The Buzz about ADP-Ribosylation Toxins from Paenibacillus larvae, the Causative Agent of American Foulbrood in Honey Bees
Abstract
:1. Introduction
2. ADP-Ribosylating Toxins of P. larvae
2.1. Plx1, a Phage Born Toxin of P. larvae
2.2. C3-Like Toxins with B-Subunits
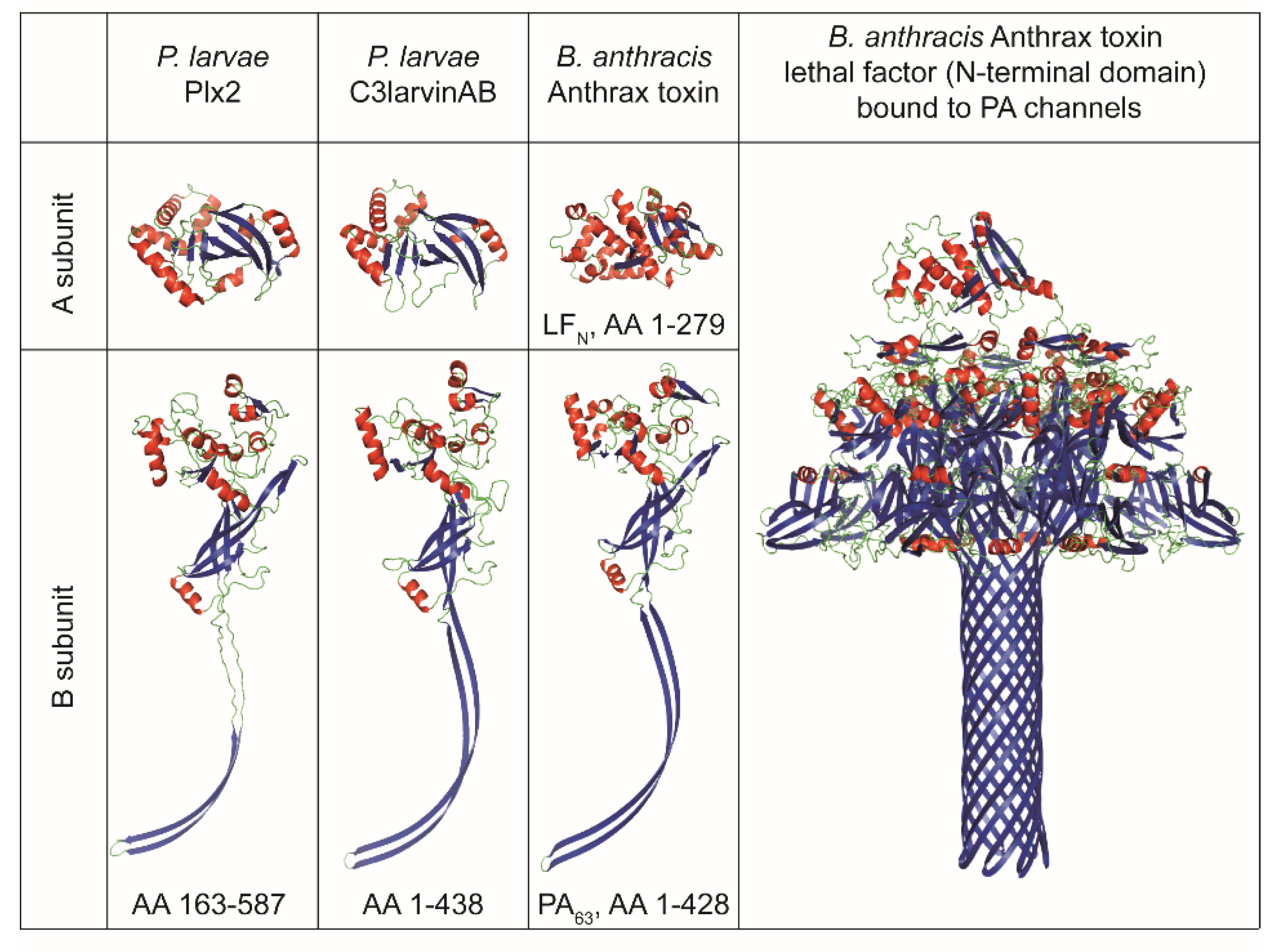
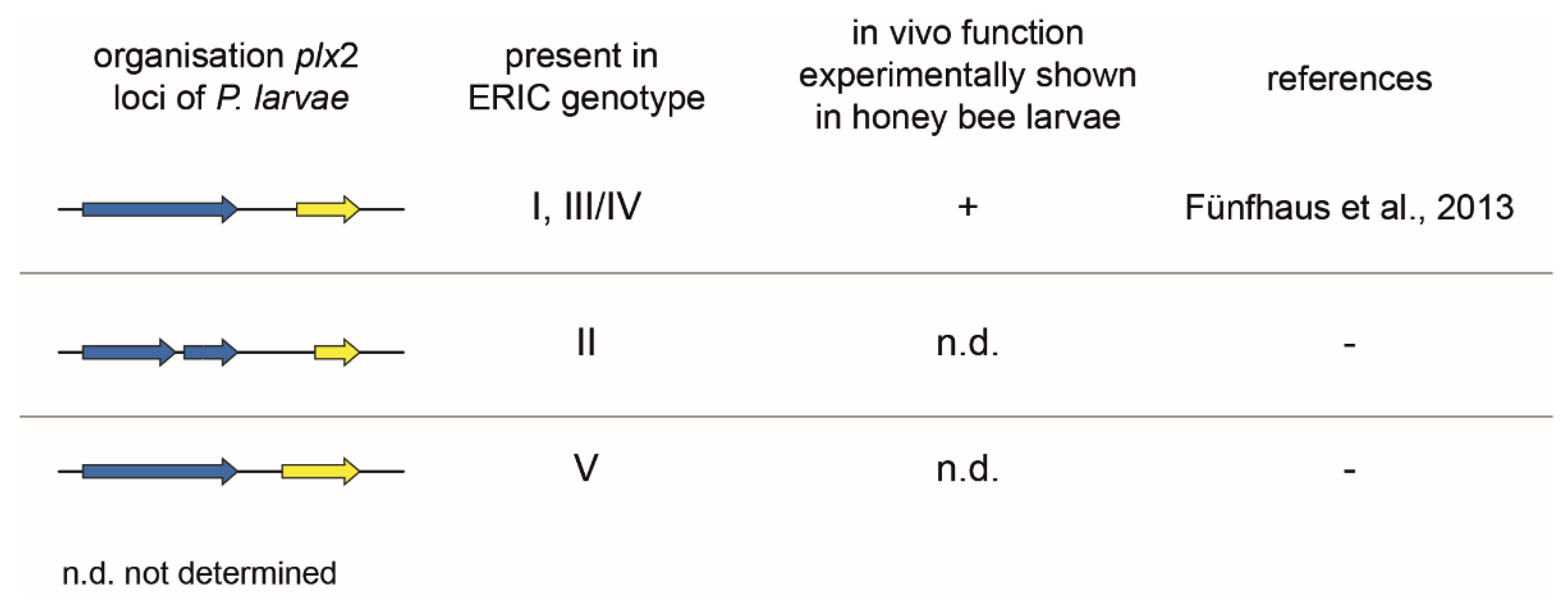
2.2.1. Binary C3-Like Toxin Plx2
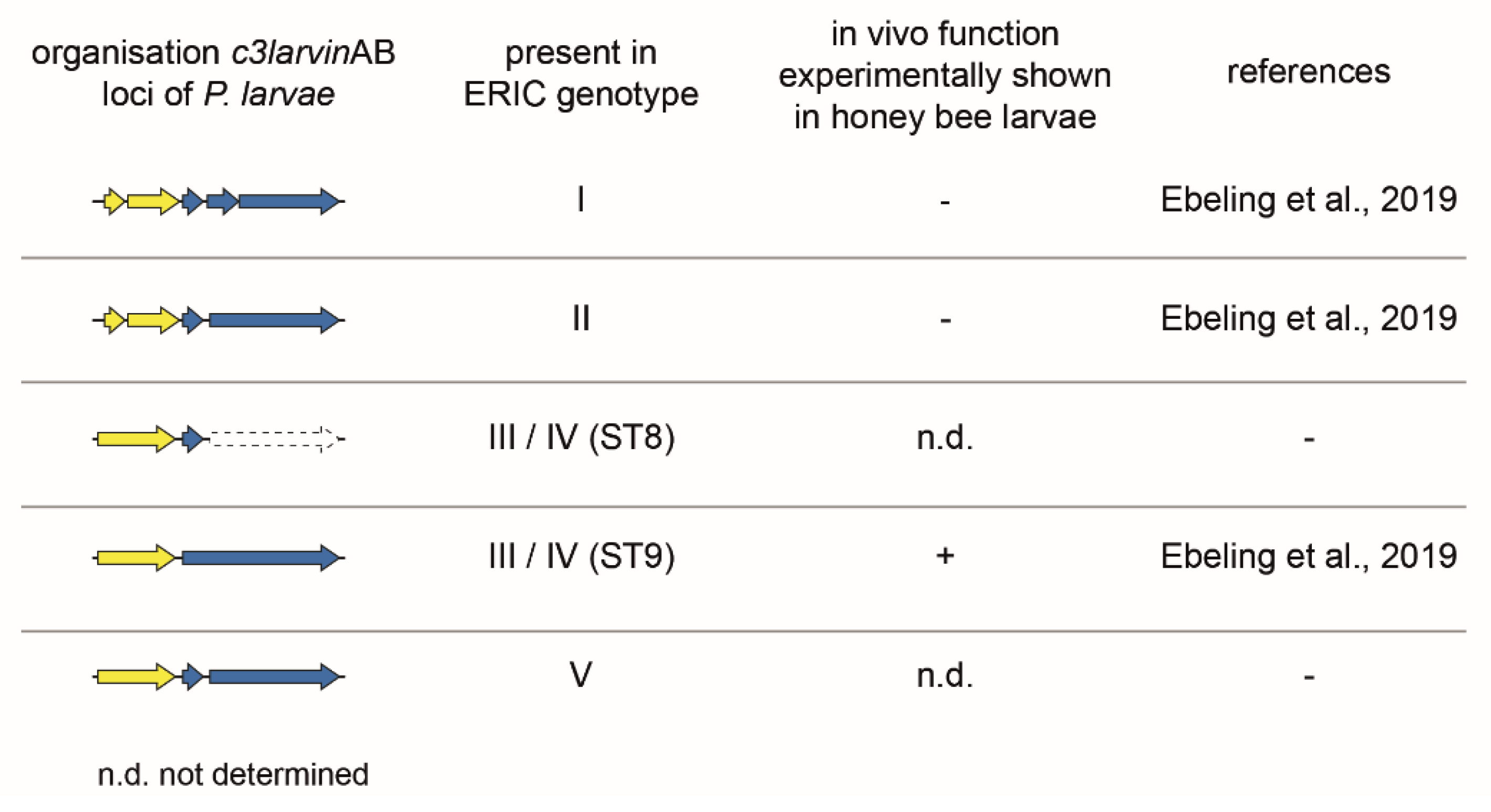
2.2.2. Binary C3-Like Toxin C3larvinAB
3. Outlook
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Collier, R.J.; Pappenheimer, A.M., Jr. Studies on the mode of action of diphtheria toxin: II. Effect of toxin on amino acid incorporation in cell-free systems. J. Exp. Med. 1964, 120, 1019–1039. [Google Scholar] [CrossRef]
- Honjo, T.; Nishizuka, Y.; Kato, I.; Hayaishi, O. Adenosine diphosphate ribosylation of aminoacyl transferase II and inhibition of protein synthesis by diphtheria toxin. J. Biol. Chem. 1971, 246, 4251–4260. [Google Scholar] [CrossRef]
- Genersch, E.; Forsgren, E.; Pentikäinen, J.; Ashiralieva, A.; Rauch, S.; Kilwinski, J.; Fries, I. Reclassification of Paenibacillus larvae subsp. pulvifaciens and Paenibacillus larvae subsp. larvae as Paenibacillus larvae without subspecies differentiation. Int. J. Syst. Evol. Microbiol. 2006, 56, 501–511. [Google Scholar] [CrossRef] [Green Version]
- Morrissey, B.J.; Helgason, T.; Poppinga, L.; Fünfhaus, A.; Genersch, E.; Budge, G.E. Biogeography of Paenibacillus larvae, the causative agent of American foulbrood, using a new multilocus sequence typing scheme. Environ. Microbiol. 2014, 17, 1414–1424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Genersch, E.; Ashiralieva, A.; Fries, I. Strain- and genotype-specific differences in virulence of Paenibacillus larvae subsp. larvae, a bacterial pathogen causing American foulbrood disease in honeybees. Appl. Environ. Microbiol. 2005, 71, 7551–7555. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rauch, S.; Ashiralieva, A.; Hedtke, K.; Genersch, E. Negative correlation between individual-insect-level virulence and colony-level virulence of Paenibacillus larvae, the etiological agent of American foulbrood of honeybees. Appl. Environ. Microbiol. 2009, 75, 3344–3347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Djukic, M.; Brzuszkiewicz, E.; Fünfhaus, A.; Voss, J.; Gollnow, K.; Poppinga, L.; Liesegang, H.; Garcia-Gonzalez, E.; Genersch, E.; Daniel, R. How to kill the honey bee larva: Genomic potential and virulence mechanisms of Paenibacillus larvae. PLoS ONE 2014, 9, e90914. [Google Scholar] [CrossRef]
- Beims, H.; Bunk, B.; Erler, S.; Mohr, K.I.; Spröer, C.; Pradella, S.; Günther, G.; Rohde, M.; Von Der Ohe, W.; Steinert, M. Discovery of Paenibacillus larvae ERIC V: Phenotypic and genomic comparison to genotypes ERIC I-IV reveal different inventories of virulence factors which correlate with epidemiological prevalences of American Foulbrood. Int. J. Med. Microbiol. 2020, 310, 151394. [Google Scholar] [CrossRef]
- Ebeling, J.; Knispel, H.; Fünfhaus, A.; Genersch, E. The biological role of the enigmatic C3larvinAB toxin of the honey bee pathogenic bacterium Paenibacillus larvae. Environ. Microbiol. 2019, 21, 3091–3106. [Google Scholar] [CrossRef] [PubMed]
- Yue, D.; Nordhoff, M.; Wieler, L.H.; Genersch, E. Fluorescence in situ-hybridization (FISH) analysis of the interactions between honey bee larvae and Paenibacillus larvae, the causative agent of American Foulbrood of honeybees (Apis mellifera). Environ. Microbiol. 2008, 10, 1612–1620. [Google Scholar] [CrossRef]
- Garcia-Gonzalez, E.; Genersch, E. Honey bee larval peritrophic matrix degradation during infection with Paenibacillus larvae, the aetiological agent of American foulbrood of honey bees, is a key step in pathogenesis. Environ. Microbiol. 2013, 15, 2894–2901. [Google Scholar] [CrossRef]
- Garcia-Gonzalez, E.; Poppinga, L.; Fünfhaus, A.; Hertlein, G.; Hedtke, K.; Jakubowska, A.; Genersch, E. Paenibacillus larvae chitin-degrading protein PlCBP49 is a key virulence factor in American Foulbrood of honey bees. PLoS Path. 2014, 10, e1004284. [Google Scholar] [CrossRef]
- Fünfhaus, A.; Ashiralieva, A.; Borriss, R.; Genersch, E. Use of suppression subtractive hybridization to identify genetic differences between differentially virulent genotypes of Paenibacillus larvae, the etiological agent of American Foulbrood of honey bees. Environ. Microbiol. Rep. 2009, 1, 240–250. [Google Scholar] [CrossRef] [PubMed]
- Fünfhaus, A.; Genersch, E. Proteome analysis of Paenibacillus larvae reveals the existence of a putative S-layer protein. Environ. Microbiol. Rep. 2012, 4, 194–202. [Google Scholar] [CrossRef]
- Poppinga, L.; Janesch, B.; Fünfhaus, A.; Sekot, G.; Garcia-Gonzalez, E.; Hertlein, G.; Hedtke, K.; Schäffer, C.; Genersch, E. Identification and functional analysis of the S-layer protein SplA of Paenibacillus larvae, the causative agent of American Foulbrood of honey bees. PLoS Path. 2012, 8, e1002716. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fünfhaus, A.; Poppinga, L.; Genersch, E. Identification and characterization of two novel toxins expressed by the lethal honey bee pathogen Paenibacillus larvae, the causative agent of American Foulbrood. Environ. Microbiol. 2013, 15, 2951–2965. [Google Scholar] [CrossRef]
- Müller, S.; Garcia-Gonzalez, E.; Genersch, E.; Süssmuth, R.D. Involvement of secondary metabolites in the pathogenesis of the American Foulbrood of honey bees caused by Paenibacillus larvae. Nat. Prod. Rep. 2015, 32, 765–778. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Müller, S.; Garcia-Gonzalez, E.; Mainz, A.; Hertlein, G.; Heid, N.C.; Mösker, E.; van den Elst, H.; Overkleeft, H.S.; Genersch, E.; Süssmuth, R.D. Paenilamicin-structure and biosynthesis of a hybrid non-ribosomal peptide/polyketide antibiotic from the bee pathogen Paenibacillus larvae. Angew. Chem. Int. Ed. Engl. 2014, 53, 10547–10828. [Google Scholar] [CrossRef]
- Hertlein, G.; Müller, S.; Garcia-Gonzalez, E.; Poppinga, L.; Süssmuth, R.D.; Genersch, E. Production of the catechol type siderophore bacillibactin by the honey bee pathogen Paenibacillus larvae. PLoS ONE 2014, 9, e108272. [Google Scholar] [CrossRef] [Green Version]
- Garcia-Gonzalez, E.; Müller, S.; Hertlein, G.; Heid, N.; Süssmuth, R.D.; Genersch, E. Biological effects of paenilamicin, a secondary metabolite antibiotic produced by the honey bee pathogenic bacterium Paenibacillus larvae. Microbiology 2014, 3, 642–656. [Google Scholar] [CrossRef] [Green Version]
- Krska, D.; Ravulapalli, R.; Fieldhouse, R.J.; Lugo, M.R.; Merrill, A.R. C3larvin toxin, an ADP-ribosyltransferase from Paenibacillus larvae. J. Biol. Chem. 2015, 290, 1639–1653. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turner, M.; Tremblay, O.; Heney, K.A.; Lugo, M.R.; Ebeling, J.; Genersch, E.; Merrill, A.R. Characterization of C3larvinA, a novel RhoA-targeting ADP-ribosyltransferase toxin produced by the honey bee pathogen, Paenibacillus larvae. Biosci. Rep. 2020, 40, BSR20193405. [Google Scholar] [CrossRef]
- Thanabalu, T.; Hindley, J.; Jackson-Yap, J.; Berry, C. Cloning, sequencing, and expression of a gene encoding a 100-kilodalton mosquitocidal toxin from Bacillus sphaericus SSII-1. J. Bacteriol. 1991, 173, 2776–2785. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Masignani, V.; Pizza, M.; Rappuoli, R. Common features of ADP—ribosyltransferases. In Handbook of Experimental Pharmacology; Springer International Publishing: Berlin/Heidelberg, Germany, 2000; Volume 145, pp. 21–44. [Google Scholar]
- Carpusca, I.; Jank, T.; Aktories, K. Bacillus sphaericus mosquitocidal toxin (MTX) and pierisin: The enigmatic offspring from the family of ADP-ribosyltransferases. Mol. Microbiol. 2006, 62, 621–630. [Google Scholar] [CrossRef]
- Matsumoto, Y.; Nakano, T.; Yamamoto, M.; Matsushima-Hibiya, Y.; Odagiri, K.-I.; Yata, O.; Koyama, K.; Sugimura, T.; Wakabayashi, K. Distribution of cytotoxic and DNA ADP-ribosylating activity in crude extracts from butterflies among the family Pieridae. Proc. Natl. Acad. Sci. USA 2008, 105, 2516–2520. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [Green Version]
- Orth, J.H.; Schorch, B.; Boundy, S.; Ffrench-Constant, R.; Kubick, S.; Aktories, K. Cell-free synthesis and characterization of a novel cytotoxic pierisin-like protein from the cabbage butterfly Pieris rapae. Toxicon 2011, 57, 199–207. [Google Scholar] [CrossRef]
- Takamura-Enya, T.; Watanabe, M.; Koyama, K.; Sugimura, T.; Wakabayashi, K. Mono(ADPribosyl)ation of the N2 amino groups of guanine residues in DNA by pierisin-2, from the cabbage butterfly, Pieris brassicae. Biochem. Biophys. Res. Commun. 2004, 323, 579–582. [Google Scholar] [CrossRef]
- Takamura-Enya, T.; Watanabe, M.; Totsuka, Y.; Kanazawa, T.; Matsushima-Hibiya, Y.; Koyama, K.; Sugimura, T.; Wakabayashi, K. Mono(ADP-ribosyl)ation of 2′-deoxyguanosine residue in DNA by an apoptosis-inducing protein, pierisin-1, from cabbage butterfly. Proc. Natl. Acad. Sci. USA 2001, 98, 12414–12419. [Google Scholar] [CrossRef] [Green Version]
- Watanabe, M.; Enomoto, S.; Takamura-Enya, T.; Nakano, T.; Koyama, K.; Sugimura, T.; Wakabayashi, K. Enzymatic properties of pierisin-1 and its N-terminal domain, a guanine-specific ADP-ribosyltransferase from the cabbage butterfly. J. Biochem. 2004, 135, 471–477. [Google Scholar] [CrossRef]
- Lyons, B.; Ravulapalli, R.; LaNoue, J.; Lugo, M.R.; Dutta, D.; Carlin, S.; Merrill, A.R. Scabin, a novel DNA-acting ADP-ribosyltransferase from Streptomyces scabies. J. Biol. Chem. 2016, 291, 11198–11215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef] [Green Version]
- Rutenber, E.; Ready, M.; Robertus, J.D. Structure and evolution of ricin B chain. Nat. Cell Biol. 1987, 326, 624–626. [Google Scholar] [CrossRef]
- Polito, L.; Bortolotti, M.; Battelli, M.G.; Calafato, G.; Bolognesi, A. Ricin: An ancient story for a timeless plant toxin. Toxins 2019, 11, 324. [Google Scholar] [CrossRef] [Green Version]
- Hazes, B.; Read, R.J. A mosquitocidal toxin with a ricin-like cell-binding domain. Nat. Struct. Mol. Biol. 1995, 2, 358–359. [Google Scholar] [CrossRef] [PubMed]
- Hazes, B. The (QxW)3domain: A flexible lectin scaffold. Protein Sci. 1996, 5, 1490–1501. [Google Scholar] [CrossRef] [Green Version]
- Carpusca, I.; Schirmer, J.; Aktories, K. Two-site autoinhibition of the ADP-ribosylating mosquitocidal toxin (MTX) from Bacillus sphaericus by its 70-kDa ricin-like binding domain. Biochemistry 2004, 43, 12009–12019. [Google Scholar] [CrossRef] [PubMed]
- Matsushima-Hibiya, Y.; Watanabe, M.; Hidari, K.I.-P.J.; Miyamoto, D.; Suzuki, Y.; Kasama, T.; Kanazawa, T.; Koyama, K.; Sugimura, T.; Wakabayashi, K. Identification of glycosphingolipid receptors for pierisin-1, a guanine-specific ADP-ribosylating toxin from the cabbage butterfly. J. Biol. Chem. 2003, 278, 9972–9978. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stamereilers, C.; Fajardo, C.P.; Walker, J.K.; Mendez, K.N.; Castro-Nallar, E.; Grose, J.H.; Hope, S.; Tsourkas, P.K. Genomic analysis of 48 Paenibacillus larvae bacteriophages. Viruses 2018, 10, 377. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Freeman, V.J. Studies on the virulence of bacteriophage-infected strains of Corynebacterium diphtheriae. J. Bacteriol. 1951, 61, 675–688. [Google Scholar] [CrossRef] [Green Version]
- Groman, N.B. Evidence for the active role of bacteriophage in the conversion of nontoxigenic Corynebacterium diphtheriae to toxin production. J. Bacteriol. 1955, 69, 9–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Groman, N.B. The relation of bacteriophage to the change of Corynebacterium diphtheriae from avirulence to virulence. Science 1953, 117, 297–299. [Google Scholar] [CrossRef] [PubMed]
- Holmes, R.K.; Barksdale, L. Comparative studies with tox+ and tox− Corynebacteriophages. J. Virol. 1970, 5, 783–794. [Google Scholar] [CrossRef] [Green Version]
- Miller, P.A.; Pappenheimer, A.M., Jr.; Doolittle, W.F. Phage-host relationships in certain strains of Corynebacterium diphtheriae. Virology 1966, 29, 410–425. [Google Scholar] [CrossRef]
- Fourel, G.; Phalipon, A.; Kaczorek, M. Evidence for direct regulation of diphtheria toxin gene transcription by an Fe2+-dependent DNA-binding repressor, DtoxR, in Corynebacterium diphtheriae. Infect. Immun. 1989, 57, 3221–3225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laird, W.; Groman, N. Orientation of the tox gene in the prophage of corynebacteriophage beta. J. Virol. 1976, 19, 228–231. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eklund, M.W.; Poysky, F.T.; Reed, S.M.; Smith, C.A. Bacteriophage and the toxigenicity of Clostridium botulinum type C. Science 1971, 172, 480–482. [Google Scholar] [CrossRef]
- Waldor, M.K.; Mekalanos, J.J. Lysogenic conversion by a filamentous phage encoding cholera toxin. Science 1996, 272, 1910–1914. [Google Scholar] [CrossRef] [Green Version]
- Scotland, S.M.; Smith, H.R.; Willshaw, A.G.; Rowe, B. Vero cytotoxin production in strain of Escherichia coli is determined by genes carried on bacteriophage. Lancet 1983, 2, 216. [Google Scholar] [CrossRef]
- Yamamoto, M.; Takahashi-Nakaguchi, A.; Wakabayashi, K.; Matsushima-Hibiya, Y.; Nakano, T.; Totsuka, Y.; Imanishi, S.; Mitsuhashi, J.; Watanabe, M.; Nakagama, H.; et al. Nucleotide sequence and chromosomal localization of the gene for pierisin-1, a DNA ADP-ribosylating protein, in the cabbage butterfly Pieris rapae. Genetics 2011, 139, 1251–1258. [Google Scholar] [CrossRef]
- Nakano, T.; Takahashi-Nakaguchi, A.; Yamamoto, M.; Watanabe, M. Pierisins and CARP-1: ADP-Ribosylation of DNA by ARTCs in butterflies and shellfish. Curr. Top. Microbiol. Immunol. 2014, 384, 127–149. [Google Scholar] [CrossRef]
- Tsourkas, P.K. Paenibacillus larvae bacteriophages: Obscure past, promising future. Microb. Genom. 2020, 6, e000329. [Google Scholar] [CrossRef] [PubMed]
- Ebeling, J.; Fünfhaus, A.; Knispel, H.; Krska, D.; Ravulapalli, R.; Heney, K.A.; Lugo, M.R.; Merrill, A.R.; Genersch, E. Characterization of the toxin Plx2A, a RhoA-targeting ADP-ribosyltransferase produced by the honey bee pathogen Paenibacillus larvae. Environ. Microbiol. 2017, 19, 5100–5116. [Google Scholar] [CrossRef]
- Holbourn, K.P.; Shone, C.C.; Acharya, K.R. A family of killer toxins: Exploring the mechanism of ADP-ribosylating toxins. FEBS J. 2006, 273, 4579–4593. [Google Scholar] [CrossRef]
- Kimura, K.; Kubota, T.; Ohishi, I.; Isogai, E.; Isogai, H.; Fujii, N. The gene for component-II of botulinum C2 toxin. Veter. Microbiol. 1998, 62, 27–34. [Google Scholar] [CrossRef]
- Stubbs, S.; Rupnik, M.; Gibert, M.; Brazier, J.; Duerden, B.; Popoff, M. Production of actin-specific ADP-ribosyltransferase (binary toxin) by strains of Clostridium difficile. FEMS Microbiol. Lett. 2000, 186, 307–312. [Google Scholar] [CrossRef]
- Perelle, S.; Gibert, M.; Boquet, P.; Popoff, M.R. Characterization of Clostridium perfringens iota-toxin genes and expression in Escherichia coli. Infect. Immun. 1993, 61, 5147–5156. [Google Scholar] [CrossRef] [Green Version]
- Welkos, S.; Lowe, J.; Eden-McCutchan, F.; Vodkin, M.; Leppla, S.; Schmidt, J. Sequence and analysis of the DNA encoding protective antigen of Bacillus anthracis. Gene 1988, 69, 287–300. [Google Scholar] [CrossRef]
- Pezard, C.; Berche, P.; Mock, M. Contribution of individual toxin components to virulence of Bacillus anthracis. Infect. Immun. 1991, 59, 3472–3477. [Google Scholar] [CrossRef] [Green Version]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Milne, J.; Furlong, D.; Hanna, P.; Wall, J.; Collier, R. Anthrax protective antigen forms oligomers during intoxication of mammalian cells. J. Biol. Chem. 1994, 269, 20607–20612. [Google Scholar] [CrossRef]
- Kintzer, A.F.; Thoren, K.L.; Sterling, H.J.; Dong, K.C.; Feld, G.K.; Tang, I.I.; Zhang, T.T.; Williams, E.R.; Berger, J.M.; Krantz, B.A. The protective antigen component of anthrax toxin forms functional octameric complexes. J. Mol. Biol. 2009, 392, 614–629. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Pentelute, B.L.; Collier, R.J.; Zhou, Z.H. Atomic structure of anthrax protective antigen pore elucidates toxin translocation. Nat. Cell Biol. 2015, 521, 545–549. [Google Scholar] [CrossRef]
- Xu, X.; Godoy-Ruiz, R.; Grishaev, A.; Neu, H.M.; Michel, S.L.J.; Yu, W.; Beckett, D.; Rustandi, R.R.; Lancaster, C.; Loughney, J.W.; et al. Structure of the cell-binding component of the Clostridium difficile binary toxin reveals a di-heptamer macromolecular assembly. Proc. Natl. Acad. Sci. USA 2020, 117, 1049–1058. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamada, T.; Yoshida, T.; Kawamoto, A.; Mitsuoka, K.; Iwasaki, K.; Tsuge, H. Cryo-EM structures reveal translocational unfolding in the clostridial binary iota toxin complex. Nat. Struct. Mol. Biol. 2020, 27, 288–296. [Google Scholar] [CrossRef]
- Duesbery, N.S.; Webb, C.P.; Leppla, S.H.; Gordon, V.M.; Klimpel, K.R.; Copeland, T.D.; Ahn, N.G.; Oskarsson, M.K.; Fukasawa, K.; Paull, K.D.; et al. Proteolytic inactivation of MAP-kinase-kinase by anthrax lethal factor. Science 1998, 280, 734–737. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.-J.; Guo, Q. The adenylyl cyclase activity of anthrax edema factor. Mol. Asp. Med. 2009, 30, 423–430. [Google Scholar] [CrossRef] [Green Version]
- Bragg, T.S.; Robertson, D.L. Nucleotide sequence and analysis of the lethal factor gene (lef) from Bacillus anthracis. Gene 1989, 81, 45–54. [Google Scholar] [CrossRef]
- Feld, G.K.; Thoren, K.L.; Kintzer, A.F.; Sterling, H.J.; Tang, I.I.; Greenberg, S.G.; Williams, E.R.; Krantz, B.A. Structural basis for the unfolding of anthrax lethal factor by protective antigen oligomers. Nat. Struct. Mol. Biol. 2010, 17, 1383–1390. [Google Scholar] [CrossRef]
- Hardenbrook, N.J.; Liu, S.; Zhou, K.; Ghosal, K.; Zhou, Z.H.; Krantz, B.A. Atomic structures of anthrax toxin protective antigen channels bound to partially unfolded lethal and edema factors. Nat. Commun. 2020, 11, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Fahrer, J.; Kuban, J.; Heine, K.; Rupps, G.; Kaiser, E.; Felder, E.; Benz, R.; Barth, H. Selective and specific internalization of clostridial C3 ADP-ribosyltransferases into macrophages and monocytes. Cell. Microbiol. 2010, 12, 233–247. [Google Scholar] [CrossRef] [PubMed]
- Lugo, M.R.; Merrill, A.R. An in-silico sequence-structure-function analysis of the N-terminal lobe in CT group bacterial ADP-ribosyltransferase toxins. Toxins 2019, 11, 365. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turner, M.; Heney, K.A.; Merrill, A.R. The N-terminus of Paenibacillus larvae C3larvinA modulates catalytic efficiency. Biosci. Rep. 2021, 41. [Google Scholar] [CrossRef]
- Barth, H.; Hofmann, F.; Olenik, C.; Just, I.; Aktories, K. The N-terminal part of the enzyme component (C2I) of the binary Clostridium botulinum C2 toxin interacts with the binding component C2II and functions as a carrier system for a Rho ADP-ribosylating C3-like fusion toxin. Infect. Immun. 1998, 66, 1364–1369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turk, E.B.; Wong, T.Y.; Schwarzenbacher, R.; Jarrell, E.T.; Leppla, S.H.; Collier, R.J.; Liddington, R.C.; Cantley, L.C. The structural basis for substrate and inhibitor selectivity of the anthrax lethal factor. Nat. Struct. Mol. Biol. 2003, 11, 60–66. [Google Scholar] [CrossRef]
- Nielsen, M.; Lundegaard, C.; Lund, O.; Petersen, T.N. CPHmodels-3.0—Remote homology modeling using structure-guided sequence profiles. Nucleic Acids Res. 2010, 38, W576–W581. [Google Scholar] [CrossRef] [Green Version]
- Metcalf, D.S.; Weese, J.S. Binary toxin locus analysis in Clostridium difficile. J. Med. Microbiol. 2011, 60, 1137–1145. [Google Scholar] [CrossRef] [Green Version]
- Rigden, D.J.; Mello, L.V.; Galperin, M.Y. The PA14 domain, a conserved all-β domain in bacterial toxins, enzymes, adhesins and signaling molecules. Trends Biochem. Sci. 2004, 29, 336–339. [Google Scholar] [CrossRef]
- Madaule, P.; Axel, R. A novel ras-related gene family. Cell 1985, 41, 31–40. [Google Scholar] [CrossRef]
- Jantsch-Plunger, V.; Gönczy, P.; Romano, A.; Schnabel, H.; Hamill, D.; Schnabel, R.; Hyman, A.A.; Glotzer, M. CYK-4: A Rho family GTPase activating protein (GAP) required for central spindle formation and cytokinesis. J. Cell Biol. 2000, 149, 1391–1404. [Google Scholar] [CrossRef] [Green Version]
- Kamijo, K.; Ohara, N.; Abe, M.; Uchimura, T.; Hosoya, H.; Lee, J.S.; Miki, T. Dissecting the role of Rho-mediated signaling in contractile ring formation. Mol. Biol. Cell 2006, 17, 43–55. [Google Scholar] [CrossRef] [Green Version]
- Maddox, A.S.; Oegema, K. Closing the GAP: A role for a RhoA GAP in cytokinesis. Mol. Cell 2003, 11, 846–848. [Google Scholar] [CrossRef]
- Chardin, P.; Boquet, P.; Madaule, P.; Popoff, M.R.; Rubin, E.J.; Gill, D.M. The mammalian G protein rhoC is ADP-ribosylated by Clostridium botulinum exoenzyme C3 and affects actin microfilaments in Vero cells. EMBO J. 1989, 8, 1087–1092. [Google Scholar] [CrossRef] [PubMed]
- Ridley, A.J.; Hall, A. The small GTP-binding protein Rho regulates the assembly of focal adhesions and actin stress fibers in response to growth factors. Cell 1992, 70, 389–399. [Google Scholar] [CrossRef]
- Mabuchi, I.; Hamaguchi, Y.; Fujimoto, H.; Morii, N.; Mishima, M.; Narumiya, S. A Rho-like protein is involved in the organisation of the contractile ring in dividing sand dollar eggs. Zygote 1993, 1, 325–331. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.-D.; Diao, M.-Q.; Niu, G.-J.; Wang, X.-W.; Zhao, X.-F.; Wang, J.-X. A small GTPase, RhoA, inhibits bacterial infection through integrin mediated phagocytosis in invertebrates. Front. Immunol. 2018, 9, 1928. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barth, H.; Preiss, J.C.; Hofmann, F.; Aktories, K. Characterization of the catalytic site of the ADP-ribosyltransferase Clostridium botulinum C2 toxin by site-directed mutagenesis. J. Biol. Chem. 1998, 273, 29506–29511. [Google Scholar] [CrossRef] [Green Version]
- Wells, J.N.; Bergendahl, L.T.; Marsh, J.A. Operon gene order is optimized for ordered protein complex assembly. Cell Rep. 2016, 14, 679–685. [Google Scholar] [CrossRef] [Green Version]



| Plx2B | C3larvinB | PA | Ib | C2II | CdtB | |
|---|---|---|---|---|---|---|
| Plx2B | - | 40.36% | 31.24% | 37.87% | 34.89% | 38.54% |
| C3larvinB | 40.36% | - | 39.45% | 36.55% | 35.11% | 39.69% |
| PA | 31.24% | 39.45% | - | 35.86% | 35.51% | 34.46% |
| Ib | 37.87% | 36.55% | 35.86% | - | 43.43% | 79.93% |
| C2II | 34.89% | 35.11% | 35.51% | 43.43% | - | 44.76% |
| CdtB | 38.54% | 39.69% | 34.46% | 79.93% | 44.76% | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ebeling, J.; Fünfhaus, A.; Genersch, E. The Buzz about ADP-Ribosylation Toxins from Paenibacillus larvae, the Causative Agent of American Foulbrood in Honey Bees. Toxins 2021, 13, 151. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins13020151
Ebeling J, Fünfhaus A, Genersch E. The Buzz about ADP-Ribosylation Toxins from Paenibacillus larvae, the Causative Agent of American Foulbrood in Honey Bees. Toxins. 2021; 13(2):151. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins13020151
Chicago/Turabian StyleEbeling, Julia, Anne Fünfhaus, and Elke Genersch. 2021. "The Buzz about ADP-Ribosylation Toxins from Paenibacillus larvae, the Causative Agent of American Foulbrood in Honey Bees" Toxins 13, no. 2: 151. https://0-doi-org.brum.beds.ac.uk/10.3390/toxins13020151





