.uk/10.3390/vetsci8070128
Chicago/Turabian StyleBerber, Engin, Nurettin Çanakoğlu, İbrahim Sözdutmaz, Emrah Simsek, Neslihan Sursal, Gencay Ekinci, Serkan Kökkaya, Ebru Arıkan, Pınar Ambarcıoğlu, Ayşe Gençay Göksu, and et al. 2021. "Seasonal and Age-Associated Pathogen Distribution in Newborn Calves with Diarrhea Admitted to ICU" Veterinary Sciences 8, no. 7: 128. https://0-doi-org.brum.beds.ac.uk/10.3390/vetsci8070128