The Implications of ncRNAs in the Development of Human Diseases
Abstract
:1. Introduction
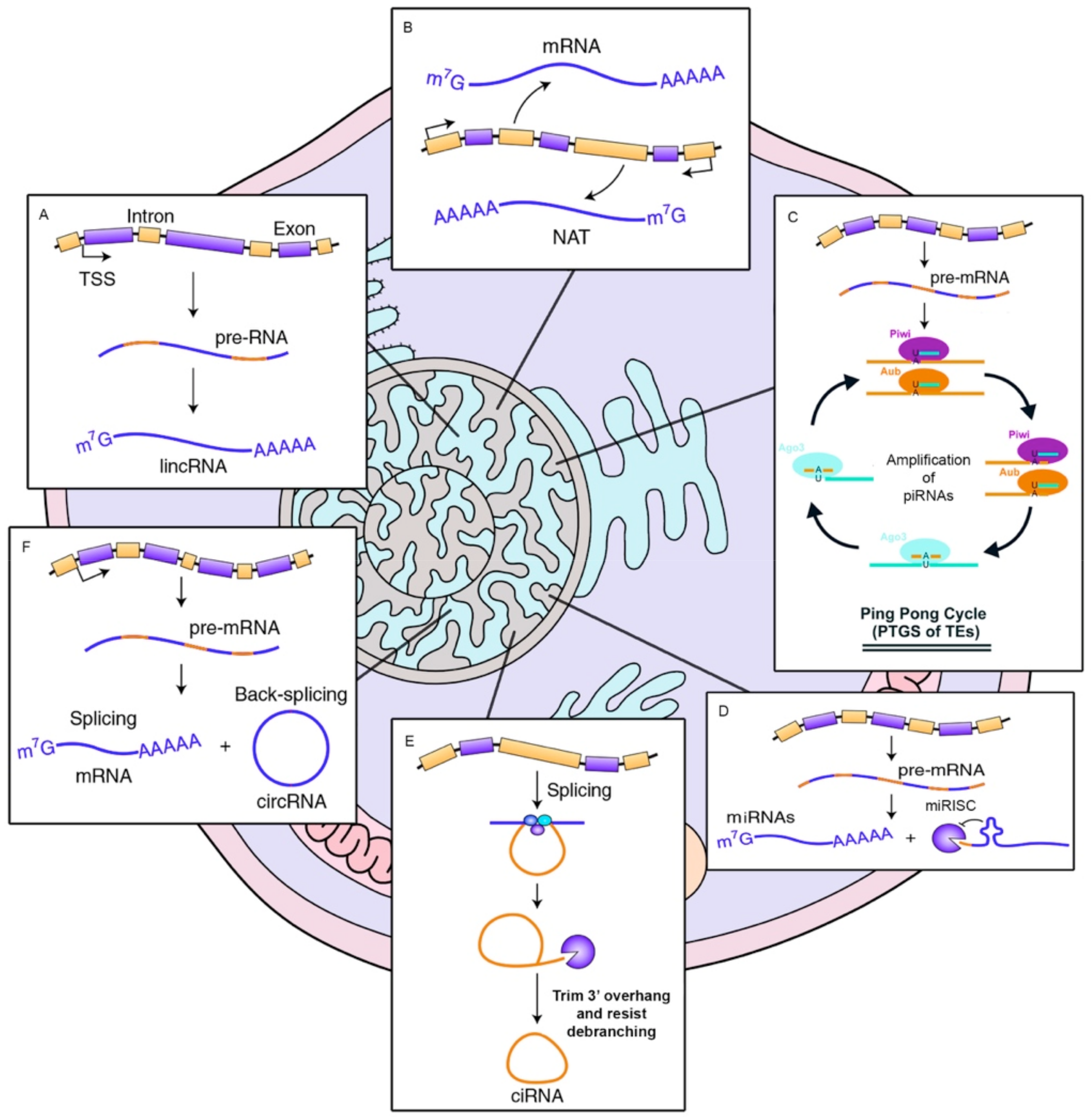
2. Types of Non-Coding RNAs, Biogenesis and Their Cellular Functions
2.1. miRNAs
2.2. lncRNAs
2.3. PiRNAs
2.4. circRNAs
3. Involvement in Human Diseases
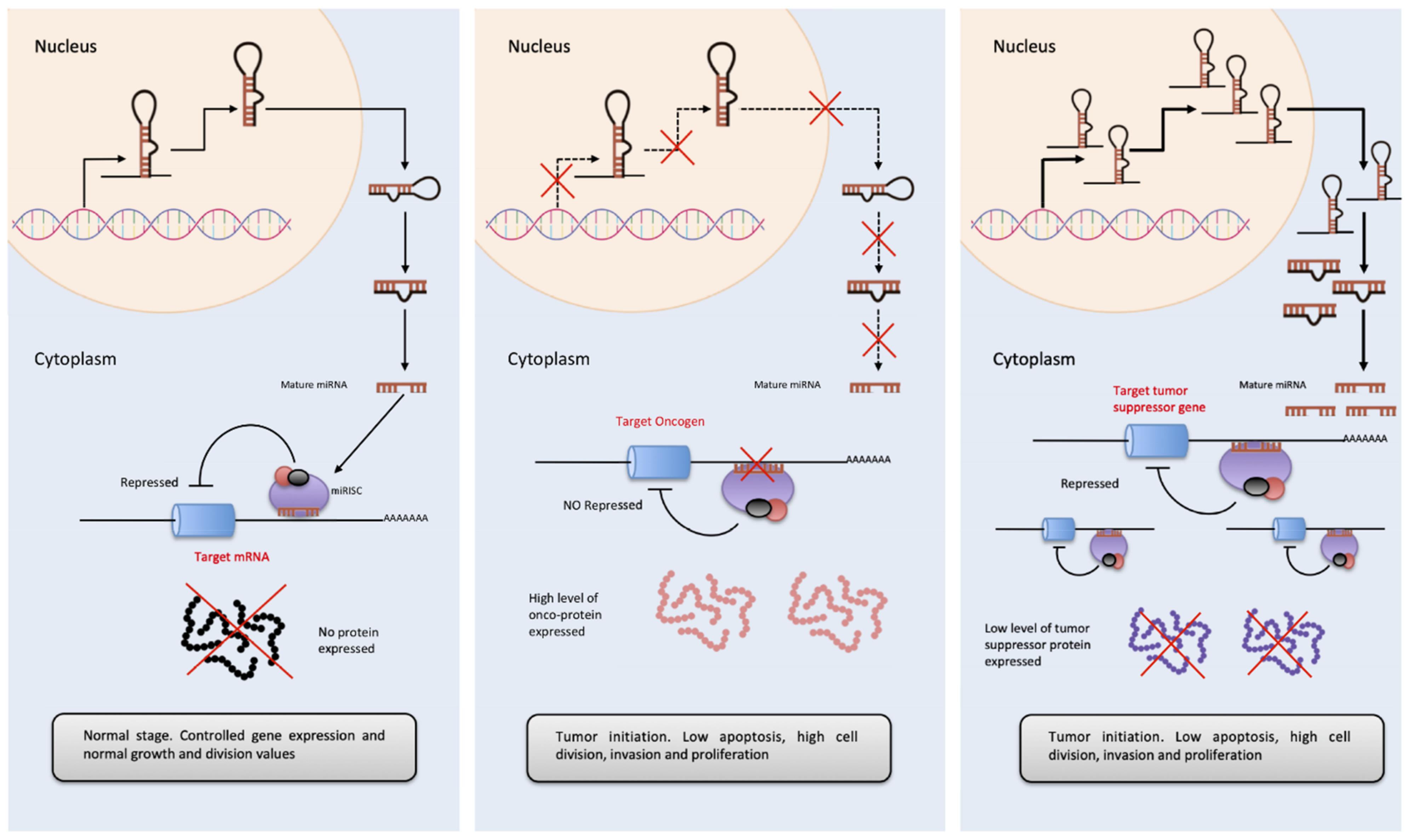
3.1. Non-Coding RNAs in Cancer
3.2. Neurodegenerative Diseases
3.2.1. Cardiovascular Diseases
3.2.2. Other Diseases
4. Bioinformatics Approaches for Identification of Novel Targets in Diseases
5. Importance and Future Perspectives
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Dunham, I.; Kundaje, A.; Aldred, S.F.; Collins, P.J.; Davis, C.; Doyle, F.; Epstein, C.B.; Frietze, S.; Harrow, J.; Kaul, R.; et al. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- Cech, T.R.; Steitz, J.A. The noncoding RNA revolution-trashing old rules to forge new ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [Green Version]
- Szell, M.; Bata-Csorgo, Z.; Kemeny, L. The enigmatic world of mRNA-like ncRNAs: Their role in human evolution and in human diseases. Semin. Cancer Biol. 2008, 18, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Taft, R.J.; Pheasant, M.; Mattick, J.S. The relationship between non-protein-coding DNA and eukaryotic complexity. Bioessays 2007, 29, 288–299. [Google Scholar] [CrossRef]
- Wilusz, J.E.; Freier, S.M.; Spector, D.L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 2008, 135, 919–932. [Google Scholar] [CrossRef] [Green Version]
- Schimmel, P. The emerging complexity of the tRNA world: Mammalian tRNAs beyond protein synthesis. Nat. Rev. Mol. Cell Biol. 2018, 19, 45–58. [Google Scholar] [CrossRef]
- Lafontaine, D.L.; Tollervey, D. The function and synthesis of ribosomes. Nat. Rev. Mol. Cell Biol. 2001, 2, 514–520. [Google Scholar] [CrossRef]
- Pekarsky, Y.; Balatti, V.; Palamarchuk, A.; Rizzotto, L.; Veneziano, D.; Nigita, G.; Rassenti, L.Z.; Pass, H.I.; Kipps, T.J.; Liu, C.G.; et al. Dysregulation of a family of short noncoding RNAs, tsRNAs, in human cancer. Proc. Natl. Acad. Sci. USA 2016, 113, 5071–5076. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuscu, C.; Kumar, P.; Kiran, M.; Su, Z.L.; Malik, A.; Dutta, A. tRNA fragments (tRFs) guide Ago to regulate gene expression post-transcriptionally in a Dicer-independent manner. RNA 2018, 24, 1093–1105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Melton, C.; Reuter, J.A.; Spacek, D.V.; Snyder, M. Recurrent somatic mutations in regulatory regions of human cancer genomes. Nat. Genet. 2015, 47, 710–716. [Google Scholar] [CrossRef]
- Bartel, D.P. Metazoan MicroRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef] [Green Version]
- Batista, P.J.; Chang, H.Y. Long noncoding RNAs: Cellular address codes in development and disease. Cell 2013, 152, 1298–1307. [Google Scholar] [CrossRef] [Green Version]
- Hanly, D.J.; Esteller, M.; Berdasco, M. Interplay between long non-coding RNAs and epigenetic machinery: Emerging targets in cancer? Philos. Trans. R. Soc. Lond. B Biol. Sci. 2018, 373, 20170074. [Google Scholar] [CrossRef]
- Ishizu, H.; Siomi, H.; Siomi, M.C. Biology of PIWI-interacting RNAs: New insights into biogenesis and function inside and outside of germlines. Genes Dev. 2012, 26, 2361–2373. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sivagurunathan, S.; Arunachalam, J.P.; Chidambaram, S. PIWI-like protein, HIWI2 is aberrantly expressed in retinoblastoma cells and affects cell-cycle potentially through OTX2. Cell Mol. Biol. Lett. 2017, 22, 17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Andres-Leon, E.; Nunez-Torres, R.; Rojas, A.M. miARma-Seq: A comprehensive tool for miRNA, mRNA and circRNA analysis. Sci. Rep. 2016, 6, 25749. [Google Scholar] [CrossRef] [Green Version]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef]
- Cai, X.; Hagedorn, C.H.; Cullen, B.R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 2004, 10, 1957–1966. [Google Scholar] [CrossRef] [Green Version]
- Denli, A.M.; Tops, B.B.; Plasterk, R.H.; Ketting, R.F.; Hannon, G.J. Processing of primary microRNAs by the Microprocessor complex. Nature 2004, 432, 231–235. [Google Scholar] [CrossRef]
- Okada, C.; Yamashita, E.; Lee, S.J.; Shibata, S.; Katahira, J.; Nakagawa, A.; Yoneda, Y.; Tsukihara, T. A high-resolution structure of the pre-microRNA nuclear export machinery. Science 2009, 326, 1275–1279. [Google Scholar] [CrossRef]
- Yoda, M.; Kawamata, T.; Paroo, Z.; Ye, X.; Iwasaki, S.; Liu, Q.; Tomari, Y. ATP-dependent human RISC assembly pathways. Nat. Struct. Mol. Biol. 2010, 17, 17–23. [Google Scholar] [CrossRef] [Green Version]
- Adams, D.; Gonzalez-Duarte, A.; O’Riordan, W.D.; Yang, C.C.; Ueda, M.; Kristen, A.V.; Tournev, I.; Schmidt, H.H.; Coelho, T.; Berk, J.L.; et al. Patisiran, an RNAi Therapeutic, for Hereditary Transthyretin Amyloidosis. N. Engl. J. Med. 2018, 379, 11–21. [Google Scholar] [CrossRef]
- Ahadi, A.; Khoury, S.; Losseva, M.; Tran, N. A comparative analysis of lncRNAs in prostate cancer exosomes and their parental cell lines. Genom Data 2016, 9, 7–9. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.S.; Maurin, T.; Robine, N.; Rasmussen, K.D.; Jeffrey, K.L.; Chandwani, R.; Papapetrou, E.P.; Sadelain, M.; O’Carroll, D.; Lai, E.C. Conserved vertebrate mir-451 provides a platform for Dicer-independent, Ago2-mediated microRNA biogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 15163–15168. [Google Scholar] [CrossRef] [Green Version]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [Green Version]
- Bethune, J.; Artus-Revel, C.G.; Filipowicz, W. Kinetic analysis reveals successive steps leading to miRNA-mediated silencing in mammalian cells. EMBO Rep. 2012, 13, 716–723. [Google Scholar] [CrossRef] [Green Version]
- Ma, F.; Lin, P.; Chen, Q.; Lu, X.; Zhang, Y.E.; Wu, C.I. Direct measurement of pervasive weak repression by microRNAs and their role at the network level. BMC Genom. 2018, 19, 362. [Google Scholar] [CrossRef] [Green Version]
- Chi, S.W.; Zang, J.B.; Mele, A.; Darnell, R.B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 2009, 460, 479–486. [Google Scholar] [CrossRef] [Green Version]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Spruce, T.; Pernaute, B.; Di-Gregorio, A.; Cobb, B.S.; Merkenschlager, M.; Manzanares, M.; Rodriguez, T.A. An early developmental role for miRNAs in the maintenance of extraembryonic stem cells in the mouse embryo. Dev. Cell 2010, 19, 207–219. [Google Scholar] [CrossRef] [Green Version]
- Alvarez-Garcia, I.; Miska, E.A. MicroRNA functions in animal development and human disease. Development 2005, 132, 4653–4662. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kung, J.T.; Colognori, D.; Lee, J.T. Long noncoding RNAs: past, present, and future. Genetics 2013, 193, 651–669. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D.; Huarte, M.; Zuk, O.; Carey, B.W.; Cassady, J.P.; et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009, 458, 223–227. [Google Scholar] [CrossRef]
- Cabili, M.N.; Trapnell, C.; Goff, L.; Koziol, M.; Tazon-Vega, B.; Regev, A.; Rinn, J.L. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes. Dev. 2011, 25, 1915–1927. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Yang, C.; Gu, Y.; Li, C.; Zhang, H.; Zhang, W.; Wang, X.; Wu, N.; Zheng, C. Knockdown of the lncRNA SNHG8 inhibits cell growth in Epstein-Barr virus-associated gastric carcinoma. Cell. Mol. Biol. Lett. 2018, 23, 17. [Google Scholar] [CrossRef]
- Wu, Y.; Shao, A.; Wang, L.; Hu, K.; Yu, C.; Pan, C.; Zhang, S. The Role of lncRNAs in the Distant Metastasis of Breast Cancer. Front. Oncol. 2019, 9, 407. [Google Scholar] [CrossRef] [Green Version]
- Martens, J.A.; Wu, P.Y.; Winston, F. Regulation of an intergenic transcript controls adjacent gene transcription in Saccharomyces cerevisiae. Genes. Dev. 2005, 19, 2695–2704. [Google Scholar] [CrossRef] [Green Version]
- Engreitz, J.M.; Haines, J.E.; Perez, E.M.; Munson, G.; Chen, J.; Kane, M.; McDonel, P.E.; Guttman, M.; Lander, E.S. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 2016, 539, 452–455. [Google Scholar] [CrossRef]
- Wu, Z.; Liu, X.; Liu, L.; Deng, H.; Zhang, J.; Xu, Q.; Cen, B.; Ji, A. Regulation of lncRNA expression. Cell. Mol. Biol. Lett. 2014, 19, 561–575. [Google Scholar] [CrossRef]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef] [Green Version]
- Carrieri, C.; Cimatti, L.; Biagioli, M.; Beugnet, A.; Zucchelli, S.; Fedele, S.; Pesce, E.; Ferrer, I.; Collavin, L.; Santoro, C.; et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature 2012, 491, 454–457. [Google Scholar] [CrossRef] [PubMed]
- Orom, U.A.; Derrien, T.; Beringer, M.; Gumireddy, K.; Gardini, A.; Bussotti, G.; Lai, F.; Zytnicki, M.; Notredame, C.; Huang, Q.; et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 2010, 143, 46–58. [Google Scholar] [CrossRef] [Green Version]
- Andrews, S.J.; Rothnagel, J.A. Emerging evidence for functional peptides encoded by short open reading frames. Nat. Rev. Genet. 2014, 15, 193–204. [Google Scholar] [CrossRef] [PubMed]
- Housman, G.; Ulitsky, I. Methods for distinguishing between protein-coding and long noncoding RNAs and the elusive biological purpose of translation of long noncoding RNAs. Biochim. Biophys. Acta-Gene Regul. Mech. 2016, 1859, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Faghihi, M.A.; Zhang, M.; Huang, J.; Modarresi, F.; Van der Brug, M.P.; Nalls, M.A.; Cookson, M.R.; St-Laurent, G., 3rd; Wahlestedt, C. Evidence for natural antisense transcript-mediated inhibition of microRNA function. Genome Biol. 2010, 11, R56. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Modarresi, F.; Faghihi, M.A.; Lopez-Toledano, M.A.; Fatemi, R.P.; Magistri, M.; Brothers, S.P.; van der Brug, M.P.; Wahlestedt, C. Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat. Biotechnol. 2012, 30, 453–459. [Google Scholar] [CrossRef] [Green Version]
- Wan, G.; Zhou, W.; Hu, Y.; Ma, R.; Jin, S.; Liu, G.; Jiang, Q. Transcriptional Regulation of lncRNA Genes by Histone Modification in Alzheimer’s Disease. Biomed. Res. Int. 2016, 2016, 3164238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Engreitz, J.M.; Ollikainen, N.; Guttman, M. Long non-coding RNAs: Spatial amplifiers that control nuclear structure and gene expression. Nat. Rev. Mol. Cell Biol. 2016, 17, 756–770. [Google Scholar] [CrossRef] [Green Version]
- Aravin, A.A.; Sachidanandam, R.; Bourc’his, D.; Schaefer, C.; Pezic, D.; Toth, K.F.; Bestor, T.; Hannon, G.J. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol. Cell 2008, 31, 785–799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brennecke, J.; Aravin, A.A.; Stark, A.; Dus, M.; Kellis, M.; Sachidanandam, R.; Hannon, G.J. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 2007, 128, 1089–1103. [Google Scholar] [CrossRef] [Green Version]
- Lee, E.; Iskow, R.; Yang, L.; Gokcumen, O.; Haseley, P.; Luquette, L.J., 3rd; Lohr, J.G.; Harris, C.C.; Ding, L.; Wilson, R.K.; et al. Landscape of somatic retrotransposition in human cancers. Science 2012, 337, 967–971. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rojas-Rios, P.; Simonelig, M. piRNAs and PIWI proteins: Regulators of gene expression in development and stem cells. Development 2018, 145. [Google Scholar] [CrossRef] [Green Version]
- Han, B.W.; Zamore, P.D. piRNAs. Curr. Biol. 2014, 24, R730–R733. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luteijn, M.J.; Ketting, R.F. PIWI-interacting RNAs: From generation to transgenerational epigenetics. Nat. Rev. Genet. 2013, 14, 523–534. [Google Scholar] [CrossRef] [Green Version]
- Ross, R.J.; Weiner, M.M.; Lin, H. PIWI proteins and PIWI-interacting RNAs in the soma. Nature 2014, 505, 353–359. [Google Scholar] [CrossRef] [Green Version]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol. Cell 2017, 66, 22–37. [Google Scholar] [CrossRef] [Green Version]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Lopez-Jimenez, E.; Rojas, A.M.; Andres-Leon, E. RNA sequencing and Prediction Tools for Circular RNAs Analysis. Adv. Exp. Med. Biol. 2018, 1087, 17–33. [Google Scholar] [CrossRef]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jeck, W.R.; Sharpless, N.E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 2014, 32, 453–461. [Google Scholar] [CrossRef]
- Zaphiropoulos, P.G. Circular RNAs from transcripts of the rat cytochrome P450 2C24 gene: Correlation with exon skipping. Proc. Natl. Acad. Sci. USA 1996, 93, 6536–6541. [Google Scholar] [CrossRef] [Green Version]
- Chen, B.J.; Mills, J.D.; Takenaka, K.; Bliim, N.; Halliday, G.M.; Janitz, M. Characterization of circular RNAs landscape in multiple system atrophy brain. J. Neurochem. 2016, 139, 485–496. [Google Scholar] [CrossRef]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Du, W.W.; Yang, W.; Liu, E.; Yang, Z.; Dhaliwal, P.; Yang, B.B. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016, 44, 2846–2858. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gruner, H.; Cortes-Lopez, M.; Cooper, D.A.; Bauer, M.; Miura, P. CircRNA accumulation in the aging mouse brain. Sci. Rep. 2016, 6, 38907. [Google Scholar] [CrossRef] [PubMed]
- Karreth, F.A.; Tay, Y.; Perna, D.; Ala, U.; Tan, S.M.; Rust, A.G.; DeNicola, G.; Webster, K.A.; Weiss, D.; Perez-Mancera, P.A.; et al. In Vivo Identification of Tumor-Suppressive PTEN ceRNAs in an Oncogenic BRAF-Induced Mouse Model of Melanoma. Cell 2011, 147, 382–395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, C.; Zhang, J.; Qi, H.P.; Feng, C.C.; Chen, Y.P.; Cao, Y.G.; Ba, L.N.; Ai, B.; Wang, Q.Y.; Huang, W.; et al. The global view of mRNA-related ceRNA cross-talks across cardiovascular diseases. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [Green Version]
- Ghoussaini, M.; Mountjoy, E.; Carmona, M.; Peat, G.; Schmidt, E.M.; Hercules, A.; Fumis, L.; Miranda, A.; Carvalho-Silva, D.; Buniello, A.; et al. Open Targets Genetics: Systematic identification of trait-associated genes using large-scale genetics and functional genomics. Nucleic Acids Res. 2021, 49, D1311–D1320. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Tong, Y.; Zhang, H.M.; Wang, K.; Hu, T.; Shan, G.; Sun, J.; Guo, A.Y. Genome-Wide Identification of SNPs in MicroRNA Genes and the SNP Effects on MicroRNA Target Binding and Biogenesis. Hum. Mutat. 2012, 33, 254–263. [Google Scholar] [CrossRef] [PubMed]
- Tanic, M.; Andres, E.; Rodriguez-Pinilla, S.M.; Marquez-Rodas, I.; Cebollero-Presmanes, M.; Fernandez, V.; Osorio, A.; Benitez, J.; Martinez-Delgado, B. MicroRNA-based molecular classification of non-BRCA1/2 hereditary breast tumours. Br. J. Cancer 2013, 109, 2724–2734. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Freedman, M.L.; Monteiro, A.N.; Gayther, S.A.; Coetzee, G.A.; Risch, A.; Plass, C.; Casey, G.; De Biasi, M.; Carlson, C.; Duggan, D.; et al. Principles for the post-GWAS functional characterization of cancer risk loci. Nat. Genet. 2011, 43, 513–518. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andres-Leon, E.; Cases, I.; Alonso, S.; Rojas, A.M. Novel miRNA-mRNA interactions conserved in essential cancer pathways. Sci. Rep. 2017, 7, 46101. [Google Scholar] [CrossRef] [PubMed]
- Esquela-Kerscher, A.; Slack, F.J. Oncomirs—microRNAs with a role in cancer. Nat. Rev. Cancer 2006, 6, 259–269. [Google Scholar] [CrossRef] [PubMed]
- Hammond, S.M. MicroRNAs as oncogenes. Curr. Opin. Genet. Dev. 2006, 16, 4–9. [Google Scholar] [CrossRef]
- Hossain, A.; Kuo, M.T.; Saunders, G.F. Mir-17-5p regulates breast cancer cell proliferation by inhibiting translation of AIB1 mRNA. Mol. Cell Biol. 2006, 26, 8191–8201. [Google Scholar] [CrossRef] [Green Version]
- Akao, Y.; Nakagawa, Y.; Naoe, T. MicroRNAs 143 and 145 are possible common onco-microRNAs in human cancers. Oncol. Rep. 2006, 16, 845–850. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, H.I.; Yamagata, K.; Sugimoto, K.; Iwamoto, T.; Kato, S.; Miyazono, K. Modulation of microRNA processing by p53. Nature 2009, 460, 529–533. [Google Scholar] [CrossRef]
- Kent, O.A.; Chivukula, R.R.; Mullendore, M.; Wentzel, E.A.; Feldmann, G.; Lee, K.H.; Liu, S.; Leach, S.D.; Maitra, A.; Mendell, J.T. Repression of the miR-143/145 cluster by oncogenic Ras initiates a tumor-promoting feed-forward pathway. Genes Dev. 2010, 24, 2754–2759. [Google Scholar] [CrossRef] [Green Version]
- Yuan, F.; Sun, R.F.; Li, L.J.; Jin, B.; Wang, Y.Y.; Liang, Y.D.; Che, G.L.; Gao, L.B.; Zhang, L. A functional variant rs353292 in the flanking region of miR-143/145 contributes to the risk of colorectal cancer. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [Green Version]
- Janaszak-Jasiecka, A.; Bartoszewska, S.; Kochan, K.; Piotrowski, A.; Kalinowski, L.; Kamysz, W.; Ochocka, R.J.; Bartoszewski, R.; Collawn, J.F. miR-429 regulates the transition between Hypoxia-Inducible Factor (HIF)1A and HIF3A expression in human endothelial cells. Sci. Rep. 2016, 6, 22775. [Google Scholar] [CrossRef]
- Lu, Y.; Zhao, X.; Liu, Q.; Li, C.; Graves-Deal, R.; Cao, Z.; Singh, B.; Franklin, J.L.; Wang, J.; Hu, H.; et al. lncRNA MIR100HG-derived miR-100 and miR-125b mediate cetuximab resistance via Wnt/beta-catenin signaling. Nat. Med. 2017, 23, 1331–1341. [Google Scholar] [CrossRef]
- Sevignani, C.; Calin, G.A.; Nnadi, S.C.; Shimizu, M.; Davuluri, R.V.; Hyslop, T.; Demant, P.; Croce, C.M.; Siracusa, L.D. MicroRNA genes are frequently located near mouse cancer susceptibility loci. Proc. Natl. Acad. Sci. USA 2007, 104, 8017–8022. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, H.; Maruyama, R.; Yamamoto, E.; Kai, M. Epigenetic alteration and microRNA dysregulation in cancer. Front. Genet. 2013, 4, 258. [Google Scholar] [CrossRef] [Green Version]
- O’Donnell, K.A.; Wentzel, E.A.; Zeller, K.I.; Dang, C.V.; Mendell, J.T. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005, 435, 839–843. [Google Scholar] [CrossRef]
- Viswanathan, S.R.; Powers, J.T.; Einhorn, W.; Hoshida, Y.; Ng, T.L.; Toffanin, S.; O’Sullivan, M.; Lu, J.; Phillips, L.A.; Lockhart, V.L.; et al. Lin28 promotes transformation and is associated with advanced human malignancies. Nat. Genet. 2009, 41, 843–848. [Google Scholar] [CrossRef] [Green Version]
- Yang, Q.; Jie, Z.; Ye, S.; Li, Z.; Han, Z.; Wu, J.; Yang, C.; Jiang, Y. Genetic variations in miR-27a gene decrease mature miR-27a level and reduce gastric cancer susceptibility. Oncogene 2014, 33, 193–202. [Google Scholar] [CrossRef]
- Hon, C.C.; Ramilowski, J.A.; Harshbarger, J.; Bertin, N.; Rackham, O.J.L.; Gough, J.; Denisenko, E.; Schmeier, S.; Poulsen, T.M.; Severin, J.; et al. An atlas of human long non-coding RNAs with accurate 5 ‘ ends. Nature 2017, 543, 199–204. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.; Liu, S.; Ye, F.; Shen, Y.; Tie, Y.; Zhu, J.; Wei, L.; Jin, Y.; Fu, H.; Wu, Y.; et al. Long Noncoding RNA MEG3 Interacts with p53 Protein and Regulates Partial p53 Target Genes in Hepatoma Cells. PLoS ONE 2015, 10, e0139790. [Google Scholar] [CrossRef]
- Zhang, J.; Yao, T.; Wang, Y.; Yu, J.; Liu, Y.; Lin, Z. Long noncoding RNA MEG3 is downregulated in cervical cancer and affects cell proliferation and apoptosis by regulating miR-21. Cancer Biol. Ther. 2016, 17, 104–113. [Google Scholar] [CrossRef] [Green Version]
- Ying, L.; Huang, Y.; Chen, H.; Wang, Y.; Xia, L.; Chen, Y.; Liu, Y.; Qiu, F. Downregulated MEG3 activates autophagy and increases cell proliferation in bladder cancer. Mol. Biosyst. 2013, 9, 407–411. [Google Scholar] [CrossRef]
- Conte, F.; Fiscon, G.; Chiara, M.; Colombo, T.; Farina, L.; Paci, P. Role of the long non-coding RNA PVT1 in the dysregulation of the ceRNA-ceRNA network in human breast cancer. PLoS ONE 2017, 12, e0171661. [Google Scholar] [CrossRef] [Green Version]
- Perry, M.M.; Tsitsiou, E.; Austin, P.J.; Lindsay, M.A.; Gibeon, D.S.; Adcock, I.M.; Chung, K.F. Role of non-coding RNAs in maintaining primary airway smooth muscle cells. Respir. Res. 2014, 15, 58. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.R.; Lin, J.; Ye, Y.Q.; Oba, T.; Gentile, E.; Lian, J.; Wang, J.; Zhao, Y.; Gu, J.; Wistuba, I.I.; et al. Serum MicroRNA-150 Predicts Prognosis for Early-Stage Non-Small Cell Lung Cancer and Promotes Tumor Cell Proliferation by Targeting Tumor Suppressor Gene SRCIN1. Clin. Pharmacol. Ther. 2018, 103, 1061–1073. [Google Scholar] [CrossRef]
- Ghisi, M.; Corradin, A.; Basso, K.; Frasson, C.; Serafin, V.; Mukherjee, S.; Mussolin, L.; Ruggero, K.; Bonanno, L.; Guffanti, A.; et al. Modulation of microRNA expression in human T-cell development: Targeting of NOTCH3 by miR-150. Blood 2011, 117, 7053–7062. [Google Scholar] [CrossRef]
- George, J.; Lim, J.S.; Jang, S.J.; Cun, Y.P.; Ozretic, L.; Kong, G.; Leenders, F.; Lu, X.; Fernandez-Cuesta, L.; Bosco, G.; et al. Comprehensive genomic profiles of small cell lung cancer. Nature 2015, 524, 47–53. [Google Scholar] [CrossRef]
- Sun, Q.; Wang, Y.Z.; Fan, J.Y.; Li, Z.H.; Zhang, J.H.; Wang, L.J.; Fan, X.K.; Ji, M.M.; Zhu, M.; Dai, J.C.; et al. Association of expression quantitative trait loci for long noncoding RNAs with lung cancer risk in Asians. Mol. Carcinog. 2019, 58, 1303–1313. [Google Scholar] [CrossRef]
- Siddiqi, S.; Matushansky, I. Piwis and piwi-interacting RNAs in the epigenetics of cancer. J. Cell. Biochem. 2012, 113, 373–380. [Google Scholar] [CrossRef]
- Fu, A.; Jacobs, D.I.; Zhu, Y. Epigenome-wide analysis of piRNAs in gene-specific DNA methylation. RNA Biol. 2014, 11, 1301–1312. [Google Scholar] [CrossRef] [Green Version]
- Huang, G.; Hu, H.; Xue, X.; Shen, S.; Gao, E.; Guo, G.; Shen, X.; Zhang, X. Altered expression of piRNAs and their relation with clinicopathologic features of breast cancer. Clin. Transl. Oncol. 2013, 15, 563–568. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, D.I.; Qin, Q.; Lerro, M.C.; Fu, A.; Dubrow, R.; Claus, E.B.; DeWan, A.T.; Wang, G.L.; Lin, H.F.; Zhu, Y. PIWI-Interacting RNAs in Gliomagenesis: Evidence from Post-GWAS and Functional Analyses. Cancer Epidemiol. Biomark. Prev. 2016, 25, 1073–1080. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rupaimoole, R.; Calin, G.A.; Lopez-Berestein, G.; Sood, A.K. miRNA Deregulation in Cancer Cells and the Tumor Microenvironment. Cancer Discov. 2016, 6, 235–246. [Google Scholar] [CrossRef] [Green Version]
- Guo, J.U.; Agarwal, V.; Guo, H.; Bartel, D.P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014, 15, 409. [Google Scholar] [CrossRef] [PubMed]
- Salzman, J.; Chen, R.E.; Olsen, M.N.; Wang, P.L.; Brown, P.O. Cell-type specific features of circular RNA expression. PLoS Genet. 2013, 9, e1003777. [Google Scholar] [CrossRef]
- Bachmayr-Heyda, A.; Reiner, A.T.; Auer, K.; Sukhbaatar, N.; Aust, S.; Bachleitner-Hofmann, T.; Mesteri, I.; Grunt, T.W.; Zeillinger, R.; Pils, D. Correlation of circular RNA abundance with proliferation--exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci. Rep. 2015, 5, 8057. [Google Scholar] [CrossRef] [PubMed]
- Kadakkuzha, B.M.; Liu, X.A.; McCrate, J.; Shankar, G.; Rizzo, V.; Afinogenova, A.; Young, B.; Fallahi, M.; Carvalloza, A.C.; Raveendra, B.; et al. Transcriptome analyses of adult mouse brain reveal enrichment of lncRNAs in specific brain regions and neuronal populations. Front. Cell. Neurosci. 2015, 9, 63. [Google Scholar] [CrossRef] [Green Version]
- Rybak-Wolf, A.; Stottmeister, C.; Glazar, P.; Jens, M.; Pino, N.; Giusti, S.; Hanan, M.; Behm, M.; Bartok, O.; Ashwal-Fluss, R.; et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol. Cell 2015, 58, 870–885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- You, X.; Vlatkovic, I.; Babic, A.; Will, T.; Epstein, I.; Tushev, G.; Akbalik, G.; Wang, M.; Glock, C.; Quedenau, C.; et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat. Neurosci. 2015, 18, 603–610. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis, G.M.; Haas, M.A.; Pocock, R. MicroRNAs: Not “Fine-Tuners” but Key Regulators of Neuronal Development and Function. Front. Neurol. 2015, 6, 245. [Google Scholar] [CrossRef] [Green Version]
- Ng, S.Y.; Johnson, R.; Stanton, L.W. Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J. 2012, 31, 522–533. [Google Scholar] [CrossRef] [Green Version]
- Schratt, G. microRNAs at the synapse. Nat. Rev. Neurosci. 2009, 10, 842–849. [Google Scholar] [CrossRef]
- Junn, E.; Lee, K.W.; Jeong, B.S.; Chan, T.W.; Im, J.Y.; Mouradian, M.M. Repression of alpha-synuclein expression and toxicity by microRNA-7. Proc. Natl. Acad. Sci. USA 2009, 106, 13052–13057. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bak, M.; Silahtaroglu, A.; Moller, M.; Christensen, M.; Rath, M.F.; Skryabin, B.; Tommerup, N.; Kauppinen, S. MicroRNA expression in the adult mouse central nervous system. RNA 2008, 14, 432–444. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elstner, M.; Morris, C.M.; Heim, K.; Lichtner, P.; Bender, A.; Mehta, D.; Schulte, C.; Sharma, M.; Hudson, G.; Goldwurm, S.; et al. Single-Cell Expression Profiling of Dopaminergic Neurons Combined with Association Analysis Identifies Pyridoxal Kinase as Parkinson’s Disease Gene. Ann. Neurol. 2009, 66, 792–798. [Google Scholar] [CrossRef]
- Giraldez, T.; Dominguez, J.; Alvarez de la Rosa, D. ENaC in the brain--future perspectives and pharmacological implications. Curr. Mol. Pharmacol. 2013, 6, 44–49. [Google Scholar] [CrossRef]
- Mortezaei, Z.; Lanjanian, H.; Masoudi-Nejad, A. Candidate novel long noncoding RNAs, MicroRNAs and putative drugs for Parkinson’s disease using a robust and efficient genome-wide association study. Genomics 2017, 109, 158–164. [Google Scholar] [CrossRef]
- Liu, Y.; Chen, Y.Y.; Liu, H.; Yao, C.J.; Zhu, X.X.; Chen, D.J.; Yang, J.; Lu, Y.J.; Cao, J.Y. Association between ubiquitin carboxy-terminal hydrolase-L1 S18Y variant and risk of Parkinson’s disease: The impact of ethnicity and onset age. Neurol. Sci. 2015, 36, 179–188. [Google Scholar] [CrossRef]
- Lukiw, W.J. Circular RNA (circRNA) in Alzheimer’s disease (AD). Front. Genet. 2013, 4, 307. [Google Scholar] [CrossRef] [Green Version]
- Zhang, S.; Zhu, D.; Li, H.; Li, H.; Feng, C.; Zhang, W. Characterization of circRNA-Associated-ceRNA Networks in a Senescence-Accelerated Mouse Prone 8 Brain. Mol. Ther. 2017, 25, 2053–2061. [Google Scholar] [CrossRef] [Green Version]
- Wei, C.W.; Luo, T.; Zou, S.S.; Wu, A.S. The Role of Long Noncoding RNAs in Central Nervous System and Neurodegenerative Diseases. Front. Behav. Neurosci. 2018, 12, 175. [Google Scholar] [CrossRef]
- Massone, S.; Vassallo, I.; Fiorino, G.; Castelnuovo, M.; Barbieri, F.; Borghi, R.; Tabaton, M.; Robello, M.; Gatta, E.; Russo, C.; et al. 17A, a novel non-coding RNA, regulates GABA B alternative splicing and signaling in response to inflammatory stimuli and in Alzheimer disease. Neurobiol. Dis. 2011, 41, 308–317. [Google Scholar] [CrossRef] [PubMed]
- Hebert, S.S.; De Strooper, B. Alterations of the microRNA network cause neurodegenerative disease. Trends Neurosci. 2009, 32, 199–206. [Google Scholar] [CrossRef]
- Vilardo, E.; Barbato, C.; Ciotti, M.; Cogoni, C.; Ruberti, F. MicroRNA-101 regulates amyloid precursor protein expression in hippocampal neurons. J. Biol. Chem. 2010, 285, 18344–18351. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, Z.J.; Xue, W.W.; Tao, L.; Zhu, F. Identification of Key Long Non-Coding RNAs in the Pathology of Alzheimer’s Disease and their Functions Based on Genome-Wide Associations Study, Microarray, and RNA-seq Data. J. Alzheimers Dis. 2019, 68, 339–355. [Google Scholar] [CrossRef]
- Kataoka, M.; Huang, Z.P.; Wang, D.Z. Build a braveheart: The missing linc (RNA). Circ. Res. 2013, 112, 1532–1534. [Google Scholar] [CrossRef] [Green Version]
- Hou, J.; Long, H.; Zhou, C.; Zheng, S.; Wu, H.; Guo, T.; Wu, Q.; Zhong, T.; Wang, T. Long noncoding RNA Braveheart promotes cardiogenic differentiation of mesenchymal stem cells in vitro. Stem Cell Res. Ther. 2017, 8, 4. [Google Scholar] [CrossRef] [Green Version]
- Viereck, J.; Kumarswamy, R.; Foinquinos, A.; Xiao, K.; Avramopoulos, P.; Kunz, M.; Dittrich, M.; Maetzig, T.; Zimmer, K.; Remke, J.; et al. Long noncoding RNA Chast promotes cardiac remodeling. Sci. Transl. Med. 2016, 8, 326ra322. [Google Scholar] [CrossRef] [PubMed]
- Stępień, E.; Costa, M.C.; Kurc, S.; Drozdz, A.; Cortez-Dias, N.; Enguita, F.J. The circulating non-coding RNA landscape for biomarker research: Lessons and prospects from cardiovascular diseases. Acta Pharmacol. Sin. 2018, 39, 1085–1099. [Google Scholar] [CrossRef]
- Jarinova, O.; Stewart, A.F.R.; Roberts, R.; Wells, G.; Lau, P.; Naing, T.; Buerki, C.; McLean, B.W.; Cook, R.C.; Parker, J.S.; et al. Functional Analysis of the Chromosome 9p21.3 Coronary Artery Disease Risk Locus. Arterioscler. Thromb. Vasc. Biol. 2009, 29, 1671–U1670. [Google Scholar] [CrossRef] [Green Version]
- Arslan, S.; Berkan, O.; Lalem, T.; Ozbilum, N.; Goksel, S.; Korkmaz, O.; Cetin, N.; Devaux, Y.; Cardiolinc, N. Long non-coding RNAs in the atherosclerotic plaque. Atherosclerosis 2017, 266, 176–181. [Google Scholar] [CrossRef]
- Nikpay, M.; Beehler, K.; Valsesia, A.; Hager, J.; Harper, M.E.; Dent, R.; McPherson, R. Genome-wide identification of circulating-miRNA expression quantitative trait loci reveals the role of severalmiRNAs in the regulation of cardiometabolic phenotypes. Cardiovasc. Res. 2019, 115, 1629–1645. [Google Scholar] [CrossRef]
- Li, Y.; Qiu, C.; Tu, J.; Geng, B.; Yang, J.; Jiang, T.; Cui, Q. HMDD v2.0: A database for experimentally supported human microRNA and disease associations. Nucleic Acids Res. 2014, 42, D1070–D1074. [Google Scholar] [CrossRef]
- Xu, S.; Kamato, D.; Little, P.J.; Nakagawa, S.; Pelisek, J.; Jin, Z.G. Targeting epigenetics and non-coding RNAs in atherosclerosis: from mechanisms to therapeutics. Pharmacol. Ther. 2019, 196, 15–43. [Google Scholar] [CrossRef] [PubMed]
- Eken, S.M.; Jin, H.; Chernogubova, E.; Li, Y.; Simon, N.; Sun, C.; Korzunowicz, G.; Busch, A.; Backlund, A.; Osterholm, C.; et al. MicroRNA-210 Enhances Fibrous Cap Stability in Advanced Atherosclerotic Lesions. Circ. Res. 2017, 120, 633–644. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Li, D.Y.; Chernogubova, E.; Sun, C.; Busch, A.; Eken, S.M.; Saliba-Gustafsson, P.; Winter, H.; Winski, G.; Raaz, U.; et al. Local Delivery of miR-21 Stabilizes Fibrous Caps in Vulnerable Atherosclerotic Lesions. Mol. Ther. 2018, 26, 1040–1055. [Google Scholar] [CrossRef] [Green Version]
- Bye, A.; Rosjo, H.; Nauman, J.; Silva, G.J.; Follestad, T.; Omland, T.; Wisloff, U. Circulating microRNAs predict future fatal myocardial infarction in healthy individuals—The HUNT study. J. Mol. Cell. Cardiol. 2016, 97, 162–168. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsumoto, S.; Sakata, Y.; Suna, S.; Nakatani, D.; Usami, M.; Hara, M.; Kitamura, T.; Hamasaki, T.; Nanto, S.; Kawahara, Y.; et al. Circulating p53-responsive microRNAs are predictive indicators of heart failure after acute myocardial infarction. Circ. Res. 2013, 113, 322–326. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.O.; Dong, R.; Zhang, Y.; Zhang, J.L.; Luo, Z.; Zhang, J.; Chen, L.L.; Yang, L. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 2016, 26, 1277–1287. [Google Scholar] [CrossRef] [Green Version]
- Yan, Y.; Zhang, B.; Liu, N.; Qi, C.; Xiao, Y.; Tian, X.; Li, T.; Liu, B. Circulating Long Noncoding RNA UCA1 as a Novel Biomarker of Acute Myocardial Infarction. Biomed. Res. Int. 2016, 2016, 8079372. [Google Scholar] [CrossRef] [Green Version]
- Du, W.W.; Zhang, C.; Yang, W.N.; Yong, T.Q.; Awan, F.M.; Yang, B.B. Identifying and Characterizing circRNA-Protein Interaction. Theranostics 2017, 7, 4183–4191. [Google Scholar] [CrossRef]
- Grieco, F.A.; Schiavo, A.A.; Brozzi, F.; Juan-Mateu, J.; Bugliani, M.; Marchetti, P.; Eizirik, D. The microRNAs miR-211-5p and miR-204-5p modulate ER stress in human beta cells. J. Mol. Endocrinol. 2019, 63, 139–149. [Google Scholar] [CrossRef]
- Zhao, X.; Mohan, R.; Ozcan, S.; Tang, X. MicroRNA-30d induces insulin transcription factor MafA and insulin production by targeting mitogen-activated protein 4 kinase 4 (MAP4K4) in pancreatic beta-cells. J. Biol. Chem. 2012, 287, 31155–31164. [Google Scholar] [CrossRef] [Green Version]
- El Ouaamari, A.; Baroukh, N.; Martens, G.A.; Lebrun, P.; Pipeleers, D.; van Obberghen, E. miR-375 targets 3′-phosphoinositide-dependent protein kinase-1 and regulates glucose-induced biological responses in pancreatic beta-cells. Diabetes 2008, 57, 2708–2717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, S.; Zhang, M.; Sun, F.; Ren, L.; He, X.; Hua, J.; Peng, S. miR-375 controls porcine pancreatic stem cell fate by targeting 3-phosphoinositide-dependent protein kinase-1 (Pdk1). Cell. Prolif. 2016, 49, 395–406. [Google Scholar] [CrossRef]
- Poy, M.N.; Eliasson, L.; Krutzfeldt, J.; Kuwajima, S.; Ma, X.; Macdonald, P.E.; Pfeffer, S.; Tuschl, T.; Rajewsky, N.; Rorsman, P.; et al. A pancreatic islet-specific microRNA regulates insulin secretion. Nature 2004, 432, 226–230. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Gu, P.; Shi, W.; Li, J.; Hao, Q.; Cao, X.; Lu, Q.; Zeng, Y. MicroRNA-29a induces insulin resistance by targeting PPARdelta in skeletal muscle cells. Int. J. Mol. Med. 2016, 37, 931–938. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, W.M.; Jeong, H.J.; Park, S.Y.; Lee, W. Induction of miR-29a by saturated fatty acids impairs insulin signaling and glucose uptake through translational repression of IRS-1 in myocytes. FEBS Lett. 2014, 588, 2170–2176. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patel, V.; Hajarnis, S.; Williams, D.; Hunter, R.; Huynh, D.; Igarashi, P. MicroRNAs regulate renal tubule maturation through modulation of Pkd1. J. Am. Soc. Nephrol. 2012, 23, 1941–1948. [Google Scholar] [CrossRef] [Green Version]
- Wang, G.; Kwan, B.C.; Lai, F.M.; Chow, K.M.; Li, P.K.; Szeto, C.C. Elevated levels of miR-146a and miR-155 in kidney biopsy and urine from patients with IgA nephropathy. Dis. Markers 2011, 30, 171–179. [Google Scholar] [CrossRef]
- Luo, X.B.; Yang, W.L.; Ye, D.Q.; Cui, H.J.; Zhang, Y.; Hirankarn, N.; Qian, X.X.; Tang, Y.J.; Lau, Y.L.; de Vries, N.; et al. A Functional Variant in MicroRNA-146a Promoter Modulates Its Expression and Confers Disease Risk for Systemic Lupus Erythematosus. Plos Genet. 2011, 7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xue, Z.X.; Cui, C.J.; Liao, Z.J.; Xia, S.W.; Zhang, P.J.; Qin, J.L.; Guo, Q.; Chen, S.; Fu, Q.; Yin, Z.H.; et al. Identification of LncRNA Linc00513 Containing Lupus-Associated Genetic Variants as a Novel Regulator of Interferon Signaling Pathway. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Barabasi, A.L.; Gulbahce, N.; Loscalzo, J. Network medicine: A network-based approach to human disease. Nat. Rev. Genet. 2011, 12, 56–68. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Y.; Xu, G.; Yang, Y.T.; Xu, Z.; Chen, X.; Shi, B.; Xie, D.; Lu, Z.J.; Wang, P. POSTAR2: Deciphering the post-transcriptional regulatory logics. Nucleic Acids Res. 2019, 47, D203–D211. [Google Scholar] [CrossRef] [Green Version]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, D68–D73. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quek, X.C.; Thomson, D.W.; Maag, J.L.; Bartonicek, N.; Signal, B.; Clark, M.B.; Gloss, B.S.; Dinger, M.E. lncRNAdb v2.0: Expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res. 2015, 43, D168–D173. [Google Scholar] [CrossRef] [PubMed]
- Glazar, P.; Papavasileiou, P.; Rajewsky, N. circBase: A database for circular RNAs. RNA 2014, 20, 1666–1670. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sai Lakshmi, S.; Agrawal, S. piRNABank: A web resource on classified and clustered Piwi-interacting RNAs. Nucleic Acids Res. 2008, 36, D173–D177. [Google Scholar] [CrossRef] [PubMed]
- Teng, X.; Chen, X.; Xue, H.; Tang, Y.; Zhang, P.; Kang, Q.; Hao, Y.; Chen, R.; Zhao, Y.; He, S. NPInter v4.0: An integrated database of ncRNA interactions. Nucleic Acids Res. 2020, 48, D160–D165. [Google Scholar] [CrossRef]
- Wang, W.J.; Wang, Y.M.; Hu, Y.; Lin, Q.; Chen, R.; Liu, H.; Cao, W.Z.; Zhu, H.F.; Tong, C.; Li, L.; et al. HDncRNA: A comprehensive database of non-coding RNAs associated with heart diseases. Database J. Biol. Databases Curation 2018, 2018. [Google Scholar] [CrossRef] [Green Version]
- Huang, Z.; Shi, J.; Gao, Y.; Cui, C.; Zhang, S.; Li, J.; Zhou, Y.; Cui, Q. HMDD v3.0: A database for experimentally supported human microRNA-disease associations. Nucleic Acids Res. 2019, 47, D1013–D1017. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, Q.; Wang, Y.; Hao, Y.; Juan, L.; Teng, M.; Zhang, X.; Li, M.; Wang, G.; Liu, Y. miR2Disease: A manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 2009, 37, D98–D104. [Google Scholar] [CrossRef] [Green Version]
- Ruepp, A.; Kowarsch, A.; Theis, F. PhenomiR: MicroRNAs in human diseases and biological processes. Methods Mol. Biol. 2012, 822, 249–260. [Google Scholar] [CrossRef] [PubMed]
- Sarver, A.L.; Sarver, A.E.; Yuan, C.; Subramanian, S. OMCD: OncomiR Cancer Database. BMC Cancer 2018, 18, 1223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, B.; Ding, Q.; Han, H.; Wu, D. miRCancer: A microRNA-cancer association database constructed by text mining on literature. Bioinformatics 2013, 29, 638–644. [Google Scholar] [CrossRef]
- Bao, Z.; Yang, Z.; Huang, Z.; Zhou, Y.; Cui, Q.; Dong, D. LncRNADisease 2.0: An updated database of long non-coding RNA-associated diseases. Nucleic Acids Res. 2019, 47, D1034–D1037. [Google Scholar] [CrossRef]
- Gao, Y.; Wang, P.; Wang, Y.; Ma, X.; Zhi, H.; Zhou, D.; Li, X.; Fang, Y.; Shen, W.; Xu, Y.; et al. Lnc2Cancer v2.0: Updated database of experimentally supported long non-coding RNAs in human cancers. Nucleic Acids Res. 2019, 47, D1028–D1033. [Google Scholar] [CrossRef]
- Yao, D.; Zhang, L.; Zheng, M.; Sun, X.; Lu, Y.; Liu, P. Circ2Disease: A manually curated database of experimentally validated circRNAs in human disease. Sci. Rep. 2018, 8, 11018. [Google Scholar] [CrossRef] [Green Version]
- Xia, S.; Feng, J.; Chen, K.; Ma, Y.; Gong, J.; Cai, F.; Jin, Y.; Gao, Y.; Xia, L.; Chang, H.; et al. CSCD: A database for cancer-specific circular RNAs. Nucleic Acids Res. 2018, 46, D925–D929. [Google Scholar] [CrossRef] [Green Version]
- Ghosal, S.; Das, S.; Sen, R.; Basak, P.; Chakrabarti, J. Circ2Traits: A comprehensive database for circular RNA potentially associated with disease and traits. Front. Genet. 2013, 4, 283. [Google Scholar] [CrossRef] [Green Version]
- Muhammad, A.; Waheed, R.; Khan, N.A.; Jiang, H.; Song, X. piRDisease v1.0: A manually curated database for piRNA associated diseases. Database J. Biol. Databases Curation 2019, 2019. [Google Scholar] [CrossRef] [Green Version]
- Hindorff, L.A.; Sethupathy, P.; Junkins, H.A.; Ramos, E.M.; Mehta, J.P.; Collins, F.S.; Manolio, T.A. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 2009, 106, 9362–9367. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Xue, Y.W.; Amin, M.T.; Yang, Y.B.; Yang, J.J.; Zhang, W.; Yang, W.Q.; Niu, X.H.; Zhang, H.Y.; Gong, J. ncRNA-eQTL: A database to systematically evaluate the effects of SNPs on non-coding RNA expression across cancer types. Nucleic Acids Res. 2020, 48, D956–D963. [Google Scholar] [CrossRef]
- Gao, Y.; Li, X.; Shang, S.P.; Guo, S.; Wang, P.; Sun, D.L.; Gan, J.; Sun, J.; Zhang, Y.K.; Wang, J.W.; et al. LincSNP 3.0: An updated database for linking functional variants to human long non-coding RNAs, circular RNAs and their regulatory elements. Nucleic Acids Res. 2021, 49, D1244–D1250. [Google Scholar] [CrossRef] [PubMed]
- Pinero, J.; Ramirez-Anguita, J.M.; Sauch-Pitarch, J.; Ronzano, F.; Centeno, E.; Sanz, F.; Furlong, L.I. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 2020, 48, D845–D855. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Ma, R.; Ma, W.; Chen, J.; Yang, J.; Xi, Y.; Cui, Q. LncDisease: A sequence based bioinformatics tool for predicting lncRNA-disease associations. Nucleic Acids Res. 2016, 44, e90. [Google Scholar] [CrossRef] [PubMed]
- Kristen, A.V.; Ajroud-Driss, S.; Conceicao, I.; Gorevic, P.; Kyriakides, T.; Obici, L. Patisiran, an RNAi therapeutic for the treatment of hereditary transthyretin-mediated amyloidosis. Neurodegener. Dis. Manag. 2019, 9, 5–23. [Google Scholar] [CrossRef] [Green Version]
- Yang, J. Patisiran for the treatment of hereditary transthyretin-mediated amyloidosis. Expert Rev. Clin. Pharmacol. 2019, 12, 95–99. [Google Scholar] [CrossRef]
- Titze-de-Almeida, S.S.; Soto-Sanchez, C.; Fernandez, E.; Koprich, J.B.; Brotchie, J.M.; Titze-de-Almeida, R. The Promise and Challenges of Developing miRNA-Based Therapeutics for Parkinson’s Disease. Cells 2020, 9, 841. [Google Scholar] [CrossRef] [Green Version]
- Chakraborty, C.; Sharma, A.R.; Sharma, G.; Doss, C.G.P.; Lee, S.S. Therapeutic miRNA and siRNA: Moving from Bench to Clinic as Next Generation Medicine. Mol. Ther. Nucleic Acids 2017, 8, 132–143. [Google Scholar] [CrossRef] [Green Version]
- Hanna, J.; Hossain, G.S.; Kocerha, J. The Potential for microRNA Therapeutics and Clinical Research. Front Genet. 2019, 10, 478. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rupaimoole, R.; Slack, F.J. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017, 16, 203–221. [Google Scholar] [CrossRef] [PubMed]
- Setten, R.L.; Rossi, J.J.; Han, S.P. The current state and future directions of RNAi-based therapeutics. Nat. Rev. Drug Discov. 2019, 18, 421–446. [Google Scholar] [CrossRef] [PubMed]
- Lung, B.; Zemann, A.; Madej, M.J.; Schuelke, M.; Techritz, S.; Ruf, S.; Bock, R.; Huttenhofer, A. Identification of small non-coding RNAs from mitochondria and chloroplasts. Nucleic Acids Res. 2006, 34, 3842–3852. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olavarria, J.V.; Burzio, V.A.; Borgna, V.; Lobos-Gonzalez, L.; Araya, M.; Guevara, F.; Villota, C.; Burzio, L.O. Long Noncoding Mitochondrial RNAs (LncmtRNAs) as Targets for Cancer Therapy; InTech: London, UK, 2018. [Google Scholar] [CrossRef] [Green Version]
- Burzio, V.A.; Villota, C.; Villegas, J.; Landerer, E.; Boccardo, E.; Villa, L.L.; Martinez, R.; Lopez, C.; Gaete, F.; Toro, V.; et al. Expression of a family of noncoding mitochondrial RNAs distinguishes normal from cancer cells. Proc. Natl. Acad. Sci. USA 2009, 106, 9430–9434. [Google Scholar] [CrossRef] [Green Version]
- Tamtaji, O.R.; Behnam, M.; Pourattar, M.A.; Hamblin, M.R.; Mahjoubin-Tehran, M.; Mirzaei, H.; Asemi, Z. PIWI-interacting RNAs and PIWI proteins in glioma: Molecular pathogenesis and role as biomarkers. Cell. Commun. Signal. 2020, 18, 168. [Google Scholar] [CrossRef]
- Boeckel, N.; Dimmeler, S.; Zeiher, A.M.; Jaé, N. Circular RNA for the Diagnosis and Treatment of Cardiovascular Diseases. European Patent EP3054017A1, 10 August 2016. [Google Scholar]
- Dimmeler, S.; Jaé, N.; Boeckel, N.; Zeiher, A.M. Circular RNA for the Diagnosis of Cardiovascular and Inflammatory Diseases. WIPO. WO2016124655A1, 11 August 2016. [Google Scholar]

| ncRNA Species | General Functions and Properties |
|---|---|
| miRNA | Fine regulators of expression |
| Inhibition of gene transcription | |
| lncRNA | Interference in polymerase activity |
| Antisense RNA sequence matching | |
| Inhibition of histone acetyltransferase activity and repression of transcription | |
| Recruitment of transcriptional regulators | |
| Upregulate translation without altering mRNA levels | |
| Chromatin remodeling | |
| Enhancer | |
| Histone modifications and DNA methylation | |
| piRNA | Block transposable elements |
| Regulators of transcription acting primarily on transposable element sequences | |
| Protection against external nucleic acids | |
| circRNA | Monitor the production of linear transcripts of RNA and microRNAs |
| Sponge-like regulatory system decreasing miRNA concentration in the cytoplasm | |
| Can bind RNA-binding proteins | |
| tRNA | Protein synthesis. |
| Post- or pre-transcriptional regulators of gene expression | |
| rRNA | Ribosomal subunits (5S, 16S, 23S, 5.8S, 18S and 28S) |
| Protein synthesis |
| LncRNA Category | Definition | Example | |
|---|---|---|---|
| Genic | Genic intronic lncRNA | LncRNA overlapping a protein-coding intron at one or more nucleotides |  |
| Genic exonic lncRNA | LncRNA overlapping a protein-coding exon at one or more nucleotides |  | |
| Nested lncRNA | LncRNA genes contained entirely within protein-coding transcripts |  | |
| Overlapping lncRNA | LncRNA genes sharing one or more nucleotides with coding genes |  | |
| Intergenic | LincRNA | Long-intergenic non-coding RNA. No overlapping, no close to a protein-coding transcript |  |
| Same strand | LincRNA within 50 kb of and transcribed from the same strand and in the same direction as the nearest protein-coding transcript |  | |
| Convergent | LincRNA within 50 kb of and transcribed head-to-head with the nearest protein-coding transcript |  | |
| Divergent | LincRNA within 50 kb of and transcribed tail-to-tail with the nearest protein-coding transcript |  | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
López-Jiménez, E.; Andrés-León, E. The Implications of ncRNAs in the Development of Human Diseases. Non-Coding RNA 2021, 7, 17. https://0-doi-org.brum.beds.ac.uk/10.3390/ncrna7010017
López-Jiménez E, Andrés-León E. The Implications of ncRNAs in the Development of Human Diseases. Non-Coding RNA. 2021; 7(1):17. https://0-doi-org.brum.beds.ac.uk/10.3390/ncrna7010017
Chicago/Turabian StyleLópez-Jiménez, Elena, and Eduardo Andrés-León. 2021. "The Implications of ncRNAs in the Development of Human Diseases" Non-Coding RNA 7, no. 1: 17. https://0-doi-org.brum.beds.ac.uk/10.3390/ncrna7010017







