Microbiological Evaluation of Water Quality from Urban Watersheds for Domestic Water Supply Improvement
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Area and Sample Collection

| Site # | Site locations | Land use | Geographic positioning system (GPS) |
|---|---|---|---|
| S1 | Ice House Canyon | Open Space | N34° 15.057 min.; W117° 37.977 min; 1,447 m elevation |
| M1 | Cucamonga Creek. at OCWD Ponds | Open Space | San Bernardino County Flood Control District (SBCFCD) |
| S2 | Chino Creek at Central Ave. | Urban runoff | N33° 58.420 min.; W117° 41.302 min; 174 m elevation |
| S3 | Chino Creek at Schaefer Ave. | Urban runoff | N34° 0.246 min.; W117° 43.628 min; 207 m elevation |
| S4 | San Antonio Wash at County Drive | Urban runoff + Commercial wash out | N30° 1.543 min.; W117° 43.652 min; 222 m elevation; |
| S5 | Chino Creek. at Riverside Drive | Urban runoff | N34° 1.144 min.; W117° 44.204 min; 207 m elevation; |
| S6 | Cypress Channel at Schaefer Ave. | Agricultural Runoff | N34° 0.262 min.; W117° 39.766 min 208 m elevation; |
| S7 | Cypress Channel at Kimball Ave. | Agricultural Runoff | N33° 58.113 min.; W117° 39.624 min 177 m elevation; |
| S8 | Cypress Channel at Golf Course | Agricultural Runoff | N33° 57.057 min.; W117° 39.555 min; 160 m elevation; |
| S9 | Big League Dreams at storm drain | Urban runoff | N33° 57.364 min.; W117 °40.788 min; 163 m elevation; |
| S11ww | Cucamonga Creek at Regional Water Recycling Plant #1 | Effluent from wastewater treatment plant | N34°; 1.853 min; W117° 35.946 min; Altitude:246 m |
| S11ur | Cucamonga Creek at Regional Water Recycling Plant #1 | Urban runoff+ wastewater | N34°; 1.853 min; W117° 35.946min; Altitude:246 m. |
| S12 | Chino Creek at Pine Ave. | Urban runoff+ wastewater | N33° 56.941 min.;W117° 39.986 min;155 m elevation; |
| S13 | Inland Empire Utilities Agency (IEUA) Regional Water Recycling Plant #5 | Effluent from wastewater treatment plant | N33° 57.840 min.; W117 ° 40.826 min; 180 m elevation; |
| S14 | IEUA Carbon Canyon Waste Reclamation Facility (CCWRF) | Effluent from wastewater treatment plant | N33 ° 58.799 min.; W117° 41.655 min; 184 m elevation; |
| ST2 | Santa Ana River at Prado Dam | Urban Runoff | N33°; 54.737 min; W117° 38.711 min Altitude: 141 m. |
| C3 | Prado Park outlet | Urban Runoff+ waste water discharge | N33°; 56.402 min; W117° 38.763min, 166 m |
| ST5 | Santa Ana River at River road | Urban Runoff | N33°; 55.405 min; W117° 35.894 min Altitude:155 m. |
| M5 | OCWD (Prado)Wetlands Effluent | Wetland treated (bacteria loaded) Orange County Water District (OCWD | N33°; 54.737 min; W117° 38.711 min Altitude: 141 m. |
2.2. Enumeration of TC, FC, and E. coli from Chino Basin
2.3. Isolation of Pathogenic E. coli from Chino Basin
2.4. Pulsed-Field Gel Electrophoresis
2.5. Antimicrobial Resistance
2.6. Sampling Collection During Sand Filtration Experiment
2.7. Enumeration of Heterotrophic Bacteria and E. coli in Water Before and After Sand Filtration
2.8. Statistical Analysis
3. Results
3.1. Fecal Indicator Bacterial Concentrations in Chino Basin
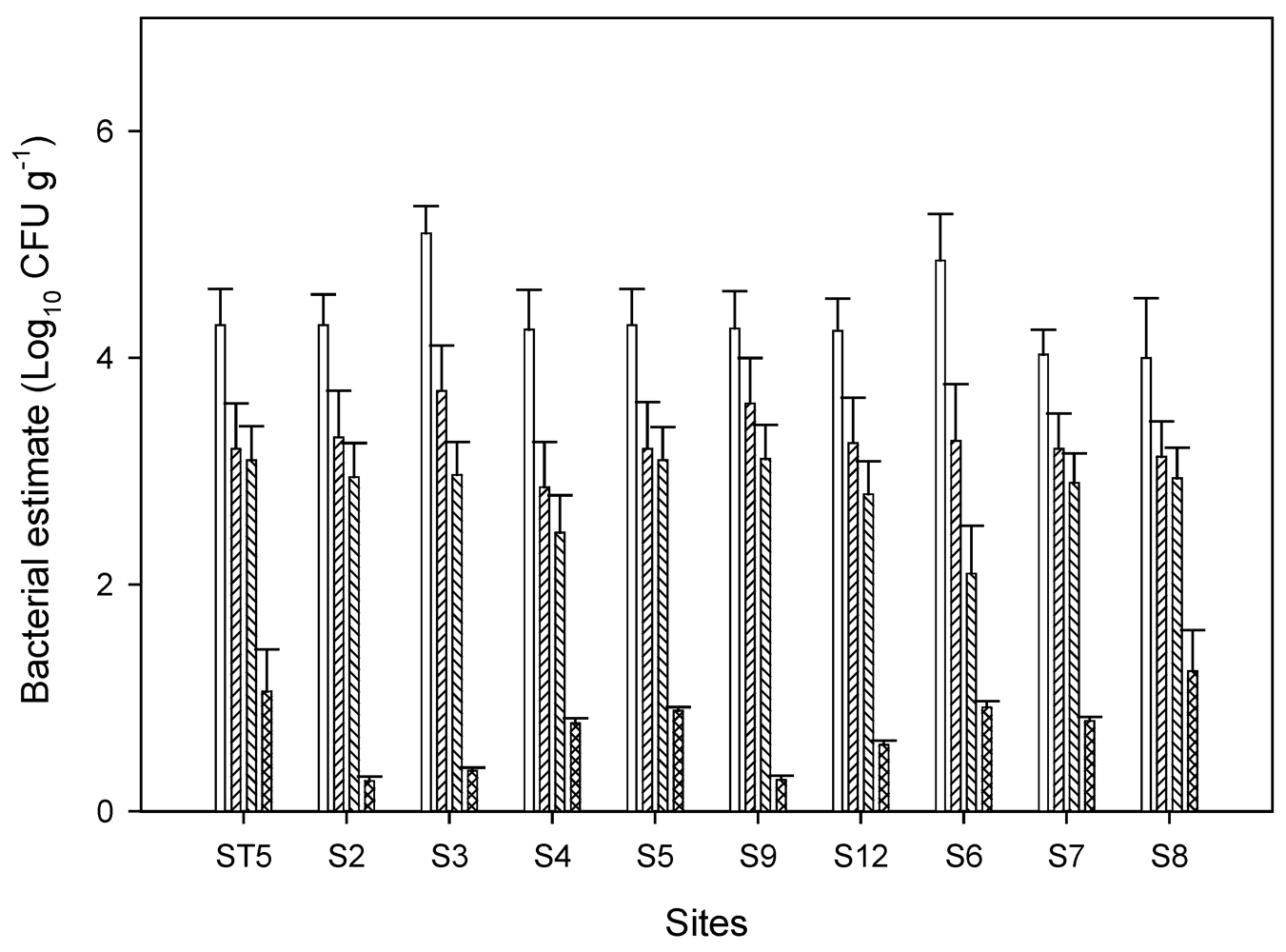
3.2. Characterization Pathogenic E. coli
| Sample Name | Amox | Amp | Ceph | Eryth | Rif | Strep | Tet | LT | STa | STb | CNF1 | CNF2 | O | H | stx2 | stx1 | eae | blaTEM | aadAI | tetA | tetB | tetC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| S8-1 | S | S | S | I | R | S | S | - | - | - | - | - | 11 | 40 | - | - | - | - | - | - | - | - |
| S8-2 | S | S | S | I | R | S | S | - | - | - | - | - | 11 | 40 | - | - | - | - | - | - | - | - |
| S8-3 | S | R | I | I | R | S | R | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | - |
| S8-4 | S | S | S | I | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | - |
| S8-5 | S | S | S | I | R | S | R | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | - |
| S8-6 | S | S | S | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + |
| ST5-1 | S | R | S | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + |
| ST5-2 | S | R | S | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + |
| ST5-3 | S | S | R | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | - |
| ST5-4 | S | R | S | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + |
| ST5-5 | S | R | S | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + |
| ST 5-6 | S | S | S | R | R | S | S | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + | |
| ST5-7 | S | S | S | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + |
| ST5-8 | S | S | S | R | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | + | - | + |
| ST 5-9 | S | S | S | R | R | S | R | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | + |
| ST5-10 | S | S | S | I | R | S | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | - |
| S12 | S | R | R | I | I | I | S | - | - | - | - | - | - | 11 | - | - | + | - | - | - | - | - |
3.3. Genetic Variations among E. coli Isolates from Watershed
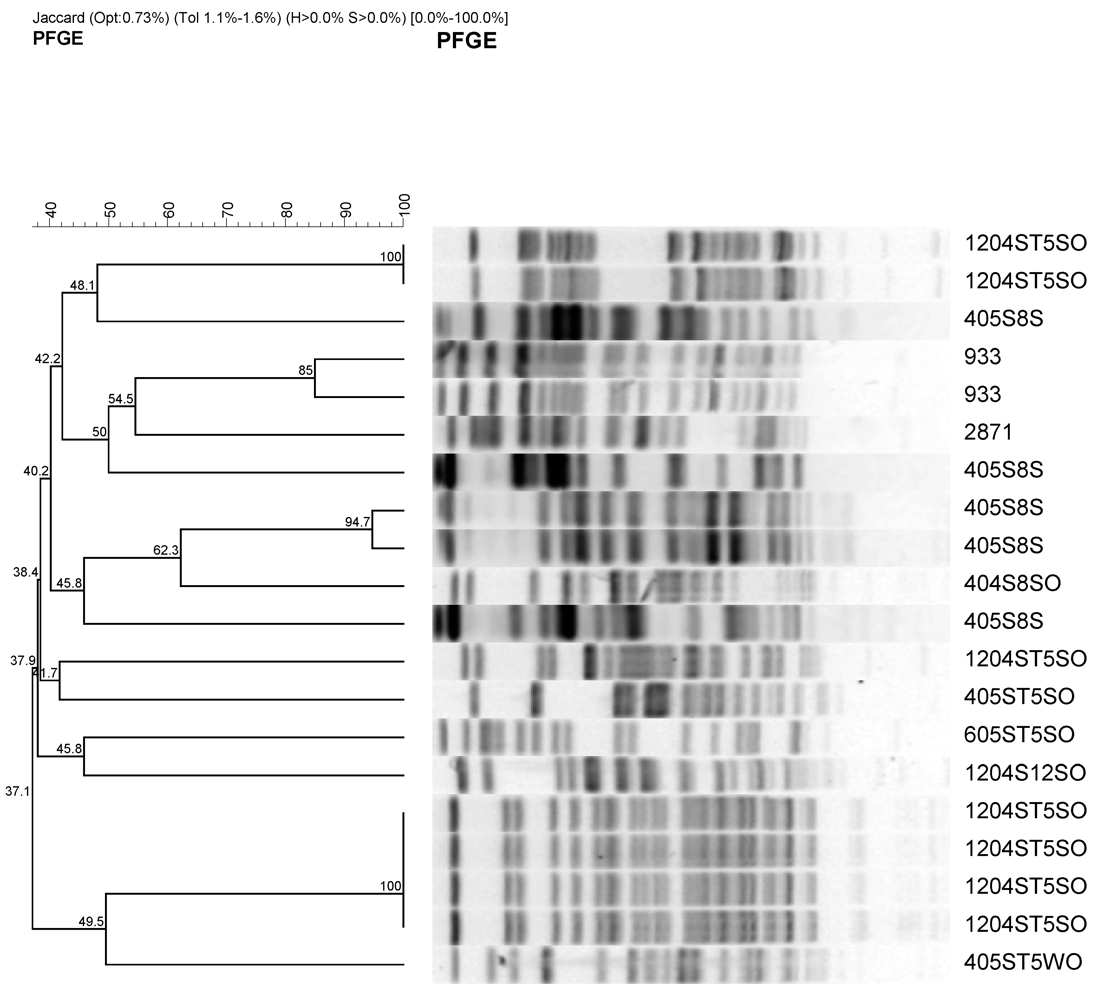
3.4. Fecal Indicator Bacterial Levels in Source Water and Aquifer Sand Material During Sand Filtration

4. Discussion
5. Conclusions
Acknowledgements
References
- Izbicki, J.A.; Pimentel, M.I.; Leddy, M.B. Microbial and Dissolved Organic Carbon Characterization of Storm Flow in the Santa Ana River at Imperial Highway, Southern California, 1999–2002; Scientific Investigation Report 2004-5116; U.S. Geological Survey: Reston, VA, USA, 2004. [Google Scholar]
- Rice, B.W. Staff Report on Bacterial Indicator Total Maximum Daily Loads in the Middle Santa Ana River Watershed; California Regional Water Quality Control Board: Santa Ana, CA, USA, 2005. [Google Scholar]
- Ishii, S.; Sadowsky, M.J. Escherichia coli in the environment: Implication for water quality and human health. Microbes Environ. 2008, 23, 101–108. [Google Scholar]
- Kaper, J.B.; Nataro, J.P.; Mobley, H.L.T. Pathogenic Escherichia coli. Nature Rev. 2004, 2, 123–140. [Google Scholar]
- Abraham, W.-R. Megacities as sources for pathogenic bacteria in rivers and their fate downstream. Int. J. Microbiol. 2011. [Google Scholar]
- Shelton, D.R.; Karns, J.S.; Higgins, J.A.; Van Kessel, J.S.; Perdue, M.L.; Belt, K.T.; Russell-Anelli, J.; DebRoy, C. Impact of microbial diversity on rapid detection of enterohemorrhagic Escherichia coli in surface waters. FEMS Microbiol. Lett. 2006, 261, 95–101. [Google Scholar]
- Hancock, D.D.; Besser, T.E.; Rice, D.H.; Herriott, D.E.; Tarr, P.I. Longitudinal study of Escherichia coli O157 in fourteen cattle herds. Epidemiol. Infect. 1997, 118, 193–195. [Google Scholar]
- Sherman, P.; Soni, R.; Karmali, M. Attaching and effacing adherence of Vero cytotoxin-producing Escherichia coli to rabbit intestinal epithelium in vivo. Infect. Immun. 1988, 56, 756–761. [Google Scholar]
- Ibekwe, A.M.; Murinda, S.E.; Graves, A.K. Genetic diversity and antimicrobial resistance of Escherichia coli from human and animal sources uncovers multiple resistances from human sources. PLoS One 2011, 6. [Google Scholar]
- Wildermuth Environmental, Inc, Optimum Basin Management Program—Chino Basin Dry-Year Yield Program; Prepared for Chino Basin Watermaster, Inland Empire Utilities Agency; Wildermuth Environmental, Inc.: Chino, CA, USA, 2003; pp. 1–141.
- Daly, C.; Gibson, W.P.; Doggett, M.; Smith, J.; Taylor, G. Up-to-date monthly climate maps for the conterminous United States. In Proceedings of the 14th Conference on Applied Climatology, 84th American Meteorological Society Annual Meeting, Seattle, WA, USA, 13–16 January 2004.
- Eaton, A.D.; Clesceri, L.S.; Greenberg, A.E.; Franson, M.A.H. Standard Method for Examination of Water and Wastewater; American Public Health Association: Washington, DC, USA, 1998. [Google Scholar]
- DebRoy, C.; Maddox, C.W. Identification of virulence attributes of gastrointestinal Escherichia coli isolates of veterinary significance. Anim. Health Res. Rev. 2001, 1, 129–140. [Google Scholar]
- Mora, A.; López, C.; Dabhi, G.; Blanco, M.; Blanco, J.E.; Alonso, M.P.; Herrera, A.; Mamani, R.; Bonacorsi, S.; Moulin-Schouleur, M.; Blanco, J. Extraintestinal pathogenic Escherichia coli O1:K1:H7/NM from human and avian origin: Detection of clonal groups B2 ST95 and D ST59 with different host distribution. BMC Microbiol. 2009, 9. [Google Scholar]
- Tenover, F.C.; Arbeit, R.D.; Goering, R.V.; Mickelsen, P.A.; Murray, B.E.; Persing, D.H.; Swaminathan, B. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J. Clin. Microbiol. 1995, 33, 2233–2239. [Google Scholar]
- Clinical and Laboratory Standard Institute, Performance Standards for Antimicrobial Susceptibility Tests—Ninth Edition: Approved Standard M02-A9; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2006.
- Gow, S.P.; Waldner, C.L.; Harel, J.; Boerlin, P. Associations between antimicrobial resistance genes in fecal generic Escherichia coli isolates from cow-calf herds in Western Canada. Appl. Environ. Microbiol. 2008, 74, 3658–3666. [Google Scholar]
- Sharma, R.; Munns, K.; Alexander, T.; Entz, T.; Mirzaagha, P.; Yanke, L.J.; Mulvey, M.; Topp, E.; McAllister, T. Diversity and distribution of commensal fecal Escherichia coli bacteria in beef cattle administered selected subtherapeutic antimicrobials in a feedlot setting. Appl. Environ. Microbiol. 2008, 74, 6178–6186. [Google Scholar]
- Ibekwe, A.M.; Lyon, S.R. Microbial characteristics through drinking water aquifer sand material. Eng. Life Sci. 2007, 7, 81–89. [Google Scholar]
- Ibekwe, A.M.; Lyon, S.R. Microbiological evaluation of fecal bacterial composition from surface water through aquifer sand material. J. Water Health 2008, 6, 411–421. [Google Scholar]
- American Public Health Association, Standard Methods for the Examination of Water and Wastewater, 19th ed; American Public Health Association: Washington, DC, USA, 1995.
- SAS Institute Inc, 2009 SAS/STAT User’s Guide; SAS Institute Inc.: Cary, NC, USA, 2009; Release 9.1.
- EPA, Ambient Water Quality Criteria for Bacteria—1986; EPA 440/5-84-002; EPA Office of Water: Washington, DC, USA, 1986.
- Ibekwe, A.M.; Lesch, S.M.; Bold, R.M.; Leddy, M.B.; Graves, A.K. Variations of indicator bacteria in a large urban watershed. In Proceedings of 21st Century Watershed Technology: Improving Water Quality and Environment Conference, Universidad EARTH, Costa Rica, 21–24 February 2010.
- Call, D.R.; Borucki, M.K.; Loge, F.J. Detection of bacterial pathogens in environmental samples using DNA microarrays. J. Microbiol. Methods 2003, 53, 235–243. [Google Scholar]
- Allen, H.K.; Donato, J.; Wang, H.H.; Cloud-Hansen, K.A.; Davies, J.; Handelsman, J. Call of the wild: Antibiotic resistance genes in natural environments. Nature Rev. Microbiol. 2010, 8, 251–259. [Google Scholar]
- Normark, B.H.; Normark, S. Evolution and spread of antibiotic resistance. J. Intern. Med. 2002, 252, 91–106. [Google Scholar]
- Jacoby, G.A. Extrachromosomal resistance in gram-negative organisms: The evolution of beta-lactamase. Trends Microbiol. 1994, 2, 357–360. [Google Scholar]
- Murinda, S.E.; Ebner, P.D.; Nguyen, L.T.; Mathew, A.G.; Oliver, S.P. Anitimicrobial resistance and class 1 integrons in pathogenic Escherichia coli from dairy farms. Foodborne Pathog. Dis. 2005, 2, 348–352. [Google Scholar]
- Tenover, F.C.; Rasheed, J.K. Genetic methods for detecting antimicrobial and antiviral resistance genes. In Manual of Clinical Microbiology, 7th; Murray, P.R., Baron, E.J., Pfaller, M.A., Tenover, F.C., Yolken, R.H., Eds.; American Society for Microbiology: Washington, DC, USA, 1998; pp. 1578–1592. [Google Scholar]
- Smith, D.L.; Harris, A.D.; Johnson, J.A.; Silbergeld, E.K.; Morris, J.G. Animal antibiotic use has an early but important impact on the emergence of antibiotic resistance in human commensal bacteria. Proc. Natl. Acad. Sci. USA 2002, 99, 6434–6439. [Google Scholar]
- Boerlin, P.; Travis, R.; Gyles, G.L.; Reid-Smith, R.; Janecko, N.; Lim, H.; Nicholson, V.; McEwen, S.A.; Friendship, R.; Archambault, M. Antimicrobial resistance and virulence genes of Escherichia coli from swine in Ontario. Appl. Environ. Microbiol. 2005, 71, 6753–6761. [Google Scholar]
- Travis, R.M.; Gyles, C.L.; Reid-Smith, R.; Poppe, C.; McEwen, S.A.; Friendship, R.; Janecko, N.; Boerlin, P. Chloramphenicol and kanamycin resistance among porcine Escherichia coli in Ontario. J. Antimicrob. Chemother. 2006, 58, 173–177. [Google Scholar]
- Aarestrup, F.M.; Kruse, H.; Tast, E.; Hammerum, A.M.; Jensen, L.B. Associations between the use of antimicrobial agents for growth promotion and the occurrence of resistance among Enterococcus faecium from broilers and pigs in Denmark, Finland, and Norway. Microbial. Drug Resist. 2000, 6, 63–70. [Google Scholar]
- Dwight, R.H.; Semenza, J.C.; Baker, D.B.; Olson, B.H. Association of urban runoff with coastal water quality in Orange County, California. Water Environ. Res. 2002, 74, 82–90. [Google Scholar]
- Reeves, R.L.; Grant, S.B.; Mrse, R.D.; Copil Oancea, C.M.; Sanders, B.F.; Boehm, A.B. Scaling and management of fecal indicator bacteria in runoff from a coastal urban watershed in southern California. Environ. Sci. Technol. 2004, 38, 2637–2648. [Google Scholar]
- Ferguson, D.M.; Moore, D.F.; Getrich, M.A.; Zhowandai, M.H. Enumeration and speciation of enterococci found in marine and intertidal sediments and coastal water in southern California. J. Appl. Microbiol. 2005, 99, 598–608. [Google Scholar]
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ibekwe, A.M.; Murinda, S.E.; Graves, A.K. Microbiological Evaluation of Water Quality from Urban Watersheds for Domestic Water Supply Improvement. Int. J. Environ. Res. Public Health 2011, 8, 4460-4476. https://0-doi-org.brum.beds.ac.uk/10.3390/ijerph8124460
Ibekwe AM, Murinda SE, Graves AK. Microbiological Evaluation of Water Quality from Urban Watersheds for Domestic Water Supply Improvement. International Journal of Environmental Research and Public Health. 2011; 8(12):4460-4476. https://0-doi-org.brum.beds.ac.uk/10.3390/ijerph8124460
Chicago/Turabian StyleIbekwe, A. Mark, Shelton E. Murinda, and Alexandria K. Graves. 2011. "Microbiological Evaluation of Water Quality from Urban Watersheds for Domestic Water Supply Improvement" International Journal of Environmental Research and Public Health 8, no. 12: 4460-4476. https://0-doi-org.brum.beds.ac.uk/10.3390/ijerph8124460




