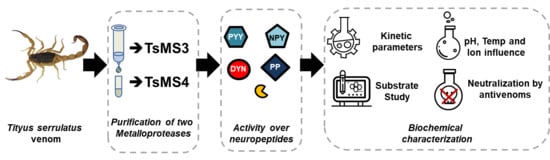
Purification and Biochemical Characterization of TsMS 3 and TsMS 4: Neuropeptide-Degrading Metallopeptidases in the Tityus serrulatus Venom
Abstract
:1. Introduction
2. Results
2.1. Isolation of TsMS 3 and TsMS 4
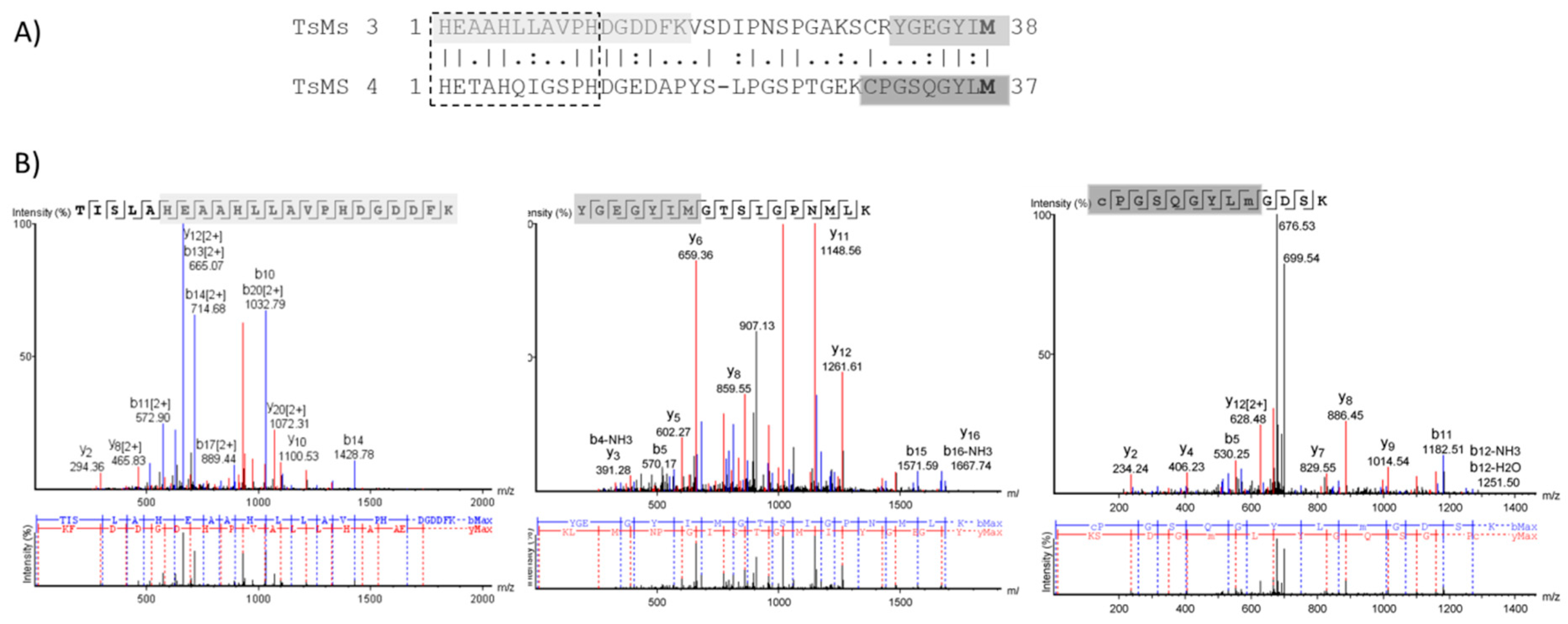
2.2. Bioinformatic Analysis
2.3. Comparative Analysis of Metalloserrulases Biochemical Parameters
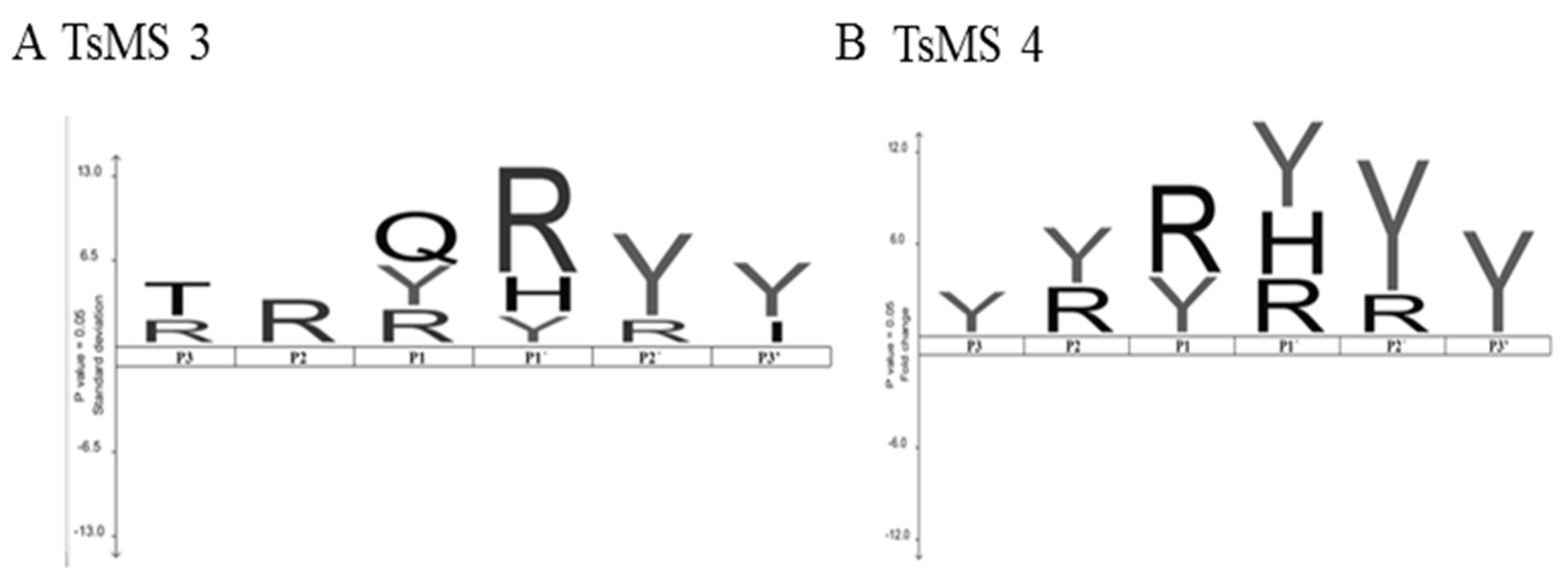
2.4. Determination of Cleavage Sites
2.5. Kinetic Parameters Determination
2.6. In Vitro Neutralization Assay of the Activity of Metalloserrulases by Commercial Antivenoms
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Reagents
5.2. Venoms and Antivenoms
5.3. Purification of Metalloserrulases 3 and 4 from Tityus serrulatus Venom
5.3.1. Chromatographic Steps
5.3.2. Screening Using FRET Substrate
5.3.3. Screening Using Human Neuropeptides
5.4. Characterization of Isolated Proteases
5.4.1. SDS-PAGE—In Gel Digestion and Mass Spectrometry
5.4.2. Biochemical Characterization: Effect of pH, Cation Concentration and Temperature
5.4.3. Analysis of The Cleavage Sites
5.4.4. Kinetic Parameters for The Hydrolysis of FRET Substrates by TsMS 4
5.5. Sera Neutralization Assays
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- SINAN. Acidente Por Animais Peçonhentos—Notificações Registradas No Sistema De Informação De Agravos De Notificação—Brasil. Available online: http://tabnet.datasus.gov.br/cgi/deftohtm.exe?sinannet/cnv/animaisbr.def;access (accessed on 10 October 2018).
- Szilagyi-Zecchin, V.J.; Fernandes, A.L.; Castagna, C.; Voltolini, J. Abundance of scorpions Tityus serrulatus and Tityus bahiensis associated with climate in urban area (Scorpiones, Buthidae). Indian J. Arachnol. 2012, 1, 15–23. [Google Scholar]
- Bucaretchi, F.; Baracat, E.C.; Nogueira, R.J.; Chaves, A.; Zambrone, F.A.; Fonseca, M.R.; Tourinho, F.S. A comparative study of severe scorpion envenomation in children caused by Tityus bahiensis and Tityus serrulatus. Rev. Inst. Med. Trop. Sao Paulo 1995, 37, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Bucaretchi, F.; Fernandes, L.C.; Fernandes, C.B.; Branco, M.M.; Prado, C.C.; Vieira, R.J.; De Capitani, E.M.; Hyslop, S. Clinical consequences of Tityus bahiensis and Tityus serrulatus scorpion stings in the region of Campinas, southeastern Brazil. Toxicon 2014, 89, 17–25. [Google Scholar] [CrossRef]
- Casewell, N.R.; Wüster, W.; Vonk, F.J.; Harrison, R.A.; Fry, B.G. Complex cocktails: The evolutionary novelty of venoms. Trends Ecol. Evol. 2013, 28, 219–229. [Google Scholar] [CrossRef]
- Waheed, H.; Moin, S.F.; Choudhary, M.I. Snake Venom: From Deadly Toxins to Life-saving Therapeutics. Curr. Med. Chem. 2017, 24, 1874–1891. [Google Scholar] [CrossRef]
- Calvete, J.J.; Sanz, L.; Angulo, Y.; Lomonte, B.; Gutiérrez, J.M. Venoms, venomics, antivenomics. FEBS Lett. 2009, 583, 1736–1743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Paes Leme, A.F.; Prezoto, B.C.; Yamashiro, E.T.; Bertholim, L.; Tashima, A.K.; Klitzke, C.F.; Camargo, A.C.; Serrano, S.M. Bothrops protease A, a unique highly glycosylated serine proteinase, is a potent, specific fibrinogenolytic agent. J. Thromb. Haemost. 2008, 6, 1363–1372. [Google Scholar] [CrossRef] [PubMed]
- White, J. Snake venoms and coagulopathy. Toxicon 2005, 45, 951–967. [Google Scholar] [CrossRef]
- Cologna, C.T.; Marcussi, S.; Giglio, J.R.; Soares, A.M.; Arantes, E.C. Tityus serrulatus scorpion venom and toxins: An overview. Protein Pept. Lett. 2009, 16, 920–932. [Google Scholar] [CrossRef] [PubMed]
- Morgenstern, D.; Rohde, B.H.; King, G.F.; Tal, T.; Sher, D.; Zlotkin, E. The tale of a resting gland: Transcriptome of a replete venom gland from the scorpion Hottentotta judaicus. Toxicon 2011, 57, 695–703. [Google Scholar] [CrossRef] [PubMed]
- Alvarenga, É.R.; Mendes, T.M.; Magalhaes, B.F.; Siqueira, F.F.; Dantas, A.E.; Barroca, T.M.; Horta, C.C.; Kalapothakis, E. Transcriptome analysis of the Tityus serrulatus scorpion venom gland. Open J. Genet. 2012, 2, 210. [Google Scholar] [CrossRef]
- Almeida, D.D.; Scortecci, K.C.; Kobashi, L.S.; Agnez-Lima, L.F.; Medeiros, S.R.; Silva-Junior, A.A.; Junqueira-de-Azevedo, I.e.L.; Fernandes-Pedrosa, M.e.F. Profiling the resting venom gland of the scorpion Tityus stigmurus through a transcriptomic survey. BMC Genom. 2012, 13, 362. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, U.C.; Candido, D.M.; Dorce, V.A.; Junqueira-de-Azevedo, I.e.L. The transcriptome recipe for the venom cocktail of Tityus bahiensis scorpion. Toxicon 2015, 95, 52–61. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, U.C.; Nishiyama, M.Y.; Dos Santos, M.B.V.; Santos-da-Silva, A.P.; Chalkidis, H.M.; Souza-Imberg, A.; Candido, D.M.; Yamanouye, N.; Dorce, V.A.C.; Junqueira-de-Azevedo, I.L.M. Proteomic endorsed transcriptomic profiles of venom glands from Tityus obscurus and T. serrulatus scorpions. PLoS ONE 2018, 13, e0193739. [Google Scholar] [CrossRef]
- Carmo, A.O.; Oliveira-Mendes, B.B.; Horta, C.C.; Magalhães, B.F.; Dantas, A.E.; Chaves, L.M.; Chávez-Olórtegui, C.; Kalapothakis, E. Molecular and functional characterization of metalloserrulases, new metalloproteases from the Tityus serrulatus venom gland. Toxicon 2014, 90, 45–55. [Google Scholar] [CrossRef]
- Hooper, N.M. Families of zinc metalloproteases. FEBS Lett. 1994, 354, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Fletcher, P.L.; Fletcher, M.D.; Weninger, K.; Anderson, T.E.; Martin, B.M. Vesicle-associated membrane protein (VAMP) cleavage by a new metalloprotease from the Brazilian scorpion Tityus serrulatus. J. Biol. Chem. 2010, 285, 7405–7416. [Google Scholar] [CrossRef] [PubMed]
- Zornetta, I.; Scorzeto, M.; Mendes Dos Reis, P.V.; De Lima, M.E.; Montecucco, C.; Megighian, A.; Rossetto, O. Electrophysiological Characterization of the Antarease Metalloprotease from Tityus serrulatus Venom. Toxins 2017, 9, 81. [Google Scholar] [CrossRef] [PubMed]
- Ortiz, E.; Rendón-Anaya, M.; Rego, S.C.; Schwartz, E.F.; Possani, L.D. Antarease-like Zn-metalloproteases are ubiquitous in the venom of different scorpion genera. Biochim. Biophys. Acta 2014, 1840, 1738–1746. [Google Scholar] [CrossRef]
- Cajado-Carvalho, D.; Kuniyoshi, A.K.; Duzzi, B.; Iwai, L.K.; Oliveira, Ú.; Junqueira de Azevedo, I.L.; Kodama, R.T.; Portaro, F.V. Insights into the Hypertensive Effects of Tityus serrulatus Scorpion Venom: Purification of an Angiotensin-Converting Enzyme-Like Peptidase. Toxins 2016, 8, 348. [Google Scholar] [CrossRef] [PubMed]
- Verano-Braga, T.; Dutra, A.A.; León, I.R.; Melo-Braga, M.N.; Roepstorff, P.; Pimenta, A.M.; Kjeldsen, F. Moving pieces in a venomic puzzle: Unveiling post-translationally modified toxins from Tityus serrulatus. J. Proteome Res. 2013, 12, 3460–3470. [Google Scholar] [CrossRef]
- Venancio, E.J.; Portaro, F.C.; Kuniyoshi, A.K.; Carvalho, D.C.; Pidde-Queiroz, G.; Tambourgi, D.V. Enzymatic properties of venoms from Brazilian scorpions of Tityus genus and the neutralisation potential of therapeutical antivenoms. Toxicon 2013, 69, 180–190. [Google Scholar] [CrossRef] [PubMed]
- Cajado Carvalho, D.; Kuniyoshi, A.K.; Kodama, R.T.; Oliveira, A.K.; Serrano, S.M.; Tambourgi, D.V.; Portaro, F.V. Neuropeptide Y family-degrading metallopeptidases in the Tityus serrulatus venom partially blocked by commercial antivenoms. Toxicol. Sci. 2014, 142, 418–426. [Google Scholar] [CrossRef] [PubMed]
- Stein, C.; Schäfer, M.; Machelska, H. Attacking pain at its source: New perspectives on opioids. Nat. Med. 2003, 9, 1003–1008. [Google Scholar] [CrossRef]
- Tan, C.M.J.; Green, P.; Tapoulal, N.; Lewandowski, A.J.; Leeson, P.; Herring, N. The Role of Neuropeptide Y in Cardiovascular Health and Disease. Front. Physiol. 2018, 9, 1281. [Google Scholar] [CrossRef]
- Spinazzi, R.; Andreis, P.G.; Nussdorfer, G.G. Neuropeptide-Y and Y-receptors in the autocrine-paracrine regulation of adrenal gland under physiological and pathophysiological conditions (Review). Int. J. Mol. Med. 2005, 15, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Nussdorfer, G.G.; Gottardo, G. Neuropeptide-Y family of peptides in the autocrine-paracrine regulation of adrenocortical function. Horm. Metab. Res. 1998, 30, 368–373. [Google Scholar] [CrossRef]
- Portaro, F.C.; Santos, A.B.; Cezari, M.H.; Juliano, M.A.; Juliano, L.; Carmona, E. Probing the specificity of cysteine proteinases at subsites remote from the active site: Analysis of P4, P3, P2′ and P3′ variations in extended substrates. Biochem. J. 2000, 347 Pt 1, 123–129. [Google Scholar] [CrossRef]
- Grundemar, L.; Håkanson, R. Effects of various neuropeptide Y/peptide YY fragments on electrically-evoked contractions of the rat vas deferens. Br. J. Pharmacol. 1990, 100, 190–192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Colaert, N.; Helsens, K.; Martens, L.; Vandekerckhove, J.; Gevaert, K. Improved visualization of protein consensus sequences by iceLogo. Nat. Methods 2009, 6, 786–787. [Google Scholar] [CrossRef]
- Tallant, C.; García-Castellanos, R.; Baumann, U.; Gomis-Rüth, F.X. On the relevance of the Met-turn methionine in metzincins. J. Biol. Chem. 2010, 285, 13951–13957. [Google Scholar] [CrossRef]
- Dudev, T.; Lim, C. Metal selectivity in metalloproteins: Zn2+ vs. Mg2+. J. Phys. Chem. B 2001, 105, 4446–4452. [Google Scholar] [CrossRef]
- Wise, R.J.; Barr, P.J.; Wong, P.A.; Kiefer, M.C.; Brake, A.J.; Kaufman, R.J. Expression of a human proprotein processing enzyme: Correct cleavage of the von Willebrand factor precursor at a paired basic amino acid site. Proc. Natl. Acad. Sci. USA 1990, 87, 9378–9382. [Google Scholar] [CrossRef]
- Langenegger, N.; Koua, D.; Schürch, S.; Heller, M.; Nentwig, W.; Kuhn-Nentwig, L. Identification of a precursor processing protease from the spider. J. Biol. Chem. 2018, 293, 2079–2090. [Google Scholar] [CrossRef]
- Saraf, R.; Mahmood, F.; Amir, R.; Matyal, R. Neuropeptide Y is an angiogenic factor in cardiovascular regeneration. Eur. J. Pharmacol. 2016, 776, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Śliwińska-Mossoń, M.; Marek, G.; Milnerowicz, H. The role of pancreatic polypeptide in pancreatic diseases. Adv. Clin. Exp. Med. 2017, 26, 1447–1455. [Google Scholar] [CrossRef] [PubMed]
- Inyushkin, A.N. Effects of leucine-enkephalin on potassium currents in neurons in the rat respiratory center in vitro. Neurosci. Behav. Physiol. 2007, 37, 739–746. [Google Scholar] [CrossRef] [PubMed]
- Tsunoo, A.; Yoshii, M.; Narahashi, T. Block of calcium channels by enkephalin and somatostatin in neuroblastoma-glioma hybrid NG108-15 cells. Proc. Natl. Acad. Sci. USA 1986, 83, 9832–9836. [Google Scholar] [CrossRef] [PubMed]
- Cabot, P.J.; Carter, L.; Schäfer, M.; Stein, C. Methionine-enkephalin-and Dynorphin A-release from immune cells and control of inflammatory pain. Pain 2001, 93, 207–212. [Google Scholar] [CrossRef]
- Zoccal, K.F.; Sorgi, C.A.; Hori, J.I.; Paula-Silva, F.W.; Arantes, E.C.; Serezani, C.H.; Zamboni, D.S.; Faccioli, L.H. Opposing roles of LTB4 and PGE2 in regulating the inflammasome-dependent scorpion venom-induced mortality. Nat. Commun. 2016, 7, 10760. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trevisan-Silva, D.; Gremski, L.H.; Chaim, O.M.; da Silveira, R.B.; Meissner, G.O.; Mangili, O.C.; Barbaro, K.C.; Gremski, W.; Veiga, S.S.; Senff-Ribeiro, A. Astacin-like metalloproteases are a gene family of toxins present in the venom of different species of the brown spider (genus Loxosceles). Biochimie 2010, 92, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Cajado-Carvalho, D.; Galvão, J.; Kuniyoshi, A.K.; Carneiro, P.D.S.; Paes Leme, A.F.; Pauletti, B.A.; Marengo, E.B.; Portaro, F.V. Tityus serrulatus Scorpion Venom: In Vitro Tests and Their Correlation with In Vivo Lethal Dose Assay. Toxins 2017, 9, 380. [Google Scholar] [CrossRef] [PubMed]
- Hui Wen, F.; Monteiro, W.M.; Moura da Silva, A.M.; Tambourgi, D.V.; Mendonça da Silva, I.; Sampaio, V.S.; dos Santos, M.C.; Sachett, J.; Ferreira, L.C.; Kalil, J.; et al. Snakebites and scorpion stings in the Brazilian Amazon: Identifying research priorities for a largely neglected problem. PLoS Negl. Trop. Dis. 2015, 9, e0003701. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Shevchenko, A.; Tomas, H.; Havlis, J.; Olsen, J.V.; Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 2006, 1, 2856–2860. [Google Scholar] [CrossRef]
- Ma, B.; Zhang, K.; Hendrie, C.; Liang, C.; Li, M.; Doherty-Kirby, A.; Lajoie, G. PEAKS: Powerful software for peptide de novo sequencing by tandem mass spectrometry. Rapid Commun. Mass Spectrom. 2003, 17, 2337–2342. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xin, L.; Shan, B.; Chen, W.; Xie, M.; Yuen, D.; Zhang, W.; Zhang, Z.; Lajoie, G.A.; Ma, B. PEAKS DB: De novo sequencing assisted database search for sensitive and accurate peptide identification. Mol. Cell Proteom. 2012, 11, M111.010587. [Google Scholar] [CrossRef] [PubMed]
- Stoll, V.S.; Blanchard, J.S. Buffers: Principles and practice. Methods Enzymol. 1990, 182, 24–38. [Google Scholar]
- Schechter, I.; Berger, A. On the size of the active site in proteases. I. Papain. Biochem. Biophys. Res. Commun. 1967, 27, 157–162. [Google Scholar] [CrossRef]





| Step | Fraction | Volume (µL) | Total Protein (µg) | Activity (units) | Total Activity (units/µL) | Specific Activity (units/µg) | Purification Factor | Yield (%) | EDTA Inhibition (%) |
|---|---|---|---|---|---|---|---|---|---|
| Venom | TsV | 1000 | 30,000 | 90 | 90,000 | 3000 | 1 | 100 | 100 |
| DEAE | F3 | 1000 | 0.12 | 13.1 | 13,100 | 109,167 | 36.39 | 14.6 | 100 |
| F5 | 1000 | 0.139 | 20.9 | 20,873 | 150,165 | 50.06 | 23.2 | 100 | |
| GF | F3-4 | 1000 | 0.024 | 8.9 | 8900 | 370,833 | 123.61 | 9.9 | 100 |
| F5-1 | 1000 | 0.071 | 12.7 | 12,729 | 179,276 | 59.76 | 14.1 | 100 |
| Peptide | Specific Activity (µM/µg/min) | Scissile Bonds | |
|---|---|---|---|
| TsMS 3 | TsMS 4 | ||
| NPY | 0.433 | 0.631 | YPSKP↑D↓NPGED↓↑A↑PAEDM↓A↓↑R↓↑Y↓↑Y↓↑S↓A↓L↓↑R↓↑H↓↑Y↑I↓N↓↑LIT↓R↓↑Q↓↑RY-NH2 |
| PYY | 0.359 | 0.420 | YPIKPEAPGED↓ASPEEL↓N↓R↓YYASL↓↑R↓HY↓L↓NLVT↑R↓Q↓RY-NH2 |
| PP | 0.347 | 0.430 | APLEPMYP↓G↑D↑Y↑A↑TH↑E↑Q↑RAQ↑YETQL↓↑R↑R↓↑Y↑YPIKPE↓↑PRY-NH2 |
| DYN | 1.539 | 1.303 | YGGFL↑R↓↑RIRPKLK |
| Substrate | Catalytic Constants | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| P4 | P3 | P2 | P1 | P1′ | P2′ | Km (µM) | kcat (s−1) | kcat/Km (µM−1·s−1) | ||
| Abz | G | F | L | R | R | V | EDDnp | 16.2 ± 4.3 | 491.7 ± 52.4 | 30.4 ± 8.1 |
| Abz | G | F | L | R | R | - | EDDnp | 36.6 ± 5.8 | 170.8 ± 8.4 | 4.7 ± 0.7 |
| Abz | - | F | L | R | R | V | EDDnp | 28.1 ± 2.6 | 1108.3 ± 33.2 | 39.4 ± 3.64 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cajado-Carvalho, D.; da Silva, C.C.F.; Kodama, R.T.; Mariano, D.O.C.; Pimenta, D.C.; Duzzi, B.; Kuniyoshi, A.K.; Portaro, F.V. Purification and Biochemical Characterization of TsMS 3 and TsMS 4: Neuropeptide-Degrading Metallopeptidases in the Tityus serrulatus Venom. Toxins 2019, 11, 194. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11040194
Cajado-Carvalho D, da Silva CCF, Kodama RT, Mariano DOC, Pimenta DC, Duzzi B, Kuniyoshi AK, Portaro FV. Purification and Biochemical Characterization of TsMS 3 and TsMS 4: Neuropeptide-Degrading Metallopeptidases in the Tityus serrulatus Venom. Toxins. 2019; 11(4):194. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11040194
Chicago/Turabian StyleCajado-Carvalho, Daniela, Cristiane Castilho Fernandes da Silva, Roberto Tadashi Kodama, Douglas Oscar Ceolin Mariano, Daniel Carvalho Pimenta, Bruno Duzzi, Alexandre Kazuo Kuniyoshi, and Fernanda Vieira Portaro. 2019. "Purification and Biochemical Characterization of TsMS 3 and TsMS 4: Neuropeptide-Degrading Metallopeptidases in the Tityus serrulatus Venom" Toxins 11, no. 4: 194. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins11040194