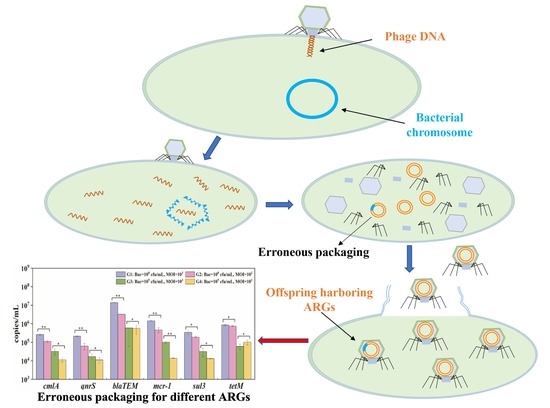
Evaluation of Potential ARG Packaging by Two Environmental T7-Like Phage during Phage-Host Interaction
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial and Phage Strains
2.2. Transmission Electron Microscopy (TEM)
2.3. Phage Testing Procedures
2.4. Genome Analysis
2.5. Potential ARG Packaging
2.6. Data Analysis
2.7. Intact ARG Testing
3. Results
3.1. Phage Characterization
3.2. Genome Annotation and Characteristics
3.3. Potential ARG Packaging
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Xiong, W.; Liu, P.; Xie, X.; Zeng, J.; Sun, Y.; Zeng, Z. Metagenomic Insights Into the Contribution of Phages to Antibiotic Resistance in Water Samples Related to Swine Feedlot Wastewater Treatment. Front. Microbiol. 2018, 9, 2474. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wright, G.D. Antibiotic resistance in the environment: A link to the clinic? Curr. Opin. Microbiol. 2010, 13, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Fruth, A.; Prager, R.; Tietze, E.; Rabsch, W.; Flieger, A. Molecular epidemiological view on Shiga toxin-producing Escherichia coli causing human disease in Germany: Diversity, prevalence, and outbreaks. Int. J. Med. Microbiol. 2015, 305, 697–704. [Google Scholar] [CrossRef] [PubMed]
- Diarra, M.S.; Malouin, F. Antibiotics in Canadian poultry productions and anticipated alternatives. Front. Microbiol. 2014, 5, 282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Kraker, M.E.; Davey, P.G.; Grundmann, H.; BURDEN Study Group. Mortality and hospital stay associated with resistant Staphylococcus aureus and Escherichia coli bacteremia: Estimating the burden of antibiotic resistance in Europe. PLoS Med. 2011, 8, e1001104. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jakobsen, L.; Spangholm, D.J.; Pedersen, K.; Jensen, L.; Emborg, H.-D.; Agersø, Y.; Aarestrup, F.M.; Hammerum, A.M.; Frimodt-Møller, N. Broiler chickens, broiler chicken meat, pigs and pork as sources of ExPEC related virulence genes and resistance in Escherichia coli isolates from community-dwelling humans and UTI patients. Int. J. Food Microbiol. 2010, 142, 264–272. [Google Scholar] [CrossRef] [PubMed]
- Jang, J.; Hur, H.-G.; Sadowsky, M.J.; Byappanahalli, M.N.; Yan, T.; Ishii, S. Environmental Escherichia coli: Ecology and public health implications—A review. J. Appl. Microbiol. 2017, 123, 570–581. [Google Scholar] [CrossRef] [Green Version]
- Alekshun, M.N.; Levy, S.B. Molecular mechanisms of antibacterial multidrug resistance. Cell 2007, 128, 1037–1050. [Google Scholar] [CrossRef] [Green Version]
- Colavecchio, A.; Cadieux, B.; Lo, A.; Goodridge, L.D. Bacteriophages Contribute to the Spread of Antibiotic Resistance Genes among Foodborne Pathogens of the Enterobacteriaceae Family—A Review. Front. Microbiol. 2017, 8, 1108. [Google Scholar] [CrossRef] [Green Version]
- Colomer-Lluch, M.; Jofre, J.; Muniesa, M. Antibiotic resistance genes in the bacteriophage DNA fraction of environmental samples. PLoS ONE 2011, 6, e17549. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marti, E.; Variatza, E.; Balcazar, J.L. The role of aquatic ecosystems as reservoirs of antibiotic resistance. Trends Microbiol. 2014, 22, 36–41. [Google Scholar] [CrossRef] [PubMed]
- Clokie, M.R.; Millard, A.D.; Letarov, A.V.; Heaphy, S. Phages in nature. Bacteriophage 2011, 1, 314–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Torres-Barceló, C. The disparate effects of bacteriophages on antibiotic-resistant bacteria. Emerg. Microbes Infect. 2018, 7, 11–12. [Google Scholar] [CrossRef]
- Ubukata, K.; Konno, M.; Fuiii, R. Transduction of drug resistance to tetracycline, chloramphenicol, macrolides, lincomycin and clindamycin with phages induced from Streptococcus pyogenes. Antiotics 1975, 28, 681–688. [Google Scholar] [CrossRef] [Green Version]
- Taylor, V.L.; Fitzpatrick, A.D.; Islam, Z.; Maxwell, K.L. The Diverse Impacts of Phage Morons on Bacterial Fitness and Virulence. Adv. Virus Res. 2019, 103, 1–31. [Google Scholar]
- Wang, M.; Liu, P.; Zhou, Q.; Tao, W.; Sun, Y.; Zeng, Z. Estimating the contribution of bacteriophage to the dissemination of antibiotic resistance genes in pig feces. Environ. Pollut. 2018, 238, 291–298. [Google Scholar] [CrossRef]
- Fernandez-Orth, D.; Miro, E.; Brown-Jaque, M.; Rodríguez-Rubio, L.; Espinal, P.; Rodriguez-Navarro, J.; González-López, J.J.; Muniesa, M.; Navarro, F. Faecal phageome of healthy individuals: Presence of antibiotic resistance genes and variations caused by ciprofloxacin treatment. J. Antimicrob. Chemother. 2019, 74, 854–864. [Google Scholar] [CrossRef]
- Subirats, J.; Sànchez-Melsió, A.; Borrego, C.M.; Balcázar, J.L.; Simonet, P. Metagenomic analysis reveals that bacteriophages are reservoirs of antibiotic resistance genes. Int. J. Antimicrob. Agents 2016, 48, 163–167. [Google Scholar] [CrossRef]
- Goh, S.; Hussain, H.; Chang, B.J.; Emmett, W.; Riley, T.V.; Mullany, P. Phage varphiC2 mediates transduction of Tn6215, encoding erythromycin resistance, between Clostridium difficile strains. MBio 2013, 4, e008401-3. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.; Zeng, X.; Yang, Q.; Yang, C. Identification of a bacteriophage from an environmental multidrug-resistant E. coli isolate and its function in horizontal transfer of ARGs. Sci. Total Environ. 2018, 639, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, R.F.; Rumi, V.; Marchisio, M.; Cejas, D.; Radice, M.; Vay, C.; Barrios, R.; Gutkind, G.O.; Di Conza, J.A. Fast and easy detection of CMY-2 in Escherichia coli by direct MALDI-TOF mass spectrometry. J. Microbiol. Methods 2018, 148, 22–28. [Google Scholar] [CrossRef] [PubMed]
- De Boer, M.; Heuer, C.; Hussein, H.; McDougall, S. Minimum inhibitory concentrations of selected antimicrobials against Escherichia coli and Trueperella pyogenes of bovine uterine origin. J. Dairy Sci. 2015, 98, 4427–4438. [Google Scholar] [CrossRef] [Green Version]
- Amarillas, L.; Rubí-Rangel, L.; Chaidez, C.; Gonzalez-Robles, A.; Lightbourn-Rojas, L.; León-Félix, J. Isolation and Characterization of phiLLS, a Novel Phage with Potential Biocontrol Agent against Multidrug-Resistant Escherichia coli. Front. Microbiol. 2017, 8, 1355. [Google Scholar] [CrossRef]
- Chen, X.; Guo, J.; Liu, Y.; Chai, S.; Ma, R.; Munguntsetseg, B. Characterization and adsorption of a Lactobacillus plantarum virulent phage. J. Dairy Sci. 2019, 102, 3879–3886. [Google Scholar] [CrossRef] [PubMed]
- Attai, H.; Brown, P.J.B. Isolation and Characterization T4- and T7-Like Phages that Infect the Bacterial Plant Pathogen Agrobacterium tumefaciens. Viruses 2019, 11, 528. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.; Yu, X.; Gu, Y.; Huang, X.; Liu, G.; Liu, X. Characterization and Genomic Study of Phage vB_EcoS-B2 Infecting Multidrug-Resistant Escherichia coli. Front. Microbiol. 2018, 9, 793. [Google Scholar] [CrossRef]
- Thurber, R.V.; Haynes, M.; Breitbart, M.; Wegley, L.; Rohwer, F. Laboratory procedures to generate viral metagenomes. Nat. Protoc. 2009, 4, 470–483. [Google Scholar] [CrossRef]
- Jia, B.; Raphenya, A.R.; Alcock, B.; Waglechner, N.; Guo, P.; Tsang, K.K.; Lago, B.A.; Dave, B.M.; Pereira, S.; Sharma, A.N.; et al. CARD 2017: Expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017, 45, D566–D573. [Google Scholar] [CrossRef]
- Li, B.; Yang, Y.; Ma, L.; Ju, F.; Guo, F.; Tiedje, J.M.; Zhang, T. Metagenomic and network analysis reveal wide distribution and co-occurrence of environmental antibiotic resistance genes. ISME J. 2015, 9, 2490–2502. [Google Scholar] [CrossRef] [Green Version]
- Xiong, W.; Wang, Y.; Sun, Y.; Ma, L.; Zeng, Q.; Jiang, X.-T.; Li, A.-D.; Zeng, Z.-L.; Zhang, T. Antibiotic-mediated changes in the fecal microbiome of broiler chickens define the incidence of antibiotic resistance genes. Microbiome 2018, 6, 34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Y.; Li, B.; Ju, F.; Zhang, T. Exploring variation of antibiotic resistance genes in activated sludge over a four-year period through a metagenomic approach. Environ. Sci. Technol. 2013, 47, 10197–10205. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Chan, P. tRNAscan-SE On-line: Integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 2016, 44, W54–W57. [Google Scholar] [CrossRef]
- Hamdi, S.; Rousseau, G.M.; Labrie, S.J.; Kourda, R.S.; Tremblay, D.M.; Moineau, S.; Slama, K.B. Characterization of Five Podoviridae Phages Infecting Citrobacter freundii. Front. Microbiol. 2016, 7, 1023. [Google Scholar] [CrossRef] [Green Version]
- Ackermann, H.W. Phage Classification and Characterization. Methods Mol. Biol. 2009, 501, 127–140. [Google Scholar] [PubMed]
- Qiao, M.; Ying, G.-G.; Singer, A.C.; Zhu, Y. Review of antibiotic resistance in China and its environment. Environ. Int. 2018, 110, 160–172. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hendrix, R.W.; Hatfull, G.F.; Ford, M.E.; Smith, M.C.; Burns, R.N. Evolutionary relationships among diverse bacteriophages and prophages All the world’s a phage. Proc. Natl. Acad. Sci. USA 1999, 96, 2192–2197. [Google Scholar] [CrossRef] [Green Version]
- Kenzaka, T.; Tani, K.; Nasu, M. High-frequency phage-mediated gene transfer in freshwater environments determined at single-cell level. ISME J. 2010, 4, 648–659. [Google Scholar] [CrossRef] [Green Version]
- Eggers, C.H.; Gray, C.M.; Preisig, A.M.; Glenn, D.M.; Pereira, J.; Ayers, R.W.; Alshahrani, M.; Acabbo, C.; Becker, M.R.; Bruenn, K.N.; et al. Phage-mediated horizontal gene transfer of both prophage and heterologous DNA by varphiBB-1, a bacteriophage of Borrelia burgdorferi. Pathog. Dis. 2016, 74, ftw107. [Google Scholar] [CrossRef] [Green Version]
- Maslanova, I.; Doškař, J.; Varga, M.; Kuntová, L.; Muzik, J.; Malúšková, D.; Růžičková, V.; Pantůček, R. Bacteriophages of Staphylococcus aureus efficiently package various bacterial genes and mobile genetic elements including SCCmec with different frequencies. Environ. Microbiol. Rep. 2013, 5, 66–73. [Google Scholar] [CrossRef]
- Lefkowitz, E.J.; Dempsey, D.M.; Hendrickson, R.C.; Orton, R.J.; Siddell, S.G.; Smith, D.B. Virus taxonomy: The database of the International Committee on Taxonomy of Viruses (ICTV). Nucleic Acids Res. 2018, 46, D708–D717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Y.; Ma, Y.; Yao, S.; Jiang, Z.; Pei, J.; Cheng, C. Characterization, Genome Sequence, and Analysis of Escherichia Phage CICC 80001, a Bacteriophage Infecting an Efficient L-Aspartic Acid Producing Escherichia coli. Food Environ. Virol. 2016, 8, 182–186. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Xu, Y.; Xu, J.; Yu, X.; Huang, X.; Liu, G.; Liu, X. Identification of novel bacteriophage vB_EcoP-EG1 with lytic activity against planktonic and biofilm forms of uropathogenic Escherichia coli. Appl. Microbiol. Biotechnol. 2019, 103, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Mao, Z.; Ji, W.; Zou, Q.; Yan, Y.; Sun, J. Bioactivity of Terminase Large Subunit Encoded by Streptococcus suis Phage. Shanghai Jiaotong Univ. Agric. Sci. 2013, 31, 70–75. [Google Scholar]
- Li, S.; Fan, H.; An, X.; Fan, H.; Jiang, H.; Chen, Y.; Tong, Y. Scrutinizing virus genome termini by high-throughput sequencing. PLoS ONE 2014, 9, e85806. [Google Scholar] [CrossRef] [Green Version]
- Breitbart, M. Global distribution of nearly identical phage-encoded DNA sequences. FEMS Microbiol. Lett. 2004, 236, 249–256. [Google Scholar] [CrossRef]
- Steven, A.C.; Trus, B.; Maizel, J.; Unser, M.; Parry, D.; Wall, J.; Hainfeld, J.; Studier, F. Molecular substructure of a viral receptor-recognition protein. The gp17 tail-fiber of bacteriophage T7. J. Mol. Biol. 1988, 200, 351–365. [Google Scholar] [CrossRef]
- Nobrega, F.L.; Vlot, M.; De Jonge, P.A.; Dreesens, L.L.; Beaumont, H.J.E.; Lavigne, R.; Dutilh, B.E.; Brouns, S.J.J. Targeting mechanisms of tailed bacteriophages. Nat. Rev. Microbiol. 2018, 16, 760–773. [Google Scholar] [CrossRef]
- Hu, M.; Zhang, H.; Gu, D.; Ma, Y.; Zhou, X. Identification of a novel bacterial receptor that binds tail tubular proteins and mediates phage infection of Vibrio parahaemolyticus. Emerg. Microbes Infect. 2020, 9, 855–867. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Novick, R. Phage-mediated intergeneric transfer of toxin genes. Science 2009, 323, 139–141. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Zeng, Z.; Jiang, F.; Zheng, Y.; Shen, H.; Macedo, N.; Sun, Y.; Sahin, O.; Li, G. Role of enterotoxigenic Escherichia coli prophage in spreading antibiotic resistance in a porcine-derived environment. Environ. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Lekunberri, I.; Subirats, J.; Borrego, C.M.; Balcázar, J.L. Exploring the contribution of bacteriophages to antibiotic resistance. Environ. Pollut. 2017, 220 Pt B, 981–984. [Google Scholar] [CrossRef]
- Liu, Y.-Y.; Wang, Y.; Walsh, T.R.; Yi, L.-X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 1611–1668. [Google Scholar] [CrossRef]
- Patterson, A.G.; Jackson, S.A.; Taylor, C.; Evans, G.B.; Salmond, G.P.; Przybilski, R.; Staals, R.H.J.; Fineran, P.C. Quorum Sensing Controls Adaptive Immunity through the Regulation of Multiple CRISPR-Cas Systems. Mol. Cell 2016, 64, 1102–1108. [Google Scholar] [CrossRef] [PubMed] [Green Version]







| Gene | Size (bp) | HZA135 | HZA50 | HZP2 | HZ2R8 |
|---|---|---|---|---|---|
| blaTEM | 247(28.7%) | + | + | + | + |
| 752(87.3%) | + | + | − | − | |
| qnrS | 428 (65.1%) | + | + | + | + |
| 619 (94.2%) | + | + | − | − | |
| cmlA | 158 (12.5%) | + | + | + | + |
| 1188 (94.3%) | + | + | − | − | |
| mcr-1 | 220 (13.5%) | + | + | + | + |
| 1495 (92.0%) | + | + | − | − | |
| tetM | 171 (8.9%) | + | + | + | − |
| 1861 (96.9%) | + | − | − | − | |
| sul3 | 128 (16.1%) | + | − | + | − |
| 714 (90.3%) | + | − | − | − | |
| ermB | 189 (25.6%) | + | − | − | − |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, J.; Liu, P.; Feng, F.; Zhang, J.; Li, F.; Wang, M.; Sun, Y. Evaluation of Potential ARG Packaging by Two Environmental T7-Like Phage during Phage-Host Interaction. Viruses 2020, 12, 1060. https://0-doi-org.brum.beds.ac.uk/10.3390/v12101060
Liu J, Liu P, Feng F, Zhang J, Li F, Wang M, Sun Y. Evaluation of Potential ARG Packaging by Two Environmental T7-Like Phage during Phage-Host Interaction. Viruses. 2020; 12(10):1060. https://0-doi-org.brum.beds.ac.uk/10.3390/v12101060
Chicago/Turabian StyleLiu, Junlin, Peng Liu, Fenglin Feng, Junxuan Zhang, Fulin Li, Mianzhi Wang, and Yongxue Sun. 2020. "Evaluation of Potential ARG Packaging by Two Environmental T7-Like Phage during Phage-Host Interaction" Viruses 12, no. 10: 1060. https://0-doi-org.brum.beds.ac.uk/10.3390/v12101060