Introduction
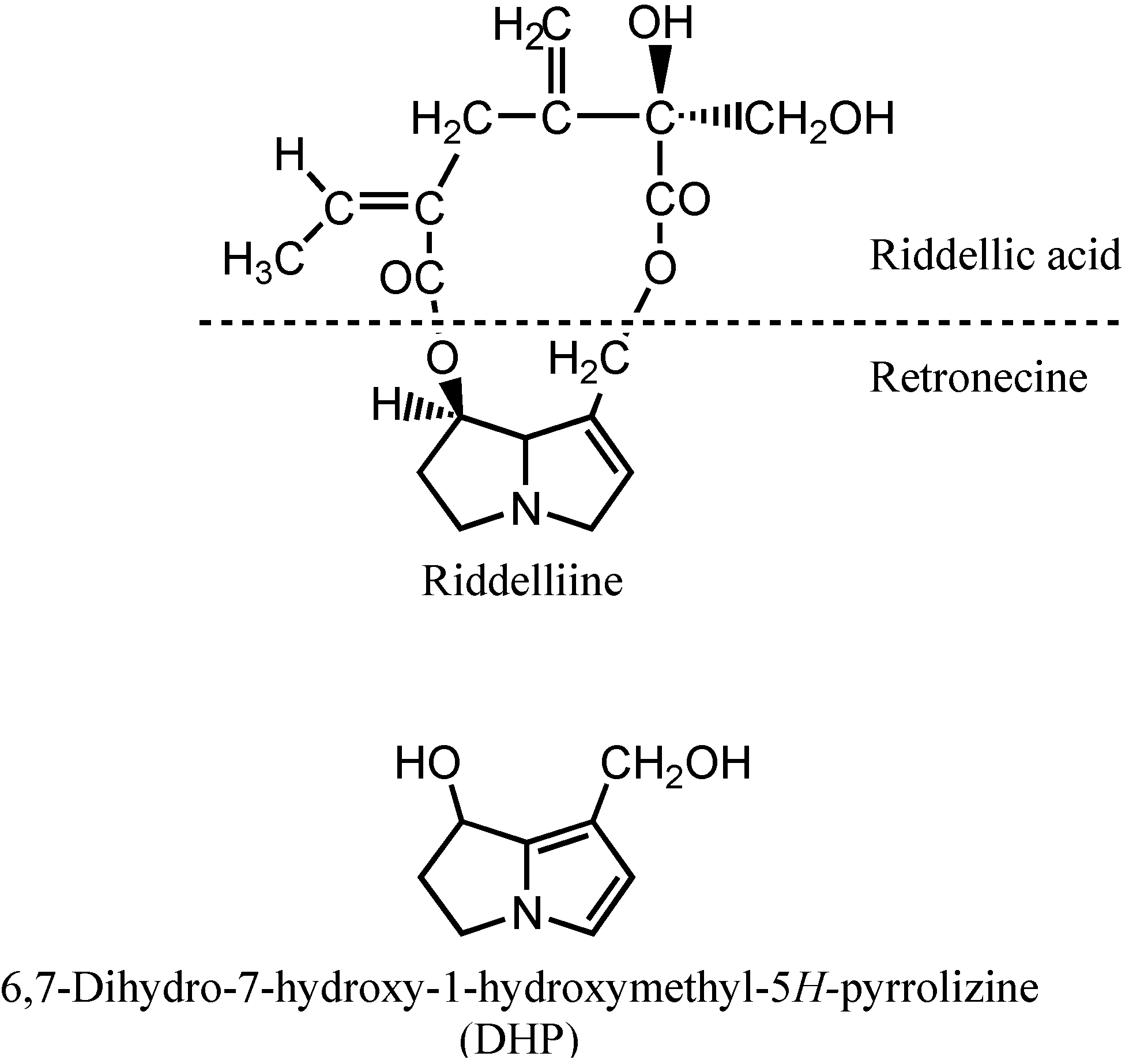
Riddelliine is a naturally occurring genotoxic pyrrolizidine alkaloid, consisting of a riddelliic acid and a retronecine base (
Scheme 1), and is widespread in rangelands of the western United States [
1,
2,
3]. Livestock have been poisoned by ingesting plants containing toxic pyrrolizidine alkaloids, including riddelliine, and resulted in tremendous economic loss [
8]. Because of its economic impact on the livestock industry and its potential for human exposure due to the contamination of human food sources, such as grains, milk, honey, and herbal teas [
2,
4,
5,
6,
7,
8,
9], riddelliine was nominated for toxicology and carcinogenesis testing by the Food and Drug Administration and conducted by National Toxicology Program (NTP). Results from NTP 2-year chronic test showed that riddelliine induced liver hemagiosarcomas and hepatocellular tumors in F344 rat and B6C3F
1 mice [
7].
Riddelliine, like other carcinogenic pyrrolizidine alkaloids, requires metabolic activation to exert its toxicity and genotoxicity [
2,
4,
10,
11,
12,
13,
14,
15]. We have previously shown that metabolism of riddelliine by rat liver microsomes resulted in the formation of (R/S)6,7-dihydro-7-hydroxy-1-hydroxymethyl-5
H-pyrrolizine (DHP) as a predominant metabolite [
4,
7]. By
32P-postlabeling/ HPLC analysis, a set of eight DHP-derived DNA adducts was detected both in vitro and in vivo [
7,
11]. These DHP-derived DNA adducts consist of a pair of epimeric DHP-3´-dGMP adducts and six DHP-derived deoxyribodinucleotides [
16]. The level of the total DHP-derived DNA adducts in liver DNA of rats fed riddelliine was dose- and time-dependent [
11].
In this study, the formation of DHP-derived DNA adducts in blood DNA of rats treated with riddelliine was investigated and the levels of the adducts formed in rat blood cells were compared with those found in liver DNA.
Scheme 1.
Chemical structures of riddelliine and its reactive metabolite 6,7-dihydro-7-hydroxy-1-(hydroxymethyl)-5H-pyrrolizine (DHP).
Scheme 1.
Chemical structures of riddelliine and its reactive metabolite 6,7-dihydro-7-hydroxy-1-(hydroxymethyl)-5H-pyrrolizine (DHP).
Materials and Methods
Materials
Riddelliine was obtained from the NTP. Retronecine, DHP, synthetic DHP-3'-dGMP DNA adduct standards, and DHP-modified calf thymus DNA were prepared in our laboratory as previously described [
14]. Nuclease P1, micrococcal nuclease (MN), spleen phosphodiesterase (SPD), calf thymus DNA, and the chemicals required for
32P-postlabeling were purchased from the Sigma Chemical Co. (St. Louis, MO). Cloned T4 polynucleotide kinase (PNK) was obtained from U.S. Biochemical Corp. (Cleveland, OH). [ γ-
32P]Adenosine 5'-triphosphate ( γ-
32P] ATP) (sp. act. >7,000 Ci/mmol) was purchased from ICN Biomedicals, Inc. (Costa Mesa, CA). All solvents used were HPLC grade.
Animals and treatments
Time course experiment. Male and female F344 rats (3 per group) were obtained from the National Center for Toxicological Research breeding colony as weanlings and maintained on a 12 hr light-dark cycle. At the age of 8 weeks, rats were dosed by oral gavage with a single dose of riddelliine at the concentration of 10 mg/kg/day in 0.1 M phosphate buffer (pH, 8.0). Control animals were gavaged with 0.1 M phosphate buffer only. Eight, 24, 48, and 168 hrs after dosing the animals were euthanized by CO2 inhalation, and blood samples were collected into heparinized tubes by cardiac puncture. Blood samples were combined within the group, and blood DNA was isolated immediately.
Dose-responsive experiment. A separate experiment was conducted using female F344 rats (3 per group) gavaged with riddelliine at concentration of 0.1 and 1.0 mg/kg body weight, respectively, for 3 consecutive days. The rats were sacrificed 24 hrs after the last dosing. Blood samples were collected by cardiac puncture and blood DNA was isolated as described. The liver tissues were removed immediately after blood was withdrawn and stored at –80oC prior to DNA isolation.
DNA isolation. Whole blood DNA was isolated and purified by using Puregene DNA isolation kit from Gentra Systems, Inc., Minneapolis, MN according to the manufacturer’s instructions. Rat liver DNA was isolated by using RecoverEase DNA Isolation kit (Stratagene, Cedar Creek, TX) according to the manufacturer’s instructions. The concentration of DNA was determined by UV spectrophotometric measurements.
32P-Postlabeling/HPLC analysis of DHP-derived DNA adducts. The analysis of DHP-derived DNA adducts was conducted by the
32P-postlabeling/HPLC method previously developed in our laboratory [
14]. Briefly, 10 µg of rat liver DNA in 10 µL Tris HCl-EDTA buffer was enzymatically hydrolyzed to the corresponding 2'-deoxyribonucleoside 3'-monophosphate at 37
oC for 20 min with 78 mU of MN and 4 mU of SPD contained in a 20 µL solution of 20 mM sodium succinate and 10 mM of calcium chloride (pH 6). After enrichment with nuclease P1, the resulting DNA hydrolysate was then postlabeled with [g-
32P]ATP (sp. act., 7000 Ci/mmol) by PNK. The
32P-labeled mixture was analyzed by reversed-phase HPLC as described previously (14). For quantification of each sample, the two epimeric DHP-3'-dGMP synthetic standards in amounts of 30 fmol that were closely matched the range of modification in the liver and blood DNA samples were also analyzed in parallel [
14].
Results
A set of eight DHP-derived DNA adducts in the DHP-modified calf thymus DNA were detected after the DNA was hydrolyzed by MN/SPD,
32P-postlabeled by PNK and analyzed by HPLC [
11,
14]. The HPLC profile of the DHP-derived DNA adduct separation from the DHP-modified calf thymus DNA is shown in
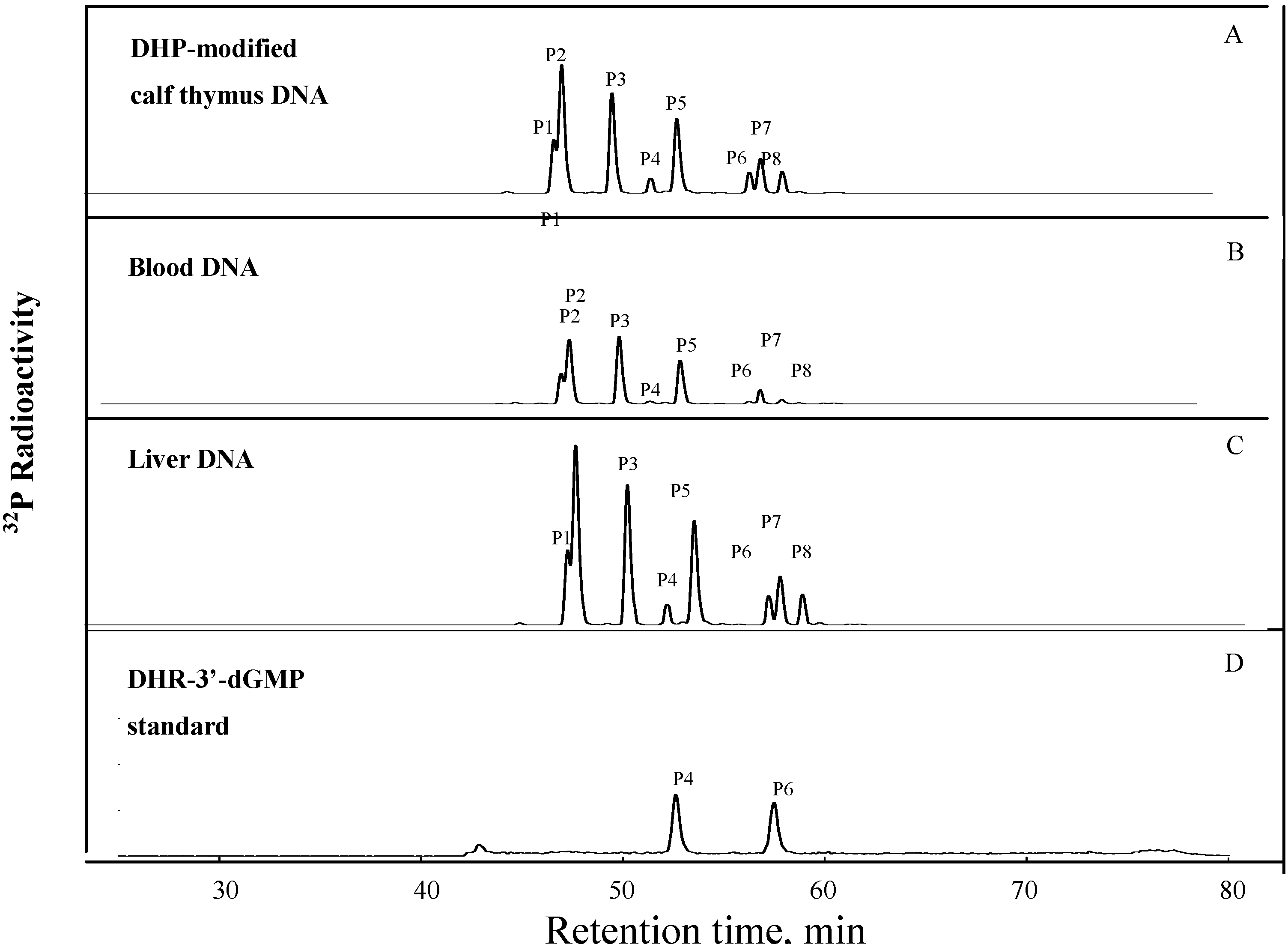
Figure 1A. Similar set of DHP-derived DNA adduct were also detected in DNA samples from blood cells (
Figure 1B) and liver DNA (
Figure 1C) of the rats treated with riddelliine. Thirty fmol of synthetic DHP-3'dGMP was used as an authentic standard for quantification of DHP-derived DNA adducts and was assayed in parellel with DNA samples (
Figure 1D). Quantification of the adduct formation was based on the comparison of the radioactivity of the DNA adduct detected in the DNA samples compared with the radioactivity obtained from the known concentration (30 fmol) of DHP-3'-dGMP standards.
Figure 1.
32P-Postlabeling/HPLC analysis of DHR-derived DNA adducts obtained from (A)reacting DHR with calf thymus DNA; (B) blood DNA from a female F344 rat treated by gavage with riddelliine; (C) liver DNA from a female F344 rat treated by gavage with riddelliine; and (D) synthetically-prepared DHR-3'-dGMP standards.
Figure 1.
32P-Postlabeling/HPLC analysis of DHR-derived DNA adducts obtained from (A)reacting DHR with calf thymus DNA; (B) blood DNA from a female F344 rat treated by gavage with riddelliine; (C) liver DNA from a female F344 rat treated by gavage with riddelliine; and (D) synthetically-prepared DHR-3'-dGMP standards.
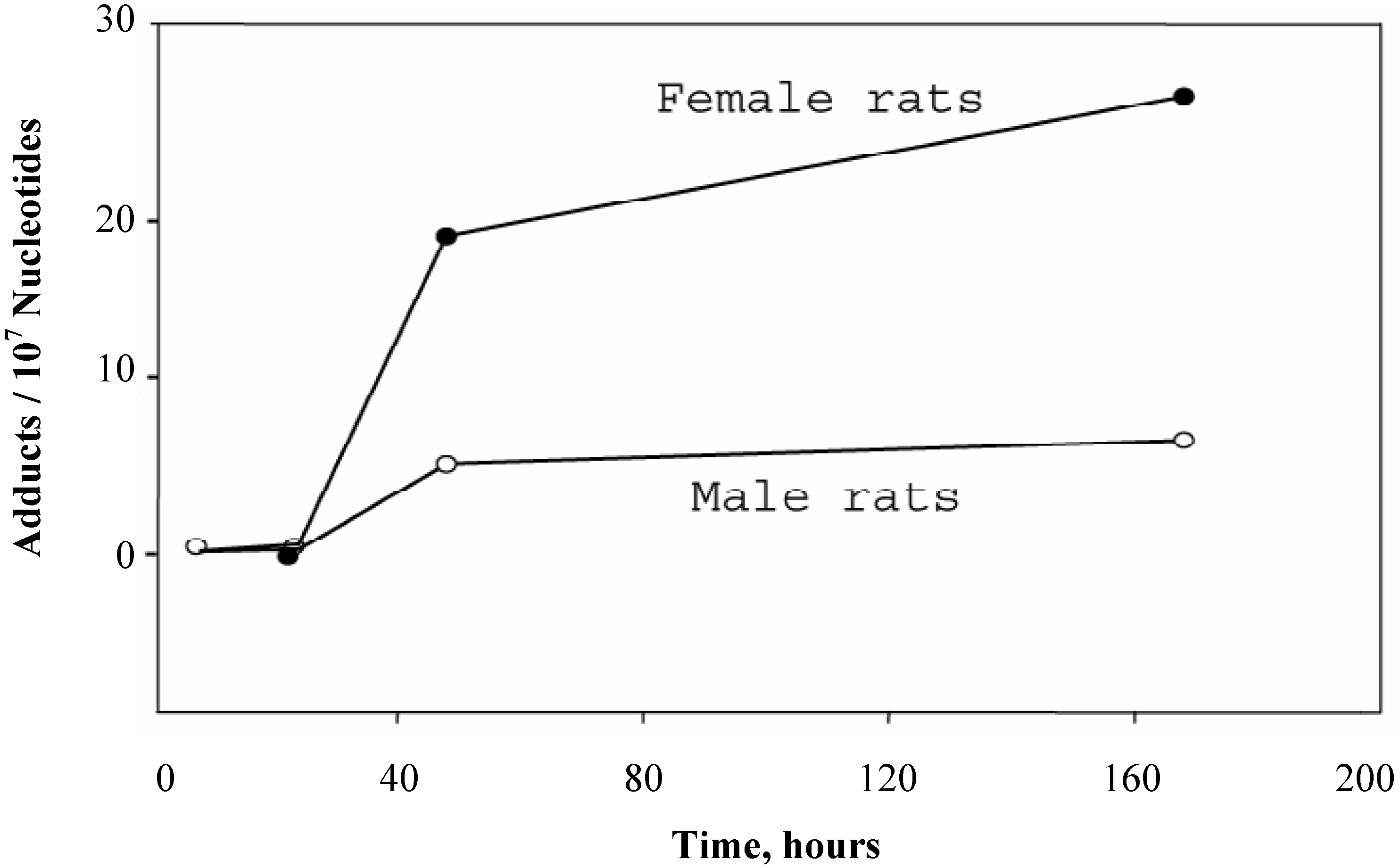
The time-course of DHP-derived DNA adduct formation and removal in blood DNA of rats treated with a single dose (10 mg/kg) of riddelliine is shown in
Figure 2. A 24-hour lag period indicated the time for the carcinogen to be metabolically activated in liver and distributed into the bloodstream. Although a toxicokinetic study (17) showed that the half life of the elimination of riddelliine in serum of female F344 rats received 10 mg/kg single dose of riddelliine was 4.2 hrs, the DHP-derived DNA adduct level in blood reached at a constant level within 2 to 7days (
Figure 2).
Figure 2 also shows that the blood DHP-derived DNA adduct level in female rats was greater than that in the male rats. During day 2 to day 7, the adduct levels in female rat blood was 4-fold greater than male rats.
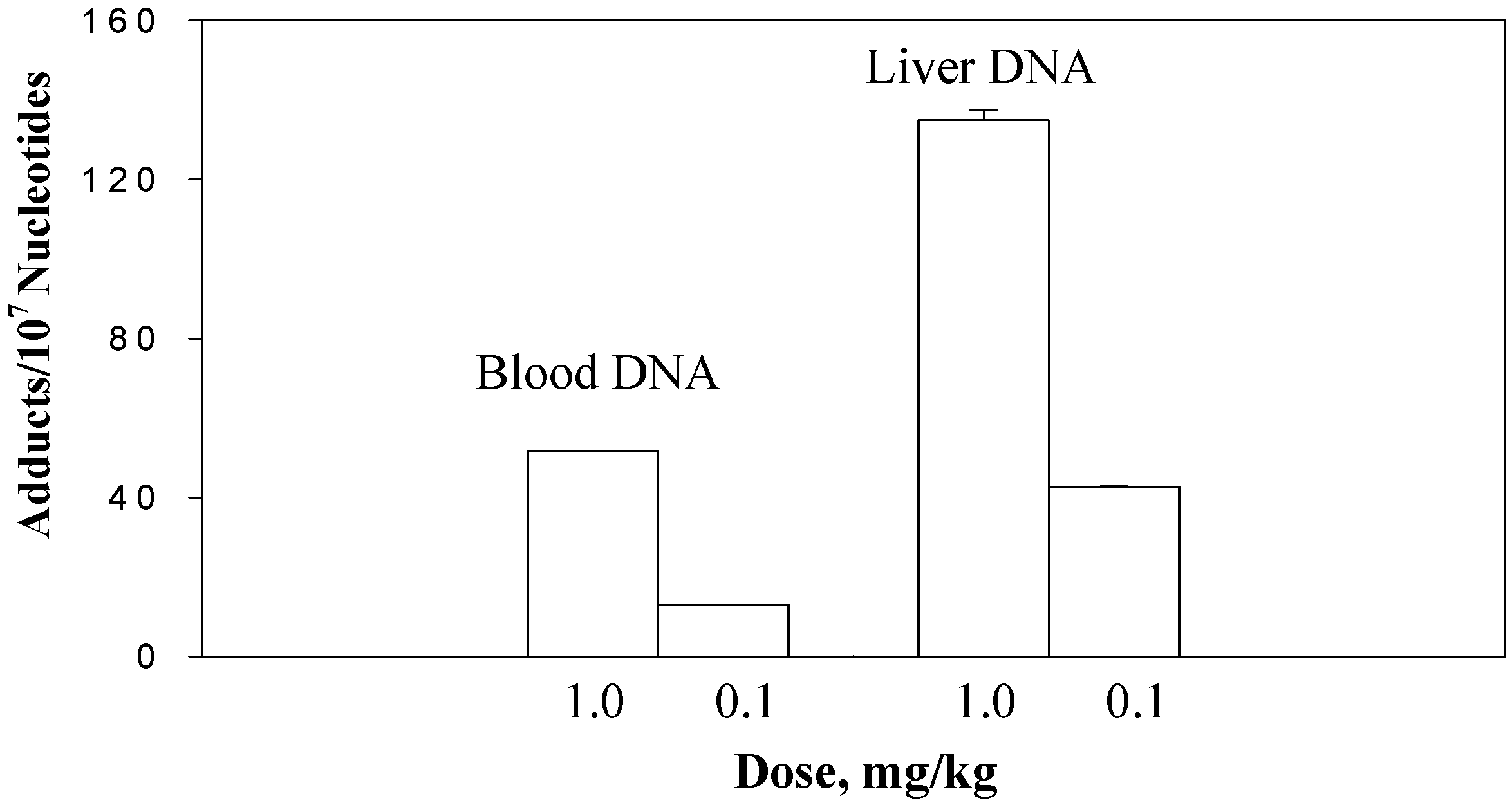
The dose-response experiment was conducted at a lower dose range, 0.1 and 1.0 mg/kg. The DHP-derived DNA adduct formation in blood and liver DNA of rats treated with riddelliine at 0.1 and 1.0 mg/kg doses is shown in
Figure 3. Results showed that DHP-derived DNA adduct formation in both blood and liver tissues was dose-dependent [
11,
14]. At the 0.1 mg/kg dose, 42.6±0.4 and 12.9 DHP-derived DNA adducts per 10
7 nucleotides were detected in the rat liver and blood DNA, respectively. The 1.0 mg/kg dose resulted in a 3- to 4-fold increase of the adduct formation (134.9±2.6 and 51.8 DHP-derived DNA adducts per 10
7 nucleotides in liver and blood DNA). The data of liver DNA adducts were expressed as means±SD from the results of three rats per group, however, the blood data were obtained from the results of the assays using combined blood samples of three rats within the same group. The formation of DHP-derived DNA adducts in liver was 3-fold greater than those in blood cells.
Figure 2.
Time course of the formation and removal of the DHP-derived DNA adducts in blood DNA samples of rats treated with a single dose of riddelliine (10 mg/kg). Blood samples were collected from 3 rats per group and combined within the group. Data from liver DNA samples were the mean ± SD (n = 3). DHP-derived DNA adducts were assayed by the 32P-postlabeling/HPLC method as described in the Method section.
Figure 2.
Time course of the formation and removal of the DHP-derived DNA adducts in blood DNA samples of rats treated with a single dose of riddelliine (10 mg/kg). Blood samples were collected from 3 rats per group and combined within the group. Data from liver DNA samples were the mean ± SD (n = 3). DHP-derived DNA adducts were assayed by the 32P-postlabeling/HPLC method as described in the Method section.
Figure 3.
The dose-responsible formation of the DHP-derived DNA adducts in the blood and liver DNA samples of rats treated with riddelliine at the doses of 1.0 and 0.1 mg/kg/day. Blood samples were collected from 3 rats per group and combined within the group. Data from liver DNA samples were the mean ± SD (n = 3). DHP-derived DNA adducts were assayed by the 32P-postlabeling/HPLC method as described in the Method section.
Figure 3.
The dose-responsible formation of the DHP-derived DNA adducts in the blood and liver DNA samples of rats treated with riddelliine at the doses of 1.0 and 0.1 mg/kg/day. Blood samples were collected from 3 rats per group and combined within the group. Data from liver DNA samples were the mean ± SD (n = 3). DHP-derived DNA adducts were assayed by the 32P-postlabeling/HPLC method as described in the Method section.
Discussion
The measurement of DNA adducts is considered a biomarker for assessing the risk of chemically induced carcinogenesis. The detection of riddelliine- or DHP-derived DNA adducts in liver DNA of rats treated with riddelliine demonstrates that riddelliine-induced tumorigenesis occurs through a genotoxic mechanism. One of the major objectives of this study is to establish a noninvasive methodology to monitor the detection of carcinogenic pyrrolizidine alkaloids in livestock as well as in humans. Our results showed that leucocyte DNA can bind with DHP to form the DHP-derived DNA adducts similar as those generated in liver, although at a lower level. The level of DHP-derived DNA adducts in blood DNA was higher than the non-target tissue, such as kidney (unpublished data) that may place riddelliine as a potential candidate for inducing leukemia in rats.
The necine-based carcinogenic pyrrolizidine alkaloids, such as riddelliine, are metabolized to form reactive pyrrolic metabolites that can bind to the cellular macromolecules, such as proteins and DNA. The primary dehydro alkaloids (dehydroriddelliine) are unstable and can be hydrolyzed rapidly to relatively stable DHP. DHP has been detected as the free form, conjugated with glutathione, cross-linking with DNA and proteins, and forming DNA adducts. Liver is the primary tissue to metabolize the pyrrolizidine alkaloids and produce the pyrrole metabolites. Previous tissue distribution studies [
18,
19] using radioisotope-labeled monocrotaline showed that red blood cells (RBCs) retained high concentrations of the radioactivity. The radioactivity associated with RBCs was significantly greater than those associated with other extrahepatic tissues, such as lung and kidney. The high concentration of pyrrolic metabolites associated with RBC proteins led the hypothesis that reactive intermediates of toxic pyrrolizidine alkaloids may be transported from liver to lung by RBCs [
17,
18]. Results from our study provided evidence that the reactive pyrrolic metabolites not only to bind with RBC proteins, but also to covalently bind with leukocyte DNA. The mechanism of the formation of DHP-derived DNA adducts in blood cells is not directly manifested from our data, however, constant interaction of leukocytes with liver cells where the reactive pyrrole metabolites are generated could cause the high level of DNA adducts in blood. A proposed metabolic activation pathway of riddelliine leading to its DHP-derived DNA adduct formation in blood cells is shown in
Scheme 2. The level of these DNA adducts and the sensitivity of the
32P-postlabeling/HPLC methodology suggest that DHP-derived DNA adducts in blood can serve as a potential biomarkers for assessing the risk from exposure to riddelliine.
Scheme 2.
Metabolic activation pathway of riddelliine to form DHP and DHP-3’-dGMP adducts.
Scheme 2.
Metabolic activation pathway of riddelliine to form DHP and DHP-3’-dGMP adducts.