Correction: Wang, Y.T., et al. Selenium Nanoparticle Synthesized by Proteus mirabilis YC801: An Efficacious Pathway for Selenite Biotransformation and Detoxification. Int. J. Mol. Sci. 2018, 19, 3809
1. Change in Main Body Paragraphs
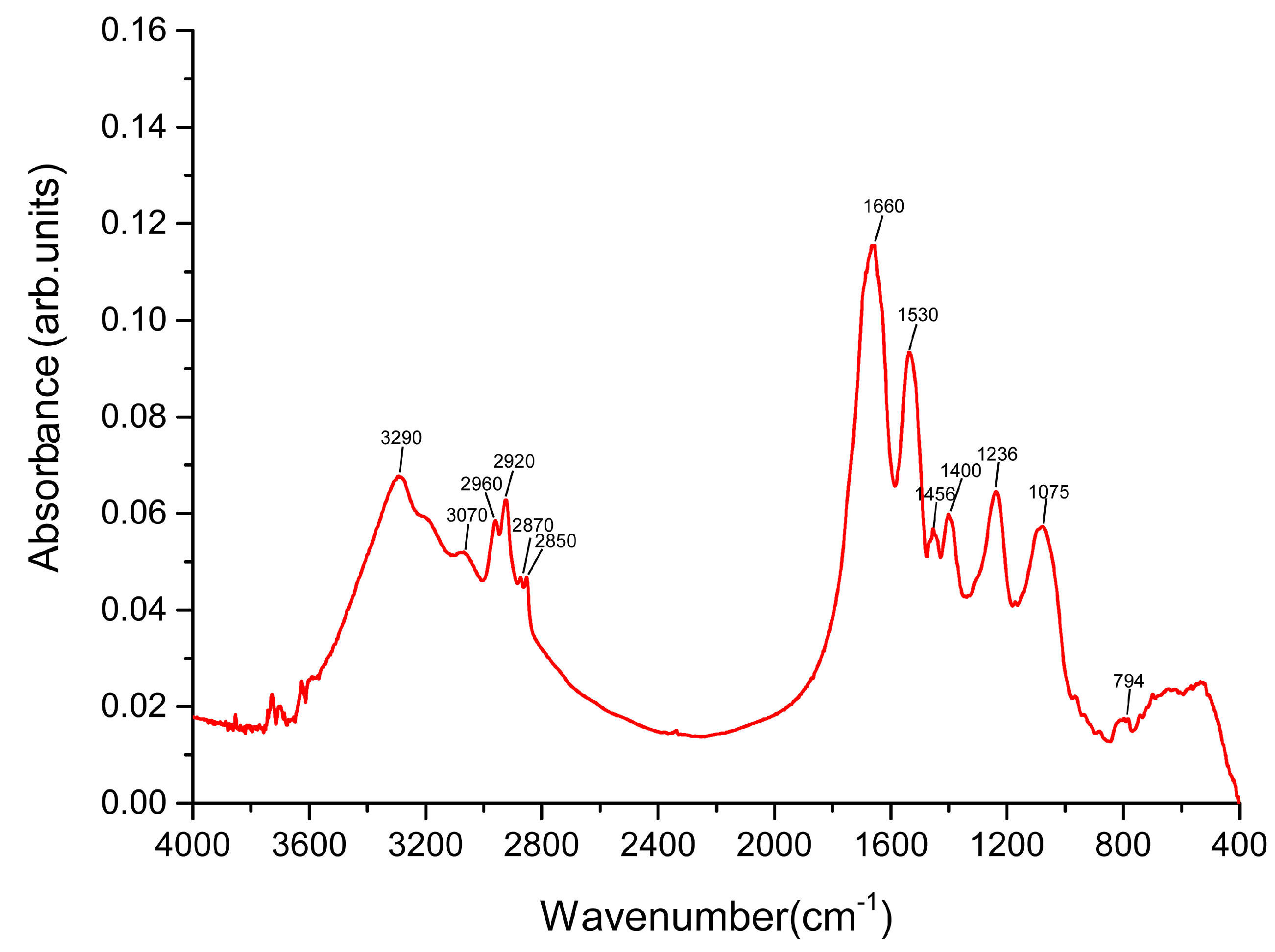
2. Change in Figure
Reference
- Wang, Y.; Shu, X.; Hou, J.; Lu, W.; Zhao, W.; Huang, S.; Wu, L. Studies on selenium nanoparticles synthesized by Proteus mirabilis YC801: An efficacious pathway for selenite biotransformation and detoxification. Int. J. Mol. Sci. 2018, 19, 3809. [Google Scholar] [CrossRef] [PubMed] [Green Version]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Y.; Shu, X.; Hou, J.; Lu, W.; Zhao, W.; Huang, S.; Wu, L. Correction: Wang, Y.T., et al. Selenium Nanoparticle Synthesized by Proteus mirabilis YC801: An Efficacious Pathway for Selenite Biotransformation and Detoxification. Int. J. Mol. Sci. 2018, 19, 3809. Int. J. Mol. Sci. 2020, 21, 2638. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21072638
Wang Y, Shu X, Hou J, Lu W, Zhao W, Huang S, Wu L. Correction: Wang, Y.T., et al. Selenium Nanoparticle Synthesized by Proteus mirabilis YC801: An Efficacious Pathway for Selenite Biotransformation and Detoxification. Int. J. Mol. Sci. 2018, 19, 3809. International Journal of Molecular Sciences. 2020; 21(7):2638. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21072638
Chicago/Turabian StyleWang, Yuting, Xian Shu, Jinyan Hou, Weili Lu, Weiwei Zhao, Shengwei Huang, and Lifang Wu. 2020. "Correction: Wang, Y.T., et al. Selenium Nanoparticle Synthesized by Proteus mirabilis YC801: An Efficacious Pathway for Selenite Biotransformation and Detoxification. Int. J. Mol. Sci. 2018, 19, 3809" International Journal of Molecular Sciences 21, no. 7: 2638. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms21072638




