Teladorsagia circumcincta 1,6-Bisphosphate Aldolase: Molecular and Biochemical Characterisation, Structure Analysis and Recognition by Immune Hosts
Abstract
:1. Introduction
2. Material and Methods
2.1. Parasite Culture and Collection
2.2. RNA Isolation and cDNA Synthesis
2.3. Cloning and Expression of T. circumcincta Recombinant TciALDO-1 in E. coli
2.4. Purification and Gel Electrophoresis
2.5. Bioinformatics
2.6. TciALDO-1 Activity (E.C. 4.2.1.11)
2.7. ELISA
2.8. Protein Modelling and Structural Analysis of TciALDO-1
2.9. Data Analysis
3. Results
3.1. TciALDO-1 Gene Sequence
3.2. Recombinant Protein Expression
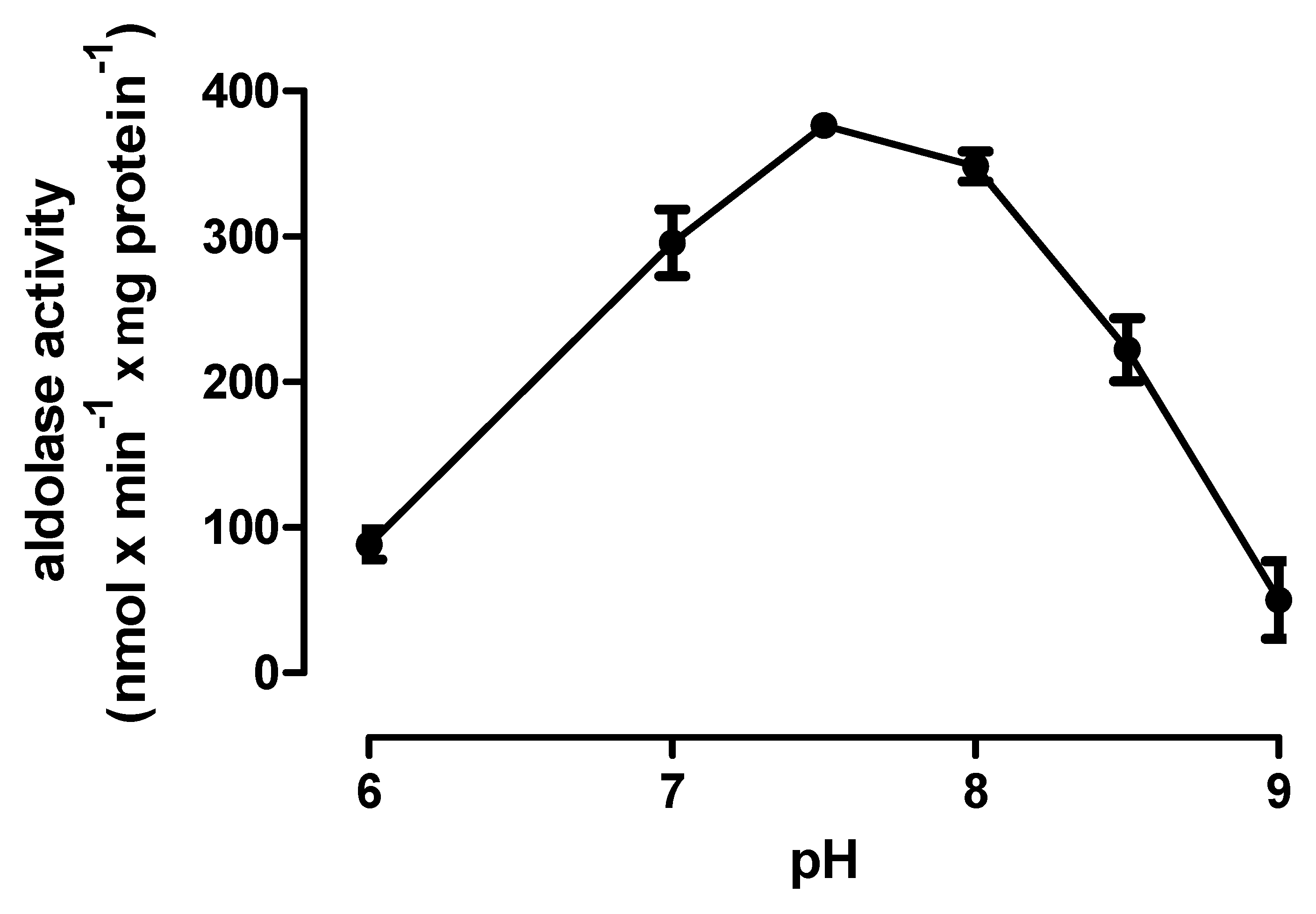
3.3. Enzyme Assays
3.4. Host Recognition
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Marsh, K.; Lebherz, H.G. Fructose-bisphosphate aldolases; an evolutionary history. Trends Biochem. Sci. 1992, 17, 110–118. [Google Scholar] [CrossRef]
- Gefflaut, T.; Blonski, C.; Perie, J.; Willson, M. Class I aldolases: Substrate specificity, mechanism, inhibitors and structural aspects. Prog. Biophys. Mol. Biol. 1995, 63, 301–340. [Google Scholar] [CrossRef]
- Sánchez, L.B.; Horner, D.S.; Moore, D.V.; Henze, K.; Embley, T.; Müller, M. Fructose-1,6-bisphosphate aldolases in amitochondriate protists constitute a single protein subfamily with eubacterial relationships. Gene 2002, 295, 51–59. [Google Scholar] [CrossRef]
- Tittmann, K. Sweet siblings with different faces: The mechanisms of FBP and F6P aldolase, transaldolase, transketolase and phosphoketolase revisited in light of recent structural data. Bioorg. Chem. 2014, 57, 263–280. [Google Scholar] [CrossRef] [PubMed]
- Inoue, T.; Yatsuki, H.; Kusakabe, T.; Joh, K.; Takasaki, Y.; Nikoh, N.; Miyata, T.; Hori, K. Caenorhabditis elegans has two isozymic forms, CE-1 and CE-2, of fructose-1,6-bisphosphate aldolase which are encoded by different genes. Arch. Biochem. Biophys. 1997, 339, 226–234. [Google Scholar] [CrossRef] [PubMed]
- Yan, R.; Xu, L.; Wang, J.; Song, X.; Li, X. Cloning and characterization of aldolase from parasitic nematode Haemonchus contortus. J. Anim. Vet. Adv. 2013, 12, 478–486. [Google Scholar]
- Kovaleva, E.S.; Masler, E.P.; Subbotin, S.A.; Chitwood, D.J. Molecular characterization of aldolase from Heterodera glycines and Globodera rostochiensis. J. Nematol. 2005, 37, 292–296. [Google Scholar]
- El-Dabaa, E.; Mei, H.; El-Sayed, A.; Karim, A.M.; Eldesoky, H.M.; Fahim, F.A.; LoVerde, P.T.; Saber, M.A. Cloning and characterization of Schistosoma mansoni fructose-1,6-bisphosphate aldolase isoenzyme. J. Parasitol. 1998, 84, 954–960. [Google Scholar] [CrossRef] [PubMed]
- Lorenzatto, K.R.; Monteiro, K.M.; Paredes, R.; Paludo, G.P.; da Fonsêca, M.M.; Galanti, N.; Zaha, A.; Ferreira, H.B. Fructose-bisphosphate aldolase and enolase from Echinococcus granulosus: Genes, expression patterns and protein interactions of two potential moonlighting proteins. Gene 2012, 506, 76–84. [Google Scholar] [CrossRef] [Green Version]
- Li, S.; Bian, M.; Wang, X.; Chen, X.; Xie, Z.; Sun, H.; Jia, F.; Liang, P.; Zhou, C.; He, L.; et al. Molecular and biochemical characterizations of three fructose-1,6-bisphosphate aldolases from Clonorchis sinensis. Mol. Biochem. Parasitol. 2014, 194, 36–43. [Google Scholar] [CrossRef]
- Hu, Q.; Xie, H.; Zhu, S.; Liao, D.; Zhan, T.; Liu, D. Cloning, expression and partial characterization of FBPA from Schistosoma japonicum, a molecule on that the fluke may develop nutrition competition and immune evasion from human. Parasitol. Res. 2015, 114, 3459–3468. [Google Scholar] [CrossRef] [PubMed]
- Prompipak, J.; Senawong, T.; Jokchaiyaphum, K.; Siriwes, K.; Nuchadomrong, S.; Laha, T.; Sripa, B.; Senawong, G. Characterization and localization of Opisthorchis viverrini fructose-1,6-bisphosphate aldolase. Parasitol. Int. 2017, 66, 413–418. [Google Scholar] [CrossRef] [PubMed]
- Reznick, A.Z.; Gershon, D. Purification of fructose-1,6-diphosphate aldolase from the free-living nematode Turbatrix aceti. Comparison of properties with those of other class I aldolases. Int. J. Biochem. 1977, 8, 53–59. [Google Scholar] [CrossRef]
- Reznick, A.Z.; Gershon, D. Age related alterations in purified fructose-1.6-diphosphate aldolase from the nematode Turbatrix aceti. Mech. Aging Dev. 1977, 6, 53–59. [Google Scholar] [CrossRef]
- Zeelon, P.; Gershon, H.; Gershon, D. Inactive enzyme molecules in aging organisms. Nematode fructose 1,6-diphosphate aldolase. Biochemistry 1973, 12, 1743–1750. [Google Scholar] [CrossRef]
- Gόmez-Arreaza, A.; Acosta, H.; Quiñones, W.; Concepción, J.L.; Michels, P.A.; Avilán, L. Extracellular functions of glycolytic enzymes of parasites: Unpredicted use of ancient proteins. Mol. Biochem. Parasitol. 2014, 193, 75–81. [Google Scholar] [CrossRef]
- Ramajo-Hernández, A.; Sánchez, R.P.; Martín, V.R.; Oleaga, A. Schistosoma bovis: Plasminogen binding in adults and the identification of plasminogen-binding proteins from the worm tegument. Exp. Parasitol. 2007, 115, 83–91. [Google Scholar]
- Marques, H.H.; Zouain, C.S.; Torres, C.B.B.; Oliveira, J.S.; Alves, J.B.; Goes, A.M. Protective effect and granuloma down-modulation promoted by RP44 antigen a fructose 1,6-bisphosphate aldolase of Schistosoma mansoni. Immunobiology 2008, 213, 437–446. [Google Scholar] [CrossRef]
- Guillou, F.; Roger, E.; Moné, Y.; Rognon, A.; Grunau, C.; Théron, A.; Mitta, G.; Coustau, C.; Gourbal, B.E. Excretory–secretory proteome of larval Schistosoma mansoni and Echinostoma caproni, two parasites of Biomphalaria glabrata. Mol. Biochem. Parasitol. 2007, 155, 45–56. [Google Scholar] [CrossRef]
- Morassutti, A.L.; Levert, K.; Pinto, P.M.; da Silva, A.J.; Wilkins, P.; Graeff-Teixeira, C. Characterization of Angiostrongylus cantonensis excretory–secretory proteins as potential diagnostic targets. Exp. Parasitol. 2012, 130, 26–31. [Google Scholar] [CrossRef] [Green Version]
- Umair, S.; Ria, C.; Knight, J.S.; Simpson, H.V. Sarcosine metabolism in Haemonchus contortus and Teladorsagia circumcincta. Exp. Parasitol. 2013, 134, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Umair, S.; Bouchet, C.L.G.; Knight, J.S.; Pernthaner, A.; Simpson, H.V. Molecular and biochemical characterisation and recognition by the immune host of the glyceraldehyde 3-phosphate dehydrogenase (GAPDH) of the abomasal nematode parasite Teladorsagia circumcincta. Exp. Parasitol. 2017, 181, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Umair, S.; Bouchet, C.L.G.; Knight, J.S.; Pernthaner, A.; Simpson, H.V. Molecular and biochemical characterisation and recognition by the immune host of the enolase of the abomasal nematode parasite Teladorsagia circumcincta. Exp. Parasitol. 2017, 172, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Umair, S.; Bouchet, C.L.G.; Deng, Q.; Palevich, N.; Simpson, H.V. Characterisation of a Teladorsagia circumcincta glutathione transferase. Mol. Biochem. Parasitol. 2020, 239, 111316. [Google Scholar] [PubMed]
- Umair, S.; Bouchet, C.; Palevich, N.; Simpson, H.V. Characterisation and structural analysis of glyoxylate cycle enzymes of Teladorsagia circumcincta. Mol. Biochem. Parasitol. 2020, 240, 111335. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Skolnick, J. Scoring function for automated assessment of protein structure template quality. Proteins 2004, 57, 702–710. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Rullmann, J.A.C.; MacArthur, M.W.; Kaptein, R.; Thornton, J.M. AQUA and PROCHECK-NMR: Programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 1996, 8, 477–486. [Google Scholar] [CrossRef]
- Wiederstein, M.; Sippl, M.J. ProSA-web: Interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 2007, 35, W407–W410. [Google Scholar] [CrossRef] [Green Version]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef] [Green Version]
- Mabiala-Bassiloua, C.G.; Arthus-Cartier, G.; Hannaert, V.; Thérisod, H.; Sygusch, J.; Thérisod, M. Mannitol bis-phosphate based inhibitors of fructose 1,6-bisphosphate aldolases. ACS Med. Chem. Lett. 2011, 2, 804–808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dalby, A.; Dauter, Z.; Littlechild, J.A. Crystal structure of human muscle aldolase complexed with fructose-1,6-bisphosphate: Mechanistic implications. Protein Sci. 1999, 8, 291–297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roslan, H.A.; Hossain, M.A.; Gerunsin, J. Molecular and 3D-structural characterization of fructose-1,6-bisphosphate aldolase derived from Metroxlon sagu. Braz. Arch. Biol. Technol. 2017, 60, 1–21. [Google Scholar] [CrossRef] [Green Version]
- Ondrovics, M.; Gasser, R.B.; Joachim, A. Recent Advances in Elucidating Nematode Moulting–Prospects of Using Oesophagostomum dentatum as a model. Adv. Parasitol. 2016, 91, 233–264. [Google Scholar] [PubMed]
- Kochman, M.; Kwiatkowska, D. Purification and properties of fructose diphosphate aldolase from Ascaris suum muscle. Arch. Biochem. Biophys. 1972, 152, 856–868. [Google Scholar] [CrossRef]
- Rhodes, M.B. Haemonchus contortus: Enzymes. II. Fructose diphosphate aldolase. Exp. Parasitol. 1972, 31, 332–340. [Google Scholar] [CrossRef]
- Sun, Z.; Shen, B.; Wu, H.; Zhou, X.; Wang, Q.; Xiao, J.; Zhang, Y. The secreted fructose-1,6-bisphosphate aldolase as a broad-spectrum vaccine candidate against pathogenic bacteria in aquaculture. Fish Shellfish Immunol. 2015, 46, 638–647. [Google Scholar] [CrossRef]






Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Umair, S.; Bouchet, C.; Palevich, N.; Simpson, H. Teladorsagia circumcincta 1,6-Bisphosphate Aldolase: Molecular and Biochemical Characterisation, Structure Analysis and Recognition by Immune Hosts. Parasitologia 2021, 1, 1-11. https://0-doi-org.brum.beds.ac.uk/10.3390/parasitologia1010001
Umair S, Bouchet C, Palevich N, Simpson H. Teladorsagia circumcincta 1,6-Bisphosphate Aldolase: Molecular and Biochemical Characterisation, Structure Analysis and Recognition by Immune Hosts. Parasitologia. 2021; 1(1):1-11. https://0-doi-org.brum.beds.ac.uk/10.3390/parasitologia1010001
Chicago/Turabian StyleUmair, Saleh, Charlotte Bouchet, Nikola Palevich, and Heather Simpson. 2021. "Teladorsagia circumcincta 1,6-Bisphosphate Aldolase: Molecular and Biochemical Characterisation, Structure Analysis and Recognition by Immune Hosts" Parasitologia 1, no. 1: 1-11. https://0-doi-org.brum.beds.ac.uk/10.3390/parasitologia1010001





