Circular and Micro RNAs from Arabidopsis thaliana Flowers Are Simultaneously Isolated from AGO-IP Libraries
Abstract
:1. Introduction
2. Results
2.1. Identification of circRNAs in AGO-IP Libraries
2.2. circRNAs with miRNA Binding Sites
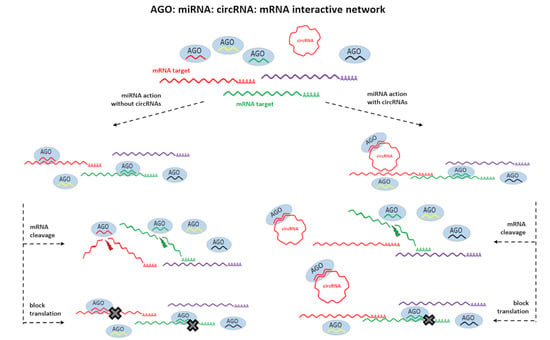
2.3. The circRNAs Harbor Reverse Complementary Sequences of miRNAs which Targeted mRNAs Present in AGO-IP Libraries
2.4. circRNAs Validation by RT-PCR and Sequencing
3. Discussion
4. Materials and Methods
4.1. mRNAseq and Small RNAs Libraries
4.2. circRNAs Identification in mRNAseq Libraries from AGO-IP
4.3. Analysis of Target mRNAs and miRNAs Counts in AGO-IP and Control Libraries
4.4. Plant Material and Growth Condition
4.5. RNA Extraction, RNase R Treatment and cDNA Synthesis
4.6. Primers Design
4.7. circRNAs and Parental mRNAs Amplification and Sequencing
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tay, Y.; Rinn, J.; Pandolfi, P.P. The multilayered complexity of ceRNA crosstalk and competition. Nature 2014, 505, 344–352. [Google Scholar] [CrossRef] [Green Version]
- Chi, S.W.; Zang, J.B.; Mele, A.; Darnell, R.B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 2009, 460, 479–486. [Google Scholar] [CrossRef]
- Licatalosi, D.D.; Mele, A.; Fak, J.J.; Ule, J.; Kayikci, M.; Chi, S.W.; Clark, T.A.; Schweitzer, A.C.; Blume, J.E.; Wang, X.; et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 2008, 456, 464–469. [Google Scholar] [CrossRef] [Green Version]
- Huntzinger, E.; Izaurralde, E. Gene silencing by microRNAs: Contributions of translational repression and mRNA decay. Nat. Rev. Genet. 2011, 12, 99–110. [Google Scholar] [CrossRef]
- Pasquinelli, A.E. MicroRNAs and their targets: Recognition, regulation and an emerging reciprocal relationship. Nat. Rev. Genet. 2012, 13, 271–282. [Google Scholar] [CrossRef]
- Poliseno, L.; Salmena, L.; Zhang, J.; Carver, B.; Haveman, W.J.; Pandolfi, P.P. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 2010, 465, 1033–1038. [Google Scholar] [CrossRef] [Green Version]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA Hypothesis: The Rosetta Stone of a Hidden RNA Language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [Green Version]
- Seitz, H. Redefining MicroRNA Targets. Curr. Biol. 2009, 19, 870–873. [Google Scholar] [CrossRef]
- Tay, Y.; Kats, L.; Salmena, L.; Weiss, D.; Tan, S.M.; Ala, U.; Karreth, F.; Poliseno, L.; Provero, P.; Di Cunto, F.; et al. Coding-Independent Regulation of the Tumor Suppressor PTEN by Competing Endogenous mRNAs. Cell 2011, 147, 344–357. [Google Scholar] [CrossRef] [Green Version]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; García, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef]
- Hansen, T.B.; Wiklund, E.D.; Bramsen, J.B.; Villadsen, S.B.; Statham, A.L.; Clark, S.J.; Kjems, J. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 2011, 30, 4414–4422. [Google Scholar] [CrossRef] [Green Version]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Capel, B.; Swain, A.; Nicolis, S.; Hacker, A.; Walter, M.; Koopman, P.; Goodfellow, P.; Lovell-Badge, R. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 1993, 73, 1019–1030. [Google Scholar] [CrossRef]
- Cocquerelle, C.; Daubersies, P.; Majérus, M.A.; Kerckaert, J.P.; Bailleul, B. Splicing with inverted order of exons occurs proximal to large introns. EMBO J. 1992, 11, 1095–1098. [Google Scholar] [CrossRef]
- Nigro, J.M.; Cho, K.R.; Fearon, E.R.; Kern, S.E.; Ruppert, J.M.; Oliner, J.D.; Kinzler, K.W.; Vogelstein, B. Scrambled exons. Cell 1991, 64, 607–613. [Google Scholar] [CrossRef]
- Kos, A.; Dijkema, R.; Arnberg, A.C.; van der Meide, P.H.; Schellekens, H. The hepatitis delta (delta) virus possesses a circular RNA. Nature 1986, 323, 558–560. [Google Scholar] [CrossRef]
- Sanger, H.L.; Klotz, G.; Riesner, D.; Gross, H.J.; Kleinschmidt, A.K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl. Acad. Sci. USA 1976, 73, 3852–3856. [Google Scholar] [CrossRef]
- Salzman, J.; Chen, R.E.; Olsen, M.N.; Wang, P.L.; Brown, P.O. Cell-Type Specific Features of Circular RNA Expression. PLoS Genet. 2013, 9, e1003777. [Google Scholar] [CrossRef]
- Ye, C.-Y.; Chen, L.; Liu, C.; Zhu, Q.-H.; Fan, L. Widespread noncoding circular RNAs in plants. New Phytol. 2015, 208, 88–95. [Google Scholar] [CrossRef]
- Salzman, J.; Gawad, C.; Wang, P.L.; Lacayo, N.; Brown, P.O. Circular RNAs Are the Predominant Transcript Isoform from Hundreds of Human Genes in Diverse Cell Types. PLoS ONE 2012, 7, e30733. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, X.-O.; Chen, T.; Xiang, J.-F.; Yin, Q.-F.; Xing, Y.-H.; Zhu, S.; Yang, L.; Chen, L.-L. Circular Intronic Long Noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef]
- Zhang, X.-O.; Wang, H.-B.; Zhang, Y.; Lu, X.; Chen, L.-L.; Yang, L. Complementary Sequence-Mediated Exon Circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef] [Green Version]
- Wang, P.L.; Bao, Y.; Yee, M.-C.; Barrett, S.P.; Hogan, G.J.; Olsen, M.N.; Dinneny, J.R.; Brown, P.O.; Salzman, J. Circular RNA Is Expressed across the Eukaryotic Tree of Life. PLoS ONE 2014, 9, e90859. [Google Scholar] [CrossRef]
- Danan, M.; Schwartz, S.; Edelheit, S.; Sorek, R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 2012, 40, 3131–3142. [Google Scholar] [CrossRef]
- Conn, V.M.; Hugouvieux, V.; Nayak, A.; Conos, S.A.; Capovilla, G.; Cildir, G.; Jourdain, A.; Tergaonkar, V.; Schmid, M.; Zubieta, C.; et al. A circRNA from SEPALLATA3 regulates splicing of its cognate mRNA through R-loop formation. Nature Plants, v. 3, n. 5, p. 17053, 18 abr. 2017. cognate mRNA through R-loop formation. Nat. Plants 2017, 3, 17053. [Google Scholar] [CrossRef]
- Darbani, B.; Noeparvar, S.; Borg, S. Identification of Circular RNAs from the Parental Genes Involved in Multiple Aspects of Cellular Metabolism in Barley. Front. Plant Sci. 2016, 7, 776. [Google Scholar] [CrossRef] [Green Version]
- Lu, T.; Cui, L.; Zhou, Y.; Zhu, C.; Fan, D.; Gong, H.; Zhao, Q.; Zhou, C.; Zhao, Y.; Lu, D.; et al. Transcriptome-wide investigation of circular RNAs in rice. RNA 2015, 21, 2076–2087. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Yang, M.; Wei, S.; Qin, F.; Zhao, H.; Suo, B. Identification of Circular RNAs and Their Targets in Leaves of Triticum aestivum L. under Dehydration Stress. Front. Plant Sci. 2017, 7, 2024. [Google Scholar] [CrossRef]
- Zuo, J.; Wang, Q.; Zhu, B.; Luo, Y.; Gao, L. Deciphering the roles of circRNAs on chilling injury in tomato. Biochem. Biophys. Res. Commun. 2016, 479, 132–138. [Google Scholar] [CrossRef]
- Rybak-Wolf, A.; Stottmeister, C.; Glažar, P.; Jens, M.; Pino, N.; Giusti, S.; Hanan, M.; Behm, M.; Bartok, O.; Ashwal-Fluss, R.; et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol. Cell 2015, 58, 870–885. [Google Scholar] [CrossRef] [Green Version]
- Westholm, J.O.; Miura, P.; Olson, S.; Shenker, S.; Joseph, B.; Sanfilippo, P.; Celniker, S.E.; Graveley, B.R.; Lai, E.C. Genome-wide Analysis of Drosophila Circular RNAs Reveals Their Structural and Sequence Properties and Age-Dependent Neural Accumulation. Cell Rep. 2014, 9, 1966–1980. [Google Scholar] [CrossRef] [Green Version]
- Suzuki, H.; Tsukahara, T. A view of pre-mRNA splicing from RNase R resistant RNAs. Int. J. Mol. Sci. 2014, 15, 9331–9342. [Google Scholar] [CrossRef]
- Burd, C.E.; Jeck, W.R.; Liu, Y.; Sanoff, H.K.; Wang, Z.; Sharpless, N.E. Expression of Linear and Novel Circular Forms of an INK4/ARF-Associated Non-Coding RNA Correlates with Atherosclerosis Risk. PLoS Genet. 2010, 6, e1001233. [Google Scholar] [CrossRef]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef]
- Talhouarne, G.J.S.; Gall, J.G. Lariat intronic RNAs in the cytoplasm of Xenopus tropicalis oocytes. RNA 2014, 20, 1476–1487. [Google Scholar] [CrossRef]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef]
- Suzuki, H.; Zuo, Y.; Wang, J.; Zhang, M.Q.; Malhotra, A.; Mayeda, A. Characterization of RNase R-digested cellular RNA source that consists of lariat and circular RNAs from pre-mRNA splicing. Nucleic Acids Res. 2006, 34, e63. [Google Scholar] [CrossRef]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. circRNA Biogenesis Competes with Pre-mRNA Splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef] [Green Version]
- Chao, C.W.; Chan, D.C.; Kuo, A.; Leder, P. The mouse formin (Fmn) gene: Abundant circular RNA transcripts and gene-targeted deletion analysis. Mol. Med. 1998, 4, 614–628. [Google Scholar] [CrossRef]
- Hentze, M.W.; Preiss, T. Circular RNAs: splicing’s enigma variations. EMBO J. 2013, 32, 923–925. [Google Scholar] [CrossRef]
- Romeo, T. Global regulation by the small RNA-binding protein CsrA and the non-coding RNA molecule CsrB. Mol. Microbiol. 1998, 29, 1321–1330. [Google Scholar] [CrossRef]
- Thomas, L.F.; Sætrom, P. Circular RNAs are depleted of polymorphisms at microRNA binding sites. Bioinformatics 2014, 30, 2243–2246. [Google Scholar] [CrossRef]
- Sablok, G.; Zhao, H.; Sun, X. Plant Circular RNAs (circRNAs): Transcriptional Regulation Beyond miRNAs in Plants. Mol. Plant 2016, 9, 192–194. [Google Scholar] [CrossRef] [Green Version]
- Carbonell, A.; Fahlgren, N.; Garcia-Ruiz, H.; Gilbert, K.B.; Montgomery, T.A.; Nguyen, T.; Cuperus, J.T.; Carrington, J.C. Functional Analysis of Three Arabidopsis ARGONAUTES Using Slicer-Defective Mutants. Plant Cell 2012, 24, 3613–3629. [Google Scholar] [CrossRef]
- Voinnet, O.; Ponce, M.R.; Vaucheret, H.; Baumberger, N.; Sarazin, A.; Clavel, M.; Micol, J.L.; Ziegler-Graff, V.; Genschik, P.; Derrien, B.; et al. A Suppressor Screen for AGO1 Degradation by the Viral F-Box P0 Protein Uncovers a Role for AGO DUF1785 in sRNA Duplex Unwinding. Plant Cell 2018, 30, 1353–1374. [Google Scholar] [Green Version]
- Lasda, E.; Parker, R. Circular RNAs: Diversity of form and function. RNA 2014, 20, 1829–1842. [Google Scholar] [CrossRef]
- Li, X.; Yang, L.; Chen, L.-L. The Biogenesis, Functions, and Challenges of Circular RNAs. Mol. Cell 2018, 71, 428–442. [Google Scholar] [CrossRef] [Green Version]
- Chu, Q.; Zhang, X.; Zhu, X.; Liu, C.; Mao, L.; Ye, C.; Zhu, Q.; Fan, L. PlantcircBase: A database for plant circular RNAs. Mol. Plant. 2017, 10, 1126–1128. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Ye, J.; Wang, L.; Li, S.; Zhang, Q.; Zhang, Q.; Tang, W.; Wang, K.; Song, K.; Sablok, G.; Sun, X.; et al. AtCircDB: A tissue-specific database for Arabidopsis circular RNAs. Brief. Bioinform. 2017, 20, 58–65. [Google Scholar] [CrossRef]
- Zhao, W.; Cheng, Y.; Zhang, C.; You, Q.; Shen, X.; Guo, W.; Jiao, Y. Genome-wide identification and characterization of circular RNAs by high throughput sequencing in soybean. Sci. Rep. 2017, 7, 5636. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Gaffo, E.; Bonizzato, A.; Kronnie, G.; Bortoluzzi, S. CirComPara: A Multi-Method Comparative Bioinformatics Pipeline to Detect and Study circRNAs from RNA-seq Data. Non Coding RNA 2017, 3, 8. [Google Scholar] [CrossRef]
- Dai, X.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server. Nucleic Acids Res. 2011, 39, W155–W159. [Google Scholar] [CrossRef]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, D68–D73. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Rozen, S.; Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 2000, 132, 365–386. [Google Scholar]
- Kulcheski, F.R.; Christoff, A.P.; Margis, R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J. Biotechnol. 2016, 238, 42–51. [Google Scholar] [CrossRef]

 ) to amplify the circRNA_At3g13990. Convergent primers (
) to amplify the circRNA_At3g13990. Convergent primers (  ) were used to amplify parental mRNA. Genomic DNA (gDNA) was used as control. Samples were analyzed on 1,5% agarose gel. (M) DNA size marker of 100 bp; cDNA: complementary DNA; cDNA*: complementary DNA produced from total RNA treated with RNase R previously to reverse transcription. bp: base pairs.
) were used to amplify parental mRNA. Genomic DNA (gDNA) was used as control. Samples were analyzed on 1,5% agarose gel. (M) DNA size marker of 100 bp; cDNA: complementary DNA; cDNA*: complementary DNA produced from total RNA treated with RNase R previously to reverse transcription. bp: base pairs.
 ) to amplify the circRNA_At3g13990. Convergent primers (
) to amplify the circRNA_At3g13990. Convergent primers (  ) were used to amplify parental mRNA. Genomic DNA (gDNA) was used as control. Samples were analyzed on 1,5% agarose gel. (M) DNA size marker of 100 bp; cDNA: complementary DNA; cDNA*: complementary DNA produced from total RNA treated with RNase R previously to reverse transcription. bp: base pairs.
) were used to amplify parental mRNA. Genomic DNA (gDNA) was used as control. Samples were analyzed on 1,5% agarose gel. (M) DNA size marker of 100 bp; cDNA: complementary DNA; cDNA*: complementary DNA produced from total RNA treated with RNase R previously to reverse transcription. bp: base pairs.

| Identification Method | CircExplorer2 | FindCirc | TestRealign | |||
|---|---|---|---|---|---|---|
| Segemehl | Star | Tophat | - | - | ||
| CircExplorer2 | Segemehl | 86 | 9 | 12 | 10 | 26 |
| Star | - | 15 | 7 | 3 | 3 | |
| Tophat | - | - | 23 | 7 | 7 | |
| FindCirc | - | - | - | 1422 | 198 | |
| TestRealign | - | - | - | - | 27,812 | |
| Library | Gene_ID | Circ_ID | Parental Gene Function | Origin | Exons | Length (nt) *** | Methods |
|---|---|---|---|---|---|---|---|
| DDH-IP | At1g02560 | circ_At1g02560 | Nuclear encoded CLP protease 5 | exonic | 2 | 123 | 5 |
| DDH-IP | At1g12080 ** | circ_At1g12080 | Vacuolar calcium-binding protein-related | exonic * | 1 | 95 | 4 |
| DDH-IP | At1g31810 | circ_At1g31810 | Formin Homology 14 | exonic | 1 | 50 | 3 |
| DDH-IP | At1g52360 | circ_At1g52360 | Coatomer beta subunit | intronic | 1 | 224 | 3 |
| DDH-IP | At2g02410 | circ_At2g02410 | K06962—uncharacterized protein (K06962) | exonic | 1 | 71 | 4 |
| DDH-IP | At2g35940 ** | circ_At2g35940 | BEL1-like homeodomain 1 | exonic | 1 | 930 | 4 |
| DDH-IP | At2g42170 ** | circ_At2g42170 | Actin family protein | exonic | 4 | 1063 | 5 |
| DDH-IP | At5g16880 | circ_At5g16880 | Target of Myb protein 1 | exonic * | 1 | 49 | 4 |
| DDH-IP | At5g56950 | circ_At5g56950 | NAP-1 Nucleosome assembly protein | intronic * | 1 | 118 | 4 |
| DAH-IP | At3g01800 | circ_At3g01800 | Ribosome recycling factor | exonic | 1 | 68 | 4 |
| DAH-IP | At3g13990 ** | circ_At3g13990 | Kinase-related protein (DUF1296) | exonic | 3 | 349 | 3 |
| DAH-IP | At5g27720 ** | circ_At5g27720 | Small nuclear ribonucleoprotein family protein | exonic | 4 | 321 | 5 |
| circRNA | circRNA Read Counts *** | miRNA | miRNA Read Counts | Inhibiton By | |||
|---|---|---|---|---|---|---|---|
| AGO-IP | Total RNA | AGO1-DDH | AGO1-DAH | Empty Vector | |||
| circ_At1g12080 ** | 175 | 13 | miR4221-5p * | 265 | 171 | 7 | Cleavage |
| miR838-3p * | 226 | 59 | 41 | Translation | |||
| - | - | - | miR397a-5p * | 1014 | 520 | 29 | Translation |
| - | - | - | miR5654-3p | 2 | 0 | 0 | Cleavage |
| circ_At2g35940 | 8 | 0 | miR8182-5p * | 2 | 9 | 0 | Translation |
| - | - | - | miR830-3p * | 29 | 13 | 0 | Cleavage |
| - | - | - | miR833a-5p * | 50 | 35 | 17 | Translation |
| - | - | - | miR8174-3p | - | - | - | Cleavage |
| circ_At2g42170 | 8 | 0 | miR831-3p * | 81 | 11 | 3 | Translation |
| - | - | - | miR838-3p * | 226 | 59 | 41 | Cleavage |
| - | - | - | miR4239-5p * | 17 | 7 | 0 | Translation |
| circ_At3g13990 ** | 17 | 0 | miR5637-5p | - | - | - | Cleavage |
| - | - | - | miR780.2-3p | - | - | - | Cleavage |
| circ_At5g27720 | 17 | 0 | miR838-3p * | 226 | 59 | 41 | Cleavage |
| Target_Access | miRNA | Expectation | Inhibition By | Lenght | Target Counts * | Function | |
|---|---|---|---|---|---|---|---|
| AgoIP | Input | ||||||
| At2g38080.1 | miR397a-5p | 1 | Cleavage | 2021 | 58 | 21 | Laccase/Diphenol oxidase |
| At5g60020.1 | 1 | Cleavage | 2049 | 33 | 18 | Laccase 17 | |
| At3g06040.1 | 3 | Cleavage | 864 | 29 | 14 | Ribosomal protein L12 | |
| At3g06470.1 | 3 | Cleavage | 1092 | 75 | 4 | GNS1/SUR4 membrane protein | |
| At3g54170.1 | miR4221-5p | 2.5 | Cleavage | 1262 | 22 | 10 | FKBP12 interacting protein 37 |
| At4g13070.1 | 2.5 | Cleavage | 1775 | 8 | 2 | RNA-binding CRS1 | |
| At5g60040.1 | 2.5 | Cleavage | 4582 | 62 | 22 | Nuclear RNA polymerase C1 | |
| At1g13350.1 | 3 | Cleavage | 2454 | 142 | 24 | Protein kinase | |
| At1g77660.1 | 3 | Cleavage | 1765 | 22 | 12 | H3K4-specific methyltransferase | |
| At2g33240.1 | 3 | Cleavage | 5313 | 36 | 12 | Myosin XI D | |
| At3g02170.1 | 3 | Cleavage | 3300 | 319 | 155 | Longifolia2 | |
| At4g14510.1 | 3 | Cleavage | 2940 | 57 | 22 | CRM family member 3B | |
| At1g31650.1 | 3 | Translation | 2255 | 164 | 28 | RHO guanyl- exchange factor 14 | |
| At2g38610.1 | 3 | Translation | 1452 | 56 | 26 | RNA-binding KH protein | |
| At2g35160.1 | miR8182-5p | 3 | Cleavage | 2798 | 20 | 9 | SU(VAR)3-9 homolog 5 |
| At4g22580.1 | 3 | Cleavage | 1628 | 39 | 10 | Exostosin family protein | |
| At1g23400.1 | 3 | Cleavage | 1822 | 81 | 24 | RNA-binding CRS1 | |
| At1g49880.1 | miR831-3p | 2.5 | Translation | 803 | 50 | 2 | FAD-linked sulfhydryl oxidase |
| At3g46060.1 | 3 | Translation | 1132 | 75 | 41 | RAS-related protein RABE1C | |
| At2g36890.1 | miR833a-5p | 2.5 | Cleavage | 971 | 6 | 1 | Myb-like DNA-binding domain |
| At3g12380.1 | miR838-3p | 2.5 | Cleavage | 2323 | 33 | 16 | Actin-related protein 5 |
| At1g21740.1 | 3 | Cleavage | 2862 | 63 | 22 | Protein of unknown function | |
| At1g64180.1 | 3 | Cleavage | 2072 | 13 | 3 | Intracellular transport protein | |
| At1g70470.1 | 3 | Cleavage | 765 | 17 | 4 | No annotated domains | |
| At4g01080.1 | 3 | Cleavage | 1583 | 98 | 33 | Trichome-birefringence like 26 | |
| At5g09460.1 | 3 | Cleavage | 2546 | 124 | 41 | Transcription Factor SAC51 | |
| At5g09461.1 | 3 | Cleavage | 2546 | 124 | 41 | Conserved peptide upstream ORF | |
| At5g20110.1 | 3 | Cleavage | 778 | 28 | 2 | Dynein light chain type 1 | |
| At5g46030.1 | 2 | Translation | 732 | 26 | 12 | Elongation factor Elf1 like | |
| At2g44430.1 | 2.5 | Translation | 2196 | 98 | 19 | DNA-binding protein | |
| At5g22640.1 | 2.5 | Translation | 2814 | 247 | 115 | MORN repeat-containing protein | |
| At5g40340.1 | 2.5 | Translation | 3096 | 624 | 75 | Tudor/PWWP/MBT protein | |
| At5g56210.1 | 2.5 | Translation | 2004 | 22 | 5 | WPP domain interacting protein 2 | |
| At5g62390.1 | 2.5 | Translation | 1859 | 349 | 152 | BCL-2-associated athanogene 7 | |
| At5g17910.1 | 3 | Translation | 4532 | 178 | 77 | No annotated domains | |
| At5g41960.1 | 3 | Translation | 874 | 9 | 4 | No annotated domains | |
| At5g57790.1 | 3 | Translation | 1407 | 29 | 12 | No annotated domains | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Frydrych Capelari, É.; da Fonseca, G.C.; Guzman, F.; Margis, R. Circular and Micro RNAs from Arabidopsis thaliana Flowers Are Simultaneously Isolated from AGO-IP Libraries. Plants 2019, 8, 302. https://0-doi-org.brum.beds.ac.uk/10.3390/plants8090302
Frydrych Capelari É, da Fonseca GC, Guzman F, Margis R. Circular and Micro RNAs from Arabidopsis thaliana Flowers Are Simultaneously Isolated from AGO-IP Libraries. Plants. 2019; 8(9):302. https://0-doi-org.brum.beds.ac.uk/10.3390/plants8090302
Chicago/Turabian StyleFrydrych Capelari, Érika, Guilherme Cordenonsi da Fonseca, Frank Guzman, and Rogerio Margis. 2019. "Circular and Micro RNAs from Arabidopsis thaliana Flowers Are Simultaneously Isolated from AGO-IP Libraries" Plants 8, no. 9: 302. https://0-doi-org.brum.beds.ac.uk/10.3390/plants8090302