Pharmacological and Epigenetic Regulators of NLRP3 Inflammasome Activation in Alzheimer’s Disease
Abstract
:1. Introduction
2. Results
2.1. Clinical Characteristics of the Individuals Enrolled in the Study
2.2. MTT Stavudine (D4T)
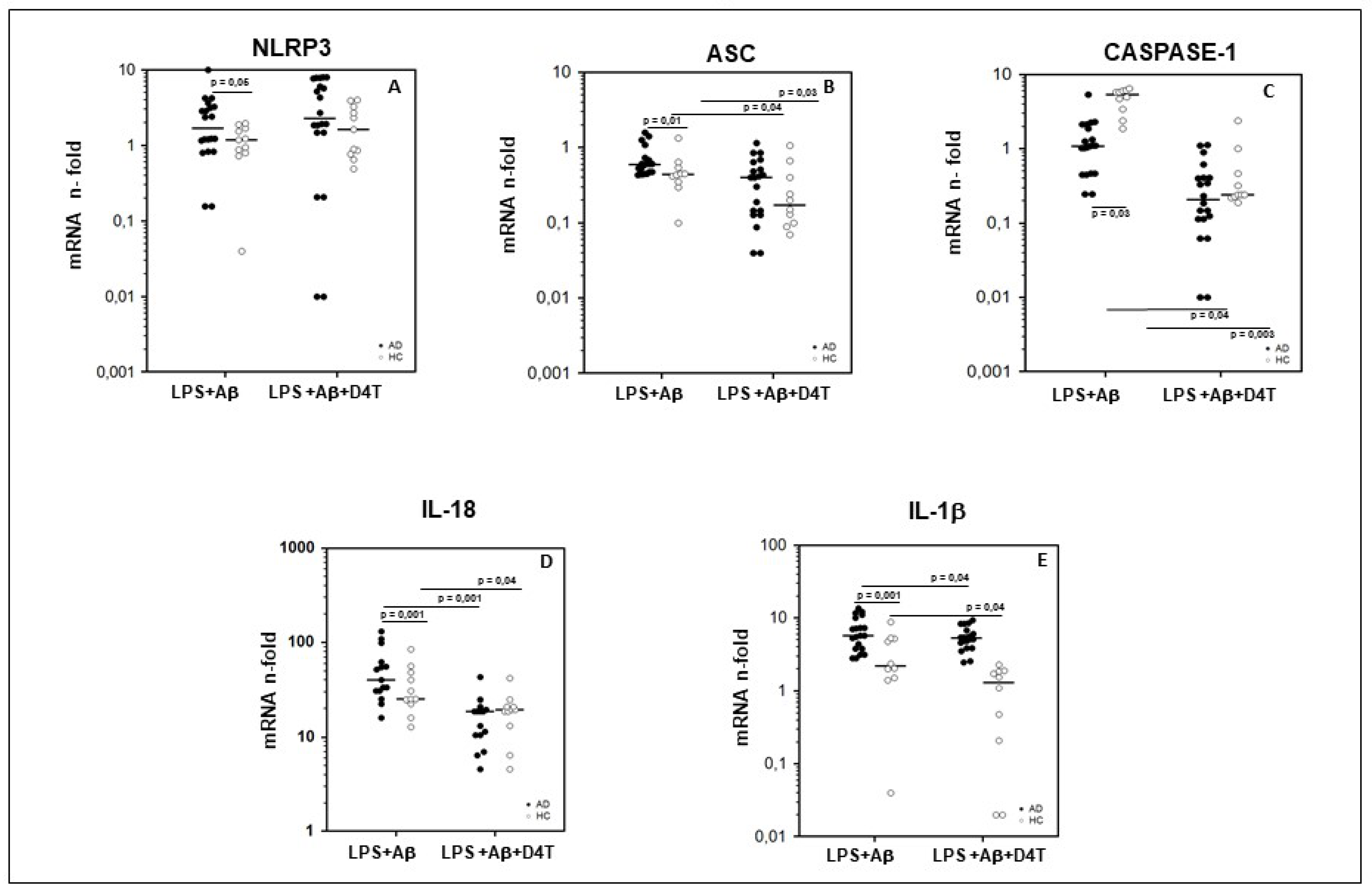
2.3. NLRP3 and Downstream Signaling of Inflammasome Gene Expression in PBMC
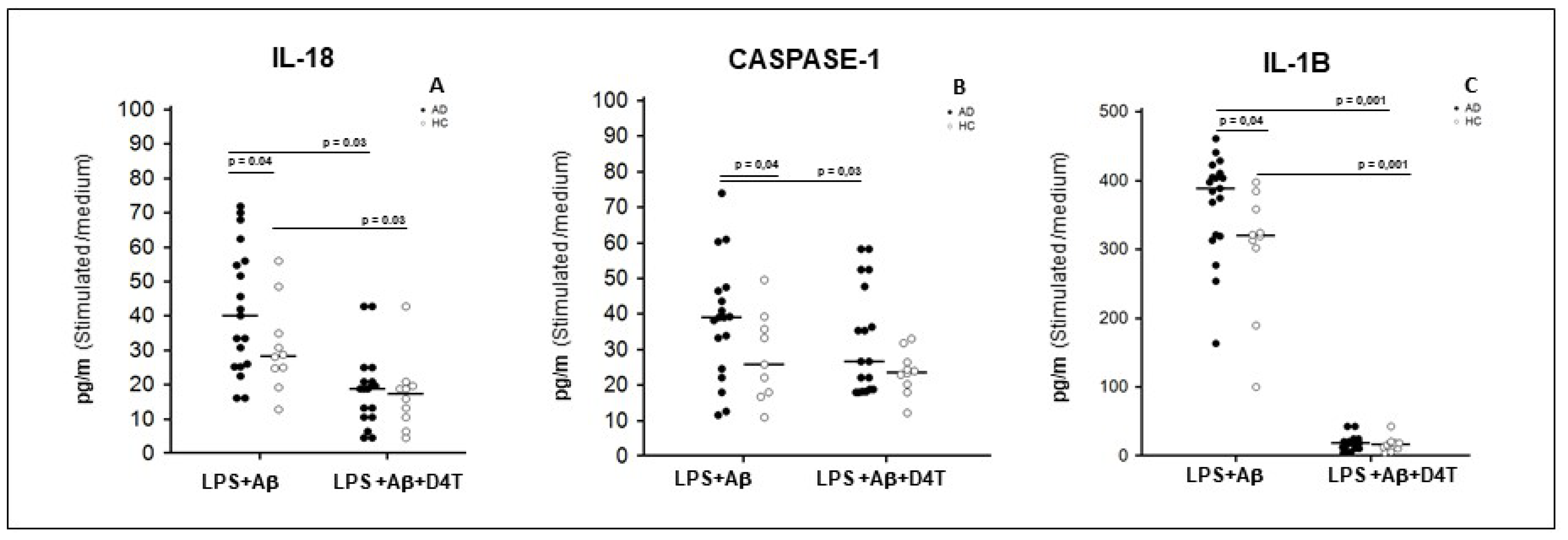
2.4. Inflammasome Related Cytokine Production in Supernatants of PBMC of AD Patients and HC
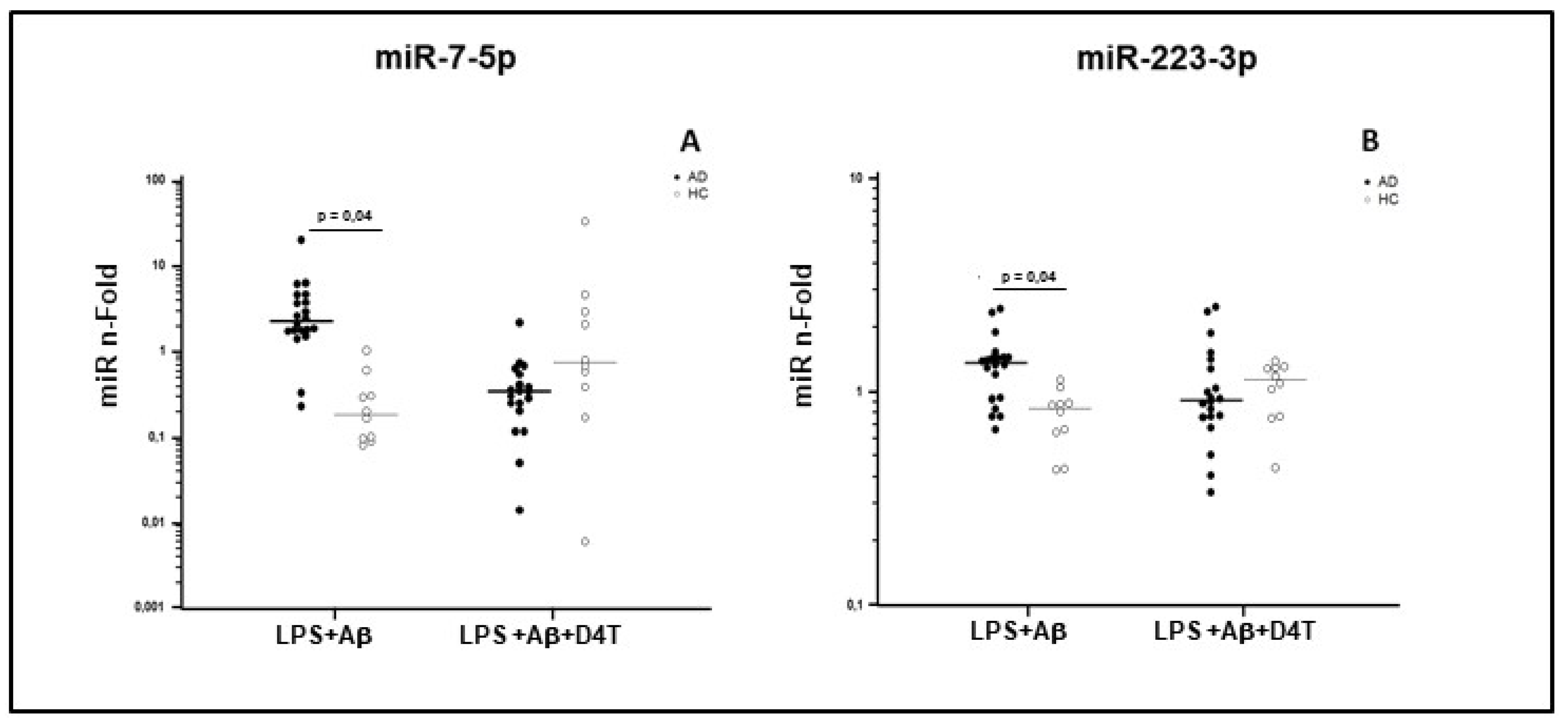
2.5. miRNAs Expression in PBMC of AD Patients and HC
3. Discussion
4. Materials and Methods
4.1. Patients and Controls
4.2. Blood Sample Collection and Cell Separation
4.3. Cell Cultures
4.4. D4T Cytotoxicity Assay
4.5. Total RNA Extraction
4.6. NLRP3-Inflammasome Pathway Quantitative Transcriptional Analysis by Real Time PCR
4.7. miRNA Quantitative Analysis by Real Time PCR
4.8. Elisa
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hardy, J.; Allsop, D. Amyloid deposition as the central event in the aetiology of Alzheimer’s disease. Trends Pharmacol. Sci. 1991, 12, 383–388. [Google Scholar] [CrossRef]
- Heppner, F.; Ransohoff, R.M.; Becher, B. Immune attack: The role of inflammation in Alzheimer disease. Nat. Rev. Neurosci. 2015, 16, 358–372. [Google Scholar] [CrossRef] [PubMed]
- Heneka, M.T.; Carson, M.J.; El Khoury, J.; Landreth, G.E.; Brosseron, F.; Feinstein, D.L.; Jacobs, A.H.; Wyss-Coray, T.; Vitorica, J.; Ransohoff, R.M.; et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 2015, 14, 388–405. [Google Scholar] [CrossRef] [Green Version]
- Fiala, M.; Zhang, L.; Gan, X.; Sherry, B.; Taub, D.; Graves, M.C.; Hama, S.; Way, D.; Weinand, M.; Lorton, D. Amyloid-beta induces chemokine secretion and monocyte migration across a human blood-brain barrier model. Mol. Med. 1998, 4, 480–489. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, Y.; Li, L.; Sun, X.-H. Monocytes and Alzheimer’s disease. Neurosci. Bull. 2011, 27, 115–122. [Google Scholar] [CrossRef] [Green Version]
- Takeshita, Y.; Ransohoff, R.M. Inflammatory cell trafficking across the blood-brain barrier: Chemokine regulation and in vitro models. Immunol. Rev. 2012, 248, 228–239. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Corraliza, I. Recruiting specialized macrophages across the borders to restore brain functions. Front. Cell. Neurosci. 2014, 8, 262. [Google Scholar] [CrossRef] [Green Version]
- Zenaro, E.; Piacentino, G.; Constantin, G. The blood-brain barrier in Alzheimer’s disease. Neurobiol. Dis. 2016, 107, 41–56. [Google Scholar] [CrossRef] [Green Version]
- Vérité, J.; Page, G.; Paccalin, M.; Julian, A.; Janet, T. Differential chemokine expression under the control of peripheral blood mononuclear cells issued from Alzheimer’s patients in a human blood brain barrier model. PLoS ONE 2018, 13, e0201232. [Google Scholar] [CrossRef] [PubMed]
- Schroder, K.; Tschopp, J. The inflammosomes. Cell 2010, 140, 821–832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martinon, F.; Burns, K.; Tschopp, J. The inflammasome: A molecular platform triggering activation of inflammatory caspases and processing of proIL-beta. J. Mol. Cell 2002, 10, 417–426. [Google Scholar] [CrossRef]
- Lamkanfi, M. Emerging inflammasome effector mechanisms. Nat. Rev. Immunol. 2011, 11, 213–220. [Google Scholar] [CrossRef]
- Broz, P.; Dixit, V.M. Inflammasomes: Mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 2016, 16, 407–420. [Google Scholar] [CrossRef] [PubMed]
- Hornung, V.; Ablasser, A.; Charrel-Dennis, M.; Bauernfeind, F.; Horvath, G.; Caffrey, D.R.; Latz, E.; Fitzgerald, K.A. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature 2009, 458, 514–518. [Google Scholar] [CrossRef] [Green Version]
- Latz, E.; Xiao, T.; Stutz, A. Activation and regulation of the inflammasomes. Nat. Rev. Immunol. 2013, 13, 397–411. [Google Scholar] [CrossRef]
- Vanaja, S.K.; Rathinam, V.A.; Fitzgerald, K.A. Mechanisms of inflammasome activation: Recent advances and novel insights. Trends Cell Biol. 2015, 25, 308–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Halle, A.; Hornung, V.; Petzold, G.C.; Stewart, C.R.; Monks, B.G.; Reinheckel, T.; Fitzgerald, K.A.; Latz, E.; Moore, K.J.; Golenbock, D.T. The NALP3 inflammasome is involved in the innate immune response to amyloid-beta. Nat. Immunol. 2008, 9, 857–865. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heneka, M.T.; Kummer, M.P.; Stutz, A.; Delekate, A.; Schwartz, S.; Vieira-Saecker, A.; Griep, A.; Axt, D.; Remus, A.; Tzeng, T.C.; et al. NLRP3 is activated in Alzheimer’s disease and contributes to pathology in APP/PS1 mice. Nature 2013, 493, 674–678. [Google Scholar] [CrossRef]
- Saresella, M.; La Rosa, F.; Piancone, F.; Zoppis, M.; Marventano, I.; Calabrese, E.; Rainone, V.; Nemni, R.; Mancuso, R.; Clerici, M. The NLRP3 and NLRP1 inflammasomes are activated in Alzheimer’s disease. Mol. Neurodegener. 2016, 11, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Fowler, B.J.; Gelfand, B.D.; Kim, Y.; Kerur, N.M.; Tarallo, V.; Hirano, Y.; Amarnath, S.; Fowler, D.H.; Radwan, M.; Young, M.T.; et al. Nucleoside reverse transcriptase inhibitors possess intrinsic anti-inflammatory activity. Science 2014, 346, 1000–1003. [Google Scholar] [CrossRef] [Green Version]
- La Rosa, F.; Saresella, M.; Marventano, I.; Piancone, F.; Ripamonti, E.; Al-Daghri, N.; Bazzini, C.; Zoia, C.P.; Conti, E.; Ferrarese, C.; et al. Stavudine Reduces NLRP3 Inflammasome Activation and Modulates Amyloid-β Autophagy. J. Alzheimers Dis. 2019, 72, 401–412. [Google Scholar] [CrossRef] [PubMed]
- Shao, B.-Z.; Xu, Z.-Q.; Han, B.-Z.; Su, D.-F.; Liu, C. NLRP3 inflammasome and its inhibitors: A review. Front. Pharmacol. 2015, 6, 262. [Google Scholar] [CrossRef] [Green Version]
- Lewis, B.P.; Burge, C.B.; Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [Green Version]
- Xie, Y.; Tang, L.; Han, Y.L.; Yang, L.; Xie, G.X.; Peng, J.W.; Tian, C.; Zhou, X.; Liu, Q.; Rong, X.M.; et al. Reduction in nitrogen fertilizer applications by the use of polymer-coated urea: Effect on maize yields and environmental impacts of nitrogen losses. J. Sci. Food Agric. 2019, 99, 2259–2266. [Google Scholar] [CrossRef]
- Gray, L.; Tachedjian, G.; Ellett, A.M.; Roche, M.; Cheng, W.-J.; Guillemin, G.; Brew, B.; Turville, S.; Wesselingh, S.L.; Gorry, P.R.; et al. The NRTIs Lamivudine, Stavudine and Zidovudine Have Reduced HIV-1 Inhibitory Activity in Astrocytes. PLoS ONE 2013, 8, e62196. [Google Scholar] [CrossRef] [Green Version]
- Cogswell, J.P.; Ward, J.; Taylor, I.A.; Waters, M.; Shi, Y.; Cannon, B.; Kelnar, K.; Kemppainen, J.; Brown, D.; Chen, C.; et al. Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. J. Alzheimers Dis. 2008, 14, 27–41. [Google Scholar] [CrossRef] [PubMed]
- Eacker, S.M.; Dawson, T.M.; Dawson, V.L. Understanding microRNAs in neurodegeneration. Nat. Rev. Neurosci. 2009, 10, 837–841. [Google Scholar] [CrossRef]
- Provost, P. Interpretation and applicability of microRNA data to the context of Alzheimer’s and age-related diseases. Aging 2010, 2, 166–169. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sonntag, K.C. MicroRNAs and deregulated gene expression networks in neurodegeneration. Brain Res. 2010, 1338, 48–57. [Google Scholar] [CrossRef] [Green Version]
- Miñones-Moyano, E.; Porta, S.; Escaramis, G.; Rabionet, R.; Iraola, S.; Kagerbauer, B.; Espinosa-Parrilla, Y.; Ferrer, I.; Estivill, X.; Martì, E. MicroRNA profiling of Parkinson’s disease brains identifies early down regulation of miR-34b/c which modulate mi-tochondrial function. Hum. Mol. Genet. 2011, 20, 3067–3078. [Google Scholar] [CrossRef] [PubMed]
- Filipowicz, W.; Bhattacharyya, S.N.; Sonenberg, N. Mechanisms of post-transcriptional regulation by mi-croRNAs: Are the answers in sight? Nat. Rev. Genet. 2008, 9, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Tezcan, G.; Martynova, E.V.; Gilazieva, Z.E.; McIntyre, A.; Rizvanov, A.A.; Khaiboullina, S.F. MicroRNA Post-transcrip-tional Regulation of the NLRP3 Inflammasome in Immunopathologies. Front. Pharmacol. 2019, 1, 451. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, H.Z.; Ong, K.L.; Seeher, K.; Armstrong, N.J.; Thalamuthu, A.; Brodaty, H.; Schadev, P.; Mather, K. Circulating microRNAs as biomarkers of Alz-heimer’s disease: Asystematic review. J. Alzheimers Dis. 2015, 49, 755–766. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Zhang, J. Circulating microRNAs: Potential and emergingbiomarkers for diagnosis of cardiovascular and cere-bro-vascular diseases. Biomed. Res. Int. 2015, 2015, 730535. [Google Scholar] [PubMed]
- Mancuso, R.; Agostini, S.; Hernis, A.; Zanzottera, M.; Bianchi, A.; Clerici, M. Circulatory miR-223–3 p Discriminates Between Parkinson’s and Alzheimer’s Patients. Sci. Rep. 2019, 9, 9393. [Google Scholar] [CrossRef]
- Jia, L.H.; Liu, Y.N. Downregulated serum miR-223 servers as biomarker in Alzheimer’s disease. Cell Biochem. Funct. 2016, 34, 233–237. [Google Scholar] [CrossRef]
- Chenyang, H.; Li, G.; Yi, Y.; Qiaobing, G.; Heping, S.; Yongjia, S.; Qingcai, J. Mechanism of mi-croRNA-22 in regulating neuroinflammation in Alzheimer’s disease. Brain Behav. 2020, 10, e01627. [Google Scholar] [CrossRef]
- Junichi, S. Molecular network analysis of human microRNA targetome: From cancers to Alzheimer’s disease. BioData Min. 2012, 5, 17. [Google Scholar] [CrossRef] [Green Version]
- Dorhoi, A.; Iannaccone, M.; Farinacci, M.; Fae, K.C.; Schreiber, J.; Moura-Alves, P.; Nouailles, G.; Mollenkopf, H.J.; Oberbeck-Muller, D.; Jorg, S.; et al. MicroRNA-223 controls sus-ceptibility to tuberculosis by regulating lung neutrophil recruitment. J. Clin. Investig. 2013, 123, 4836–4848. [Google Scholar] [CrossRef] [Green Version]
- Awad, F.; Assrawi, E.; Jumeau, C.; Lavialle, S.G.; Cobret, L.; Duquesnoy, P.; Piterboth, W.; Thomas, L.; Stankovic-Stojanovic, K.; Louvrier, C.; et al. Impact of human monocyte and mac-rophage polarization on NLR expression and NLRP3 inflammasome activation. PLoS ONE 2017, 12, e0175336. [Google Scholar] [CrossRef]
- Long, F.Q.; Kou, C.X.; Li, K.; Wu, Q.; Wang, Q. MiR-223–3 p inhibits rTp17-induced inflammasome activation and pyroptosis by targeting NLRP3. J. Cell Mol. Med. 2020, 24, 14405–14414. [Google Scholar] [CrossRef]
- Krol, J.; Loedige, I.; Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 2010, 11, 597–610. [Google Scholar] [CrossRef]
- Yates, L.A.; Norbury, C.J.; Gilbert, R.J. The long and short of microRNA. Cell 2013, 153, 516–519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKhann, G.; Drachman, D.; Folstein, M.; Katzman, R.; Price, D.; Stadlan, E.M. Clinical diagnosis of Alzheimer’s disease: Report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 1984, 34, 939. [Google Scholar] [CrossRef] [Green Version]
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders DSM-IV-R. 1994. Available online: http://www.psychiatryonline.com/DSMPDF/dsm-iv.pdf (accessed on 20 May 2021).
- Folstein, M.F.; Folstein, S.E.; McHugh, P.R. “Mini-mental state”. A practical method for grading the cognitive state of patients for the clinician. J. Psychiatr. Res. 1975, 12, 189–198. [Google Scholar] [CrossRef]
- Hughes, C.P.; Berg, L.; Danziger, W.L.; Coben, L.A.; Martin, R.L. A New Clinical Scale for the Staging of Dementia. Br. J. Psychiatry 1982, 140, 566–572. [Google Scholar] [CrossRef]
- Galimberti, D.; Bonsi, R.; Fenoglio, C.; Serpente, M.; Cioffi, S.M.G.; Fumagalli, G.; Arighi, A.; Ghezzi, L.; Arcaro, M.; Mercurio, M.; et al. Inflammatory molecules in Frontotemporal Dementia: Cerebrospinal fluid signature of progranulin mutation carriers. Brain Behav. Immun. 2015, 49, 182–187. [Google Scholar] [CrossRef] [PubMed]
- Ligthart, G.J.; Corberand, J.X.; Fournier, C.; Galanaud, P.; Hijmans, W.; Kennes, B.; Müller-Hermelink, H.K.; Steinmann, G.G. Admission criteria for immuno immunogerontological studies in man: The SENIEUR protocol. Mech. Ageing Dev. 1984, 28, 47–55. [Google Scholar] [CrossRef]
- Koch, W.; Teipel, S.; Mueller, S.; Benninghoff, J.; Wagner, M.; Bokde, A.L.W.; Hampel, H.; Coates, U.; Reiser, M.; Meindl, T. Diagnostic power of default mode network resting state fMRI in the detection of Alzheimer’s disease. Neurobiol. Aging 2012, 33, 466–478. [Google Scholar] [CrossRef]
- Mosmann, T. Rapid colorimetric assay for cellular growth and survival: Application to proliferation and cytotoxicity assays. J. Immunol. Methods 1983, 65, 55–63. [Google Scholar] [CrossRef]
- Fleige, S.; Pfaffl, M.W. RNA integrity and the effect on the real-time qRT-PCR performance. Mol. Asp. Med. 2006, 27, 126–139. [Google Scholar] [CrossRef] [PubMed]
| AD | HC | |
|---|---|---|
| N | 20 | 10 |
| Gender (M:F) | 05:05 | 04:06 |
| Age (years) | 76 a ± 5.9 b | 72 a ± 6.2 b |
| Level of education (years) | 8.25 a ± 2.71 b | 7.62 a ± 3.62 b |
| MMSE (Baseline) | 21 a ± 2.9 b | >28 |
| APOE e-4 cariers (%) | 20 a | 20 a |
| Amyloid-β (pg/mL) | 486 a ± 109 b | _ |
| Total-tau (pg/mL) | 747 a ± 206 b | _ |
| P-tau (pg/mL) | 84 a ± 44 b | _ |
| AD | HC | ||||||
|---|---|---|---|---|---|---|---|
| Condition | N | Median | IQR | N | Median | IQR | |
| miR-7 | LPS+Aβ | 20 | 2.170 | 1.625–4.455 | 10 | 0.100 | 0.083–0.823 |
| LPS+Aβ+D4T | 0.355 | 0.250–0.640 | 0.810 | 0.225–4.185 | |||
| miR-223 | LPS+Aβ | 20 | 1.424 | 0.964–1.511 | 10 | 0.865 | 0.659–0.920 |
| LPS+Aβ+D4T | 0.993 | 0.737–1.396 | 1.087 | 0.755–1.281 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
La Rosa, F.; Mancuso, R.; Agostini, S.; Piancone, F.; Marventano, I.; Saresella, M.; Hernis, A.; Fenoglio, C.; Galimberti, D.; Scarpini, E.; et al. Pharmacological and Epigenetic Regulators of NLRP3 Inflammasome Activation in Alzheimer’s Disease. Pharmaceuticals 2021, 14, 1187. https://0-doi-org.brum.beds.ac.uk/10.3390/ph14111187
La Rosa F, Mancuso R, Agostini S, Piancone F, Marventano I, Saresella M, Hernis A, Fenoglio C, Galimberti D, Scarpini E, et al. Pharmacological and Epigenetic Regulators of NLRP3 Inflammasome Activation in Alzheimer’s Disease. Pharmaceuticals. 2021; 14(11):1187. https://0-doi-org.brum.beds.ac.uk/10.3390/ph14111187
Chicago/Turabian StyleLa Rosa, Francesca, Roberta Mancuso, Simone Agostini, Federica Piancone, Ivana Marventano, Marina Saresella, Ambra Hernis, Chiara Fenoglio, Daniela Galimberti, Elio Scarpini, and et al. 2021. "Pharmacological and Epigenetic Regulators of NLRP3 Inflammasome Activation in Alzheimer’s Disease" Pharmaceuticals 14, no. 11: 1187. https://0-doi-org.brum.beds.ac.uk/10.3390/ph14111187






