Contemporary Distribution, Estimated Age, and Prehistoric Migrations of Old World Monkey Retroviruses
Abstract
:1. Introduction
2. Materials and Methods
2.1. Survey Methodology
2.2. Identifying Endogenous Retrovirus Sequences in OWM Genomes
2.3. Analysis of Proviral Sequences
3. Results
3.1. Exogenous Retroviruses in OWM Species
- Sampling is not systematic; some species remain untested because they may be difficult to reach.
- Sampling may not be optimal: it is done at the wrong age (for instance SIV is a sexually transmitted infection with juveniles normally being negative for the virus), the wrong type of sample is taken, or the viral load is below the detection level.
- The distinction between an exogenous and endogenous retrovirus is not always clear (e.g., murine leukemia virus, MuLV, feline leukemia virus, FeLV, and koala retrovirus, KoRV, have both infectious and endogenous variants). Moreover, every type of retrovirus has the capacity to enter the mammalian germ line, so Mendelian inheritance is not a distinguishing characteristic [42,43,44].
3.1.1. Simian Foamy Virus (SFV)
3.1.2. Simian Immunodeficiency Virus (SIV)
3.1.3. Simian Type D retrovirus (SRV)
3.1.4. Simian T-Lymphotropic Virus (STLV)
3.2. Endogenous Retrovirus Integrations Predating OWM Speciation
3.3. Endogenous Retroviruses Specific for OWM
- Not all infected species may still roam the earth
- Not all infected species may contain germ line integrations
- Not all species have had their genomes sequenced
- Not all proviral sequences are likely due to bona fide viral infection and integration; they may for instance be acquired by hybridization between species
- Proviruses may have been completely or partially lost from the germ line, making identification challenging
- The quality of a genome assembly may be insufficient for provirus detection
- The distinction between endogenous and exogenous retroviruses is not always clear (see above, and the comment in [129])
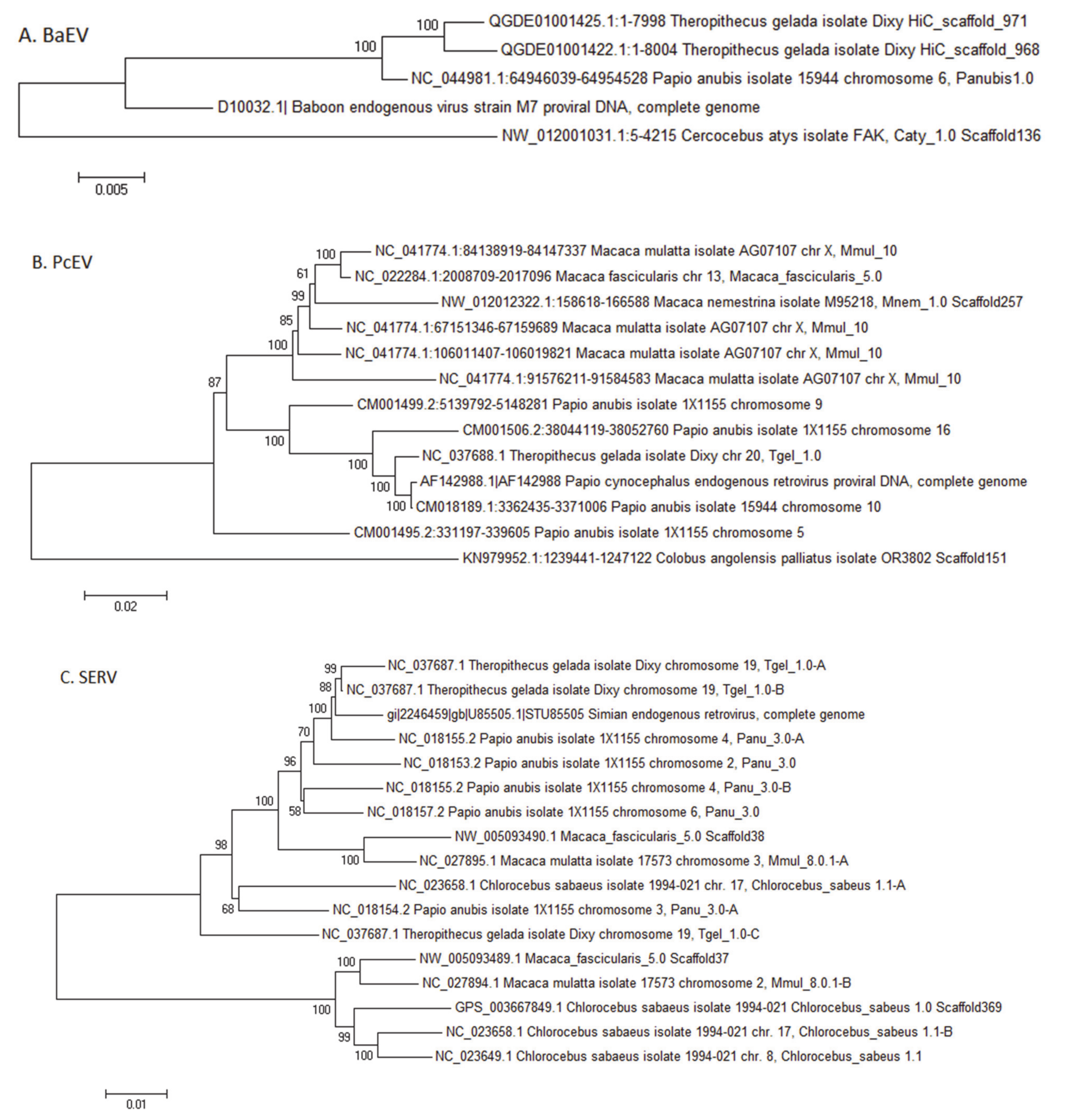
3.3.1. Baboon Endogenous Virus (BaEV)
3.3.2. Papio Cynocephalus Endogenous Virus (PcEV)
3.3.3. Simian Endogenous Retrovirus (SERV)
3.3.4. Simian Endogenous Retrovirus-K1 (SERV-K1)/RhERV-K
4. Discussion
5. Conclusions
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
Appendix B

References
- Elton, S. Environmental correlates of the cercopithecoid radiations. Folia Primatol. 2007, 78, 344–364. [Google Scholar] [CrossRef] [PubMed]
- Steiper, M.E.; Young, N.M. Primate molecular divergence dates. Mol. Phylogenet. Evol. 2006, 41, 384–394. [Google Scholar] [CrossRef] [PubMed]
- Stevens, N.J.; Seiffert, E.R.; O’Connor, P.M.; Roberts, E.M.; Schmitz, M.D.; Krause, C.; Gorscak, E.; Ngasala, S.; Hieronymus, T.L.; Temu, J. Palaeontological evidence for an Oligocene divergence between Old World monkeys and apes. Nature 2013, 497, 611–614. [Google Scholar] [CrossRef]
- Raaum, R.L.; Sterner, K.N.; Noviello, C.M.; Stewart, C.-B.; Disotell, T.R. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J. Hum. Evol. 2005, 48, 237–257. [Google Scholar] [CrossRef] [PubMed]
- Hayward, A. Origin of the retroviruses: When, where, and how? Curr. Opin. Virol. 2017, 25, 23–27. [Google Scholar] [CrossRef] [PubMed]
- Lerat, E.; Capy, P. Retrotransposons and retroviruses: Analysis of the envelope gene. Mol. Biol. Evol. 1999, 16, 1198–1207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koonin, E.V.; Dolja, V.V. Virus world as an evolutionary network of viruses and capsidless selfish elements. Microbiol. Mol. Biol. Rev. 2014, 78, 278–303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cordonnier, A.; Casella, J.F.; Heidmann, T. Isolation of novel human endogenous retrovirus-like elements with foamy virus-related pol sequence. J. Virol. 1995, 69, 5890–5897. [Google Scholar] [CrossRef] [Green Version]
- Zhu, T.; Korber, B.T.; Nahmias, A.J.; Hooper, E.; Sharp, P.M.; Ho, D.D. An African HIV-1 sequence from 1959 and implications for the origin of the epidemic. Nature 1998, 391, 594–597. [Google Scholar] [CrossRef]
- Worobey, M.; Gemmel, M.; Teuwen, D.E.; Haselkorn, T.; Kunstman, K.; Bunce, M.; Muyembe, J.-J.; Kabongo, J.-M.M.; Kalengayi, R.M.; Van Marck, E.; et al. Direct evidence of extensive diversity of HIV-1 in Kinshasa by 1960. Nature 2008, 455, 661–664. [Google Scholar] [CrossRef] [Green Version]
- Faria, N.R.; Rambaut, A.; Suchard, M.A.; Baele, G.; Bedford, T.; Ward, M.J.; Tatem, A.J.; Sousa, J.D.; Arinaminpathy, N.; Pépin, J.; et al. HIV epidemiology. The early spread and epidemic ignition of HIV-1 in human populations. Science 2014, 346, 56–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hirsch, V.M.; Olmsted, R.A.; Murphey-Corb, M.; Purcell, R.H.; Johnson, P.R. An African primate lentivirus (SIVsmclosely related to HIV-2. Nature 1989, 339, 389–392. [Google Scholar] [CrossRef] [PubMed]
- Plantier, J.-C.; Leoz, M.; Dickerson, J.E.; De Oliveira, F.; Cordonnier, F.; Lemée, V.; Damond, F.; Robertson, D.L.; Simon, F. A new human immunodeficiency virus derived from gorillas. Nat. Med. 2009, 15, 871–872. [Google Scholar] [CrossRef]
- D’Arc, M.; Ayouba, A.; Esteban, A.; Learn, G.H.; Boué, V.; Liegeois, F.; Etienne, L.; Tagg, N.; Leendertz, F.H.; Boesch, C.; et al. Origin of the HIV-1 group O epidemic in western lowland gorillas. Proc. Natl. Acad. Sci. USA 2015, 112, E1343–E1352. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, F.; Bailes, E.; Robertson, D.L.; Chen, Y.; Rodenburg, C.M.; Michael, S.F.; Cummins, L.B.; Arthur, L.O.; Peeters, M.; Shaw, G.M.; et al. Origin of HIV-1 in the chimpanzee Pan troglodytes troglodytes. Nature 1999, 397, 436–441. [Google Scholar] [CrossRef]
- Huet, T.; Cheynier, R.; Meyerhans, A.; Roelants, G.; Wain-Hobson, S. Genetic organization of a chimpanzee lentivirus related to HIV-1. Nature 1990, 345, 356–359. [Google Scholar] [CrossRef]
- Roques, P.; Robertson, D.L.; Souquière, S.; Apetrei, C.; Nerrienet, E.; Barré-Sinoussi, F.; Müller-Trutwin, M.; Simon, F. Phylogenetic characteristics of three new HIV-1 N strains and implications for the origin of group N. AIDS 2004, 18, 1371–1381. [Google Scholar] [CrossRef]
- Chen, Z.; Telfier, P.; Gettie, A.; Reed, P.; Zhang, L.; Ho, D.D.; Marx, P.A. Genetic characterization of new West African simian immunodeficiency virus SIVsm: Geographic clustering of household-derived SIV strains with human immunodeficiency virus type 2 subtypes and genetically diverse viruses from a single feral sooty mangabey troop. J. Virol. 1996, 70, 3617–3627. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Luckay, A.; Sodora, D.L.; Telfer, P.; Reed, P.; Gettie, A.; Kanu, J.M.; Sadek, R.F.; Yee, J.; Ho, D.D.; et al. Human immunodeficiency virus type 2 (HIV-2) seroprevalence and characterization of a distinct HIV-2 genetic subtype from the natural range of simian immunodeficiency virus-infected sooty mangabeys. J. Virol. 1997, 71, 3953–3960. [Google Scholar] [CrossRef] [Green Version]
- Santiago, M.L.; Range, F.; Keele, B.F.; Li, Y.; Bailes, E.; Bibollet-Ruche, F.; Fruteau, C.; Noë, R.; Peeters, M.; Brookfield, J.F.Y.; et al. Simian immunodeficiency virus infection in free-ranging sooty mangabeys (cercocebus atys atys) from the Taï Forest, Côte d’Ivoire: Implications for the origin of epidemic human immunodeficiency virus type 2. J. Virol. 2005, 79, 12515–12527. [Google Scholar] [CrossRef] [Green Version]
- Van Dooren, S.; Salemi, M.; Vandamme, A.-M. Dating the origin of the African human T-cell lymphotropic virus type-I (HTLV-I) subtypes. Mol. Biol. Evol. 2001, 18, 661–671. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Dooren, S.; Verschoor, E.J.; Fagrouch, Z.; Vandamme, A.-M. Phylogeny of primate T lymphotropic virus type 1 (PTLV-1) including various new Asian and African non-human primate strains. Infect. Genet. Evol. 2007, 7, 374–381. [Google Scholar] [CrossRef] [PubMed]
- Pecon-Slattery, J.; Franchini, G.; Gessain, A. Genomic evolution, patterns of global dissemination, and interspecies transmission of human and simian T-cell leukemia/lymphotropic viruses. Genome Res. 1999, 9, 525–540. [Google Scholar]
- Reid, M.J.; Switzer, W.M.; Schillaci, M.A.; Ragonnet-Cronin, M.; Joanisse, I.; Caminiti, K.; Lowenberger, C.; Galdikas, B.M.F.; Sandstrom, P.A.; Brooks, J.I. Detailed phylogenetic analysis of primate T-lymphotropic virus type 1 (PTLV-1) sequences from orangutans (Pongo pygmaeus) reveals new insights into the evolutionary history of PTLV-1 in Asia. Infect. Genet. Evol. 2016, 43, 434–450. [Google Scholar] [CrossRef]
- Hron, T.; Elleder, D.; Gifford, R.J. Deltaretroviruses have circulated since at least the Paleogene and infected a broad range of mammalian species. Retrovirology 2019, 16, 33. [Google Scholar] [CrossRef]
- Salemi, M.; Desmyter, J.; Vandamme, A.-M. Tempo and mode of human and simian T-lymphotropic virus (HTLV/STLV) evolution revealed by analyses of full-genome sequences. Mol. Biol. Evol. 2000, 17, 374–386. [Google Scholar] [CrossRef]
- Afonso, P.V.; Cassar, O.; Gessain, A. Molecular epidemiology, genetic variability and evolution of HTLV-1 with special emphasis on African genotypes. Retrovirology 2019, 16, 39. [Google Scholar] [CrossRef] [Green Version]
- Switzer, W.M.; Salemi, M.; Qari, S.H.; Jia, H.; Gray, R.R.; Katzourakis, A.; Marriott, S.J.; Pryor, K.N.; Wolfe, N.D.; Burke, D.S.; et al. Ancient, independent evolution and distinct molecular features of the novel human T-lymphotropic virus type 4. Retrovirology 2009, 6, 9. [Google Scholar] [CrossRef] [Green Version]
- Zheng, H.; Wolfe, N.D.; Sintasath, D.M.; Tamoufe, U.; Lebreton, M.; Djoko, C.F.; Diffo, J.L.D.; Pike, B.L.; Heneine, W.; Switzer, W.M. Emergence of a novel and highly divergent HTLV-3 in a primate hunter in Cameroon. Virology 2010, 401, 137–145. [Google Scholar] [CrossRef] [Green Version]
- Filippone, C.; Betsem, E.; Tortevoye, P.; Cassar, O.; Bassot, S.; Froment, A.; Fontanet, A.; Gessain, A. A Severe Bite from a Nonhuman Primate Is a Major Risk Factor for HTLV-1 Infection in Hunters From Central Africa. Clin. Infect. Dis. 2015, 60, 1667–1676. [Google Scholar] [CrossRef] [Green Version]
- Pinto-Santini, D.M.; Stenbak, C.R.; Linial, M.L. Foamy virus zoonotic infections. Retrovirology 2017, 14, 55. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heneine, W.; Schweizer, M.; Sandstrom, P.; Folks, T. Human infection with foamy viruses. Curr. Top Microbiol. Immunol. 2003, 277, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Mouinga-Ondémé, A.; Kazanji, M. Simian foamy virus in non-human primates and cross-species transmission to humans in Gabon: An emerging zoonotic disease in Central Africa? Viruses 2013, 5, 1536–1552. [Google Scholar] [CrossRef] [Green Version]
- Power, M.D.; Marx, P.A.; Bryant, M.L.; Gardner, M.B.; Barr, P.J.; Luciw, P.A. Nucleotide sequence of SRV-1, a type D simian acquired immune deficiency syndrome retrovirus. Science 1986, 231, 1567–1572. [Google Scholar] [CrossRef] [PubMed]
- Gardner, M.B. The history of simian AIDS. J. Med. Primatol. 1996, 25, 148–157. [Google Scholar] [CrossRef]
- Lerche, N.W.; Switzer, W.M.; Yee, J.L.; Shanmugam, V.; Rosenthal, A.N.; Chapman, L.E.; Folks, T.M.; Heneine, W. Evidence of infection with simian Type D retrovirus in persons occupationally exposed to nonhuman primates. J. Virol. 2001, 75, 1783–1789. [Google Scholar] [CrossRef] [Green Version]
- Mang, R.; Goudsmit, J.; Van Der Kuyl, A.C. Novel endogenous Type C retrovirus in baboons: Complete sequence, providing evidence for baboon endogenous virus gag-pol ancestry. J. Virol. 1999, 73, 7021–7026. [Google Scholar] [CrossRef] [Green Version]
- Van Der Kuyl, A.C.; Mang, R.; Dekker, J.T.; Goudsmit, J. Complete nucleotide sequence of simian endogenous type D retrovirus with intact genome organization: Evidence for ancestry to simian retrovirus and baboon endogenous virus. J. Virol. 1997, 71, 3666–3676. [Google Scholar] [CrossRef] [Green Version]
- Kato, S.; Matsuo, K.; Nishimura, N.; Takahashi, N.; Takano, T. The entire nucleotide sequence of baboon endogenous virus DNA: A chimeric genome structure of murine type C and simian type D retroviruses. Jpn. J. Genet. 1987, 62, 127–137. [Google Scholar] [CrossRef] [Green Version]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katzourakis, A.; Tristem, M.; Pybus, O.G.; Gifford, R.J. Discovery and analysis of the first endogenous lentivirus. Proc. Natl. Acad. Sci. USA 2007, 104, 6261–6265. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Farkasova, H.; Hron, T.; Paces, J.; Hulva, P.; Benda, P.; Gifford, R.J.; Elleder, D. Discovery of an endogenous Deltaretrovirus in the genome of long-fingered bats (Chiroptera: Miniopteridae). Proc. Natl. Acad. Sci. USA 2017, 114, 3145–3150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, G.-Z.; Worobey, M. An endogenous foamy virus in the aye-aye (Daubentonia madagascariensis). J. Virol. 2012, 86, 7696–7698. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khan, A.S.; Bodem, J.; Buseyne, F.; Gessain, A.; Johnson, W.; Kuhn, J.H.; Kuzmak, J.; Lindemann, D.; Linial, M.L.; Löchelt, M.; et al. Spumaretroviruses: Updated taxonomy and nomenclature. Virology 2018, 516, 158–164. [Google Scholar] [CrossRef]
- Falcone, V.; Leupold, J.; Clotten, J.; Urbanyi, E.; Herchenröder, O.; Spatz, W.; Volk, B.; Böhm, N.; Toniolo, A.; Neumann-Haefelin, D.; et al. Sites of simian foamy virus persistence in naturally infected African green monkeys: Latent provirus is ubiquitous, whereas viral replication is restricted to the oral mucosa. Virology 1999, 257, 7–14. [Google Scholar] [CrossRef] [Green Version]
- Murray, S.; Picker, L.J.; Axthelm, M.K.; Hudkins, K.; Alpers, C.E.; Linial, M.L. Replication in a superficial epithelial cell niche explains the lack of pathogenicity of primate foamy virus infections. J. Virol. 2008, 82, 5981–5985. [Google Scholar] [CrossRef] [Green Version]
- Aiewsakun, P.; Katzourakis, A. Marine origin of retroviruses in the early Palaeozoic Era. Nat. Commun. 2017, 8, 13954. [Google Scholar] [CrossRef]
- Shankar, A.; Sibley, S.D.; Goldberg, T.L.; Switzer, W.M. Molecular analysis of the complete genome of a simian foamy virus infecting hylobates pileatus (pileated gibbon) reveals ancient co-evolution with lesser apes. Viruses 2019, 11, 605. [Google Scholar] [CrossRef] [Green Version]
- Reid, M.J.C.; Switzer, W.M.; Schillaci, M.A.; Klegarth, A.R.; Campbell, E.; Ragonnet-Cronin, M.; Joanisse, I.; Caminiti, K.; Lowenberger, C.; Galdikas, B.M.F.; et al. Bayesian inference reveals ancient origin of simian foamy virus in orangutans. Infect. Genet. Evol. 2017, 51, 54–66. [Google Scholar] [CrossRef]
- Ghersi, B.M.; Jia, H.; Aiewsakun, P.; Katzourakis, A.; Mendoza, P.; Bausch, D.G.; Kasper, M.R.; Montgomery, J.M.; Switzer, W.M. Wide distribution and ancient evolutionary history of simian foamy viruses in New World primates. Retrovirology 2015, 12, 89. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katzourakis, A.; Aiewsakun, P.; Jia, H.; Wolfe, N.D.; Lebreton, M.; Yoder, A.D.; Switzer, W.M. Discovery of prosimian and afrotherian foamy viruses and potential cross species transmissions amidst stable and ancient mammalian co-evolution. Retrovirology 2014, 11, 61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Broussard, S.R.; Comuzzie, A.G.; Leighton, K.L.; Leland, M.; Whitehead, E.M.; Allan, J.S. Characterization of new simian foamy viruses from African nonhuman primates. Virology 1997, 237, 349–359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goldberg, T.L.; Sintasath, D.M.; Chapman, C.A.; Cameron, K.M.; Karesh, W.B.; Tang, S.; Wolfe, N.D.; Rwego, I.; Ting, N.; Switzer, W.M. Coinfection of Ugandan Red Colobus (Procolobus [Piliocolobus] rufomitratus tephrosceles) with Novel, Divergent Delta-, Lenti-, and Spumaretroviruses. J. Virol. 2009, 83, 11318–11329. [Google Scholar] [CrossRef] [Green Version]
- Hussain, A.I.; Shanmugam, V.; Bhullar, V.B.; Beer, B.E.; Vallet, D.; Gautier-Hion, A.; Wolfe, N.D.; Karesh, W.B.; Kilbourn, A.M.; Tooze, Z.; et al. Screening for simian foamy virus infection by using a combined antigen Western blot assay: Evidence for a wide distribution among Old World primates and identification of four new divergent viruses. Virology 2003, 309, 248–257. [Google Scholar] [CrossRef]
- Meiering, C.D.; Linial, M.L. Historical perspective of foamy virus epidemiology and infection. Clin. Microbiol. Rev. 2001, 14, 165–176. [Google Scholar] [CrossRef] [Green Version]
- Aghokeng, A.F.; Ayouba, A.; Mpoudi-Ngole, E.; Loul, S.; Liegeois, F.; Delaporte, E.; Peeters, M. Extensive survey on the prevalence and genetic diversity of SIVs in primate bushmeat provides insights into risks for potential new cross-species transmissions. Infect. Genet. Evol. 2010, 10, 386–396. [Google Scholar] [CrossRef] [Green Version]
- Liegeois, F.; Courgnaud, V.; Switzer, W.M.; Murphy, H.W.; Loul, S.; Aghokeng, A.; Pourrut, X.; Mpoudi-Ngolé, E.; Delaporte, E.; Peeters, M. Molecular characterization of a novel simian immunodeficiency virus lineage (SIVtal) from northern talapoins (Miopithecus ogouensis). Virology 2006, 349, 55–65. [Google Scholar] [CrossRef] [Green Version]
- Locatelli, S.; Lafay, B.; Liegeois, F.; Ting, N.; Delaporte, E.; Peeters, M. Full molecular characterization of a simian immunodeficiency virus, SIVwrcpbt from Temminck’s red colobus (Piliocolobus badius temminckii) from Abuko Nature Reserve, The Gambia. Virology 2008, 376, 90–100. [Google Scholar] [CrossRef]
- Steve, A.-M.; Ahidjo, A.; Placide, M.-K.; Caroline, F.; Mukulumanya, M.; Simon-Pierre, N.-K.; Octavie, L.-M.; Valentin, M.-A.; Jean-Jacques, M.-T.; Eric, D.; et al. High prevalences and a wide genetic diversity of simian retroviruses in non-human primate bushmeat in rural areas of the democratic republic of congo. Ecohealth 2017, 14, 100–114. [Google Scholar] [CrossRef] [Green Version]
- Vandewoude, S.; Apetrei, C. Going wild: Lessons from naturally occurring T-lymphotropic lentiviruses. Clin. Microbiol. Rev. 2006, 19, 728–762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takemura, T.; Ekwalanga, M.; Bikandou, B.; Ido, E.; Yamaguchi-Kabata, Y.; Ohkura, S.; Harada, H.; Takehisa, J.; Ichimura, H.; Parra, H.-J.; et al. A novel simian immunodeficiency virus from black mangabey (Lophocebus aterrimus) in the Democratic Republic of Congo. J. Gen. Virol. 2005, 86, 1967–1971. [Google Scholar] [CrossRef] [PubMed]
- Guzman, R.E.; Kerlin, R.L.; Zimmerman, T.E. Histologic lesions in cynomolgus monkeys (Macaca fascicularis) naturally infected with simian retrovirus type D: Comparison of seropositive, virus-positive, and uninfected animals. Toxicol. Pathol. 1999, 27, 672–677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moazed, T.C.; Thouless, M.E. Viral persistence of simian type D retrovirus (SRV-2/W) in naturally infected pigtailed macaques (Macaca nemestrina). J. Med. Primatol. 1993, 22, 382–389. [Google Scholar] [CrossRef] [PubMed]
- Nandi, J.S.; Tikute, S.A.; Chhangani, A.K.; Potdar, V.A.; Tiwari-Mishra, M.; Ashtekar, R.A.; Kumari, J.; Walimbe, A.; Mohnot, S. Natural infection by simian retrovirus-6 (SRV-6) in Hanuman langurs (Semnopithecus entellus) from two different geographical regions of India. Virology 2003, 311, 192–201. [Google Scholar] [CrossRef] [Green Version]
- Nandi, J.S.; Van Dooren, S.; Chhangani, A.K.; Mohnot, S.M. New simian β retroviruses from rhesus monkeys (Macaca mulatta) and langurs (Semnopithecus entellus) from Rajasthan, India. Virus Genes 2006, 33, 107–116. [Google Scholar] [CrossRef]
- Nandi, J.S.; Bhavalkar-Potdar, V.; Tikute, S.; Raut, C.G. A Novel Type D Simian Retrovirus Naturally Infecting the Indian Hanuman Langur (Semnopithecus entellus). Virology 2000, 277, 6–13. [Google Scholar] [CrossRef] [Green Version]
- Ahuka-Mundeke, S.; Mbala-Kingebeni, P.; Liegeois, F.; Ayouba, A.; Lunguya-Metila, O.; Demba, D.; Bilulu, G.; Mbenzo-Abokome, V.; Inogwabini, B.-I.; Muyembe-Tamfum, J.-J.; et al. Identification and molecular characterization of new simian T cell lymphotropic viruses in nonhuman primates bushmeat from the Democratic Republic of Congo. AIDS Res. Hum. Retroviruses 2012, 28, 628–635. [Google Scholar] [CrossRef] [Green Version]
- Ayouba, A.; Duval, L.; Liégeois, F.; Ngin, S.; Ahuka-Mundeke, S.; Switzer, W.M.; Delaporte, E.; Ariey, F.; Peeters, M.; Nerrienet, E. Nonhuman primate retroviruses from Cambodia: High simian foamy virus prevalence, identification of divergent STLV-1 strains and no evidence of SIV infection. Infect. Genet. Evol. 2013, 18, 325–334. [Google Scholar] [CrossRef]
- LeBreton, M.; Switzer, W.M.; Djoko, C.F.; Gillis, A.; Jia, H.; Sturgeon, M.M.; Shankar, A.; Zheng, H.; Nkeunen, G.; Tamoufe, U.; et al. A gorilla reservoir for human T-lymphotropic virus type 4. Emerg. Microbes Infect. 2014, 3, e7. [Google Scholar] [CrossRef]
- Leendertz, S.A.J.; Junglen, S.; Hedemann, C.; Goffe, A.; Calvignac-Spencer, S.; Boesch, C.; Leendertz, F.H. High prevalence, coinfection rate, and genetic diversity of retroviruses in wild red colobus monkeys (Piliocolobus badius badius) in Taï National Park, Côte d’Ivoire. J. Virol. 2010, 84, 7427–7436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mossoun, A.; Calvignac-Spencer, S.; Anoh, A.E.; Pauly, M.S.; Driscoll, D.; Michel, A.O.; Nazaire, L.G.; Pfister, S.; Sabwe, P.; Thiesen, U.; et al. Bushmeat hunting and zoonotic transmission of simian T-lymphotropic virus 1 in tropical west and central Africa. J. Virol. 2017, 91, e02479-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nerrienet, E.; Meertens, L.; Kfutwah, A.; Foupouapouognigni, Y.; Ayouba, A.; Gessain, A. Simian T cell leukaemia virus type I subtype B in a wild-caught gorilla (Gorilla gorilla gorilla) and chimpanzee (Pan troglodytes vellerosus) from Cameroon. J. Gen. Virol. 2004, 85, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Saksena, N.K.; Hervé, V.; Durand, J.P.; Leguenno, B.; Diop, O.M.; Digoutte, J.P.; Mathiot, C.; Muller-Trutwin, M.; Love, J.L.; Dube, S.; et al. Seroepidemiologic, Molecular, and Phylogenetic Analyses of Simian T-Cell Leukemia Viruses (STLV-I) from Various Naturally infected Monkey Species from Central and Western Africa. Virology 1994, 198, 297–310. [Google Scholar] [CrossRef]
- Van Brussel, M.; Salemi, M.; Liu, H.-F.; Goubau, P.; Desmyter, J.; Vandamme, A.-M. The discovery of two new divergent STLVs has implications for the evolution and epidemiology of HTLVs. Rev. Med. Virol. 1999, 9, 155–170. [Google Scholar] [CrossRef]
- Jégado, B.; Kashanchi, F.; Dutartre, H.; Mahieux, R. STLV-1 as a model for studying HTLV-1 infection. Retrovirology 2019, 16, 41. [Google Scholar] [CrossRef]
- Courgnaud, V.; Van Dooren, S.; Liegeois, F.; Pourrut, X.; Abela, B.; Loul, S.; Mpoudi-Ngole, E.; Vandamme, A.; Delaporte, E.; Peeters, M. Simian T-Cell Leukemia Virus (STLV) Infection in Wild Primate Populations in Cameroon: Evidence for Dual STLV Type 1 and Type 3 Infection in Agile Mangabeys (Cercocebus agilis). J. Virol. 2004, 78, 4700–4709. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muniz, C.P.; Jia, H.; Shankar, A.; Troncoso, L.L.; Augusto, A.M.; Farias, E.; Pissinatti, A.; Fedullo, L.P.; Santos, A.F.; Soares, M.A.; et al. An expanded search for simian foamy viruses (SFV) in Brazilian New World primates identifies novel SFV lineages and host age-related infections. Retrovirology 2015, 12, 94. [Google Scholar] [CrossRef]
- Jin, M.J.; Rogers, J.; Phillips-Conroy, J.E.; Allan, J.S.; Desrosiers, R.C.; Shaw, G.M.; Sharp, P.M.; Hahn, B.H. Infection of a yellow baboon with simian immunodeficiency virus from African green monkeys: Evidence for cross-species transmission in the wild. J. Virol. 1994, 68, 8454–8460. [Google Scholar] [CrossRef] [Green Version]
- Van Rensburg, E.J.; Mwenda, J.; Robson, B.A.; Stander, T.; Laten, J.D.; Chege, G.K.; Engelbrecht, S. Simian immunodeficiency viruses (SIVs) from eastern and southern Africa: Detection of a SIVagm variant from a chacma baboon. J. Gen. Virol. 1998, 79, 1809–1814. [Google Scholar] [CrossRef] [Green Version]
- Nyamota, R.; Owino, V.; Murungi, E.K.; Villinger, J.; Otiende, M.; Masiga, D.; Thuita, J.; Lekolool, I.; Jeneby, M.M. Broad diversity of simian immunodeficiency virus infecting Chlorocebus species (African green monkey) and evidence of cross-species infection in Papio anubis (olive baboon) in Kenya. J. Med. Primatol. 2020, 49, 165–178. [Google Scholar] [CrossRef] [PubMed]
- Bibollet-Ruche, F.; Galat-Luong, A.; Cuny, G.; Sarni-Manchado, P.; Galat, G.; Durand, J.-P.; Pourrut, X.; Veas, F. Simian immunodeficiency virus infection in a patas monkey (Erythrocebus patas): Evidence for cross-species transmission from African green monkeys (Cercopithecus aethiops sabaeus) in the wild. J. Gen. Virol. 1996, 77, 773–781. [Google Scholar] [CrossRef] [PubMed]
- Gifford, R.J.; Katzourakis, A.; Tristem, M.; Pybus, O.G.; Winters, M.; Shafer, R.W. A transitional endogenous lentivirus from the genome of a basal primate and implications for lentivirus evolution. Proc. Natl. Acad. Sci. USA 2008, 105, 20362–20367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gilbert, C.; Maxfield, D.G.; Goodman, S.M.; Feschotte, C. Parallel germline infiltration of a lentivirus in two malagasy lemurs. PLoS Genet. 2009, 5, e1000425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, G.-Z.; Worobey, M. A primitive endogenous lentivirus in a colugo: Insights into the early evolution of lentiviruses. Mol. Biol. Evol. 2015, 32, 211–215. [Google Scholar] [CrossRef]
- Gifford, R.J. Viral evolution in deep time: Lentiviruses and mammals. Trends Genet. 2012, 28, 89–100. [Google Scholar] [CrossRef]
- Raehtz, K.; Pandrea, I.; Apetrei, C. The well-tempered SIV infection: Pathogenesis of SIV infection in natural hosts in the wild, with emphasis on virus transmission and early events post-infection that may contribute to protection from disease progression. Infect. Genet. Evol. 2016, 46, 308–323. [Google Scholar] [CrossRef] [Green Version]
- Ma, D.; Jasinska, A.; Kristoff, J.; Grobler, J.P.; Turner, T.; Jung, Y.; Schmitt, C.; Raehtz, K.; Feyertag, F.; Sosa, N.M.; et al. SIVagm infection in wild African green monkeys from South Africa: Epidemiology, natural history, and evolutionary considerations. PLOS Pathog. 2013, 9, e1003011. [Google Scholar] [CrossRef] [Green Version]
- Apetrei, C. The history of SIVS and AIDS: Epidemiology, phylogeny and biology of isolates from naturally SIV infected non-human primates (NHP) in Africa. Front. Biosci. 2004, 9, 225–254. [Google Scholar] [CrossRef] [Green Version]
- Sharp, P.M.; Bailes, E.; Gao, F.; Beer, B.E.; Hirsch, V.M.; Hahn, B.H. Origins and evolution of AIDS viruses: Estimating the time-scale. Biochem. Soc. Trans. 2000, 28, 275–282. [Google Scholar] [CrossRef] [Green Version]
- Wertheim, J.O.; Worobey, M. Dating the age of the SIV lineages that gave rise to HIV-1 and HIV-2. PLoS Comput. Biol. 2009, 5, e1000377. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Worobey, M.; Telfer, P.; Souquière, S.; Hunter, M.; Coleman, C.A.; Metzger, M.J.; Reed, P.; Makuwa, M.; Hearn, G.; Honarvar, S.; et al. Island biogeography reveals the deep history of SIV. Science 2010, 329, 1487. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fleagle, J.G.; Gilbert, C.C. The biogeography of primate evolution: The role of plate tectonics, climate and chance. In Primate Biogeography; Lehman, S.M., Fleagle, J.G., Eds.; Springer: New York, NY, USA, 2006; pp. 375–418. [Google Scholar] [CrossRef]
- Rook, L.; Martínez-Navarro, B.; Howell, F.C. Occurrence of Theropithecus sp. in the Late Villafranchian of Southern Italy and implication for Early Pleistocene “out of Africa” dispersals. J. Hum. Evol. 2004, 47, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Winney, B.; Hammond, R.L.; Macasero, W.; Flores, B.; Boug, A.; Biquand, V.; Biquand, S.; Bruford, M.W. Crossing the Red Sea: Phylogeography of the hamadryas baboon, Papio hamadryas hamadryas. Mol. Ecol. 2004, 13, 2819–2827. [Google Scholar] [CrossRef] [PubMed]
- Kopp, G.H.; Roos, C.; Butynski, T.M.; Wildman, D.E.; Alagaili, A.N.; Groeneveld, L.F.; Zinner, D. Out of Africa, but how and when? The case of hamadryas baboons (Papio hamadryas). J. Hum. Evol. 2014, 76, 154–164. [Google Scholar] [CrossRef] [Green Version]
- Wertheim, J.O.; Worobey, M. A Challenge to the Ancient Origin of SIVagm Based on African Green Monkey Mitochondrial Genomes. PLOS Pathog. 2007, 3, e95. [Google Scholar] [CrossRef] [Green Version]
- Yoshikawa, R.; Okamoto, M.; Sakaguchi, S.; Nakagawa, S.; Miura, T.; Hirai, H.; Miyazawa, T. Simian Retrovirus 4 Induces Lethal Acute Thrombocytopenia in Japanese Macaques. J. Virol. 2015, 89, 3965–3975. [Google Scholar] [CrossRef] [Green Version]
- Zao, C.-L.; Tomanek, L.; Cooke, A.; Berger, R.; Yang, L.; Xie, C.; Chen, S.; Shi, C.; Rong, R. A novel simian retrovirus subtype discovered in cynomolgus monkeys (Macaca fascicularis). J. Gen. Virol. 2016, 97, 3017–3023. [Google Scholar] [CrossRef]
- Sommerfelt, M.A.; Harkestad, N.; Hunter, E. The endogenous langur type D retrovirus PO-1-Lu and its exogenous counterparts in macaque and langur monkeys. Virology 2003, 315, 275–282. [Google Scholar] [CrossRef] [Green Version]
- Todaro, G.J.; Benveniste, R.E.; Sherr, C.J.; Schlom, J.; Schidlovsky, G.; Stephenson, J.R. Isolation and characterization of a new type D retrovirus from the Asian primate, Presbytis obscurus (spectacled langur). Virology 1978, 84, 189–194. [Google Scholar] [CrossRef]
- Sonigo, P.; Barker, C.; Hunter, E.; Wain-Hobson, S. Nucleotide sequence of Mason-Pfizer monkey virus: An immunosuppressive D-type retrovirus. Cell 1986, 45, 375–385. [Google Scholar] [CrossRef]
- Montiel, N.A. An updated review of simian betaretrovirus (SRV) in macaque hosts. J. Med. Primatol. 2010, 39, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Hayami, M.; Komuro, A.; Nozawa, K.; Shotake, T.; Ishikawa, K.-I.; Yamamoto, K.; Ishida, T.; Honjo, S.; Hinuma, Y. Prevalence of antibody to adult T-cell leukemia virus-associated antigens (ATLA) in Japanese monkeys and other non-human primates. Int. J. Cancer 1984, 33, 179–183. [Google Scholar] [CrossRef] [PubMed]
- Vandamme, A.-M.; Liu, H.-F.; Goubau, P.; Desmyter, J. Primate T-Lymphotropic Virus Type I LTR Sequence Variation and Its Phylogenetic Analysis: Compatibility with an African Origin of PTLV-I. Virology 1994, 202, 212–223. [Google Scholar] [CrossRef] [PubMed]
- Vandamme, A.-M.; Liu, H.-F.; Van Brussel, M.; De Meurichy, W.; Desmyter, J.; Goubau, P. The presence of a divergent T-lymphotropic virus in a wild-caught pygmy chimpanzee (Pan paniscus) supports an African origin for the human T-lymphotropic/simian T-lymphotropic group of viruses. J. Gen. Virol. 1996, 77, 1089–1099. [Google Scholar] [CrossRef] [PubMed]
- Mahieux, R.; Gessain, A. HTLV-3/STLV-3 and HTLV-4 viruses: Discovery, epidemiology, serology and molecular aspects. Viruses 2011, 3, 1074–1090. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, N.D.; Heneine, W.; Carr, J.K.; Garcia, A.D.; Shanmugam, V.; Tamoufe, U.; Torimiro, J.N.; Prosser, A.T.; Lebreton, M.; Mpoudi-Ngole, E.; et al. Emergence of unique primate T-lymphotropic viruses among central African bushmeat hunters. Proc. Natl. Acad. Sci. USA 2005, 102, 7994–7999. [Google Scholar] [CrossRef] [Green Version]
- Eciminale, V.; Erende, F.; Ebertazzoni, U.; Romanelli, M. HTLV-1 and HTLV-2: Highly similar viruses with distinct oncogenic properties. Front. Microbiol. 2014, 5, 398. [Google Scholar] [CrossRef] [Green Version]
- Wattel, E.; Vartanian, J.P.; Pannetier, C.; Wain-Hobson, S. Clonal expansion of human T-cell leukemia virus type I-infected cells in asymptomatic and symptomatic carriers without malignancy. J. Virol. 1995, 69, 2863–2868. [Google Scholar] [CrossRef] [Green Version]
- Vandamme, A.-M. The simian origins of the pathogenic human T-cell lymphotropic virus type I. Trends Microbiol. 1998, 6, 477–483. [Google Scholar] [CrossRef]
- Switzer, W.M.; Qari, S.H.; Wolfe, N.D.; Burke, D.S.; Folks, T.M.; Heneine, W. Ancient origin and molecular features of the novel human T-lymphotropic virus type 3 revealed by complete genome analysis. J. Virol. 2006, 80, 7427–7438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mager, D.L.; Freeman, J. HERV-H endogenous retroviruses: Presence in the new world branch but amplification in the old world primate lineage. Virology 1995, 213, 395–404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, A.H.-S.; Kim, D.-S.; Huh, J.-W.; Ahn, K.; Yi, J.M.; Lee, J.-R.; Hirai, H. Molecular characterization of the HERV-W env gene in humans and primates: Expression, FISH, phylogeny, and evolution. Mol. Cells 2008, 26, 53–60. [Google Scholar] [PubMed]
- Yi, J.-M.; Kim, T.-H.; Huh, J.-W.; Park, K.S.; Jang, S.B.; Kim, H.-M.; Kim, A.H.-S. Human endogenous retroviral elements belonging to the HERV-S family from human tissues, cancer cells, and primates: Expression, structure, phylogeny and evolution. Gene 2004, 342, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Kim, A.H.-S.; Yi, J.-M.; Hirai, H.; Huh, J.-W.; Jeong, M.-S.; Jang, S.-B.; Kim, C.-G.; Saitou, N.; Hyun, B.-H.; Lee, W.-H. Human Endogenous Retrovirus (HERV)-R family in primates: Chromosomal location, gene expression, and evolution. Gene 2006, 370, 34–42. [Google Scholar] [CrossRef]
- Lee, J.-W.; Kim, H.-S. Endogenous retrovirus HERV-I LTR family in primates: Sequences, phylogeny, and evolution. Arch. Virol. 2006, 151, 1651–1658. [Google Scholar] [CrossRef]
- Yi, J.-M.; Kim, H.-S. Molecular evolution of the HERV-E family in primates. Arch. Virol. 2006, 151, 1107–1116. [Google Scholar] [CrossRef]
- Subramanian, R.P.; Wildschutte, J.H.; Russo, C.; Coffin, J.M. Identification, characterization, and comparative genomic distribution of the HERV-K (HML-2) group of human endogenous retroviruses. Retrovirology 2011, 8, 90. [Google Scholar] [CrossRef] [Green Version]
- Romano, C.M.; De Melo, F.L.; Corsini, M.A.B.; Holmes, E.C.; Zanotto, P.M.D.A. Demographic histories of ERV-K in humans, chimpanzees and rhesus Monkeys. PLoS ONE 2007, 2, e1026. [Google Scholar] [CrossRef]
- Reus, K.; Mayer, J.; Sauter, M.; Zischler, H.; Müller-Lantzsch, N.; Meese, E. HERV-K(OLD): Ancestor Sequences of the Human Endogenous Retrovirus Family HERV-K(HML-2). J. Virol. 2001, 75, 8917–8926. [Google Scholar] [CrossRef] [Green Version]
- Van Der Kuyl, A.C.; Dekker, J.T.; Goudsmit, J. Full-length proviruses of baboon endogenous virus (BaEV) and dispersed BaEV reverse transcriptase retroelements in the genome of baboon species. J. Virol. 1995, 69, 5917–5924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, H.L.; Leon, E.J.; Wallace, L.; Nimiyongskul, F.A.; Buechler, M.B.; Newman, L.P.; Castrovinci, P.A.; Johnson, R.P.; Gifford, R.J.; Jones, R.B.; et al. Identification and spontaneous immune targeting of an endogenous retrovirus K envelope protein in the Indian rhesus macaque model of human disease. Retrovirology 2016, 13, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, H.; Ma, Y.; Ma, W.; Williams, D.K.; Galvin, T.A.; Khan, A.S. Chemical induction of endogenous retrovirus particles from the vero cell line of African green monkeys. J. Virol. 2011, 85, 6579–6588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cohen, M.; Rein, A.; Stephens, R.M.; O’Connell, C.; Gilden, R.V.; Shure, M.; Nicolson, M.O.; McAllister, R.M.; Davidson, N. Baboon endogenous virus genome: Molecular cloning and structural characterization of nondefective viral genomes from DNA of a baboon cell strain. Proc. Natl. Acad. Sci. USA 1981, 78, 5207–5211. [Google Scholar] [CrossRef] [Green Version]
- Maze, E.A.; Ham, C.; Kelly, J.; Ussher, L.; Almond, N.; Towers, G.J.; Berry, N.; Belshaw, R. Variable baseline papio cynocephalus endogenous retrovirus (PcEV) expression is upregulated in acutely SIV-infected macaques and correlated to STAT1 expression in the spleen. Front. Immunol. 2019, 10, 901. [Google Scholar] [CrossRef]
- Macfarlane, C.M.; Badge, R.M. Genome-wide amplification of proviral sequences reveals new polymorphic HERV-K(HML-2) proviruses in humans and chimpanzees that are absent from genome assemblies. Retrovirology 2015, 12, 35. [Google Scholar] [CrossRef] [Green Version]
- Sakuma, C.; Sekizuka, T.; Kuroda, M.; Kasai, F.; Saito, K.; Ikeda, M.; Yamaji, T.; Osada, N.; Hanada, K. Novel endogenous simian retroviral integrations in Vero cells: Implications for quality control of a human vaccine cell substrate. Sci. Rep. 2018, 8, 644. [Google Scholar] [CrossRef]
- Vandamme, A.-M. BaEV is a relic from an ancient retrovirus that crossed species barriers. Trends Microbiol. 1996, 4, 478. [Google Scholar] [CrossRef]
- Van Der Kuyl, A.C.; Dekker, J.T.; Goudsmit, J. Distribution of baboon endogenous virus among species of African monkeys suggests multiple ancient cross-species transmissions in shared habitats. J. Virol. 1995, 69, 7877–7887. [Google Scholar] [CrossRef] [Green Version]
- Mang, R.; Maas, J.; Van Der Kuyl, A.C.; Goudsmit, J. Papio cynocephalus endogenous retrovirus among old world monkeys: Evidence for coevolution and ancient cross-species transmissions. J. Virol. 2000, 74, 1578–1586. [Google Scholar] [CrossRef] [Green Version]
- Ikeda, M.; Satomura, K.; Sekizuka, T.; Hanada, K.; Endo, T.; Osada, N. Comprehensive phylogenomic analysis reveals a novel cluster of simian endogenous retroviral sequences in Colobinae monkeys. Am. J. Primatol. 2018, 80, e22882. [Google Scholar] [CrossRef] [PubMed]
- Van Der Kuyl, A.C.; Dekker, J.T.; Goudsmit, J. Primate genus miopithecus: Evidence for the existence of species and subspecies of dwarf guenons based on cellular and endogenous viral sequences. Mol. Phylogenetics Evol. 2000, 14, 403–413. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.; Davidson, N.; Gilden, R.V.; McAllister, R.M.; Nicolson, M.O.; Stephens, R.M. The baboon endogenous virus genome. II. Provirus sequence variations in baboon cell DNA. Nucleic Acids Res. 1980, 8, 4423–4440. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holloway, J.R.; Williams, Z.H.; Freeman, M.M.; Bulow, U.; Coffin, J.M. Gorillas have been infected with the HERV-K (HML-2) endogenous retrovirus much more recently than humans and chimpanzees. Proc. Natl. Acad. Sci. USA 2019, 116, 1337–1346. [Google Scholar] [CrossRef] [Green Version]
- Magiorkinis, G.; Blanco-Melo, D.; Belshaw, R. The decline of human endogenous retroviruses: Extinction and survival. Retrovirology 2015, 12, 81. [Google Scholar] [CrossRef] [Green Version]
- Mailund, T.; Munch, K.; Schierup, M.H. Lineage sorting in apes. Annu. Rev. Genet. 2014, 48, 519–535. [Google Scholar] [CrossRef]
- Roos, C.; Kothe, M.; Alba, D.M.; Delson, E.; Zinner, D. The radiation of macaques out of Africa: Evidence from mitogenome divergence times and the fossil record. J. Hum. Evol. 2019, 133, 114–132. [Google Scholar] [CrossRef]
- Hughes, J.K.; Elton, S.; O’Regan, H.J. Theropithecus and ‘Out of Africa’ dispersal in the Plio-Pleistocene. J. Hum. Evol. 2008, 54, 43–77. [Google Scholar] [CrossRef]
- Elton, S.; O’Regan, H.J. Macaques at the margins: The biogeography and extinction of Macaca sylvanus in Europe. Quat. Sci. Rev. 2014, 96, 117–130. [Google Scholar] [CrossRef] [Green Version]
- Hawkins, J.A.; Adams, W.V., Jr.; Wilson, B.H.; Issel, C.J.; Roth, E.E. Transmission of equine infectious anemia virus by Tabanus fuscicostatus. J. Am. Veter-Med. Assoc. 1976, 168, 63–64. [Google Scholar]
- Kohara, J.; Takeuchi, M.; Hirano, Y.; Sakurai, Y.; Takahashi, T. Vector control efficacy of fly nets on preventing bovine leukemia virus transmission. J. Veter. Med. Sci. 2018, 80, 1524–1527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Williams, K.C.; Burdo, T.H. HIV and SIV infection: The role of cellular restriction and immune responses in viral replication and pathogenesis. Apmis 2009, 117, 400–412. [Google Scholar] [CrossRef] [PubMed]
- Bialik, O.M.; Frank, M.; Betzler, C.; Zammit, R.; Waldmann, N.D. Two-step closure of the Miocene Indian Ocean Gateway to the Mediterranean. Sci. Rep. 2019, 9, 8842. [Google Scholar] [CrossRef] [PubMed]
- Parrish, S.W.; Brown, A.E.; Chanbancherd, P.; Gettayacamin, M.; Parrish, J.H. Transmission of STLV in a closed colony of macaques. Am. J. Primatol. 2004, 63, 103–109. [Google Scholar] [CrossRef] [PubMed]
- Murata, M.; Yasunaga, J.-I.; Washizaki, A.; Seki, Y.; Kuramitsu, M.; Tan, W.K.; Hu, A.; Okuma, K.; Hamaguchi, I.; Mizukami, T.; et al. Frequent horizontal and mother-to-child transmission may contribute to high prevalence of STLV-1 infection in Japanese macaques. Retrovirology 2020, 17, 15. [Google Scholar] [CrossRef]
- Afonso, P.V.; Fagrouch, Z.; Deijs, M.; Niphuis, H.; Bogers, W.; Gessain, A.; Van Der Hoek, L.; Verschoor, E.J. Absence of accessory genes in a divergent simian T-lymphotropic virus type 1 isolated from a bonnet macaque (Macaca radiata). PLOS Neglected Trop. Dis. 2019, 13, e0007521. [Google Scholar] [CrossRef]
- Merceron, G.; Kaiser, T.M.; Kostopoulos, D.S.; Schulz-Kornas, E. Ruminant diets and the Miocene extinction of European great apes. Proc. R. Soc. B Biol. Sci. 2010, 277, 3105–3112. [Google Scholar] [CrossRef] [Green Version]
- Merceron, G.; Kostopoulos, D.S.; De Bonis, L.; Fourel, F.; Koufos, G.D.; Lécuyer, C.; Martineau, F. Stable isotope ecology of Miocene bovids from northern Greece and the ape/monkey turnover in the Balkans. J. Hum. Evol. 2013, 65, 185–198. [Google Scholar] [CrossRef]
- Nelson, S.V.; Rook, L. Isotopic reconstructions of habitat change surrounding the extinction of Oreopithecus, the last European ape. Am. J. Phys. Anthr. 2016, 160, 254–271. [Google Scholar] [CrossRef]
- Kostopoulos, D.S.; Guy, F.; Kynigopoulou, Z.; Koufos, G.D.; Valentin, X.; Merceron, G. A 2Ma old baboon-like monkey from Northern Greece and new evidence to support the Paradolichopithecus—Procynocephalus synonymy (Primates: Cercopithecidae). J. Hum. Evol. 2018, 121, 178–192. [Google Scholar] [CrossRef]
- Radović, P.; Lindal, J.; Marković, Z.; Alaburić, S.; Roksandic, M. First record of a fossil monkey (Primates, Cercopithecidae) from the Late Pliocene of Serbia. J. Hum. Evol. 2019, 137, 102681. [Google Scholar] [CrossRef]
- Sinha, A.; Johnson, W.E. Retroviruses of the RDR superinfection interference group: Ancient origins and broad host distribution of a promiscuous Env gene. Curr. Opin. Virol. 2017, 25, 105–112. [Google Scholar] [CrossRef]
- Malicorne, S.; Vernochet, C.; Cornelis, G.; Mulot, B.; Delsuc, F.; Heidmann, O.; Heidmann, T.; Dupressoir, A. Genome-Wide Screening of Retroviral Envelope Genes in the Nine-Banded Armadillo (Dasypus novemcinctus, Xenarthra) Reveals an Unfixed Chimeric Endogenous Betaretrovirus Using the ASCT2 Receptor. J. Virol. 2016, 90, 8132–8149. [Google Scholar] [CrossRef] [Green Version]
- Van Der Kuyl, A.C.; Berkhout, B. Viruses in the reproductive tract: On their way to the germ line? Virus Res. 2020, 286, 198101. [Google Scholar] [CrossRef]
- Marin, M.; Tailor, C.S.; Nouri, A.; Kabat, D. Sodium-Dependent neutral amino acid transporter type 1 is an auxiliary receptor for baboon endogenous retrovirus. J. Virol. 2000, 74, 8085–8093. [Google Scholar] [CrossRef] [Green Version]
- Shimode, S.; Nakaoka, R.; Shogen, H.; Miyazawa, T. Characterization of feline ASCT1 and ASCT2 as RD-114 virus receptor. J. Gen. Virol. 2013, 94, 1608–1612. [Google Scholar] [CrossRef]
- Nethe, M.; Berkhout, B.; Van Der Kuyl, A.C. Retroviral superinfection resistance. Retrovirology 2005, 2, 52. [Google Scholar] [CrossRef] [Green Version]
- Sugimoto, J.; Sugimoto, M.; Bernstein, H.B.; Jinno, Y.; Schust, D.J. A novel human endogenous retroviral protein inhibits cell-cell fusion. Sci. Rep. 2013, 3, srep01462. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]

| Retrovirus Species Genus | African OWM Species + | Asian OWM Species + | Other Primate Species + | References | ||
|---|---|---|---|---|---|---|
| Cercopithecinae | Colobinae | Cercopithecinae | Colobinae | |||
| Simian foamy virus (SFV) 1 Simii-spumavirus | Cercopithecini: Chlorocebus Erythrocebus Papionini: Cercocebus Lophocebus Macaca Mandrillus Papio Theropithecus | Colobus Procolobus | Papionini: Macaca | Pygathrix Trachypithecus | Gorilla Hylobates Pan Pongo | [50,53,54,55,56] |
| Simian immuno-deficiency virus (SIV) 2 Lentivirus | Cercopithecini: Allenopithecus Cercopithecus Chlorocebus Miopithecus Papionini: Cercocebus Lophocebus Mandrillus | Colobus Piliocolobus Procolobus | None | None | Gorilla Homo (HIV) Pan | [54,57,58,59,60,61,62] |
| Simian type D retrovirus 3 (SRV) Betaretrovirus | None | None | Papionini: Macaca | Semnopithecus? | None | [63,64,65,66,67] |
| Simian T-lymphotropic virus (STLV) Deltavirus | Cercopithecini: Allenopithecus Cercopithecus Chlorocebus Erythrocebus Miopithecus Papionini: Cercocebus Lophocebus Macaca Mandrillus Papio | Piliocolobus Procolobus | Papionini: Macaca | Presbytis | Gorilla Homo (HTLV) Hylobates Pan Pongo | [22,24,54,68,69,70,71,72,73,74,75,76,77] |
| Retrovirus Species Genus | African OWM Species + | Asian OWM Species + | References | ||
|---|---|---|---|---|---|
| Cercopithecinae | Colobinae | Cercopithecinae | Colobinae | ||
| Baboon endogenous virus (BaEV) Gammaretrovirus | Cercopithecini: Chlorocebus Papionini: Cercocebus Mandrillus Lophocebus Papio Theropithecus | None | None | None | [130] Figure A1 |
| Papio cynocephalus endogous virus (PcEV) Gammaretrovirus | Papionini: Lophocebus Papio Theropithecus | Colobus | Papionini: Macaca | None | [131] Figure A1 |
| Simian endogenous retrovirus (SERV) Betaretrovirus (Simian type D) | Cercopithecini: Cercopithecus Chlorocebus Erythrocebus Miopithecus Papionini: Macaca | None | Papionini: Macaca | Rhinopithecus | [38,132,133] Figure A1 |
| Simian endogenous retrovirus-K1 (SERV-K1/ RhERV-K) Betaretrovirus | Papionini: Cercocebus Mandrillus Papio Theropithecus | Colobus | Papionini: Macaca | Trachypithecus | Figure A2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
van der Kuyl, A.C. Contemporary Distribution, Estimated Age, and Prehistoric Migrations of Old World Monkey Retroviruses. Epidemiologia 2021, 2, 46-67. https://0-doi-org.brum.beds.ac.uk/10.3390/epidemiologia2010005
van der Kuyl AC. Contemporary Distribution, Estimated Age, and Prehistoric Migrations of Old World Monkey Retroviruses. Epidemiologia. 2021; 2(1):46-67. https://0-doi-org.brum.beds.ac.uk/10.3390/epidemiologia2010005
Chicago/Turabian Stylevan der Kuyl, Antoinette C. 2021. "Contemporary Distribution, Estimated Age, and Prehistoric Migrations of Old World Monkey Retroviruses" Epidemiologia 2, no. 1: 46-67. https://0-doi-org.brum.beds.ac.uk/10.3390/epidemiologia2010005





