A cis-Acting Mutation in the PxABCG1 Promoter Is Associated with Cry1Ac Resistance in Plutella xylostella (L.)
Abstract
:1. Introduction
2. Results
2.1. Cloning and Analysis of the PxABCG1 Promoter in Bt-Susceptible and Bt-Resistant Strains
2.2. PxABCG1 Promoter Activity Differs between Bt-Susceptible and Bt-Resistant Strains
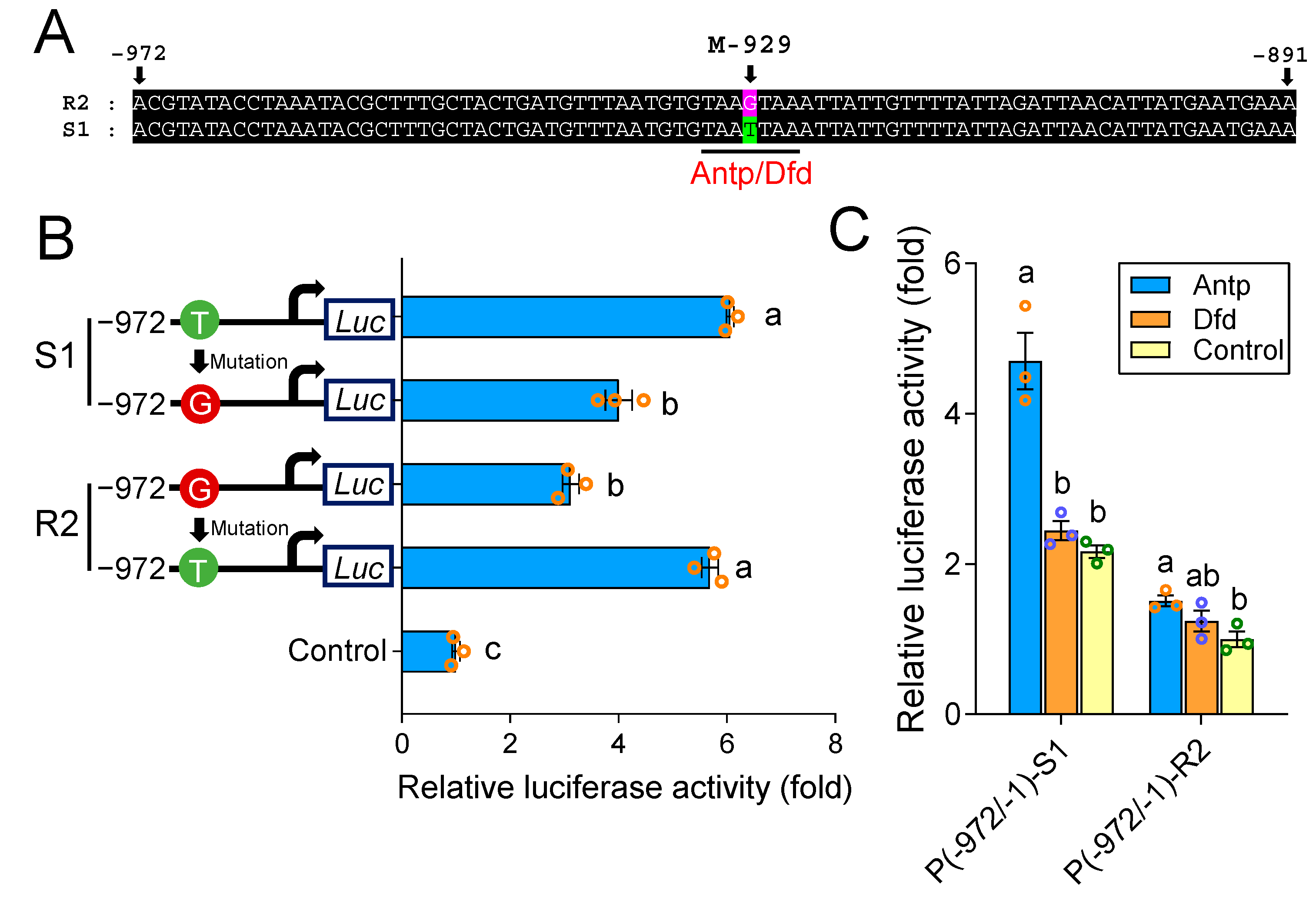
2.3. A cis-Acting Mutation in the Binding Site of Antp Reduces PxABCG1 Promoter Activity
2.4. A cis-Acting Mutation Causes Antp to Fail to Bind to the Promoter and Regulate PxABCG1
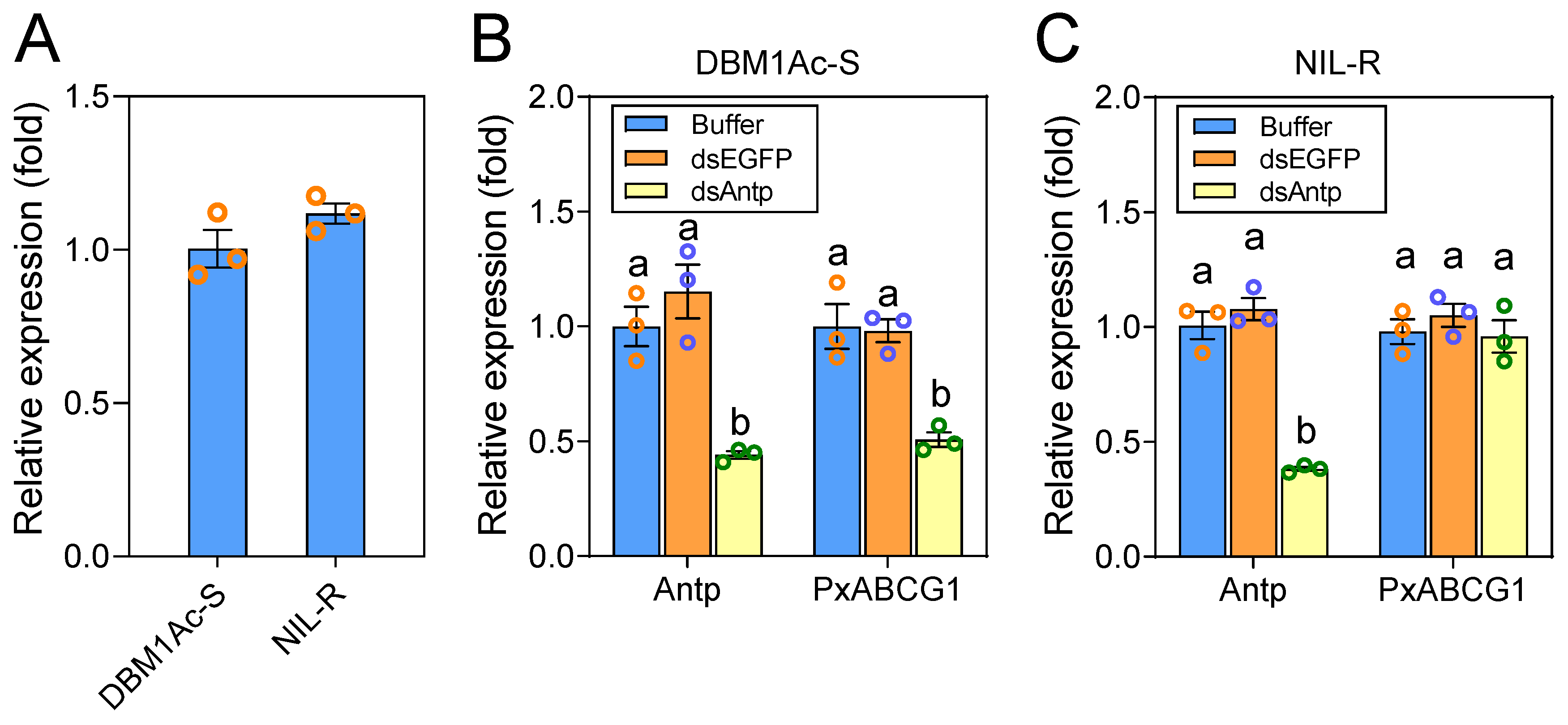
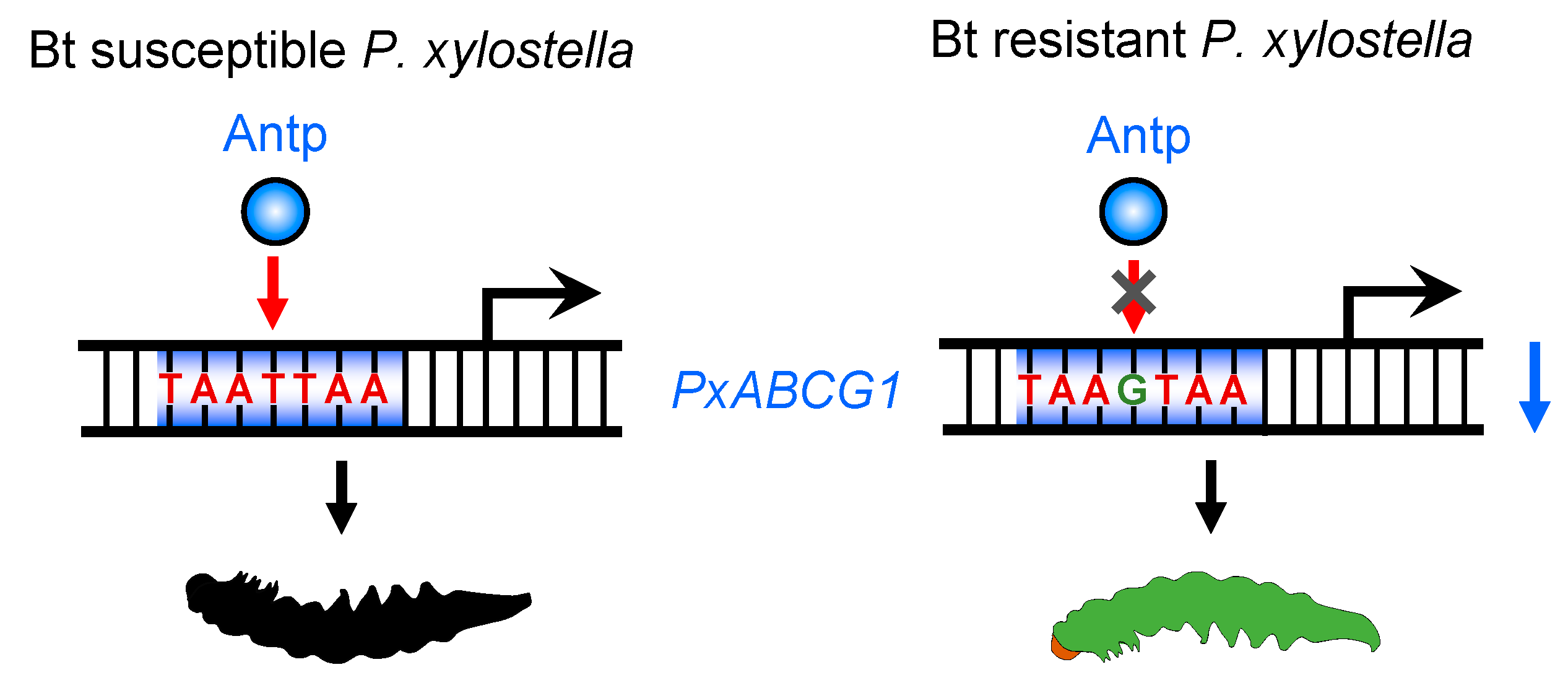
2.5. Antp-Mediated Positive Regulation of PxABCG1 In Vivo Is Eliminated in the Bt-Resistant Strain
3. Discussion
4. Materials and Methods
4.1. Insect Strains and Cell Line
4.2. Toxin Preparation and Bioassay
4.3. Extraction of DNA and RNA
4.4. Cloning of the Promoter and TFs
4.5. Bioinformatic Analysis
4.6. Dual-Luciferase Assay
4.7. Y1H Assay
4.8. qPCR Analysis
4.9. RNAi
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bravo, A.; Likitvivatanavong, S.; Gill, S.S.; Soberón, M. Bacillus thuringiensis: A Story of a Successful Bioinsecticide. Insect Biochem. Mol. Biol. 2011, 41, 423–431. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sanahuja, G.; Banakar, R.; Twyman, R.M.; Capell, T.; Christou, P. Bacillus thuringiensis: A Century of Research, Development and Commercial Applications. Plant Biotechnol. J. 2011, 9, 283–300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sanchis, V. From Microbial Sprays to Insect-Resistant Transgenic Plants: History of the Biopesticide Bacillus thuringiensis. A Review. Agron. Sustain. Dev. 2011, 31, 217–231. [Google Scholar] [CrossRef]
- Jurat-Fuentes, J.L.; Heckel, D.G.; Ferre, J. Mechanisms of Resistance to Insecticidal Proteins from Bacillus thuringiensis. Annu. Rev. Entomol. 2021, 66, 121–140. [Google Scholar] [CrossRef]
- ISAAA. Global Status of Commercialized Biotech/GM Crops in 2019: Biotech Crops Drive Socio-Economic Development and Sustainable Environment in the New Frontieri. In ISAAA Brief No. 55; Cornell University: Ithaca, NY, USA, 2019. [Google Scholar]
- Tabashnik, B.E.; Brévault, T.; Carrière, Y. Insect Resistance to Bt Crops: Lessons from the First Billion Acres. Nat. Biotechnol. 2013, 31, 510–521. [Google Scholar] [CrossRef] [PubMed]
- Tabashnik, B.E.; Carrière, Y. Surge in Insect Resistance to Transgenic Crops and Prospects for Sustainability. Nat. Biotechnol. 2017, 35, 926–935. [Google Scholar] [CrossRef] [PubMed]
- Pardo-López, L.; Soberón, M.; Bravo, A. Bacillus thuringiensis Insecticidal Three-Domain Cry Toxins: Mode of Action, Insect Resistance and Consequences for Crop Protection. FEMS Microbiol. Rev. 2013, 37, 3–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jurat-Fuentes, J.L.; Crickmore, N. Specificity Determinants for Cry Insecticidal Proteins: Insights from Their Mode of Action. J. Invertebr. Pathol. 2017, 142, 5–10. [Google Scholar] [CrossRef] [Green Version]
- de Bortoli, C.P.; Jurat-Fuentes, J.L. Mechanisms of Resistance to Commercially Relevant Entomopathogenic Bacteria. Curr. Opin. Insect Sci. 2019, 33, 56–62. [Google Scholar] [CrossRef]
- Pigott, C.R.; Ellar, D.J. Role of Receptors in Bacillus thuringiensis Crystal Toxin Activity. Microbiol. Mol. Biol. Rev. 2007, 71, 255–281. [Google Scholar] [CrossRef] [Green Version]
- Wu, C.; Chakrabarty, S.; Jin, M.; Liu, K.; Xiao, Y. Insect ATP-Binding Cassette (ABC) Transporters: Roles in Xenobiotic Detoxification and Bt Insecticidal Activity. Int. J. Mol. Sci. 2019, 20, 2829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adang, M.J.; Crickmore, N.; Jurat-Fuentes, J.L. Diversity of Bacillus thuringiensis Crystal Toxins and Mechanism of Action. Adv. Insect Physiol. 2014, 47, 39–87. [Google Scholar]
- Guo, Z.; Kang, S.; Zhu, X.; Xia, J.; Wu, Q.; Wang, S.; Xie, W.; Zhang, Y. Down-Regulation of a Novel ABC Transporter Gene (Pxwhite) Is Associated with Cry1Ac Resistance in the Diamondback Moth, Plutella xylostella (L.). Insect Biochem. Mol. Biol. 2015, 59, 30–40. [Google Scholar] [CrossRef] [PubMed]
- Ewart, G.D.; Howells, A.J. ABC Transporters Involved in Transport of Eye Pigment Precursors in Drosophila melanogaster. In ABC Transporters: Biochemical, Cellular, and Molecular Aspects, Methods Enzymol; Ambudkar, S.V., Gottesman, M.M., Eds.; Academic Press: San Diego, CA, USA, 1998; Volume 292, pp. 213–224. [Google Scholar]
- Tarr, P.T.; Tarling, E.J.; Bojanic, D.D.; Edwards, P.A.; Baldan, A. Emerging New Paradigms for ABCG Transporters. Biochim. Biophys. Acta 2009, 1791, 584–593. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kerr, I.D.; Haider, A.J.; Gelissen, I.C. The ABCG Family of Membrane-Associated Transporters: You Don’t Have to Be Big to Be Mighty. Br. J. Pharmacol. 2011, 164, 1767–1779. [Google Scholar] [CrossRef] [Green Version]
- Frambach, S.J.C.M.; de Haas, R.; Smeitink, J.A.M.; Rongen, G.A.; Russel, F.G.M.; Schirris, T.J.J. Brothers in Arms: ABCA1-and ABCG1-Mediated Cholesterol Efflux as Promising Targets in Cardiovascular Disease Treatment. Pharmacol. Rev. 2020, 72, 152–190. [Google Scholar] [CrossRef]
- Dermauw, W.; Van Leeuwen, T. The ABC Gene Family in Arthropods: Comparative Genomics and Role in Insecticide Transport and Resistance. Insect Biochem. Mol. Biol. 2014, 45, 89–110. [Google Scholar] [CrossRef]
- Zhou, J.; He, W.Y.; Wang, W.N.; Yang, C.W.; Wang, L.; Xin, Y.; Wu, J.; Cai, D.X.; Liu, Y.; Wang, A.L. Molecular Cloning and Characterization of an ATP-Binding Cassette (ABC) Transmembrane Transporter from the White Shrimp Litopenaeus vannamei. Comp. Biochem. Physiol. 2009, 150, 450–458. [Google Scholar] [CrossRef]
- Chang, Z.Q.; Li, J.; Liu, P.; Kuo, M.M.C.; He, Y.Y.; Chen, P.; Li, J.T. cDNA Cloning and Expression Profile Analysis of an ATP-Binding Cassette Transporter in the Hepatopancreas and Intestine of Shrimp Fenneropenaeus Chinensis. Aquaculture 2012, 356, 250–255. [Google Scholar] [CrossRef]
- Zhai, Q.Q.; Li, J.; Chang, Z.Q. cDNA Cloning, Characterization and Expression Analysis of ATP-Binding Cassette Transmembrane Transporter in Exopalaemon carinicauda. Aquac. Res. 2017, 48, 4143–4154. [Google Scholar] [CrossRef]
- Guo, Z.; Kang, S.; Sun, D.; Gong, L.; Zhou, J.; Qin, J.; Guo, L.; Zhu, L.; Bai, Y.; Ye, F.; et al. MAPK-Dependent Hormonal Signaling Plasticity Contributes to Overcoming Bacillus thuringiensis Toxin Action in an Insect Host. Nat. Commun. 2020, 11, 3003. [Google Scholar] [CrossRef]
- Signor, S.A.; Nuzhdin, S.V. The Evolution of Gene Expression in cis and Trans. Trends Genet. 2018, 34, 532–544. [Google Scholar] [CrossRef]
- Daborn, P.J.; Yen, J.L.; Bogwitz, M.R.; Le Goff, G.; Feil, E.; Jeffers, S.; Tijet, N.; Perry, T.; Heckel, D.; Batterham, P.; et al. A Single P450 Allele Associated with Insecticide Resistance in Drosophila. Science 2002, 297, 2253–2356. [Google Scholar] [CrossRef]
- Schmidt, J.M.; Good, R.T.; Appleton, B.; Sherrard, J.; Raymant, G.C.; Bogwitz, M.R.; Martin, J.; Daborn, P.J.; Goddard, M.E.; Batterham, P.; et al. Copy Number Variation and Transposable Elements Feature in Recent, Ongoing Adaptation at the Cyp6g1 Locus. PLoS Genet. 2010, 6, e1000998. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Itokawa, K.; Komagata, O.; Kasai, S.; Tomita, T. A Single Nucleotide Change in a Core Promoter Is Involved in the Progressive Overexpression of the Duplicated CYP9M10 Haplotype Lineage in Culex quinquefasciatus. Insect Biochem. Mol. Biol. 2015, 66, 96–102. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bass, C.; Zimmer, C.T.; Riveron, J.M.; Wilding, C.S.; Wondji, C.S.; Kaussmann, M.; Field, L.M.; Williamson, M.S.; Nauen, R. Gene Amplification and Microsatellite Polymorphism Underlie a Recent Insect Host Shift. Proc. Natl. Acad. Sci. USA 2013, 110, 19460–19465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pu, J.; Sun, H.; Wang, J.; Wu, M.; Wang, K.; Denholm, I.; Han, Z. Multiple cis-Acting Elements Involved in Up-Regulation of a Cytochrome P450 Gene Conferring Resistance to Deltamethrin in Smal Brown Planthopper, Laodelphax striatellus (Fallén). Insect Biochem. Mol. Biol. 2016, 78, 20–28. [Google Scholar] [CrossRef]
- Weedall, G.D.; Mugenzi, L.M.J.; Menze, B.D.; Tchouakui, M.; Ibrahim, S.S.; Amvongo-Adjia, N.; Irving, H.; Wondji, M.J.; Tchoupo, M.; Djouaka, R.; et al. A Cytochrome P450 Allele Confers Pyrethroid Resistance on a Major African Malaria Vector, Reducing Insecticide-Treated Bednet Efficacy. Sci. Transl. Med. 2019, 11, eaat7386. [Google Scholar] [CrossRef] [Green Version]
- Mugenzi, L.M.J.; Menze, B.D.; Tchouakui, M.; Wondji, M.J.; Irving, H.; Tchoupo, M.; Hearn, J.; Weedall, G.D.; Riveron, J.M.; Wondji, C.S. cis-Regulatory CYP6P9b P450 Variants Associated with Loss of Insecticide-Treated Bed Net Efficacy against Anopheles funestus. Nat. Commun. 2019, 10, 4652. [Google Scholar] [CrossRef] [Green Version]
- Hu, B.; Huang, H.; Hu, S.; Ren, M.; Wei, Q.; Tian, X.; Elzaki, M.E.A.; Bass, C.; Su, J.; Reddy Palli, S. Changes in Both Trans- and cis-Regulatory Elements Mediate Insecticide Resistance in a Lepidopteron Pest, Spodoptera exigua. PLoS Genet. 2021, 17, e1009403. [Google Scholar] [CrossRef]
- Wilding, C.S. Regulating Resistance: Cncc:Maf, Antioxidant Response Elements and the Overexpression of Detoxification Genes in Insecticide Resistance. Curr. Opin. Insect Sci. 2018, 27, 89–96. [Google Scholar] [CrossRef]
- Zhou, J.; Guo, Z.; Kang, S.; Qin, J.; Gong, L.; Sun, D.; Guo, L.; Zhu, L.; Bai, Y.; Zhang, Z.; et al. Reduced Expression of the P-Glycoprotein Gene Pxabcb1 Is Linked to Resistance to Bacillus thuringiensis Cry1Ac Toxin in Plutella xylostella (L.). Pest Manag. Sci. 2020, 76, 712–720. [Google Scholar] [CrossRef]
- Guo, Z.; Kang, S.; Chen, D.; Wu, Q.; Wang, S.; Xie, W.; Zhu, X.; Baxter, S.W.; Zhou, X.; Jurat-Fuentes, J.L.; et al. MAPK Signaling Pathway Alters Expression of Midgut ALP and ABCC Genes and Causes Resistance to Bacillus thuringiensis Cry1Ac Toxin in Diamondback Moth. PLoS Genet. 2015, 11, e1005124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, Z.; Sun, D.; Kang, S.; Zhou, J.; Gong, L.; Qin, J.; Guo, L.; Zhu, L.; Bai, Y.; Luo, L.; et al. CRISPR/Cas9-Mediated Knockout of Both the Pxabcc2 and Pxabcc3 Genes Confers High-Level Resistance to Bacillus thuringiensis Cry1Ac Toxin in the Diamondback Moth, Plutella xylostella (L.). Insect Biochem. Mol. Biol. 2019, 107, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Qin, J.; Zhou, X.; Zhang, Y. Insect Transcription Factors: A Landscape of Their Structures and Biological Functions in Drosophila and Beyond. Int. J. Mol. Sci. 2018, 19, 3691. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Whitehead, A.; Crawford, D.L. Variation Within and Among Species in Gene Expression: Raw Material for Evolution. Mol. Ecol. 2006, 15, 1197–1211. [Google Scholar] [CrossRef]
- Uusi-Heikkilä, S.; Sävilammi, T.; Leder, E.; Arlinghaus, R.; Primmer, C.R. Rapid, Broad-Scale Gene Expression Evolution in Experimentally Harvested Fish Populations. Mol. Ecol. 2017, 26, 3954–3967. [Google Scholar] [CrossRef] [PubMed]
- Stern, D.L.; Orgogozo, V. The Loci of Evolution: How Predictable Is Genetic Evolution? Evolution 2008, 62, 2155–2177. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rebeiz, M.; Williams, T.M. Using Drosophila Pigmentation Traits to Study the Mechanisms of cis-Regulatory Evolution. Curr. Opin. Insect Sci. 2017, 19, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Wittkopp, P.J.; Kalay, G. cis-Regulatory Elements: Molecular Mechanisms and Evolutionary Processes Underlying Divergence. Nat. Rev. Genet. 2011, 13, 59–69. [Google Scholar] [CrossRef] [PubMed]
- Akam, M. Hox and HOM: Homologous Gene Clusters in Insects and Vertebrates. Cell 1989, 57, 347–349. [Google Scholar] [CrossRef]
- Yu, M.; Zhan, J.; Zhang, H. HOX Family Transcription Factors: Related Signaling Pathways and Post-Translational Modifications in Cancer. Cell. Signal. 2020, 66, 109469. [Google Scholar] [CrossRef] [PubMed]
- Schneuwly, S.; Klemenz, R.; Gehring, W.J. Redesigning the Body Plan of Drosophila by Ectopic Expression of the Homoeotic Gene Antennapedia. Nature 1987, 325, 816–818. [Google Scholar] [CrossRef] [PubMed]
- Kimoto, M.; Tsubota, T.; Uchino, K.; Sezutsu, H.; Takiya, S. Hox Transcription Factor Antp Regulates Sericin-1 Gene Expression in the Terminal Differentiated Silk Gland of Bombyx mori. Dev. Biol. 2014, 386, 64–71. [Google Scholar] [CrossRef]
- Tsubota, T.; Tomita, S.; Uchino, K.; Kimoto, M.; Takiya, S.; Kajiwara, H.; Yamazaki, T.; Sezutsu, H. A Hox Gene, Antennapedia, Regulates Expression of Multiple Major Silk Protein Genes in the Silkworm Bombyx mori. J. Biol. Chem. 2016, 291, 7087–7096. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saenko, S.V.; Marialva, M.S.; Beldade, P. Involvement of the Conserved Hox Gene Antennapedia in the Development and Evolution of a Novel Trait. EvoDevo 2011, 2, 9. [Google Scholar] [CrossRef] [Green Version]
- Tomoyasu, Y.; Wheeler, S.R.; Denell, R.E. Ultrabithorax Is Required for Membranous Wing Identity in the Beetle Tribolium castaneum. Nature 2005, 433, 643–647. [Google Scholar] [CrossRef]
- Tomoyasu, Y. Ultrabithorax and the Evolution of Insect Forewing/Hindwing Differentiation. Curr. Opin. Insect Sci. 2017, 19, 8–15. [Google Scholar] [CrossRef] [Green Version]
- Sun, D.; Guo, Z.; Liu, Y.; Zhang, Y. Progress and Prospects of CRISPR/Cas Systems in Insects and Other Arthropods. Front. Physiol. 2017, 8, 608. [Google Scholar] [CrossRef]
- Wolter, F.; Schindele, P.; Puchta, H. Plant Breeding at the Speed of Light: The Power of CRISPR/Cas to Generate Directed Genetic Diversity at Multiple Sites. BMC Plant Biol. 2019, 19, 176. [Google Scholar] [CrossRef] [Green Version]
- Soyk, S.; Lemmon, Z.H.; Oved, M.; Fisher, J.; Liberatore, K.L.; Park, S.J.; Goren, A.; Jiang, K.; Ramos, A.; van der Knaap, E.; et al. Bypassing Negative Epistasis on Yield in Tomato Imposed by a Domestication Gene. Cell 2017, 169, 1142–1155. [Google Scholar] [CrossRef] [Green Version]
- Mohamad Ishak, N.S.; Nong, Q.D.; Matsuura, T.; Kato, Y.; Watanabe, H. Co-Option of the Bzip Transcription Factor Vrille as the Activator of Doublesex1 in Environmental Sex Determination of the Crustacean Daphnia magna. PLoS Genet. 2017, 13, e1006953. [Google Scholar] [CrossRef]
- Qin, J.; Guo, L.; Ye, F.; Kang, S.; Sun, D.; Zhu, L.; Bai, Y.; Cheng, Z.; Xu, L.; Ouyang, C.; et al. MAPK-Activated Transcription Factor PxJun Suppresses Pxabcb1 Expression and Confers Resistance to Bacillus thuringiensis Cry1Ac Toxin in Plutella xylostella (L.). Appl. Environ. Microbiol. 2021, 87, e00466-21. [Google Scholar] [CrossRef]
- Zhu, X.; Lei, Y.; Yang, Y.; Baxter, S.W.; Li, J.; Wu, Q.; Wang, S.; Xie, W.; Guo, Z.; Fu, W.; et al. Construction and Characterisation of Near-Isogenic Plutella xylostella (Lepidoptera: Plutellidae) Strains Resistant to Cry1Ac Toxin. Pest Manag. Sci. 2015, 71, 225–233. [Google Scholar] [CrossRef]
- Guo, Z.; Kang, S.; Zhu, X.; Wu, Q.; Wang, S.; Xie, W.; Zhang, Y. The Midgut Cadherin-Like Gene Is Not Associated with Resistance to Bacillus thuringiensis Toxin Cry1Ac in Plutella xylostella (L.). J. Invertebr. Pathol. 2015, 126, 21–30. [Google Scholar] [CrossRef]
- Guo, Z.; Gong, L.; Kang, S.; Zhou, J.; Sun, D.; Qin, J.; Guo, L.; Zhu, L.; Bai, Y.; Bravo, A.; et al. Comprehensive Analysis of Cry1Ac Protoxin Activation Mediated by Midgut Proteases in Susceptible and Resistant Plutella xylostella (L.). Pestic. Biochem. Physiol. 2020, 163, 23–30. [Google Scholar] [CrossRef]
- Xie, W.; Lei, Y.; Fu, W.; Yang, Z.; Zhu, X.; Guo, Z.; Wu, Q.; Wang, S.; Xu, B.; Zhou, X.; et al. Tissue-Specific Transcriptome Profiling of Plutella xylostella Third Instar Larval Midgut. Int. J. Biol. Sci. 2012, 8, 1142–1155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, Z.; Kang, S.; Zhu, X.; Xia, J.; Wu, Q.; Wang, S.; Xie, W.; Zhang, Y. The novel ABC Transporter ABCH1 Is a Potential Target for RNAi-Based Insect Pest Control and Resistance Management. Sci. Rep. 2015, 5, 13728. [Google Scholar] [CrossRef] [PubMed] [Green Version]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qin, J.; Ye, F.; Xu, L.; Zhou, X.; Crickmore, N.; Zhou, X.; Zhang, Y.; Guo, Z. A cis-Acting Mutation in the PxABCG1 Promoter Is Associated with Cry1Ac Resistance in Plutella xylostella (L.). Int. J. Mol. Sci. 2021, 22, 6106. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22116106
Qin J, Ye F, Xu L, Zhou X, Crickmore N, Zhou X, Zhang Y, Guo Z. A cis-Acting Mutation in the PxABCG1 Promoter Is Associated with Cry1Ac Resistance in Plutella xylostella (L.). International Journal of Molecular Sciences. 2021; 22(11):6106. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22116106
Chicago/Turabian StyleQin, Jianying, Fan Ye, Linzheng Xu, Xuguo Zhou, Neil Crickmore, Xiaomao Zhou, Youjun Zhang, and Zhaojiang Guo. 2021. "A cis-Acting Mutation in the PxABCG1 Promoter Is Associated with Cry1Ac Resistance in Plutella xylostella (L.)" International Journal of Molecular Sciences 22, no. 11: 6106. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22116106







