Unique Biological Activity and Potential Role of Monomeric Laminin-γ2 as a Novel Biomarker for Hepatocellular Carcinoma: A Review
Abstract
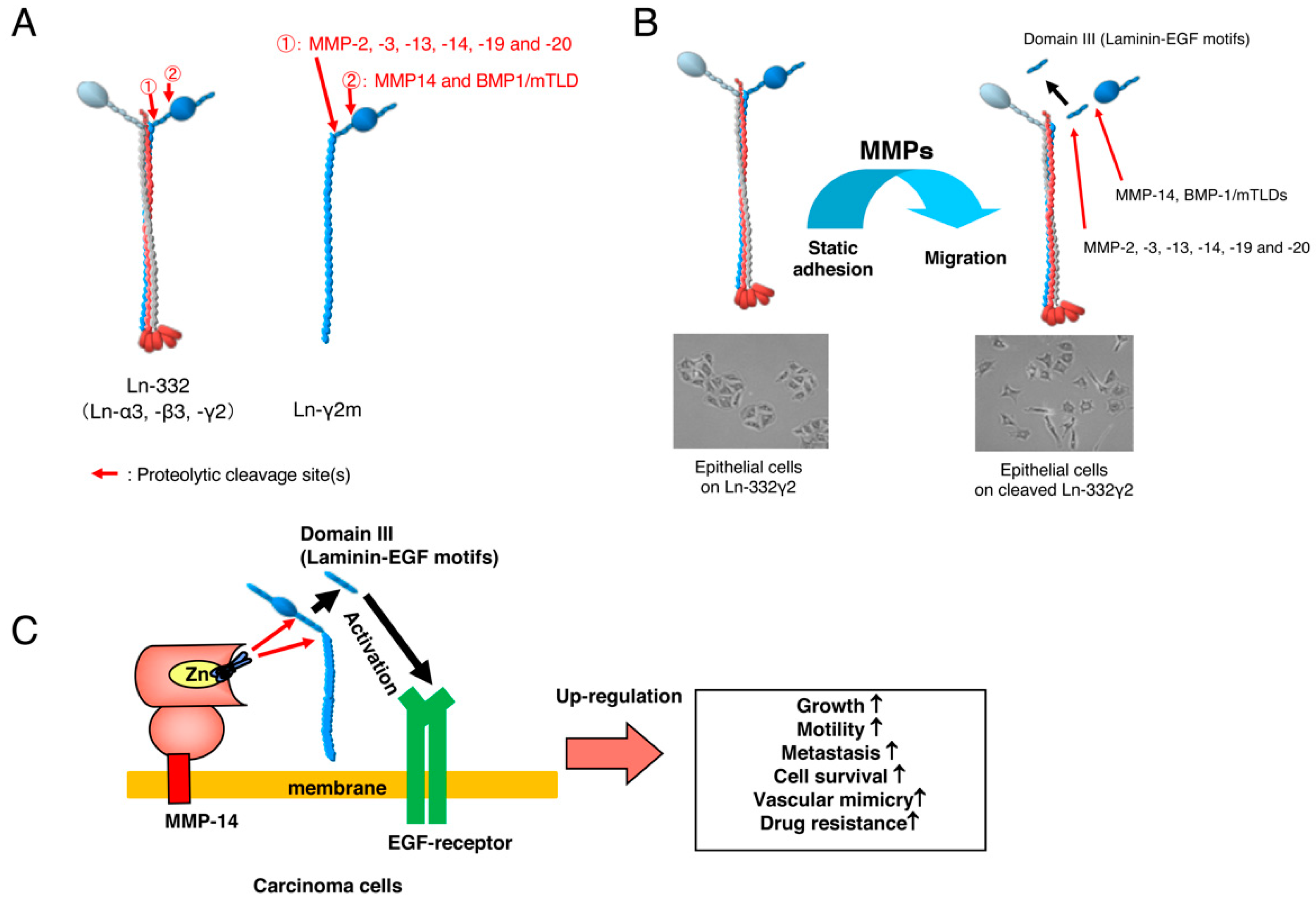
:1. Laminin-332
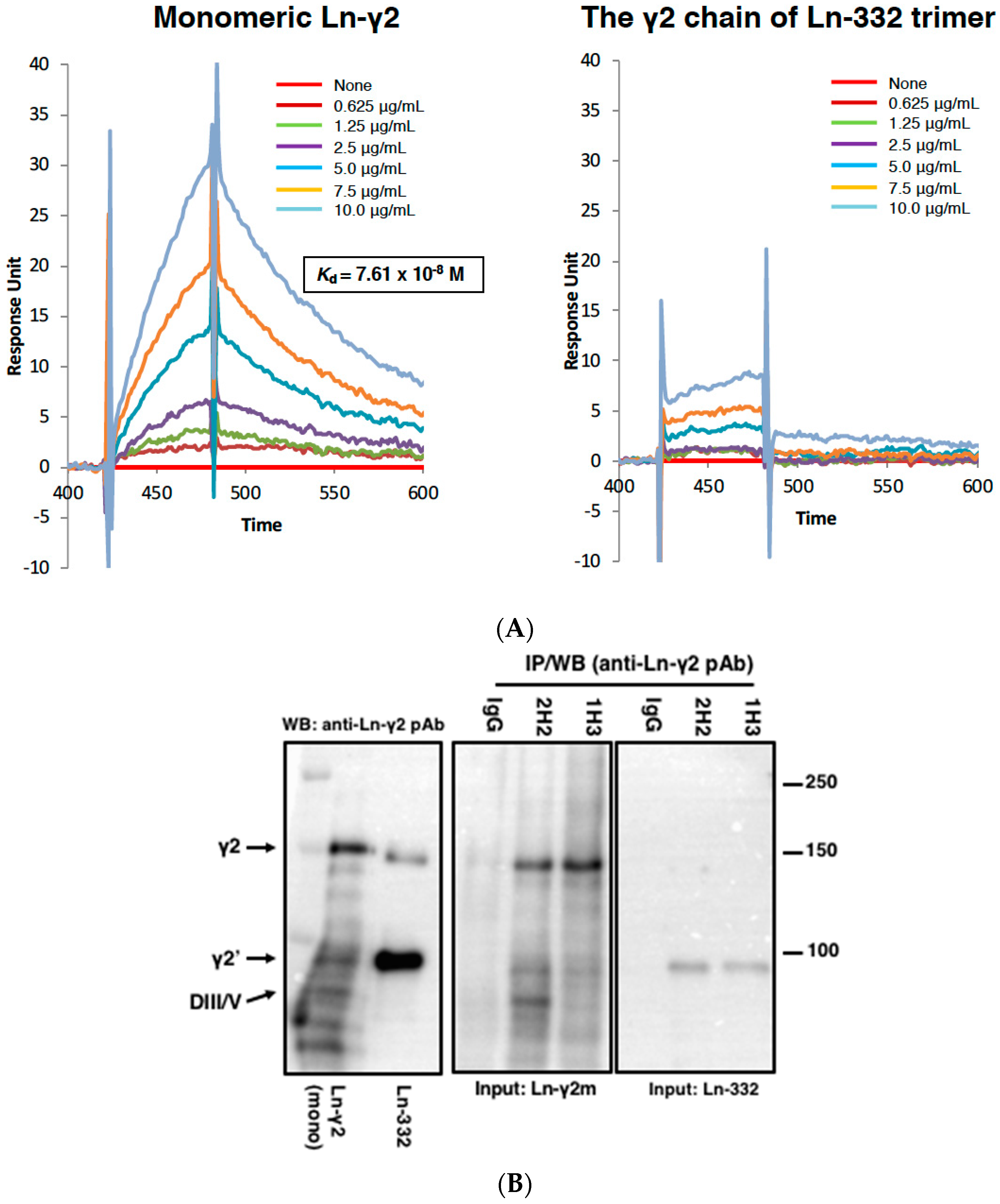
2. Monomeric Laminin-γ2
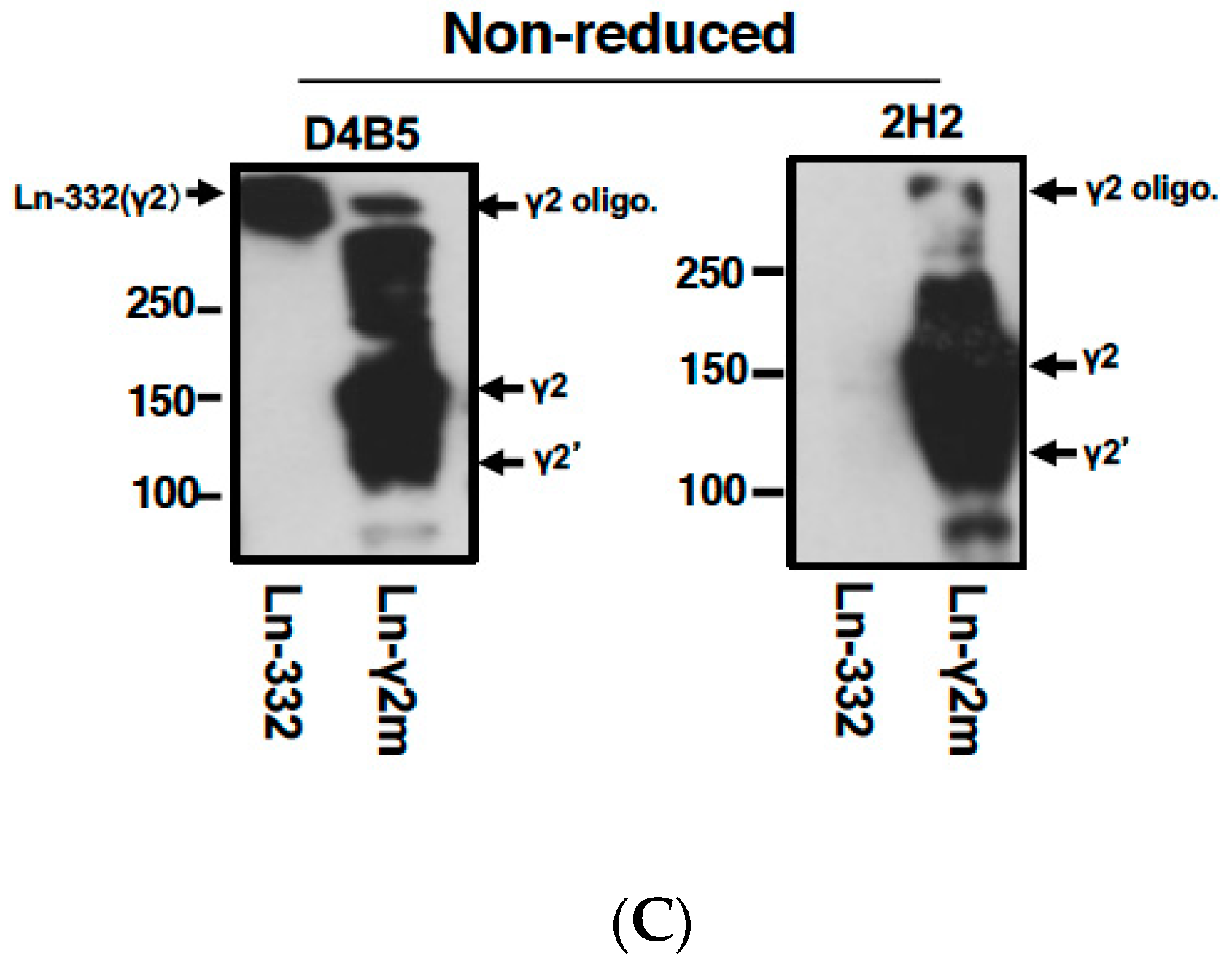
3. Development of a Specific Antibody against Monomeric Ln-γ2 Chain
4. Detection of Ln-γ2 or Its N-Terminal Domain Fragment in Serum Specimens of Patients with Cancer
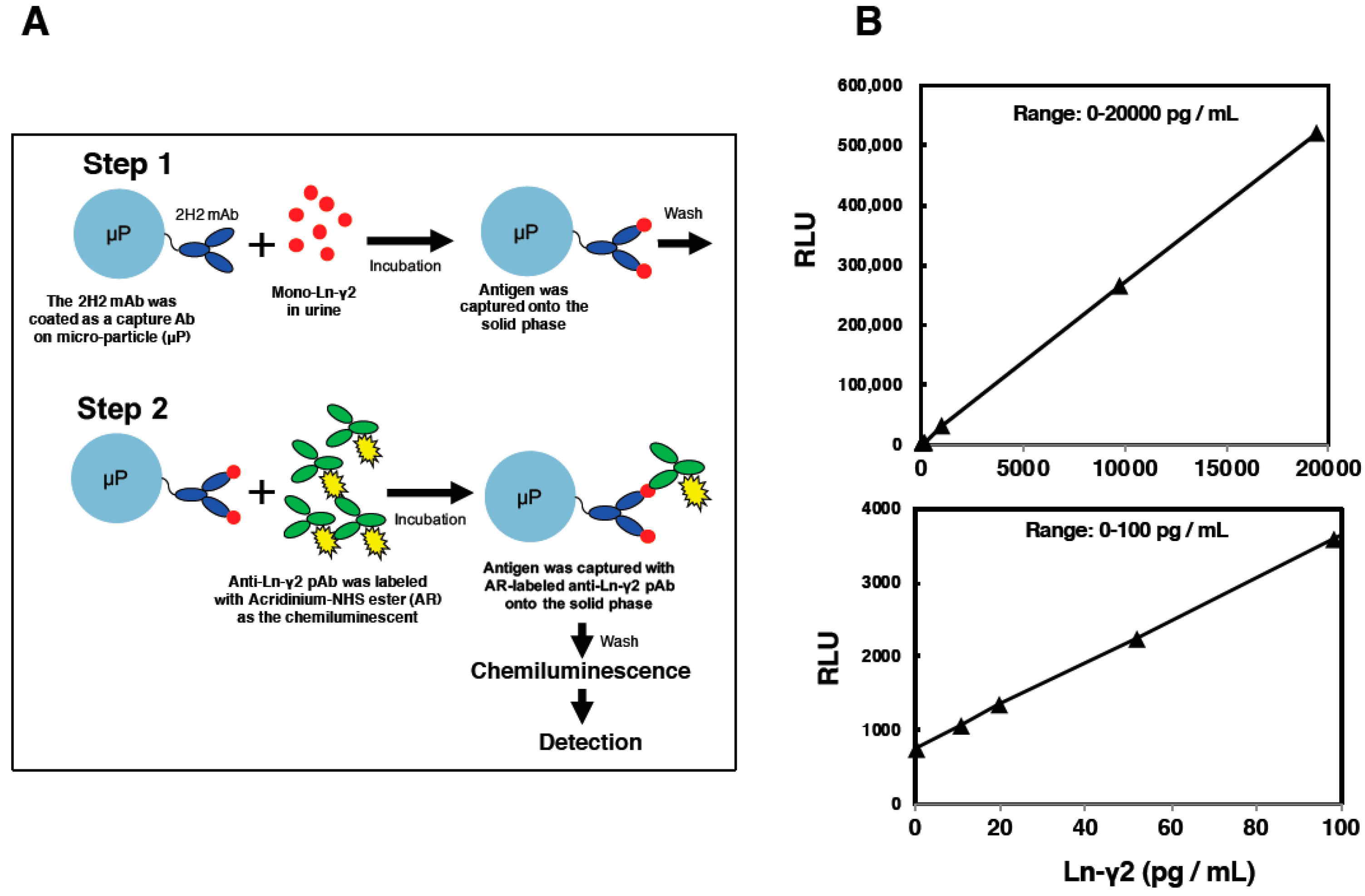
5. Establishment of an Automated Chemiluminescent Immunoassay (CLIA)
6. Hepatocellular Carcinoma Surveillance and Biomarkers
7. Serum Monomeric Ln-γ2 as a Novel Biomarker for Hepatocellular Carcinoma
8. Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AFP | α-fetoprotein |
| BM | basement membrane |
| PIVKA-II | prothrombin induced by Vitamin K Absence II |
| CLIA | chemiluminescent immunoassay |
| CLD | chronic liver disease |
| ECM | extracellular matrix |
| HBV | hepatitis B virus |
| HCC | hepatocellular carcinoma |
| HCV | hepatitis C virus |
| IP | Immunoprecipitation |
| Ln-332 | laminin 332 |
| Ln-γ2 | laminin-γ2 |
| mAb | monoclonal antibody |
| pAb | polyclonal antibody |
| ROC curve AUC | receiver operating characteristic area under curve |
References
- Colognato, H.; Yurchenco, P.D. Form and function: The laminin family of heterotrimers. Dev. Dyn. 2000, 218, 213–234. [Google Scholar] [CrossRef] [Green Version]
- Miyazaki, K. Laminin-5 (laminin-332): Unique biological activity and role in tumor growth and invasion. Cancer Sci. 2006, 97, 91–98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guess, C.M.; Quaranta, V. Defining the role of laminin-332 in carcinoma. Matrix Biol. 2009, 28, 445–455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shang, M.; Koshikawa, N.; Schenk, S.; Quaranta, V. The LG3 module of laminin-5 harbors a binding site for integrin α3β1 that promotes cell adhesion, spreading, and migration. J. Biol. Chem. 2001, 276, 33045–33053. [Google Scholar] [CrossRef] [PubMed]
- Gaietta, G.; Redelmeier, T.E.; Jackson, M.R.; Tamura, R.N.; Quaranta, V. Quantitative measurement of α6β1 and α6β4 integrin internalization under cross-linking conditions: A possible role for α6 cytoplasmic domains. J. Cell Sci. 1994, 107, 3339–3349. [Google Scholar]
- Rousselle, P.; Beck, K. Laminin 332 processing impacts cellular behavior. Cell Adh. Migr. 2013, 7, 122–134. [Google Scholar] [CrossRef] [Green Version]
- Giannelli, G.; Falk-Marziller, J.; Schiraldi, O.; Stetler-Stevenson, W.G.; Quaranta, V. Induction of cell migration by matrix metalloprotease-2 cleavage of laminin-5. Science 1997, 277, 225–228. [Google Scholar] [CrossRef] [PubMed]
- Koshikawa, N.; Giannelli, G.; Cirulli, V.; Miyazaki, K.; Quaranta, V. Role of cell surface metalloprotease MT1-MMP in epithelial cell migration over laminin-5. J. Cell Biol. 2000, 148, 615–624, Erratum in 2000, 151, 479–480. [Google Scholar] [CrossRef]
- Schenk, S.; Hintermann, E.; Bilban, M.; Koshikawa, N.; Hojilla, C.; Khokha, R.; Quaranta, V. Binding to EGF receptor of a laminin-5 EGF-like fragment liberated during MMP-dependent mammary gland involution. J. Cell Biol. 2003, 161, 197–209. [Google Scholar] [CrossRef] [Green Version]
- Ogawa, T.; Tsubota, Y.; Hashimoto, J.; Kariya, Y.; Miyazaki, K. The short arm of laminin γ2 chain of laminin-5 (laminin-332) binds syndecan-1 and regulates cellular adhesion and migration by suppressing phosphorylation of integrin β4 chain. Mol. Biol. Cell 2007, 18, 1621–1633. [Google Scholar] [CrossRef]
- Carpenter, P.M.; Sivadas, P.; Hua, S.S.; Xiao, C.; Gutierrez, A.B.; Ngo, T.; Gershon, P.D. Migration of breast cancer cell lines in response to pulmonary laminin 332. Cancer Med. 2017, 6, 220–234. [Google Scholar] [CrossRef]
- Hao, J.; Jackson, L.; Calaluce, R.; McDaniel, K.; Dalkin, B.L.; Nagle, R.B. Investigation into the mechanism of the loss of laminin 5 (α3β3γ2) expression in prostate cancer. Am. J. Pathol. 2001, 158, 1129–1135. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, H.; Luo, J.; Wu, X.; Li, X.; Zhao, X.; Zhou, D.; Yu, S. Overexpression of α3, β3 and γ2 chains of laminin-332 is associated with poor prognosis in pancreatic ductal adenocarcinoma. Oncol. Lett. 2018, 16, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Koshikawa, N.; Moriyama, K.; Takamura, H.; Mizushima, H.; Nagashima, Y.; Yanoma, S.; Miyazaki, K. Overexpression of laminin γ2 chain monomer in invading gastric carcinoma cells. Cancer Res. 1999, 59, 5596–5601. [Google Scholar] [PubMed]
- Takahashi, S.; Hasebe, T.; Oda, T.; Sasaki, S.; Kinoshita, T.; Konishi, M.; Ochiai, T.; Ochiai, A. Cytoplasmic expression of laminin γ2 chain correlates with postoperative hepatic metastasis and poor prognosis in patients with pancreatic ductal adenocarcinoma. Cancer 2002, 94, 1894–1901. [Google Scholar] [CrossRef]
- Soini, Y.; Määttä, M.; Salo, S.; Tryggvason, K.; Autio-Harmainen, H. Expression of the laminin γ2 chain in pancreatic adenocarcinoma. J. Pathol. 1996, 180, 290–294. [Google Scholar] [CrossRef]
- Fukushima, N.; Sakamoto, M.; Hirohashi, S. Expression of laminin-5-γ-2 chain in intraductal papillary-mucinous and invasive ductal tumors of the pancreas. Mod. Pathol. 2001, 14, 404–409. [Google Scholar] [CrossRef]
- Yamamoto, H.; Itoh, F.; Iku, S.; Hosokawa, M.; Imai, K. Expression of the γ2 chain of laminin-5 at the invasive front is associated with recurrence and poor prognosis in human esophageal squamous cell carcinoma. Clin. Cancer Res. 2001, 7, 896–900. [Google Scholar]
- Garg, M.; Kanojia, D.; Okamoto, R.; Jain, S.; Madan, V.; Chien, W.; Sampath, A.; Ding, L.W.; Xuan, M.; Said, J.W.; et al. Laminin-5γ-2 (LAMC2) is highly expressed in anaplastic thyroid carcinoma and is associated with tumor progression, migration, and invasion by modulating signaling of EGFR. J. Clin. Endocrinol. Metab. 2014, 99, E62–E72. [Google Scholar] [CrossRef]
- Yamamoto, H.; Kitadai, Y.; Yamamoto, H.; Oue, N.; Ohdan, H.; Yasui, W.; Kikuchi, A. Laminin γ2 Mediates Wnt5a-induced invasion of gastric cancer cells. Gastroenterology 2009, 137, 242–252. [Google Scholar] [CrossRef]
- Sordat, I.; Bosman, F.T.; Dorta, G.; Rousselle, P.; Aberdam, D.; Blum, A.L.; Sordat, B. Differential expression of laminin-5 subunits and integrin receptors in human colorectal neoplasia. J. Pathol. 1998, 185, 44–52. [Google Scholar] [CrossRef]
- Kiyokawa, H.; Yasuda, H.; Oikawa, R.; Okuse, C.; Matsumoto, N.; Ikeda, H.; Watanabe, T.; Yamamoto, H.; Itoh, F.; Otsubo, T.; et al. Serum monomeric laminin-γ2 as a novel biomarker for hepatocellular carcinoma. Cancer Sci. 2017, 108, 1432–1439. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Giannelli, G.; Fransvea, E.; Bergamini, C.; Marinosci, F.; Antonaci, S. Laminin-5 chains are expressed differentially in metastatic and nonmetastatic hepatocellular carcinoma. Clin. Cancer Res. 2003, 9, 3684–3691. [Google Scholar] [PubMed]
- Kosanam, H.; Prassas, I.; Chrystoja, C.C.; Soleas, I.; Chan, A.; Dimitromanolakis, A.; Blasutig, I.M.; Ruckert, F.; Gruetzmann, R.; Pilarsky, C.; et al. Laminin, γ2 (LAMC2): A promising new putative pancreatic cancer biomarker identified by proteomic analysis of pancreatic adenocarcinoma tissues. Mol. Cell. Proteom. 2013, 12, 2820–2832. [Google Scholar] [CrossRef] [PubMed]
- Oka, T.; Yamamoto, H.; Sasaki, S.; Ii, M.; Hizaki, K.; Taniguchi, H.; Adachi, Y.; Imai, K.; Shinomura, Y. Overexpression of β3/γ2 chains of laminin-5 and MMP7 in biliary cancer. World J. Gastroenterol. 2009, 15, 3865–3873. [Google Scholar] [CrossRef] [PubMed]
- Koshikawa, N.; Minegishi, T.; Sharabi, A.; Quaranta, V.; Seiki, M. Membrane-type matrix metalloproteinase-1 (MT1-MMP) is a processing enzyme for human laminin γ2 chain. J. Biol. Chem. 2005, 280, 88–93. [Google Scholar] [CrossRef]
- Niki, T.; Kohno, T.; Iba, S.; Moriya, Y.; Takahashi, Y.; Saito, M.; Maeshima, A.; Yamada, T.; Matsuno, Y.; Fukayama, M.; et al. Frequent co-localization of Cox-2 and laminin-5 γ2 chain at the invasive front of early-stage lung adenocarcinomas. Am. J. Pathol. 2002, 160, 1129–1141. [Google Scholar] [CrossRef]
- Kang, S.G.; Ha, Y.R.; Ko, Y.H.; Kang, S.H.; Joo, K.J.; Cho, H.Y.; Park, H.S.; Kim, C.H.; Kwon, S.Y.; Kim, J.J.; et al. Effect of laminin 332 on motility and invasion in bladder cancer. Kaohsiung J. Med. Sci. 2013, 29, 422–429. [Google Scholar] [CrossRef] [Green Version]
- Mizushima, H.; Koshikawa, N.; Moriyama, K.; Takamura, H.; Nagashima, Y.; Hirahara, F.; Miyazaki, K. Wide distribution of laminin-5 γ2 chain in basement membranes of various human tissues. Horm. Res. 1998, 50 (Suppl. 2), 7–14. [Google Scholar] [CrossRef]
- Koshikawa, N.; Minegishi, T.; Nabeshima, K.; Seiki, M. Development of a new tracking tool for the human monomeric laminin-γ2 chain in vitro and in vivo. Cancer Res. 2008, 68, 530–536, Erratum in 2016, 76, 762. [Google Scholar] [CrossRef]
- Katayama, M.; Funakoshi, A.; Sumii, T.; Sanzen, N.; Sekiguchi, K. Laminin γ2-chain fragment circulating level increases in patients with metastatic pancreatic ductal cell adenocarcinomas. Cancer Lett. 2005, 225, 167–176. [Google Scholar] [CrossRef]
- Kuratomi, Y.; Sato, S.; Monji, M.; Shimazu, R.; Tanaka, G.; Yokogawa, K.; Inoue, A.; Inokuchi, A.; Katayama, M. Serum concentrations of laminin γ2 fragments in patients with head and neck squamous cell carcinoma. Head Neck 2008, 30, 1058–1063. [Google Scholar] [CrossRef] [PubMed]
- Katayama, M.; Ishizaka, A.; Sakamoto, M.; Fujishima, S.; Sekiguchi, K.; Asano, K.; Betsuyaku, T.; Kotani, T.; Ware, L.B.; Matthay, M.A.; et al. Laminin γ2 fragments are increased in the circulation of patients with early phase acute lung injury. Intensive Care Med. 2010, 36, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, M.; Karashima, T.; Kamada, M.; Yoshida, E.; Yoshimura, T.; Nojima, M.; Inoue, K.; Shuin, T.; Seiki, M.; Koshikawa, N. Development of a fully automated chemiluminescence immunoassay for urine monomeric laminin-γ2 as a promising diagnostic tool of non-muscle invasive bladder cancer. Biomark. Res. 2017, 5, 29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, B.H.; Yang, B.H.; Tang, Z.Y. Randomized controlled trial of screening for hepatocellular carcinoma. J. Cancer Res. Clin. Oncol. 2004, 130, 417–422. [Google Scholar] [CrossRef] [PubMed]
- Yeh, Y.P.; Hu, T.H.; Cho, P.Y.; Chen, H.H.; Yen, A.M.; Chen, S.L.; Chiu, S.Y.; Fann, J.C.; Su, W.W.; Fang, Y.J.; et al. Evaluation of abdominal ultrasonography mass screening for hepatocellular carcinoma in Taiwan. Hepatology 2014, 59, 1840–1849. [Google Scholar] [CrossRef] [Green Version]
- Bruix, J.; Sherman, M. Management of hepatocellular carcinoma: An update. Hepatology 2011, 53, 1020–1022. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.G.; Parkin, D.M.; Chen, Q.G.; Lu, J.H.; Shen, Q.J.; Zhang, B.C.; Zhu, Y.R. Screening for liver cancer: Results of a randomized controlled trial in Qidong, China. J. Med. Screen. 2003, 10, 204–209. [Google Scholar] [CrossRef] [PubMed]
- Bolondi, L.; Sofia, S.; Siringo, S.; Gaiani, S.; Casali, A.; Zironi, G.; Piscaglia, F.; Gramantieri, L.; Zanetti, M.; Sherman, M. Surveillance programme of cirrhotic patients for early diagnosis and treatment of hepatocellular carcinoma: A cost effectiveness analysis. Gut 2001, 48, 251–259. [Google Scholar] [CrossRef]
- Clinical Practice Guidelines for Hepatocellular Carcinoma 2013. Japan Society of Hepatology. Available online: http://www.jsh.or.jp/English/examination_en (accessed on 6 January 2019).
- Yoon, Y.J.; Han, K.H.; Kim, D.Y. Role of serum prothrombin induced by vitamin K absence or antagonist-II in the early detection of hepatocellular carcinoma in patients with chronic hepatitis B virus infection. Scand. J. Gastroenterol. 2009, 44, 861–866. [Google Scholar] [CrossRef]
- Li, C.; Zhang, Z.; Zhang, P.; Liu, J. Diagnostic accuracy of des-gamma-carboxy prothrombin versus α-fetoprotein for hepatocellular carcinoma: A systematic review. Hepatol. Res. 2014, 44, E11–E25. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, K.; Imamura, H.; Matsuyama, Y.; Hasegawa, K.; Beck, Y.; Sugawara, Y.; Makuuchi, M.; Kokudo, N. Significance of α-fetoprotein and des-γ-carboxy prothrombin in patients with hepatocellular carcinoma undergoing hepatectomy. Ann. Surg. Oncol. 2009, 16, 2795–2804. [Google Scholar] [CrossRef] [PubMed]
- Nobuoka, D.; Kato, Y.; Gotohda, N.; Takahashi, S.; Nakagohri, T.; Konishi, M.; Kinoshita, T.; Nakatsura, T. Postoperative serum α-fetoprotein level is a useful predictor of recurrence after hepatectomy for hepatocellular carcinoma. Oncol. Rep. 2010, 24, 521–528. [Google Scholar] [PubMed]
- Rani, B.; Cao, Y.; Malfettone, A.; Tomuleasa, C.; Fabregat, I.; Giannelli, G. Role of the tissue microenvironment as a therapeutic target in hepatocellular carcinoma. World J. Gastroenterol. 2014, 20, 4128–4140. [Google Scholar] [CrossRef] [PubMed]
- Giannelli, G.; Bergamini, C.; Fransvea, E.; Marinosci, F.; Quaranta, V.; Antonaci, S. Human hepatocellular carcinoma (HCC) cells require both α3β1 integrin and matrix metalloproteinases activity for migration and invasion. Lab. Investig. 2001, 81, 613–627. [Google Scholar] [CrossRef] [PubMed]
- Govaere, O.; Wouters, J.; Petz, M.; Vandewynckel, Y.P.; Van den Eynde, K.; Van den Broeck, A.; Verhulst, S.; Dollé, L.; Gremeaux, L.; Ceulemans, A.; et al. Laminin-332 sustains chemoresistance and quiescence as part of the human hepatic cancer stem cell niche. J. Hepatol. 2016, 64, 609–617. [Google Scholar] [CrossRef] [Green Version]
- Fujimoto, A.; Furuta, M.; Totoki, Y.; Tsunoda, T.; Kato, M.; Shiraishi, Y.; Tanaka, H.; Taniguchi, H.; Kawakami, Y.; Ueno, M.; et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat. Genet. 2016, 48, 500–509. [Google Scholar] [CrossRef]
- Hlubek, F.; Jung, A.; Kotzor, N.; Kirchner, T.; Brabletz, T. Expression of the invasion factor laminin γ2 in colorectal carcinomas is regulated by β-catenin. Cancer Res. 2001, 61, 8089–8093. [Google Scholar]
- Hashimoto, K.; Mori, N.; Tamesa, T.; Okada, T.; Kawauchi, S.; Oga, A.; Furuya, T.; Tangoku, A.; Oka, M.; Sasaki, K. Analysis of DNA copy number aberrations in hepatitis C virus-associated hepatocellular carcinomas by conventional CGH and array CGH. Mod. Pathol. 2004, 17, 617–622. [Google Scholar] [CrossRef] [Green Version]
- Sathyanarayana, U.G.; Maruyama, R.; Padar, A.; Suzuki, M.; Bondaruk, J.; Sagalowsky, A.; Minna, J.D.; Frenkel, E.P.; Grossman, H.B.; Czerniak, B.; et al. Molecular detection of noninvasive and invasive bladder tumor tissues and exfoliated cells by aberrant promoter methylation of laminin-5 encoding genes. Cancer Res. 2004, 64, 1425–1430. [Google Scholar] [CrossRef]
- Sathyanarayana, U.G.; Padar, A.; Huang, C.X.; Suzuki, M.; Shigematsu, H.; Bekele, B.N.; Gazdar, A.F. Aberrant promoter methylation and silencing of laminin-5-encoding genes in breast carcinoma. Clin. Cancer Res. 2003, 9, 6389–6394. [Google Scholar] [PubMed]
- Sathyanarayana, U.G.; Padar, A.; Suzuki, M.; Maruyama, R.; Shigematsu, H.; Hsieh, J.T.; Frenkel, E.P.; Gazdar, A.F. Aberrant promoter methylation of laminin-5-encoding genes in prostate cancers and its relationship to clinicopathological features. Clin. Cancer Res. 2003, 9, 6395–6400. [Google Scholar] [PubMed]
- Sathyanarayana, U.G.; Toyooka, S.; Padar, A.; Takahashi, T.; Brambilla, E.; Minna, J.D.; Gazdar, A.F. Epigenetic inactivation of laminin-5-encoding genes in lung cancers. Clin. Cancer Res. 2003, 9, 2665–2672. [Google Scholar] [PubMed]

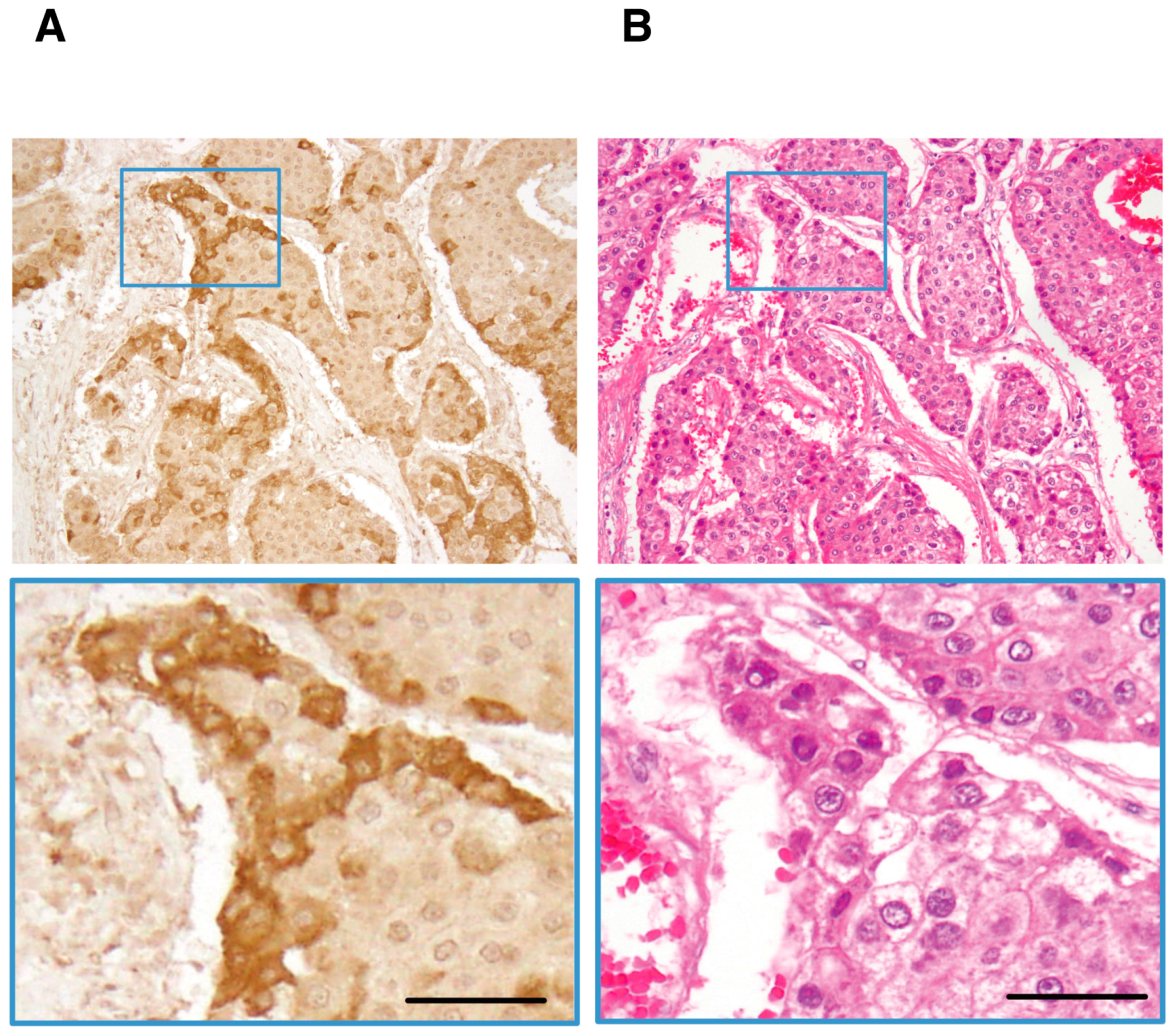



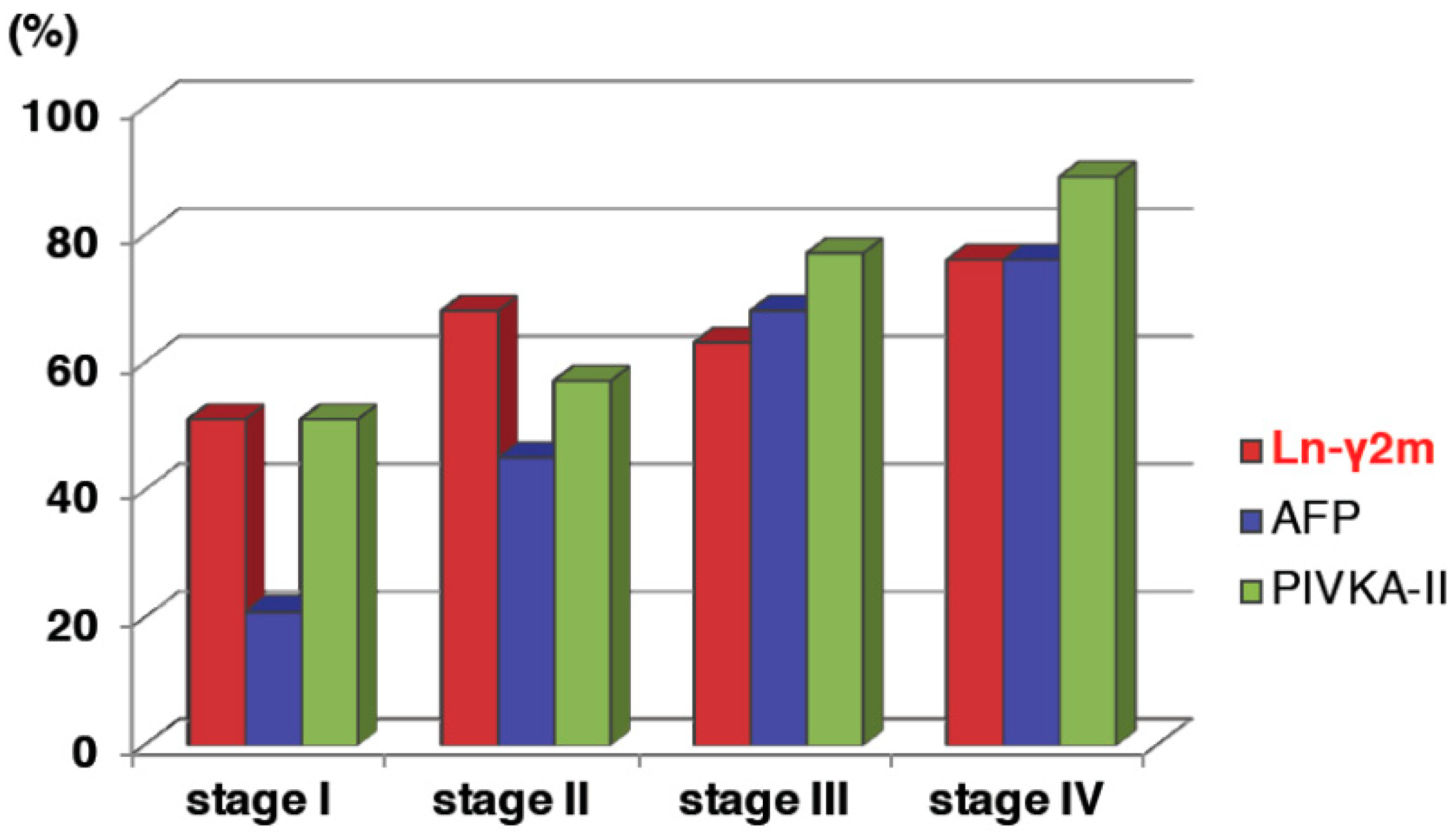
| Organ | Detection Rate by IHC | Detection Rate by Serological Assay | References |
|---|---|---|---|
| esophagus | 44% (44/100) | N/A | [18] |
| stomach | 23% (35/153) | N/A | [20] |
| colon | 65% (29/39) | N/A | [21] |
| liver | 63% (25/40) | CLIA, 63% (36/57) | [22,23] |
| pancreas | 53% (8/15) | ELISA, 72% (36/50) | [16,24] |
| biliary tract | 57% (35/61) | N/A | [25] |
| ELISA | Automated CLIA | |
|---|---|---|
| Dynamic range | Narrow *1 | Wide *2 |
| Detection sensitivity | Low | High |
| Diagnostic accuracy | Low | High |
| Background | High | Low |
| Throughput | Low | High |
| Reaction time | > 2.5 h | 30 min |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yasuda, H.; Nakagawa, M.; Kiyokawa, H.; Yoshida, E.; Yoshimura, T.; Koshikawa, N.; Itoh, F.; Seiki, M. Unique Biological Activity and Potential Role of Monomeric Laminin-γ2 as a Novel Biomarker for Hepatocellular Carcinoma: A Review. Int. J. Mol. Sci. 2019, 20, 226. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms20010226
Yasuda H, Nakagawa M, Kiyokawa H, Yoshida E, Yoshimura T, Koshikawa N, Itoh F, Seiki M. Unique Biological Activity and Potential Role of Monomeric Laminin-γ2 as a Novel Biomarker for Hepatocellular Carcinoma: A Review. International Journal of Molecular Sciences. 2019; 20(1):226. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms20010226
Chicago/Turabian StyleYasuda, Hiroshi, Masatoshi Nakagawa, Hirofumi Kiyokawa, Eisaku Yoshida, Toru Yoshimura, Naohiko Koshikawa, Fumio Itoh, and Motoharu Seiki. 2019. "Unique Biological Activity and Potential Role of Monomeric Laminin-γ2 as a Novel Biomarker for Hepatocellular Carcinoma: A Review" International Journal of Molecular Sciences 20, no. 1: 226. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms20010226





