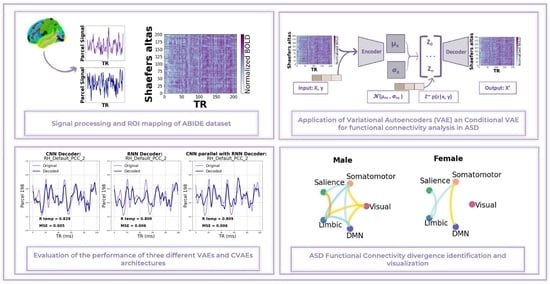
Conditional Variational Autoencoder for Functional Connectivity Analysis of Autism Spectrum Disorder Functional Magnetic Resonance Imaging Data: A Comparative Study
Abstract
:1. Introduction
2. Related Works
2.1. Traditional Approaches to FC Analysis
2.2. Application of VAEs in fMRI Domain
2.3. Application of CVAEs in fMRI Domain
2.4. Functional Connectivity in ASD
3. Materials and Methods
3.1. Dataset
3.2. Data Preprocessing
3.3. Variational Autoencoder (VAE)
3.4. Conditional VAE
3.5. Experimental Setup
3.6. VAE Performance Evaluations
3.7. Functional Connectivity Analysis
4. Results
4.1. VAE Performance Evaluations
4.2. Functional Connectivity Analysis
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ASD | Autism Spectrum Disorder |
| TD | Typically Developing |
| FC | Functional Connectivity |
| fMRI | functional Magnetic Resonance Imaging |
| BOLD | Blood-Oxygen-Level-Dependent |
| CNN | Convolutional Neural Network |
| RNN | Recurrent Neural Network |
| VAE | Variational Autoencoder |
| CVAE | Conditional Variational Autoencoder |
| DMN | Default Mode Network |
| PCC | Pearson’s Correlation Coefficient |
References
- Lord, C.; Elsabbagh, M.; Baird, G.; Veenstra-Vanderweele, J. Autism spectrum disorder. Lancet 2018, 392, 508–520. [Google Scholar] [CrossRef] [PubMed]
- Loomes, R.; Hull, L.; Mandy, W.P.L. What Is the Male-to-Female Ratio in Autism Spectrum Disorder? A Systematic Review and Meta-Analysis. J. Am. Acad. Child Adolesc. Psychiatry 2017, 56, 466–474. [Google Scholar] [CrossRef] [PubMed]
- Biswal, B.; Zerrin Yetkin, F.; Haughton, V.M.; Hyde, J.S. Functional connectivity in the motor cortex of resting human brain using echo-planar MRI. Magn. Reson. Med. 1995, 34, 537–541. [Google Scholar] [CrossRef] [PubMed]
- Hull, J.V.; Dokovna, L.B.; Jacokes, Z.J.; Torgerson, C.M.; Irimia, A.; Van Horn, J.D. Resting-state functional connectivity in autism spectrum disorders: A review. Front. Psychiatry 2017, 7, 205. [Google Scholar] [CrossRef] [PubMed]
- Just, M.A.; Cherkassky, V.L.; Keller, T.A.; Kana, R.K.; Minshew, N.J. Functional and anatomical cortical underconnectivity in autism: Evidence from an FMRI study of an executive function task and corpus callosum morphometry. Cereb. Cortex 2007, 17, 951–961. [Google Scholar] [CrossRef]
- Courchesne, E.; Pierce, K. Why the frontal cortex in autism might be talking only to itself: Local over-connectivity but long-distance disconnection. Curr. Opin. Neurobiol. 2005, 15, 225–230. [Google Scholar] [CrossRef]
- Cherkassky, V.L.; Kana, R.K.; Keller, T.A.; Just, M.A. Functional connectivity in a baseline resting-state network in autism. Neuroreport 2006, 17, 1687–1690. [Google Scholar] [CrossRef]
- Calhoun, V.D.; Liu, J.; Adalı, T. A review of group ICA for fMRI data and ICA for joint inference of imaging, genetic, and ERP data. Neuroimage 2009, 45, S163–S172. [Google Scholar] [CrossRef]
- Farahani, F.V.; Karwowski, W.; Lighthall, N.R. Application of graph theory for identifying connectivity patterns in human brain networks: A systematic review. Front. Neurosci. 2019, 13, 585. [Google Scholar] [CrossRef]
- Chen, G.; Ward, B.D.; Xie, C.; Li, W.; Chen, G.; Goveas, J.S.; Antuono, P.G.; Li, S.J. A clustering-based method to detect functional connectivity differences. NeuroImage 2012, 61, 56–61. [Google Scholar] [CrossRef]
- Hutchison, R.M.; Womelsdorf, T.; Allen, E.A.; Bandettini, P.A.; Calhoun, V.D.; Corbetta, M.; Della Penna, S.; Duyn, J.H.; Glover, G.H.; Gonzalez-Castillo, J.; et al. Dynamic functional connectivity: Promise, issues, and interpretations. Neuroimage 2013, 80, 360–378. [Google Scholar] [CrossRef] [PubMed]
- Havlicek, M.; Jan, J.; Brazdil, M.; Calhoun, V.D. Dynamic Granger causality based on Kalman filter for evaluation of functional network connectivity in fMRI data. Neuroimage 2010, 53, 65–77. [Google Scholar] [CrossRef] [PubMed]
- Stephan, K.E.; Kasper, L.; Harrison, L.M.; Daunizeau, J.; den Ouden, H.E.; Breakspear, M.; Friston, K.J. Nonlinear Dynamic Causal Models for fMRI. Neuroimage 2008, 42, 649–662. [Google Scholar] [CrossRef] [PubMed]
- Kingma, D.P.; Welling, M. Auto-Encoding Variational Bayes. arXiv 2022, arXiv:1312.6114. [Google Scholar] [CrossRef]
- Sohn, K.; Lee, H.; Yan, X. Learning Structured Output Representation using Deep Conditional Generative Models. In Advances in Neural Information Processing Systems; Cortes, C., Lawrence, N., Lee, D., Sugiyama, M., Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2015; Volume 28. [Google Scholar]
- Lee, M.H.; Smyser, C.D.; Shimony, J.S. Resting-state fMRI: A review of methods and clinical applications. Am. J. Neuroradiol. 2013, 34, 1866–1872. [Google Scholar] [CrossRef] [PubMed]
- Neha; Gandhi, T.K. Resting state fMRI analysis using seed based and ICA methods. In Proceedings of the 2016 3rd International Conference on Computing for Sustainable Global Development (INDIACom), New Delhi, India, 16–18 March 2016; pp. 2551–2554. [Google Scholar]
- Erhardt, E.B.; Rachakonda, S.; Bedrick, E.J.; Allen, E.A.; Adali, T.; Calhoun, V.D. Comparison of multi-subject ICA methods for analysis of fMRI data. Hum. Brain Mapp. 2011, 32, 2075–2095. [Google Scholar] [CrossRef]
- Almuqhim, F.; Saeed, F. ASD-SAENet: A sparse autoencoder, and deep-neural network model for detecting autism spectrum disorder (ASD) using fMRI data. Front. Comput. Neurosci. 2021, 15, 654315. [Google Scholar] [CrossRef]
- Kang, L.; Chen, J.; Huang, J.; Jiang, J. Autism spectrum disorder recognition based on multi-view ensemble learning with multi-site fMRI. Cogn. Neurodyn. 2023, 17, 345–355. [Google Scholar] [CrossRef]
- Qiang, N.; Dong, Q.; Liang, H.; Ge, B.; Zhang, S.; Sun, Y.; Zhang, C.; Zhang, W.; Gao, J.; Liu, T. Modeling and augmenting of fMRI data using deep recurrent variational auto-encoder. J. Neural Eng. 2021, 18, 0460b6. [Google Scholar] [CrossRef]
- Kim, J.H.; Zhang, Y.; Han, K.; Wen, Z.; Choi, M.; Liu, Z. Representation learning of resting state fMRI with variational autoencoder. NeuroImage 2021, 241, 118423. [Google Scholar] [CrossRef]
- Huang, H.; Hu, X.; Zhao, Y.; Makkie, M.; Dong, Q.; Zhao, S.; Guo, L.; Liu, T. Modeling task fMRI data via deep convolutional autoencoder. IEEE Trans. Med. Imaging 2017, 37, 1551–1561. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Q.; Zhu, Y.; Lu, L.; Yang, Z.; Li, Y.; Zhang, N. Fusing Structural and Functional Connectivities using Disentangled VAE for Detecting MCI. arXiv 2023, arXiv:2306.09629. [Google Scholar]
- Choi, H. Functional connectivity patterns of autism spectrum disorder identified by deep feature learning. arXiv 2017, arXiv:1707.07932. [Google Scholar] [CrossRef]
- Zhuang, P.; Schwing, A.G.; Koyejo, O. fMRI Data Augmentation Via Synthesis. In Proceedings of the 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), Venice, Italy, 8–11 April 2019; IEEE: Piscataway, NJ, USA, 2019; pp. 1783–1787. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, X.; Wa, S.; Liu, Y.; Kang, J.; Lv, C. GenU-Net++: An Automatic Intracranial Brain Tumors Segmentation Algorithm on 3D Image Series with High Performance. Symmetry 2021, 13, 2395. [Google Scholar] [CrossRef]
- Tashiro, T.; Matsubara, T.; Uehara, K. Deep neural generative model for fMRI image based diagnosis of mental disorder. IEICE Proc. Ser. 2017, 29, 700–703. [Google Scholar] [CrossRef]
- Zou, A.; Ji, J. Learning brain effective connectivity networks via controllable variational autoencoder. In Proceedings of the 2021 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Houston, TX, USA, 9–12 December 2021; pp. 284–287. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, K.; Zhou, R.; Leow, A.; Osorio, R.; Zhang, Y.; He, L. Normative Modeling via Conditional Variational Autoencoder and Adversarial Learning to Identify Brain Dysfunction in Alzheimer’s Disease. arXiv 2022, arXiv:2211.08982. [Google Scholar] [CrossRef]
- Gao, M.S.; Tsai, F.S.; Lee, C.C. Learning a Phenotypic-Attribute Attentional Brain Connectivity Embedding for ADHD Classification using rs-fMRI. In Proceedings of the 2020 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), Montreal, QC, Canada, 20–24 July 2020; pp. 5472–5475. [Google Scholar] [CrossRef]
- Broyd, S.J.; Demanuele, C.; Debener, S.; Helps, S.K.; James, C.J.; Sonuga-Barke, E.J. Default-mode brain dysfunction in mental disorders: A systematic review. Neurosci. Biobehav. Rev. 2009, 33, 279–296. [Google Scholar] [CrossRef]
- Rajmohan, V.; Mohandas, E. The Limbic System. Indian J. Psychiatry 2007, 49, 132. [Google Scholar] [CrossRef]
- Rolls, E.T. The cingulate cortex and limbic systems for emotion, action, and memory. Brain Struct. Funct. 2019, 224, 3001–3018. [Google Scholar] [CrossRef]
- Yang, Y.L.; Deng, H.X.; Xing, G.Y.; Xia, X.L.; Li, H.F. Brain functional network connectivity based on a visual task: Visual information processing-related brain regions are significantly activated in the task state. Neural Regen. Res. 2015, 10, 298–307. [Google Scholar] [CrossRef]
- Menon, V.; Uddin, L.Q. Saliency, switching, attention and control: A network model of insula function. Brain Struct. Funct. 2010, 214, 655–667. [Google Scholar] [CrossRef]
- Chand, G.B.; Dhamala, M. The salience network dynamics in perceptual decision-making. Neuroimage 2016, 134, 85–93. [Google Scholar] [CrossRef]
- Abrams, D.A.; Lynch, C.J.; Cheng, K.M.; Phillips, J.; Supekar, K.; Ryali, S.; Uddin, L.Q.; Menon, V. Underconnectivity between voice-selective cortex and reward circuitry in children with autism. Proc. Natl. Acad. Sci. USA 2013, 110, 12060–12065. [Google Scholar] [CrossRef] [PubMed]
- Lynch, C.J.; Uddin, L.Q.; Supekar, K.; Khouzam, A.; Phillips, J.; Menon, V. Default mode network in childhood autism: Posteromedial cortex heterogeneity and relationship with social deficits. Biol. Psychiatry 2013, 74, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Yerys, B.E.; Gordon, E.M.; Abrams, D.N.; Satterthwaite, T.D.; Weinblatt, R.; Jankowski, K.F.; Strang, J.; Kenworthy, L.; Gaillard, W.D.; Vaidya, C.J. Default mode network segregation and social deficits in autism spectrum disorder: Evidence from non-medicated children. NeuroImage Clin. 2015, 9, 223–232. [Google Scholar] [CrossRef] [PubMed]
- Buch, A.M.; Vértes, P.E.; Seidlitz, J.; Kim, S.H.; Grosenick, L.; Liston, C. Molecular and network-level mechanisms explaining individual differences in autism spectrum disorder. Nat. Neurosci. 2023, 26, 650–663. [Google Scholar] [CrossRef]
- Green, S.A.; Hernandez, L.; Bookheimer, S.Y.; Dapretto, M. Salience network connectivity in autism is related to brain and behavioral markers of sensory overresponsivity. J. Am. Acad. Child Adolesc. Psychiatry 2016, 55, 618–626. [Google Scholar] [CrossRef]
- Ypma, R.J.; Moseley, R.L.; Holt, R.J.; Rughooputh, N.; Floris, D.L.; Chura, L.R.; Spencer, M.D.; Baron-Cohen, S.; Suckling, J.; Bullmore, E.T.; et al. Default Mode Hypoconnectivity Underlies a Sex-Related Autism Spectrum. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 2016, 1, 364–371. [Google Scholar] [CrossRef]
- Lawrence, K.E.; Hernandez, L.M.; Bowman, H.C.; Padgaonkar, N.T.; Fuster, E.; Jack, A.; Aylward, E.; Gaab, N.; Van Horn, J.D.; Bernier, R.A.; et al. Sex differences in functional connectivity of the salience, default mode, and central executive networks in youth with ASD. Cereb. Cortex 2020, 30, 5107–5120. [Google Scholar] [CrossRef]
- Smith, R.E.; Avery, J.A.; Wallace, G.L.; Kenworthy, L.; Gotts, S.J.; Martin, A. Sex differences in resting-state functional connectivity of the cerebellum in autism spectrum disorder. Front. Hum. Neurosci. 2019, 13, 104. [Google Scholar] [CrossRef]
- Craddock, C.; Benhajali, Y.; Chu, C.; Chouinard, F.; Evans, A.; Jakab, A.; Khundrakpam, B.S.; Lewis, J.D.; Li, Q.; Milham, M.; et al. The neuro bureau preprocessing initiative: Open sharing of preprocessed neuroimaging data and derivatives. Front. Neuroinform. 2013, 7, 27. [Google Scholar]
- Schaefer, A.; Kong, R.; Gordon, E.M.; Laumann, T.O.; Zuo, X.N.; Holmes, A.J.; Eickhoff, S.B.; Yeo, B.T. Local-global parcellation of the human cerebral cortex from intrinsic functional connectivity MRI. Cereb. Cortex 2018, 28, 3095–3114. [Google Scholar] [CrossRef]
- Rumelhart, D.E.; Hinton, G.E.; Williams, R.J. Learning Internal Representations by Error Propagation. Parallel Distrib. Process. 1985, 1, 318–362. [Google Scholar]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In Advances in Neural Information Processing Systems 32; Curran Associates, Inc.: Red Hook, NY, USA, 2019; pp. 8024–8035. [Google Scholar]
- Xiao, Z.; Kreis, K.; Vahdat, A. Tackling the generative learning trilemma with denoising diffusion gans. arXiv 2021, arXiv:2112.07804. [Google Scholar] [CrossRef]









| Model | Cosine Similarity | PCC | L1 TD | L1 ASD |
|---|---|---|---|---|
| CNN | 0.9930 | 0.6551 | 0.0693 | 0.0781 |
| RNN | 0.9817 | 0.6105 | 0.0728 | 0.0819 |
| CNN and RNN | 0.9820 | 0.6356 | 0.0717 | 0.0803 |
| Model | Cosine Similarity | PCC | L1 ASD | L1 TD |
|---|---|---|---|---|
| Conditional CNN | 0.9961 | 0.7165 | 0.0643 | 0.0733 |
| Conditional RNN | 0.9818 | 0.6382 | 0.0681 | 0.077 |
| Conditional CNN and RNN | 0.9825 | 0.6558 | 0.0687 | 0.0778 |
| Model Architecture | Unconditional FC Similarity | Conditional FC Similarity |
|---|---|---|
| CNN | 0.35 | 0.70 |
| RNN | 0.66 | 0.80 |
| CNN parallel with RNN | 0.78 | 0.85 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sidulova, M.; Park, C.H. Conditional Variational Autoencoder for Functional Connectivity Analysis of Autism Spectrum Disorder Functional Magnetic Resonance Imaging Data: A Comparative Study. Bioengineering 2023, 10, 1209. https://0-doi-org.brum.beds.ac.uk/10.3390/bioengineering10101209
Sidulova M, Park CH. Conditional Variational Autoencoder for Functional Connectivity Analysis of Autism Spectrum Disorder Functional Magnetic Resonance Imaging Data: A Comparative Study. Bioengineering. 2023; 10(10):1209. https://0-doi-org.brum.beds.ac.uk/10.3390/bioengineering10101209
Chicago/Turabian StyleSidulova, Mariia, and Chung Hyuk Park. 2023. "Conditional Variational Autoencoder for Functional Connectivity Analysis of Autism Spectrum Disorder Functional Magnetic Resonance Imaging Data: A Comparative Study" Bioengineering 10, no. 10: 1209. https://0-doi-org.brum.beds.ac.uk/10.3390/bioengineering10101209