Osmolyte-Like Stabilizing Effects of Low GdnHCl Concentrations on d-Glucose/d-Galactose-Binding Protein
Abstract
:1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. Materials
3.2. Methods
3.2.1. Steady-State Fluorescence Spectroscopy
3.2.2. Time-Resolved Fluorescence Anisotropy Spectroscopy
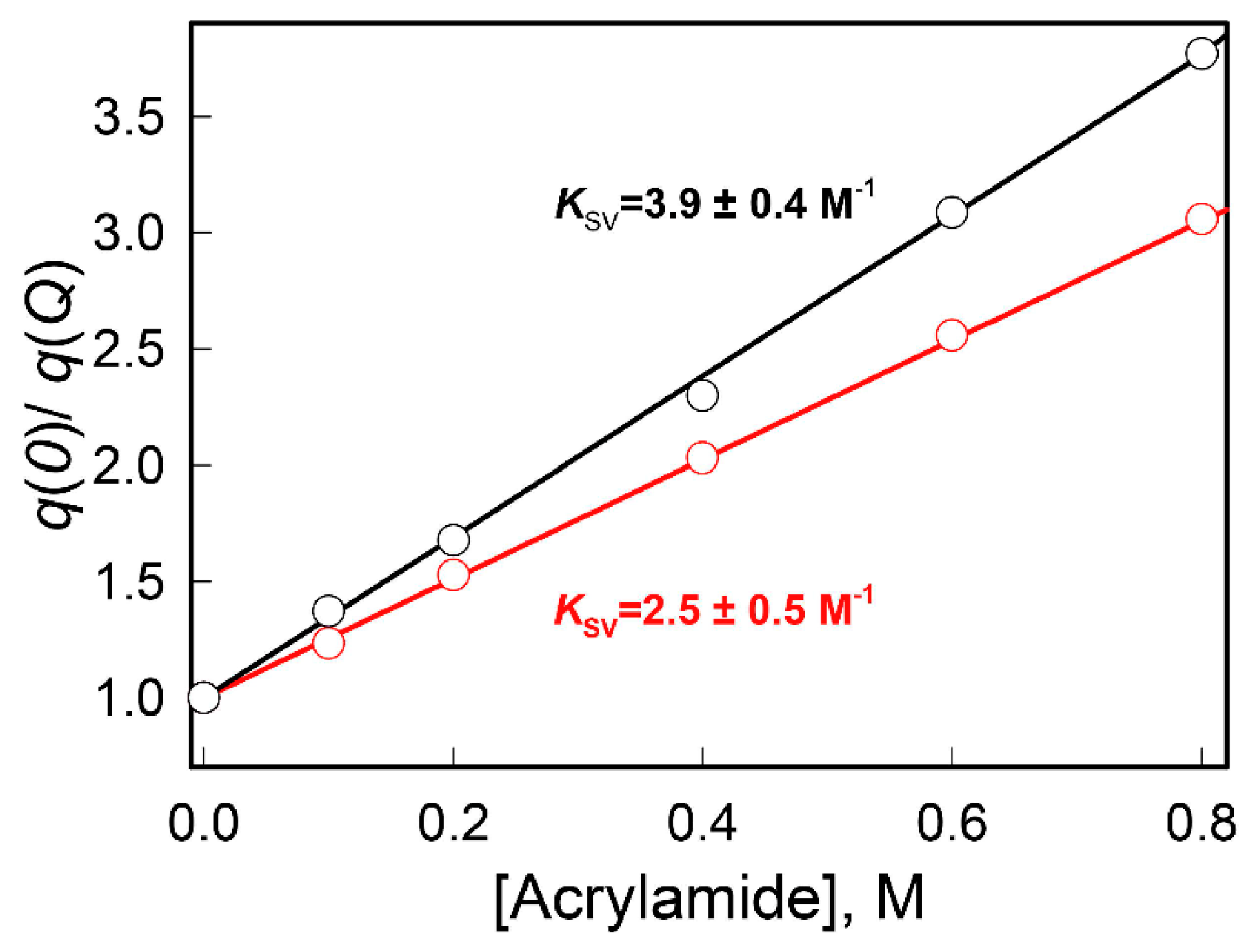
3.2.3. The Dynamic Quenching of the Intrinsic UV Fluorescence
3.2.4. Stopped-Flow Fluorescence Kinetic Experiments
3.2.5. Isothermal Titration Calorimetry
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Koshland, D.E. Application of a theory of enzyme specificity to protein synthesis. Proc. Natl. Acad. Sci. USA 1958, 44, 98–104. [Google Scholar] [CrossRef] [PubMed]
- Boehr, D.D.; Nussinov, R.; Wright, P.E. The role of dynamic conformational ensembles in biomolecular recognition. Nat. Chem. Biol. 2009, 5, 789–796. [Google Scholar] [CrossRef] [PubMed]
- Abaturov, L.V.; Varshavsky, Y.M. Equilibrium dynamics of the space structure of globular proteins. Mol. Biol. (SSSR) 1976, 12, 36–42. [Google Scholar]
- Varshavsky, Y.M. Conformational features of biopolymers. Mol. Biol. 1985, 19, 230–247. [Google Scholar]
- Maity, A.; Majumdar, S.; Priya, P.; De, P.; Saha, S.; Ghosh Dastidar, S. Adaptability in protein structures: Structural dynamics and implications in ligand design. J. Biomol. Struct. Dyn. 2015, 33, 298–321. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Ma, B.; Tsai, C.J.; Sinha, N.; Nussinov, R. Folding and binding cascades: Dynamic landscapes and population shifts. Protein Sci. 2000, 9, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Fischer, E. Einfluss der configuration auf die wirkung der enzyme. Ber. Dtsch. Chem. Ges. 1894, 27, 2985–2993. [Google Scholar] [CrossRef]
- Csermely, P.; Palotai, R.; Nussinov, R. Induced fit, conformational selection and independent dynamic segments: An extended view of binding events. Trends Biochem. Sci. 2010, 35, 539–546. [Google Scholar] [CrossRef] [PubMed]
- Grant, B.J.; Gorfe, A.A.; McCammon, J.A. Large conformational changes in proteins: Signaling and other functions. Curr. Opin. Struct. Biol. 2010, 20, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Hammes, G.G.; Chang, Y.C.; Oas, T.G. Conformational selection or induced fit: A flux description of reaction mechanism. Proc. Natl. Acad. Sci. USA 2009, 106, 13737–13741. [Google Scholar] [CrossRef] [PubMed]
- Fukami-Kobayashi, K.; Tateno, Y.; Nishikawa, K. Domain dislocation: A change of core structure in periplasmic binding proteins in their evolutionary history. J. Mol. Biol. 1999, 286, 279–290. [Google Scholar] [CrossRef] [PubMed]
- Brautigam, C.A.; Deka, R.K.; Liu, W.Z.; Norgard, M.V. The Tp0684 (MglB-2) lipoprotein of Treponema pallidum: A glucose-binding protein with divergent topology. PLoS ONE 2016, 11, e0161022. [Google Scholar] [CrossRef] [PubMed]
- Stepanenko, O.V.; Fonin, A.V.; Kuznetsova, I.M.; Turoverov, K.K. Ligand-binding proteins: Structure, stability and practical application. Protein Struct. InTech Rijeka 2012, 265–290. [Google Scholar] [CrossRef]
- Yesylevskyy, S.O.; Kharkyanen, V.N.; Demchenko, A.P. The change of protein intradomain mobility on ligand binding: Is it a commonly observed phenomenon? Biophys. J. 2006, 91, 3002–3013. [Google Scholar] [CrossRef] [PubMed]
- Yesylevskyy, S.O.; Kharkyanen, V.N.; Demchenko, A.P. Dynamic protein domains: Identification, interdependence, and stability. Biophys. J. 2006, 91, 670–685. [Google Scholar] [CrossRef] [PubMed]
- Felder, C.B.; Graul, R.C.; Lee, A.Y.; Merkle, H.P.; Sadee, W. The Venus flytrap of periplasmic binding proteins: An ancient protein module present in multiple drug receptors. AAPS Pharm. Sci. 1999, 1, E2. [Google Scholar] [CrossRef]
- Flocco, M.M.; Mowbray, S.L. The 1.9 A X-ray structure of a closed unliganded form of the periplasmic glucose/galactose receptor from Salmonella typhimurium. J. Biol. Chem. 1994, 269, 8931–8936. [Google Scholar] [PubMed]
- Ortega, G.; Castano, D.; Diercks, T.; Millet, O. Carbohydrate affinity for the glucose-galactose binding protein is regulated by allosteric domain motions. J. Am. Chem. Soc. 2012, 134, 19869–19876. [Google Scholar] [CrossRef] [PubMed]
- Luck, L.A.; Falke, J.J. 19F-NMR studies of the d-galactose chemosensory receptor. 1. Sugar binding yields a global structural change. Biochemistry 1991, 30, 4248–4256. [Google Scholar] [CrossRef] [PubMed]
- Careaga, C.L.; Sutherland, J.; Sabeti, J.; Falke, J.J. Large amplitude twisting motions of an interdomain hinge: A disulfide trapping study of the galactose-glucose binding protein. Biochemistry 1995, 34, 3048–3055. [Google Scholar] [CrossRef] [PubMed]
- Atilgan, C.; Atilgan, A.R. Perturbation-response scanning reveals ligand entry-exit mechanisms of ferric binding protein. PLoS Comput. Biol. 2009, 5, e1000544. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oswald, C.; Smits, S.H.; Hoing, M.; Sohn-Bosser, L.; Dupont, L.; Le Rudulier, D.; Schmitt, L.; Bremer, E. Crystal structures of the choline/acetylcholine substrate-binding protein ChoX from Sinorhizobium meliloti in the liganded and unliganded-closed states. J. Biol. Chem. 2008, 283, 32848–32859. [Google Scholar] [CrossRef] [PubMed]
- Bermejo, G.A.; Strub, M.P.; Ho, C.; Tjandra, N. Ligand-free open-closed transitions of periplasmic binding proteins: The case of glutamine-binding protein. Biochemistry 2010, 49, 1893–1902. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.; Lee, S.; Jeon, A.; Choi, J.M.; Lee, H.S.; Hohng, S.; Kim, H.S. A single-molecule dissection of ligand binding to a protein with intrinsic dynamics. Nat. Chem. Biol. 2013, 9, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Bucher, D.; Grant, B.J.; McCammon, J.A. Induced fit or conformational selection? The role of the semi-closed state in the maltose binding protein. Biochemistry 2011, 50, 10530–10539. [Google Scholar] [CrossRef] [PubMed]
- Silva, D.A.; Bowman, G.R.; Sosa-Peinado, A.; Huang, X. A role for both conformational selection and induced fit in ligand binding by the LAO protein. PLoS Comput. Biol. 2011, 7, e1002054. [Google Scholar] [CrossRef] [PubMed]
- Vyas, N.K.; Vyas, M.N.; Quiocho, F.A. The 3 A resolution structure of a d-galactose-binding protein for transport and chemotaxis in Escherichia coli. Proc. Natl. Acad. Sci. USA 1983, 80, 1792–1796. [Google Scholar] [CrossRef] [PubMed]
- Borrok, M.J.; Kiessling, L.L.; Forest, K.T. Conformational changes of glucose/galactose-binding protein illuminated by open, unliganded, and ultra-high-resolution ligand-bound structures. Protein Sci. 2007, 16, 1032–1041. [Google Scholar] [CrossRef] [PubMed]
- Vyas, N.K.; Vyas, M.N.; Quiocho, F.A. Sugar and signal-transducer binding sites of the Escherichia coli galactose chemoreceptor protein. Science 1988, 242, 1290–1295. [Google Scholar] [CrossRef] [PubMed]
- Aqvist, J.; Mowbray, S.L. Sugar recognition by a glucose/galactose receptor. Evaluation of binding energetics from molecular dynamics simulations. J. Biol. Chem. 1995, 270, 9978–9981. [Google Scholar] [PubMed]
- Boos, W. The galactose binding protein and its relationship to the beta-methylgalactoside permease from Escherichia coli. Eur. J. Biochem. 1969, 10, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Hazelbauer, G.L.; Adler, J. Role of the galactose binding protein in chemotaxis of Escherichia coli toward galactose. Nat. New Biol. 1971, 230, 101–104. [Google Scholar] [CrossRef] [PubMed]
- Rotman, B.; Ellis, J.H., Jr. Antibody-mediated modification of the binding properties of a protein related to galactose transport. J. Bacteriol. 1972, 111, 791–796. [Google Scholar] [PubMed]
- Miller, D.M., III; Olson, J.S.; Quiocho, F.A. The mechanism of sugar binding to the periplasmic receptor for galactose chemotaxis and transport in Escherichia coli. J. Biol. Chem. 1980, 255, 2465–2471. [Google Scholar] [PubMed]
- Kim, M.; Cho, A.E. The role of water molecules in stereoselectivity of glucose/galactose-binding protein. Sci. Rep. 2016, 6, 36807. [Google Scholar] [CrossRef] [PubMed]
- Shilton, B.H.; Flocco, M.M.; Nilsson, M.; Mowbray, S.L. Conformational changes of three periplasmic receptors for bacterial chemotaxis and transport: The maltose-, glucose/galactose- and ribose-binding proteins. J. Mol. Biol. 1996, 264, 350–363. [Google Scholar] [CrossRef] [PubMed]
- Ravindranathan, K.P.; Gallicchio, E.; Levy, R.M. Conformational equilibria and free energy profiles for the allosteric transition of the ribose-binding protein. J. Mol. Biol. 2005, 353, 196–210. [Google Scholar] [CrossRef] [PubMed]
- Messina, T.C.; Talaga, D.S. Protein free energy landscapes remodeled by ligand binding. Biophys. J. 2007, 93, 579–585. [Google Scholar] [CrossRef] [PubMed]
- Unione, L.; Ortega, G.; Mallagaray, A.; Corzana, F.; Perez-Castells, J.; Canales, A.; Jimenez-Barbero, J.; Millet, O. Unraveling the Conformational landscape of ligand binding to glucose/galactose-binding protein by paramagnetic NMR and MD simulations. ACS Chem. Biol. 2016, 11, 2149–2157. [Google Scholar] [CrossRef] [PubMed]
- Amiss, T.J.; Sherman, D.B.; Nycz, C.M.; Andaluz, S.A.; Pitner, J.B. Engineering and rapid selection of a low-affinity glucose/galactose-binding protein for a glucose biosensor. Protein Sci. 2007, 16, 2350–2359. [Google Scholar] [CrossRef] [PubMed]
- De Lorimier, R.M.; Smith, J.J.; Dwyer, M.A.; Looger, L.L.; Sali, K.M.; Paavola, C.D.; Rizk, S.S.; Sadigov, S.; Conrad, D.W.; Loew, L.; et al. Construction of a fluorescent biosensor family. Protein Sci. 2002, 11, 2655–2675. [Google Scholar] [CrossRef] [PubMed]
- Deuschle, K.; Okumoto, S.; Fehr, M.; Looger, L.L.; Kozhukh, L.; Frommer, W.B. Construction and optimization of a family of genetically encoded metabolite sensors by semirational protein engineering. Protein Sci. 2005, 14, 2304–2314. [Google Scholar] [CrossRef] [PubMed]
- Fehr, M.; Lalonde, S.; Lager, I.; Wolff, M.W.; Frommer, W.B. In vivo imaging of the dynamics of glucose uptake in the cytosol of COS-7 cells by fluorescent nanosensors. J. Biol. Chem. 2003, 278, 19127–19133. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.; Rao, G.; Tolosa, L. On the possibility of real-time monitoring of glucose in cell culture by microdialysis using a fluorescent glucose binding protein sensor. Biotechnol. Prog. 2008, 24, 691–697. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.; Tolosa, L.; Rao, G. Dual-labeled glucose binding protein for ratiometric measurements of glucose. Anal. Chem. 2004, 76, 1403–1410. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, H.V.; Pfeiffer, Z.A.; Amiss, T.J.; Sherman, D.B.; Pitner, J.B. Direct detection of glucose by surface plasmon resonance with bacterial glucose/galactose-binding protein. Biosens. Bioelectron. 2004, 19, 653–660. [Google Scholar] [CrossRef]
- Khan, F.; Gnudi, L.; Pickup, J.C. Fluorescence-based sensing of glucose using engineered glucose/galactose-binding protein: A comparison of fluorescence resonance energy transfer and environmentally sensitive dye labelling strategies. Biochem. Biophys. Res. Commun. 2008, 365, 102–106. [Google Scholar] [CrossRef] [PubMed]
- Khan, F.; Pickup, J.C. Near-infrared fluorescence glucose sensing based on glucose/galactose-binding protein coupled to 651-Blue Oxazine. Biochem. Biophys. Res. Commun. 2013, 438, 488–492. [Google Scholar] [CrossRef] [PubMed]
- Khan, F.; Saxl, T.E.; Pickup, J.C. Fluorescence intensity- and lifetime-based glucose sensing using an engineered high-Kd mutant of glucose/galactose-binding protein. Anal. Biochem. 2010, 399, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Marvin, J.S.; Hellinga, H.W. Engineering biosensors by introducing fluorescent allosteric signal transducers: Construction of a novel glucose sensor. J. Am. Chem. Soc. 1998, 120, 7–11. [Google Scholar] [CrossRef]
- Sakaguchi-Mikami, A.; Taneoka, A.; Yamoto, R.; Ferri, S.; Sode, K. Engineering of ligand specificity of periplasmic binding protein for glucose sensing. Biotechnol. Lett. 2008, 30, 1453–1460. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi-Mikami, A.; Taniguchi, A.; Sode, K.; Yamazaki, T. Construction of a novel glucose-sensing molecule based on a substrate-binding protein for intracellular sensing. Biotechnol. Bioeng. 2011, 108, 725–733. [Google Scholar] [CrossRef] [PubMed]
- Saxl, T.; Khan, F.; Ferla, M.; Birch, D.; Pickup, J. A fluorescence lifetime-based fibre-optic glucose sensor using glucose/galactose-binding protein. Analyst 2011, 136, 968–972. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saxl, T.; Khan, F.; Matthews, D.R.; Zhi, Z.L.; Rolinski, O.; Ameer-Beg, S.; Pickup, J. Fluorescence lifetime spectroscopy and imaging of nano-engineered glucose sensor microcapsules based on glucose/galactose-binding protein. Biosens. Bioelectron. 2009, 24, 3229–3234. [Google Scholar] [CrossRef] [PubMed]
- Sokolov, I.; Subba-Rao, V.; Luck, L.A. Change in rigidity in the activated form of the glucose/galactose receptor from Escherichia coli: A phenomenon that will be key to the development of biosensors. Biophys. J. 2006, 90, 1055–1063. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.; Sherman, D.B.; Amiss, T.J.; Andaluz, S.A.; Pitner, J.B. Synthesis and biosensor performance of a near-IR thiol-reactive fluorophore based on benzothiazolium squaraine. Bioconjug. Chem. 2007, 18, 1841–1846. [Google Scholar] [CrossRef] [PubMed]
- Thomas, K.J.; Sherman, D.B.; Amiss, T.J.; Andaluz, S.A.; Pitner, J.B. A long-wavelength fluorescent glucose biosensor based on bioconjugates of galactose/glucose binding protein and Nile Red derivatives. Diabetes Technol. Ther. 2006, 8, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Tolosa, L. On the design of low-cost fluorescent protein biosensors. Adv. Biochem. Eng. Biotechnol. 2010, 116, 143–157. [Google Scholar]
- Tolosa, L.; Gryczynski, I.; Eichhorn, L.R.; Dattelbaum, J.D.; Castellano, F.N.; Rao, G.; Lakowicz, J.R. Glucose sensor for low-cost lifetime-based sensing using a genetically engineered protein. Anal. Biochem. 1999, 267, 114–120. [Google Scholar] [CrossRef] [PubMed]
- Veetil, J.V.; Jin, S.; Ye, K. A glucose sensor protein for continuous glucose monitoring. Biosens. Bioelectron. 2010, 26, 1650–1655. [Google Scholar] [CrossRef] [PubMed]
- Weidemaier, K.; Lastovich, A.; Keith, S.; Pitner, J.B.; Sistare, M.; Jacobson, R.; Kurisko, D. Multi-day pre-clinical demonstration of glucose/galactose binding protein-based fiber optic sensor. Biosens. Bioelectron. 2011, 26, 4117–4123. [Google Scholar] [CrossRef] [PubMed]
- Ye, K.; Schultz, J.S. Genetic engineering of an allosterically based glucose indicator protein for continuous glucose monitoring by fluorescence resonance energy transfer. Anal. Chem. 2003, 75, 3451–3459. [Google Scholar] [CrossRef] [PubMed]
- Fonin, A.V.; Stepanenko, O.V.; Povarova, O.I.; Volova, C.A.; Philippova, E.M.; Bublikov, G.S.; Kuznetsova, I.M.; Demchenko, A.P.; Turoverov, K.K. Spectral characteristics of the mutant form GGBP/H152C of d-glucose/d-galactose-binding protein labeled with fluorescent dye BADAN: Influence of external factors. PeerJ 2014, 2, e275. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuznetsova, I.M.; Stepanenko, O.V.; Turoverov, K.K.; Zhu, L.; Zhou, J.M.; Fink, A.L.; Uversky, V.N. Unraveling multistate unfolding of rabbit muscle creatine kinase. Biochim. Biophys. Acta 2002, 1596, 138–155. [Google Scholar] [CrossRef]
- Povarova, O.I.; Kuznetsova, I.M.; Turoverov, K.K. Differences in the pathways of proteins unfolding induced by urea and guanidine hydrochloride: Molten globule state and aggregates. PLoS ONE 2010, 5, e15035. [Google Scholar] [CrossRef] [PubMed]
- Verkhusha, V.V.; Kuznetsova, I.M.; Stepanenko, O.V.; Zaraisky, A.G.; Shavlovsky, M.M.; Turoverov, K.K.; Uversky, V.N. High stability of Discosoma DsRed as compared to Aequorea EGFP. Biochemistry 2003, 42, 7879–7884. [Google Scholar] [CrossRef] [PubMed]
- Stepanenko, O.V.; Verkhusha, V.V.; Kazakov, V.I.; Shavlovsky, M.M.; Kuznetsova, I.M.; Uversky, V.N.; Turoverov, K.K. Comparative studies on the structure and stability of fluorescent proteins EGFP, zFP506, mRFP1, “dimer2”, and DsRed1. Biochemistry 2004, 43, 14913–14923. [Google Scholar] [CrossRef] [PubMed]
- Stepanenko, O.V.; Verkhusha, V.V.; Shavlovsky, M.M.; Kuznetsova, I.M.; Uversky, V.N.; Turoverov, K.K. Understanding the role of Arg96 in structure and stability of green fluorescent protein. Proteins 2008, 73, 539–551. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Bhuyan, A.K. Entropic stabilization of myoglobin by subdenaturing concentrations of guanidine hydrochloride. J. Biol. Inorg. Chem. 2009, 14, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Mayr, L.M.; Schmid, F.X. Stabilization of a protein by guanidinium chloride. Biochemistry 1993, 32, 7994–7998. [Google Scholar] [CrossRef] [PubMed]
- Rao, M.T.; Bhuyan, A.K.; Venu, K.; Sastry, V.S. Nonlinear effect of GdnHCl on hydration dynamics of proteins: A 1H magnetic relaxation dispersion study. J. Phys. Chem. B 2009, 113, 6994–7002. [Google Scholar] [CrossRef] [PubMed]
- Sen, P.; Khan, M.M.; Equbal, A.; Ahmad, E.; Khan, R.H. At very low concentrations known chaotropes act as kosmotropes for the N and B isoforms of human serum albumin. Biochem. Cell Biol. 2013, 91, 72–78. [Google Scholar] [CrossRef] [PubMed]
- Zarrine-Afsar, A.; Mittermaier, A.; Kay, L.E.; Davidson, A.R. Protein stabilization by specific binding of guanidinium to a functional arginine-binding surface on an SH3 domain. Protein Sci. 2006, 15, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Bhuyan, A.K. Protein stabilization by urea and guanidine hydrochloride. Biochemistry 2002, 41, 13386–13394. [Google Scholar] [CrossRef] [PubMed]
- Singh, L.R.; Poddar, N.K.; Dar, T.A.; Kumar, R.; Ahmad, F. Protein and DNA destabilization by osmolytes: The other side of the coin. Life Sci. 2011, 88, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Stepanenko, O.V.; Povarova, O.I.; Fonin, A.V.; Kuznetsova, I.M.; Turoverov, K.K.; Staiano, M.; Varriale, A.; D’Auria, S. New insight into protein-ligand interactions. The case of the d-galactose/d-glucose-binding protein from Escherichia coli. J. Phys. Chem. B 2011, 115, 2765–2773. [Google Scholar] [CrossRef] [PubMed]
- Stepanenko, O.V.; Fonin, A.V.; Morozova, K.S.; Verkhusha, V.V.; Kuznetsova, I.M.; Turoverov, K.K.; Staiano, M.; D’Auria, S. New insight in protein-ligand interactions. 2. Stability and properties of two mutant forms of the d-galactose/d-glucose-binding protein from E. coli. J. Phys. Chem. B 2011, 115, 9022–9032. [Google Scholar] [CrossRef] [PubMed]
- Stepanenko, O.V.; Fonin, A.V.; Stepanenko, O.V.; Staiano, M.; D’Auria, S.; Kuznetsova, I.M.; Turoverov, K.K. Tryptophan residue of the d-galactose/d-glucose-binding protein from E. Coli localized in its active center does not contribute to the change in intrinsic fluorescence upon glucose binding. J. Fluoresc. 2015, 25, 87–94. [Google Scholar] [CrossRef] [PubMed]
- Fonin, A.V.; Kuznetsova, I.M.; Turoverov, K.K. Spectral properties of BADAN in solutions with different polarities. J. Mol. Struct. 2015, 1090, 107–111. [Google Scholar] [CrossRef]
- Fonin, A.V.; Uversky, V.N.; Kuznetsova, I.M.; Turoverov, K.K. Protein folding and stability in the presence of osmolytes. Biofizika 2016, 61, 222–230. [Google Scholar] [CrossRef] [PubMed]
- Yancey, P.H. Organic osmolytes as compatible, metabolic and counteracting cytoprotectants in high osmolarity and other stresses. J. Exp. Biol. 2005, 208, 2819–2830. [Google Scholar] [CrossRef] [PubMed]
- Chebotareva, N.A.; Kurganov, B.I.; Harding, S.E.; Winzor, D.J. Effect of osmolytes on the interaction of flavin adenine dinucleotide with muscle glycogen phosphorylase b. Biophys. Chem. 2005, 113, 61–66. [Google Scholar] [CrossRef] [PubMed]
- Ignatova, Z.; Gierasch, L.M. Effects of osmolytes on protein folding and aggregation in cells. Methods Enzymol. 2007, 428, 355–372. [Google Scholar] [PubMed]
- Flores Jimenez, R.H.; Do Cao, M.A.; Kim, M.; Cafiso, D.S. Osmolytes modulate conformational exchange in solvent-exposed regions of membrane proteins. Protein Sci. 2010, 19, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Jain, R.; Sharma, D.; Kumar, S.; Kumar, R. Factor defining the effects of glycine betaine on the thermodynamic stability and internal dynamics of horse cytochrome C. Biochemistry 2014, 53, 5221–5235. [Google Scholar] [CrossRef] [PubMed]
- Street, T.O.; Krukenberg, K.A.; Rosgen, J.; Bolen, D.W.; Agard, D.A. Osmolyte-induced conformational changes in the Hsp90 molecular chaperone. Protein Sci. 2010, 19, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Cioni, P.; Bramanti, E.; Strambini, G.B. Effects of sucrose on the internal dynamics of azurin. Biophys. J. 2005, 88, 4213–4222. [Google Scholar] [CrossRef] [PubMed]
- Gianni, S.; Dogan, J.; Jemth, P. Distinguishing induced fit from conformational selection. Biophys. Chem. 2014, 189, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Vogt, A.D.; Di Cera, E. Conformational selection or induced fit? A critical appraisal of the kinetic mechanism. Biochemistry 2012, 51, 5894–5902. [Google Scholar] [CrossRef] [PubMed]
- Vogt, A.D.; Pozzi, N.; Chen, Z.; Di Cera, E. Essential role of conformational selection in ligand binding. Biophys. Chem. 2014, 186, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Olsson, T.S.; Williams, M.A.; Pitt, W.R.; Ladbury, J.E. The thermodynamics of protein-ligand interaction and solvation: Insights for ligand design. J. Mol. Biol. 2008, 384, 1002–1017. [Google Scholar] [CrossRef] [PubMed]
- Velazquez-Campoy, A.; Freire, E. Isothermal titration calorimetry to determine association constants for high-affinity ligands. Nat. Protoc. 2006, 1, 186–191. [Google Scholar] [CrossRef] [PubMed]
- Butler, S.L.; Falke, J.J. Effects of protein stabilizing agents on thermal backbone motions: A disulfide trapping study. Biochemistry 1996, 35, 10595–10600. [Google Scholar] [CrossRef] [PubMed]
- Bellissent-Funel, M.C.; Hassanali, A.; Havenith, M.; Henchman, R.; Pohl, P.; Sterpone, F.; van der Spoel, D.; Xu, Y.; Garcia, A.E. Water determines the structure and dynamics of proteins. Chem. Rev. 2016, 116, 7673–7697. [Google Scholar] [CrossRef] [PubMed]
- Timasheff, S.N. Protein-solvent preferential interactions, protein hydration, and the modulation of biochemical reactions by solvent components. Proc. Natl. Acad. Sci. USA 2002, 99, 9721–9726. [Google Scholar] [CrossRef] [PubMed]
- Kendrick, B.S.; Chang, B.S.; Arakawa, T.; Peterson, B.; Randolph, T.W.; Manning, M.C.; Carpenter, J.F. Preferential exclusion of sucrose from recombinant interleukin-1 receptor antagonist: Role in restricted conformational mobility and compaction of native state. Proc. Natl. Acad. Sci. USA 1997, 94, 11917–11922. [Google Scholar] [CrossRef] [PubMed]
- Dwyer, M.A.; Hellinga, H.W. Periplasmic binding proteins: A versatile superfamily for protein engineering. Curr. Opin. Struct. Biol. 2004, 14, 495–504. [Google Scholar] [CrossRef] [PubMed]
- Schreier, B.; Stumpp, C.; Wiesner, S.; Hocker, B. Computational design of ligand binding is not a solved problem. Proc. Natl. Acad. Sci. USA 2009, 106, 18491–18496. [Google Scholar] [CrossRef] [PubMed]
- Mizoue, L.S.; Chazin, W.J. Engineering and design of ligand-induced conformational change in proteins. Curr. Opin. Struct. Biol. 2002, 12, 459–463. [Google Scholar] [CrossRef]
- Telmer, P.G.; Shilton, B.H. Insights into the conformational equilibria of maltose-binding protein by analysis of high affinity mutants. J. Biol. Chem. 2003, 278, 34555–34567. [Google Scholar] [CrossRef] [PubMed]
- Walker, I.H.; Hsieh, P.C.; Riggs, P.D. Mutations in maltose-binding protein that alter affinity and solubility properties. Appl. Microbiol. Biotechnol. 2010, 88, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Harayama, S.; Bollinger, J.; Iino, T.; Hazelbauer, G.L. Characterization of the mgl operon of Escherichia coli by transposon mutagenesis and molecular cloning. J. Bacteriol. 1983, 153, 408–415. [Google Scholar] [PubMed]
- Rotman, B.; Ganesan, A.K.; Guzman, R. Transport systems for galactose and galactosides in Escherichia coli. II. Substrate and inducer specificities. J. Mol. Biol. 1968, 36, 247–260. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Fonin, A.V.; Sulatskaya, A.I.; Kuznetsova, I.M.; Turoverov, K.K. Fluorescence of dyes in solutions with high absorbance. Inner filter effect correction. PLoS ONE 2014, 9, e103878. [Google Scholar] [CrossRef] [PubMed]
- Lakowicz, J.R. Principles of Fluorescence Spectroscopy, 3rd ed.; Springer: New York, NY, USA, 2006; p. 954. [Google Scholar]
- Anufrieva, Y.V.; Gotlib, Y.Y.; Krakovyak, M.G.; Pautov, V.D. Polarized luminescence as used for analyses of high frequency twisting vibrations in macromolecules. Polym. Sci. USSR 1976, 18, 3132–3141. [Google Scholar] [CrossRef]
- Tcherkasskaya, O.; Ptitsyn, O.B.; Knutson, J.R. Nanosecond dynamics of tryptophans in different conformational states of apomyoglobin proteins. Biochemistry 2000, 39, 1879–1889. [Google Scholar] [CrossRef] [PubMed]
- Freire, E.; Mayorga, O.L.; Straume, M. Isothermal titration calorimetry. Anal. Chem. 1990, 62, 950A–959A. [Google Scholar] [CrossRef]
- Brown, A. Analysis of cooperativity by isothermal titration calorimetry. Int. J. Mol. Sci. 2009, 10, 3457–3477. [Google Scholar] [CrossRef] [PubMed]
- Wiseman, T.; Williston, S.; Brandts, J.F.; Lin, L.N. Rapid measurement of binding constants and heats of binding using a new titration calorimeter. Anal. Biochem. 1989, 179, 131–137. [Google Scholar] [CrossRef]
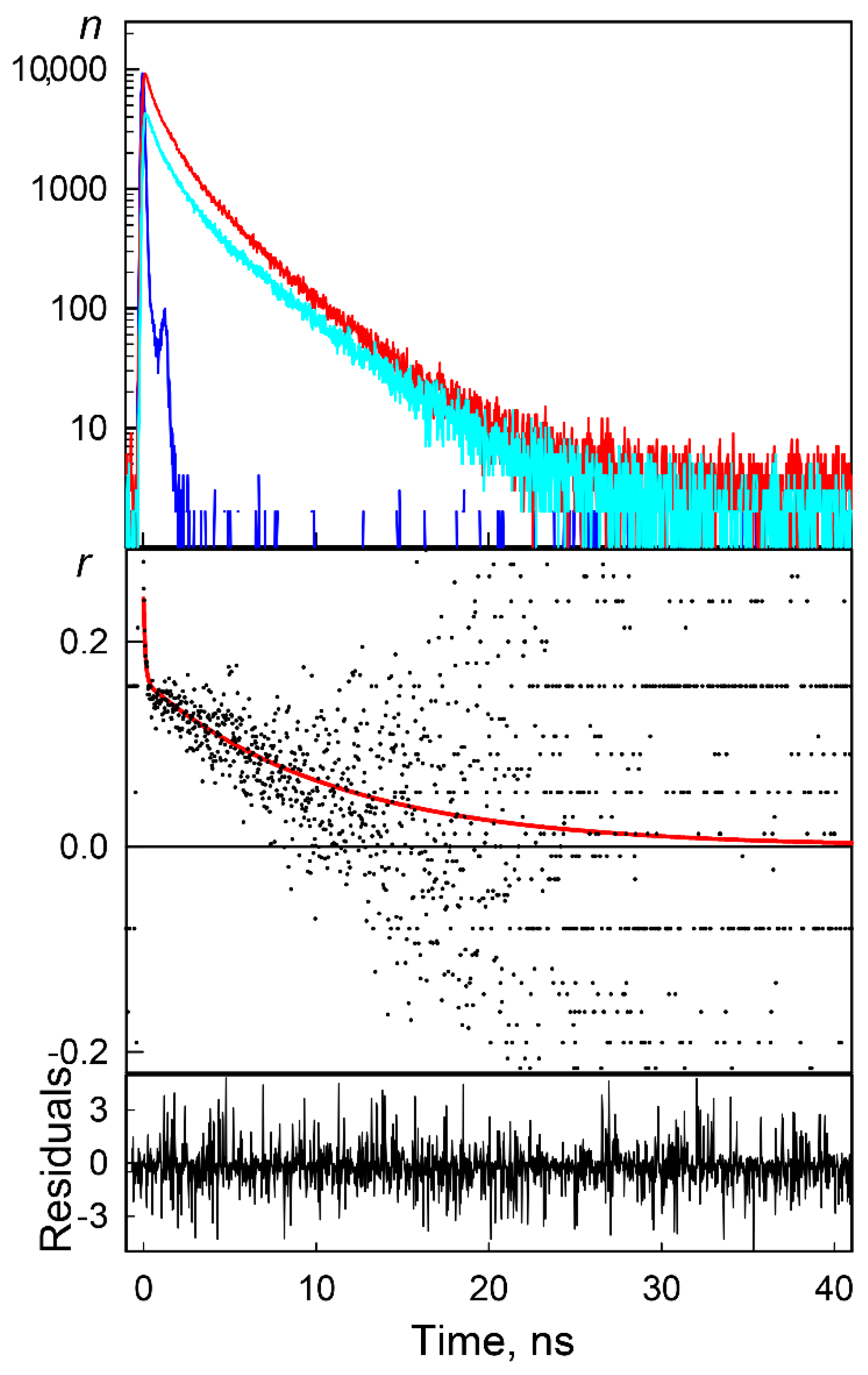


| GGBP/H152C-BADAN Apo-Form | ||||||||
| GdnHCl Concentration, M | <t>, ns | rfast | rslow | r0 | θ ** Degree | τfast, ns | τslow, ns | χ2 |
| 0.0 | 1.3 | 0.08 | 0.16 | 0.24 | 29 | 0.3 | 34 | 0.87 |
| 0.1 | 1.4 | 0.08 | 0.19 | 0.27 | 28 | 0.4 | 30 | 0.99 |
| 0.5 | 2.7 | 0.14 | 0.14 | 0.28 | 38 | 0.3 | 30 | 1.08 |
| GGBP/H152C-BADAN Holo-Form | ||||||||
| GdnHCl Concentration, M | <t>, ns | rfast | rslow | r0 | θ **, Degree | τfast, ns | τslow, ns | χ2 |
| 0.0 | 3.1 | 0.07 | 0.19 | 0.26 | 25 | 0.3 | 33 | 1.01 |
| 0.1 | 3.1 | 0.04 | 0.16 | 0.20 | 22 | 1.5 | 27 | 1.06 |
| 0.5 | 3.4 | 0.04 | 0.25 | 0.29 | 19 | 1.2 | 28 | 1.10 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fonin, A.V.; Golikova, A.D.; Zvereva, I.A.; D’Auria, S.; Staiano, M.; Uversky, V.N.; Kuznetsova, I.M.; Turoverov, K.K. Osmolyte-Like Stabilizing Effects of Low GdnHCl Concentrations on d-Glucose/d-Galactose-Binding Protein. Int. J. Mol. Sci. 2017, 18, 2008. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18092008
Fonin AV, Golikova AD, Zvereva IA, D’Auria S, Staiano M, Uversky VN, Kuznetsova IM, Turoverov KK. Osmolyte-Like Stabilizing Effects of Low GdnHCl Concentrations on d-Glucose/d-Galactose-Binding Protein. International Journal of Molecular Sciences. 2017; 18(9):2008. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18092008
Chicago/Turabian StyleFonin, Alexander V., Alexandra D. Golikova, Irina A. Zvereva, Sabato D’Auria, Maria Staiano, Vladimir N. Uversky, Irina M. Kuznetsova, and Konstantin K. Turoverov. 2017. "Osmolyte-Like Stabilizing Effects of Low GdnHCl Concentrations on d-Glucose/d-Galactose-Binding Protein" International Journal of Molecular Sciences 18, no. 9: 2008. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms18092008









