Epidemiological Characteristics of Staphylococcus Aureus in Raw Goat Milk in Shaanxi Province, China
Abstract
:1. Introduction
2. Results
2.1. Isolation and Identification of S. aureus
2.2. Antibiotic Resistance of S. Aureus Isolated from Raw Goat Milk
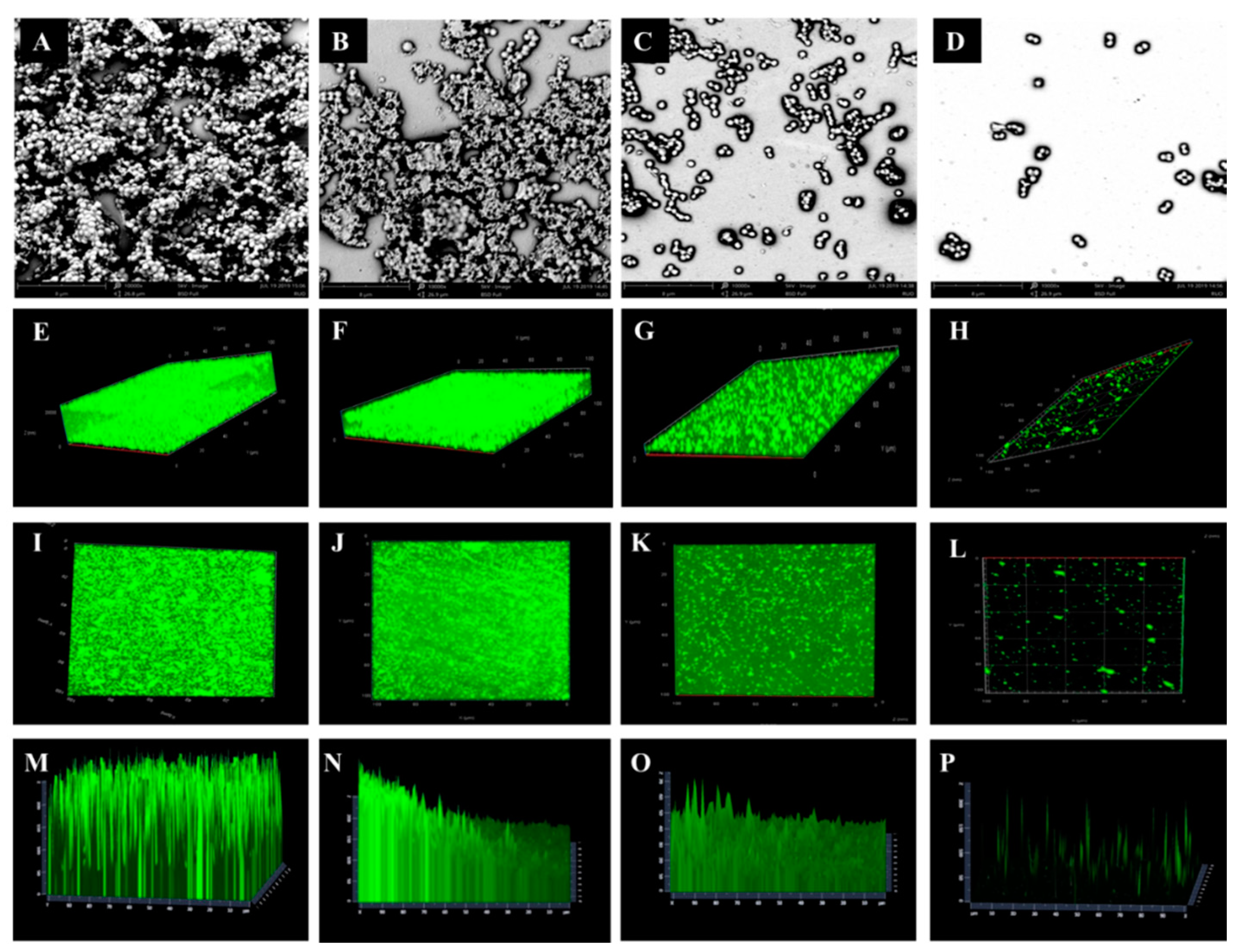
2.3. Biofilm Formation and Observation
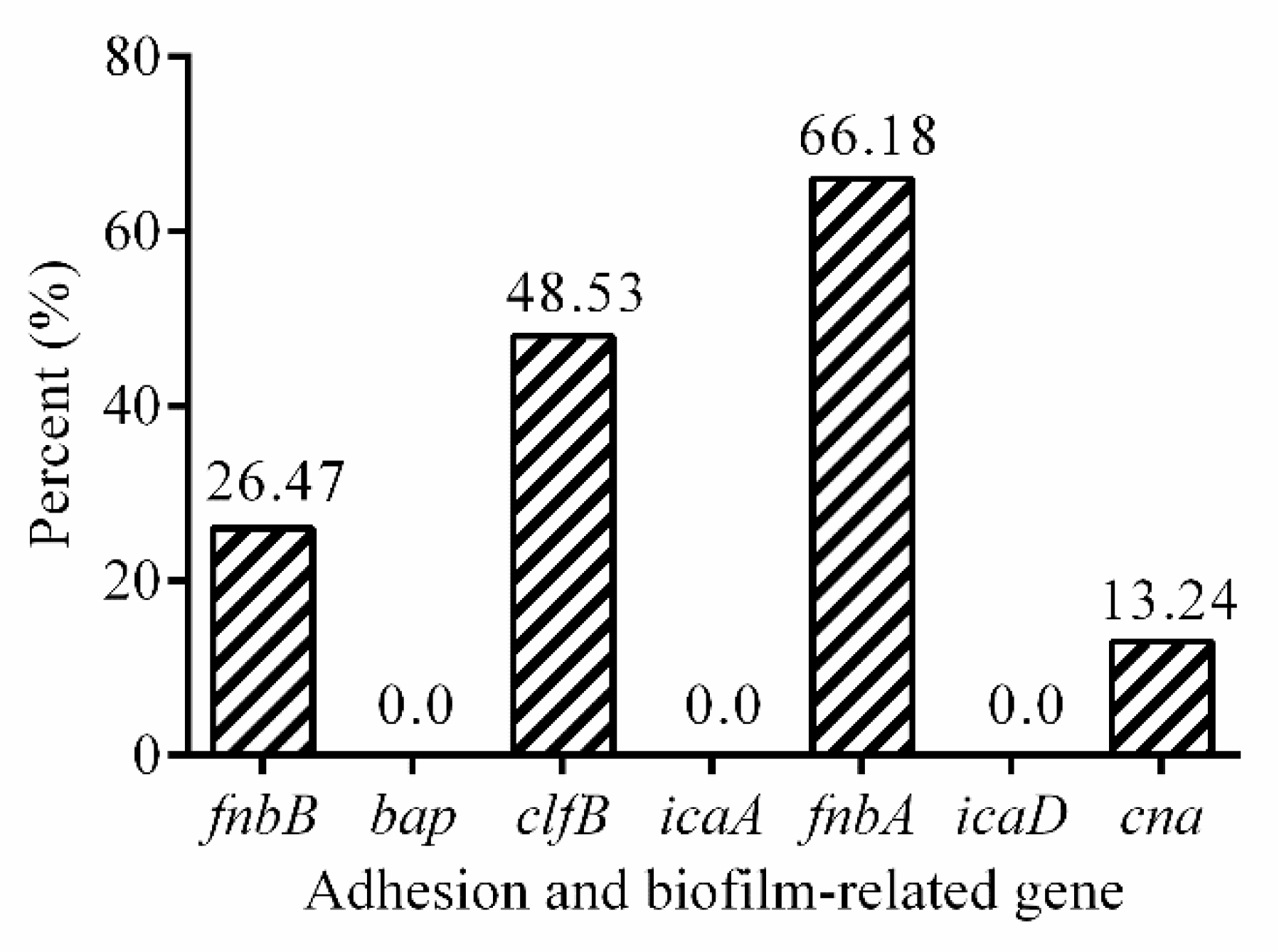
2.4. Adhesion and Biofilm-Related Genes
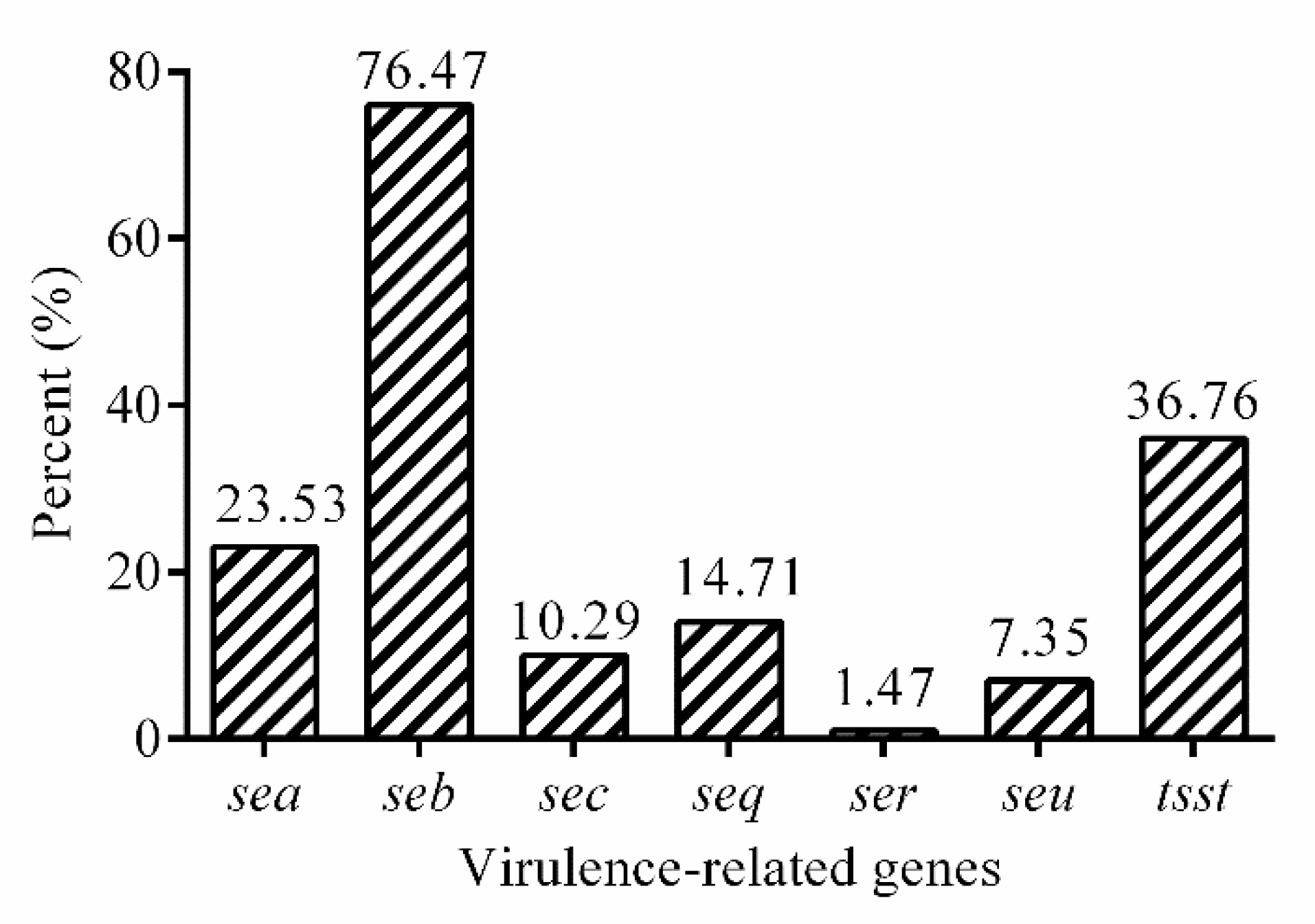
2.5. Virulence-Related Genes
2.6. The Association between Biofilm Formation and Biofilm-Related Genes
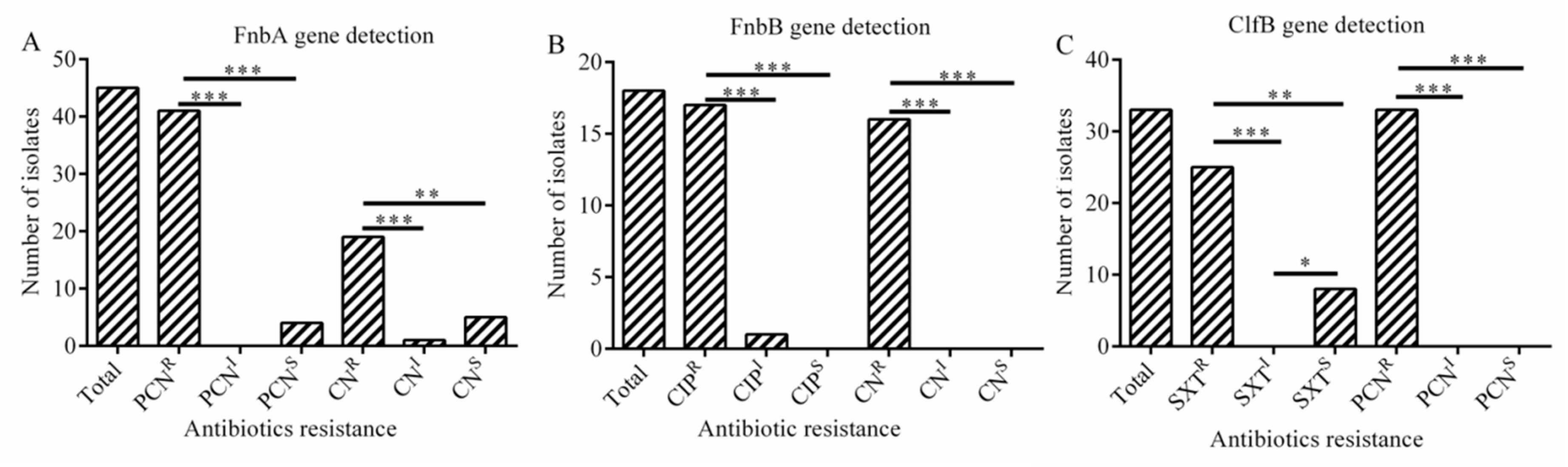
2.7. The Relationship between Antibiotic Resistance and Biofilm-Related Genes
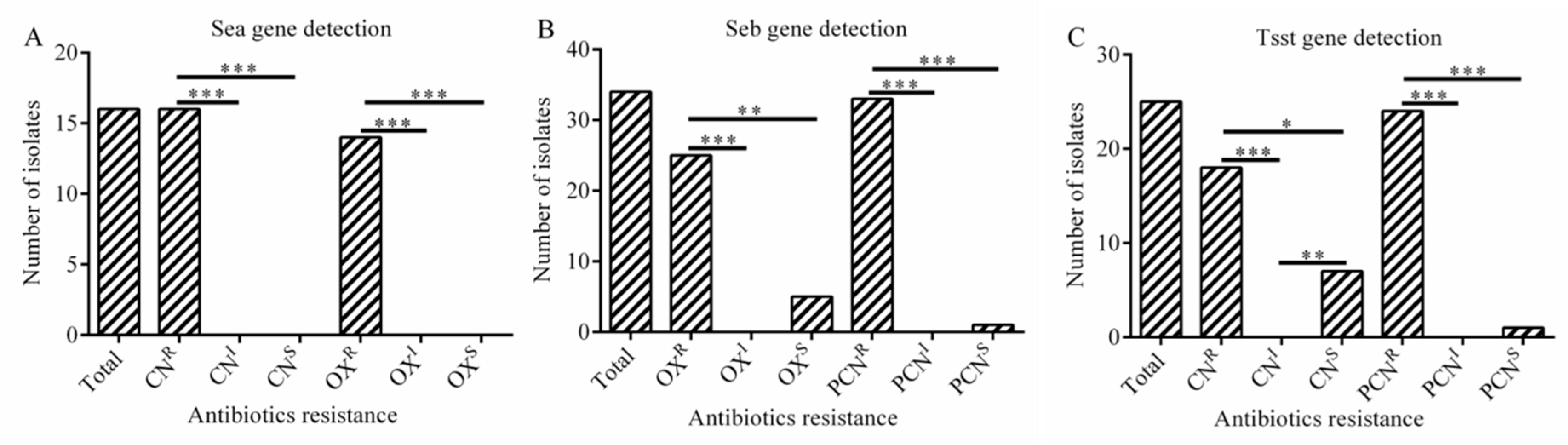
2.8. The Relationship between Antibiotic Resistance and Virulence Genes
3. Discussion
4. Materials and Methods
4.1. Isolation and Identification of Strains
4.2. Antimicrobial Susceptibility Testing
4.3. Biofilm Formation and Classification
4.4. Scanning Electron Microscope and Confocal Laser Scanning Microscopy Analysis of Biofilm
4.5. DNA Extraction and Gene Detection of PCR
4.6. Data Analysis and Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Wu, S.; Huang, J.; Wu, Q.; Zhang, J.; Zhang, F.; Yang, X.; Wu, H.; Zeng, H.; Chen, M.; Ding, Y.; et al. Staphylococcus aureus Isolated from Retail Meat and Meat Products in China: Incidence, Antibiotic Resistance and Genetic Diversity. Front. Microbiol. 2018, 9, 2767. [Google Scholar] [CrossRef] [PubMed]
- Abd-elghany, S.M.; Sallam, K.I.; Abd-elkhalek, A.; Tamure, T. Occurrence, genetic characterization and antimicrobial resistance of Salmonella isolated from chicken meat and giblets. Epidemiol. Infect. 2015, 143, 997–1003. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Sanchez, D.; Lopez-Cabo, M.; Saa-Ibusquiza, P.; Rodriguez-Herrera, J.J. Incidence and characterization of Staphylococcus aureus in fishery products marketed in Galicia (Northwest Spain). Int. J. Food Microbiol. 2012, 157, 286–296. [Google Scholar] [CrossRef] [PubMed]
- Migueis, S.; Santos, C.; Saraiva, C.; Esteves, A. Evaluation of ready to eat sashimi in northern Portugal restaurants. Food Control 2015, 47, 32–36. [Google Scholar] [CrossRef]
- Hammad, A.M.; Watanabe, W.; Fujii, T.; Shimamoto, T. Occurrence and characteristics of methicillin-resistant and susceptible Staphylococcus aureus and methicillin-resistant coagulase-negative staphylococci from Japanese retail ready-to-eat raw fish. Int. J. Food Microbiol. 2012, 156, 286–289. [Google Scholar] [CrossRef] [PubMed]
- Muscolino, D.; Giarratana, F.; Beninati, C.; Tornambene, A.; Panebianco, A.; Ziino, G. Hygienic-Sanitary Evaluation of Sushi and Sashimi Sold in Messina and Catania, Italy. Ital. J. Food Saf. 2014, 3, 1701. [Google Scholar] [CrossRef] [PubMed]
- Oliver, S.P.; Boor, K.J.; Murphy, S.C.; Murinda, S.E. Food safety hazards associated with consumption of raw milk. Foodborne Pathog. Dis. 2009, 6, 793–806. [Google Scholar] [CrossRef]
- Song, M.H.; Li, Q.Q.; Zhang, Y.; Song, J.X.; Shi, X.M.; Shi, C.L. Biofilm formation and antibiotic resistance pattern of dominant Staphylococcus aureus clonal lineages in China. J. Food Saf. 2017, 37, e12304. [Google Scholar] [CrossRef]
- Taj, Y.; Essa, F.; Aziz, F.; Kazmi, S.U. Study on biofilm-forming properties of clinical isolates of Staphylococcus aureus. J. Infect. Dev. Ctries. 2012, 6, 403–409. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilms: microbial life on surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef]
- Boles, B.R.; Horswill, A.R. Agr-mediated dispersal of Staphylococcus aureus biofilms. PLoS Pathog. 2008, 4, e1000052. [Google Scholar] [CrossRef] [PubMed]
- Otto, M. Staphylococcal Infections: Mechanisms of Biofilm Maturation and Detachment as Critical Determinants of Pathogenicity. Annu. Rev. Med. 2013, 64, 175–188. [Google Scholar] [CrossRef] [PubMed]
- Foster, T.J.; Geoghegan, J.A.; Ganesh, V.K.; Hook, M. Adhesion, invasion and evasion: the many functions of the surface proteins of Staphylococcus aureus. Nat. Rev. Microbiol. 2014, 12, 49–62. [Google Scholar] [CrossRef] [PubMed]
- Giaouris, D.; Papadopoulos, A.I.; Voutetakis, S.; Papadopoulou, S.; Seferlis, P. A power grand composite curves approach for analysis and adaptive operation of renewable energy smart grids. Clean Technol. Environ. 2015, 17, 1171–1193. [Google Scholar] [CrossRef]
- Laverty, G.; Gorman, S.P.; Gilmore, B.F. Biomolecular mechanisms of staphylococcal biofilm formation. Future Microbiol. 2013, 8, 509–524. [Google Scholar] [CrossRef] [Green Version]
- Gundogan, N.; Ataol, O. Biofilm, protease and lipase properties and antibiotic resistance profiles of staphylococci isolated from various foods. Afr. J. Microbiol. Res. 2013, 7, 3582–3588. [Google Scholar]
- De la Fuente-Nunez, C.; Reffuveille, F.; Fernandez, L.; Hancock, R.E.W. Bacterial biofilm development as a multicellular adaptation: Antibiotic resistance and new therapeutic strategies. Curr. Opin. Microbiol. 2013, 16, 580–589. [Google Scholar] [CrossRef]
- Spanu, V.; Spanu, C.; Virdis, S.; Cossu, F.; Scarano, C.; De Santis, E.P.L. Virulence factors and genetic variability of Staphylococcus aureus strains isolated from raw sheep’s milk cheese. Int. J. Food Microbiol. 2012, 153, 53–57. [Google Scholar] [CrossRef]
- Larkin, E.A.; Carman, R.J.; Krakauer, T.; Stiles, B.G. Staphylococcus aureus: The toxic presence of a pathogen extraordinaire. Curr. Med. Chem. 2009, 16, 4003–4019. [Google Scholar] [CrossRef]
- Tarekgne, E.K.; Skjerdal, T.; Skeie, S.; Rudi, K.; Porcellato, D.; Felix, B.; Narvhus, J.A. Enterotoxin Gene Profile and Molecular Characterization of Staphylococcus aureus Isolates from Bovine Bulk Milk and Milk Products of Tigray Region, Northern Ethiopia. J. Food Protect. 2016, 79, 1387–1395. [Google Scholar] [CrossRef]
- Shallcross, L.J.; Fragaszy, E.; Johnson, A.M.; Hayward, A.C. The role of the Panton-Valentine leucocidin toxin in staphylococcal disease: A systematic review and meta-analysis. Lancet Infect. Dis. 2013, 13, 43–54. [Google Scholar] [CrossRef]
- Jarraud, S.; Peyrat, M.A.; Lim, A.; Tristan, A.; Bes, M.; Mougel, C.; Etienne, J.; Vandenesch, F.; Bonneville, M.; Lina, G. Egc, a highly prevalent operon of enterotoxin gene, forms a putative nursery of superantigens in Staphylococcus aureus. J. Immunol. 2001, 166, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Miwa, N.; Kawamura, A.; Masuda, T.; Akiyama, M. An outbreak of food poisoning due to egg yolk reaction-negative Staphylococcus aureus. Int. J. Food Microbiol. 2001, 64, 361–366. [Google Scholar] [CrossRef]
- Fetsch, A.; Contzen, M.; Hartelt, K.; Kleiser, A.; Maassen, S.; Rau, J.; Kraushaar, B.; Layer, F.; Strommenger, B. Staphylococcus aureus food-poisoning outbreak associated with the consumption of ice-cream. Int. J. Food Microbiol. 2014, 187, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Skapetas, B.; Bampidis, V. Goat production in the World: Present situation and trends. Livest. Res. Rural. Dev. 2016, 28, 11. [Google Scholar]
- Xing, X.; Zhang, Y.; Wu, Q. Prevalence and characterization of Staphylococcus aureus isolated from goat milk powder processing plants. Food Control 2016, 59, 644–650. [Google Scholar] [CrossRef]
- Rong, D.; Wu, Q.; Xu, M.; Zhang, J.; Yu, S. Prevalence, Virulence Genes, Antimicrobial Susceptibility, and Genetic Diversity of Staphylococcus aureus from Retail Aquatic Products in China. Front. Microbiol. 2017, 8, 714. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, Y.; Cai, R.; Shi, L.; Li, C.; Yan, H. Prevalence of Enterotoxin Genes in Staphylococcus aureus Isolates from Pork Production. Foodborne Pathog. Dis. 2018, 15, 437–443. [Google Scholar] [CrossRef]
- Giacinti, G.; Carfora, V.; Caprioli, A.; Sagrafoli, D.; Marri, N.; Giangolini, G.; Amoruso, R.; Iurescia, M.; Stravino, F.; Dottarelli, S.; et al. Prevalence and characterization of methicillin-resistant Staphylococcus aureus carrying mecA or mecC and methicillin-susceptible Staphylococcus aureus in dairy sheep farms in central Italy. J. Dairy Sci. 2017, 100, 7857–7863. [Google Scholar] [CrossRef]
- Merz, A.; Stephen, R.; Johler, S. Staphylococcus aureus Isolates from Goat and Sheep Milk Seem to Be Closely Related and Differ from Isolates Detected from Bovine Milk. Front. Microbiol. 2016, 7, 319. [Google Scholar] [CrossRef]
- Spanu, V.; Scarano, C.; Virdis, S.; Melito, S.; Spanu, C.; De Santis, E.P. Population structure of Staphylococcus aureus isolated from bulk tank goat’s milk. Foodborne Pathog. Dis. 2013, 10, 310–315. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.; Wu, S.; Huang, J.; Wu, Q.; Zhang, F.; Zhang, J.; Wang, J.; Ding, Y.; Zhang, S.; Yang, X.; et al. Prevalence and Characterization of Staphylococcus aureus Isolated From Pasteurized Milk in China. Front. Microbiol. 2019, 10, 641. [Google Scholar] [CrossRef] [PubMed]
- Obaidat, M.M.; Bani Salman, A.E.; Roess, A.A. High prevalence and antimicrobial resistance of mecA Staphylococcus aureus in dairy cattle, sheep, and goat bulk tank milk in Jordan. Trop. Anim. Health Prod. 2018, 50, 405–412. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Li, S.; Meng, L.; Dong, L.; Zhao, S.; Lan, X.; Wang, J.; Zheng, N. Prevalence, antimicrobial susceptibility, and molecular characterization of Staphylococcus aureus isolated from dairy herds in northern China. J. Diary Sci. 2017, 100, 8796–8803. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Qu, W.; Barkema, H.W.; Nobrega, D.B.; Gao, J.; Liu, G.; De Buck, J.; Kastelic, J.P.; Sun, H.; Han, B. Antimicrobial resistance profiles of 5 common bovine mastitis pathogens in large Chinese dairy herds. J. Diary Sci. 2019, 102, 2416–2426. [Google Scholar] [CrossRef] [Green Version]
- Pekana, A.; Green, E. Antimicrobial Resistance Profiles of Staphylococcus aureus Isolated from Meat Carcasses and Bovine Milk in Abattoirs and Dairy Farms of the Eastern Cape, South Africa. Int. J. Environ. Res. Public Health 2018, 15, 2223. [Google Scholar] [CrossRef] [PubMed]
- Mistry, H.; Sharma, P.; Mahato, S.; Saravanan, R.; Kumar, P.A.; Bhandari, V. Prevalence and Characterization of Oxacillin Susceptible mecA-Positive Clinical Isolates of Staphylococcus aureus Causing Bovine Mastitis in India. PLoS ONE 2016, 11, e0162256. [Google Scholar] [CrossRef]
- Ünal, N.; Askar, I.; Macun, H.C.; Sakarya, F.; Altun, B.; Yldrm, M. Panton–Valentine leukocidin and some exotoxins of Staphylococcus aureus and antimicrobial susceptibility profiles of staphylococci isolated from milks of small ruminants. Trop. Anim. Health Prod. 2012, 44, 573–579. [Google Scholar] [CrossRef]
- Wang, W.; Lin, X.; Jiang, T.; Peng, Z.; Xu, J.; Yi, L.; Li, F.; Fanning, S.; Baloch, Z. Prevalence and Characterization of Staphylococcus aureus Cultured From Raw Milk Taken From Dairy Cows With Mastitis in Beijing, China. Front. Microbiol. 2018, 9, 1123. [Google Scholar] [CrossRef]
- Cheng, J.; Wang, Y.; Cao, Y.; Yan, W.; Niu, X.; Zhou, L.; Chen, J.; Sun, Y.; Li, C.; Zhang, X.; et al. The Distribution of 18 Enterotoxin and Enterotoxin-Like Genes in Staphylococcus aureus Strains from Different Sources in East China. Foodborne Pathog. Dis. 2016, 13, 171–176. [Google Scholar] [CrossRef]
- Akineden, O.; Hassan, A.A.; Schneider, E.; Usleber, E. Enterotoxigenic properties of Staphylococcus aureus isolated from goats’ milk cheese. Int. J. Food Microbiol. 2008, 124, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Lyra, D.G.; Sousa, F.G.; Borges, M.F.; Givisiez, P.E.; Queiroga, R.C.; Souza, E.L.; Gebreyes, W.A.; Oliveira, C.J. Enterotoxin-encoding genes in Staphylococcus spp. from bulk goat milk. Foodborne Pathog. Dis. 2013, 10, 126–130. [Google Scholar] [CrossRef] [PubMed]
- Pexara, A.; Solomakos, N.; Sergelidis, D.; Angelidis, A.S.; Govaris, A. Occurrence and antibiotic resistance of enterotoxigenic Staphylococcus aureus in raw ovine and caprine milk in Greece. Dairy Sci. Technol. 2016, 96, 345–357. [Google Scholar] [CrossRef]
- Maslankova, J.; Pilipcincova, I.; Tkacikova, L. Pheno- and Genotyping of Staphylococcus aureus Isolates of Sheep Origin. Acta Vet. Brno 2009, 78, 345–352. [Google Scholar] [CrossRef]
- McClure, J.A.; Conly, J.M.; Lau, V.; Elsayed, S.; Louie, T.; Hutchins, W.; Zhang, K. Novel multiplex PCR assay for detection of the staphylococcal virulence marker Panton-Valentine leukocidin genes and simultaneous discrimination of methicillin-susceptible from-resistant staphylococci. J. Clin. Microbiol. 2006, 44, 1141–1144. [Google Scholar] [CrossRef] [PubMed]
- Cavicchioli, V.Q.; Scatamburlo, T.M.; Yamazi, A.K.; Pieri, F.A.; Nero, L.A. Occurrence of Salmonella, Listeria monocytogenes, and enterotoxigenic Staphylococcus in goat milk from small and medium-sized farms located in Minas Gerais State, Brazil. J. Diary Sci. 2015, 98, 8386–8390. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bimanand, L.; Taherikalani, M.; Jalilian, F.A.; Sadeghifard, N.; Ghafourian, S.; Mahdavi, Z.; Mohamadi, S.; Sayehmiri, K.; Hematian, A.; Pakzad, I. Association between biofilm production, adhesion genes and drugs resistance in different SCCmec types of methicillin resistant Staphylococcus aureus strains isolated from several major hospitals of Iran. Iran J. Basic Med. Sci. 2018, 21, 400–403. [Google Scholar]
- Zmantar, T.; Makni, H.; Mahdouani, K.; Miladi, H.; Abdallah, J.F.; Bakhrouf, A.; Chaieb, K. Detection by PCR of adhesins genes and slime production in clinical Staphylococcus aureus. J. Basic Microbiol. 2010, 48, 308–314. [Google Scholar] [CrossRef]
- Uribe-Garcia, A.; Paniagua-Contreras, G.L.; Monroy-Perez, E.; Bustos-Martinez, J.; Hamdan-Partida, A.; Garzon, J.; Alanis, J.; Quezada, R.; Vaca-Paniagua, F.; Vaca, S. Frequency and expression of genes involved in adhesion and biofilm formation in Staphylococcus aureus strains isolated from periodontal lesions. J. Microbiol. Immunol. Infect. 2019, in press. [Google Scholar] [CrossRef]
- Pereyra, E.A.L.; Picech, F.; Renna, M.S.; Baravalle, C.; Andreotti, C.S.; Russi, R.; Calvinho, L.F.; Diez, C.; Dallard, B.E. Detection of Staphylococcus aureus adhesion and biofilm-producing genes and their expression during internalization in bovine mammary epithelial cells. Vet. Microbiol. 2016, 183, 69–77. [Google Scholar] [CrossRef]
- Klein, R.C.; Fabres-Klein, M.H.; Brito, M.A.V.P.; Fietto, L.G.; Ribon, A.D.O.B. Staphylococcus aureus of bovine origin: Genetic diversity, prevalence and the expression of adhesin-encoding genes. Vet. Microbiol. 2012, 160, 183–188. [Google Scholar] [CrossRef] [PubMed]
- Ciftci, A.; Findik, A.; Onuk, E.E.; Savasan, S. Detection of methicillin resistance and slime factor production of Staphylococcus aureus in bovine mastitis. Braz. J. Microbiol. 2009, 40, 254–261. [Google Scholar] [CrossRef] [PubMed]
- Gross, M.; Cramton, S.E.; Götz, F.; Peschel, A. Key Role of Teichoic Acid Net Charge in Staphylococcus aureus Colonization of Artificial Surfaces. Infect. Immun. 2001, 69, 3423–3426. [Google Scholar] [CrossRef] [PubMed]
| Antibiotic | Content (µg) | Diameter of Inhibition Zone (mm) | Susceptibility and Resistance (%) | ||||
|---|---|---|---|---|---|---|---|
| S | I | R | % | % | % | ||
| Trimethoprim-sulfamethoxazole | 25 | ≥21 | - | ≤20 | 45 (66.18) | 0 (0) | 23 (33.82) |
| Linezolid | 30 | ≥16 | 11–15 | ≤10 | 57 (83.82) | 1 (1.47) | 10 (14.71) |
| Gentamicin | 10 | ≥15 | 13–14 | ≤12 | 48 (70.59) | 1 (1.47) | 19 (27.94) |
| Ciprofloxacin | 5 | ≥21 | 16–20 | ≤15 | 43 (63.24) | 5 (7.35) | 20 (29.41) |
| Cefazolin | 30 | ≥18 | 15–17 | ≤14 | 50 (73.53) | 5 (7.35) | 13 (19.12) |
| Clindamycin | 2 | ≥21 | 15–20 | ≤14 | 40 (58.82) | 14 (20.59) | 14 (20.59) |
| Vancomycin | 30 | ≥15 | - | ≤16 | 68 (100.00) | 0 (0) | 0 (0) |
| Penicillin | 10 | ≥29 | - | ≤28 | 14 (20.59) | 0 (0) | 54 (79.41) |
| Oxacillin | 1 | ≥18 | - | ≤17 | 27 (39.71) | 0 (0) | 41 (60.29) |
| Piperacillin | 110 | ≥18 | - | ≤17 | 40 (58.82) | 0 (0) | 28 (41.18) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qian, W.; Shen, L.; Li, X.; Wang, T.; Liu, M.; Wang, W.; Fu, Y.; Zeng, Q. Epidemiological Characteristics of Staphylococcus Aureus in Raw Goat Milk in Shaanxi Province, China. Antibiotics 2019, 8, 141. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics8030141
Qian W, Shen L, Li X, Wang T, Liu M, Wang W, Fu Y, Zeng Q. Epidemiological Characteristics of Staphylococcus Aureus in Raw Goat Milk in Shaanxi Province, China. Antibiotics. 2019; 8(3):141. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics8030141
Chicago/Turabian StyleQian, Weidong, Lanfang Shen, Xinchen Li, Ting Wang, Miao Liu, Wenjing Wang, Yuting Fu, and Qiao Zeng. 2019. "Epidemiological Characteristics of Staphylococcus Aureus in Raw Goat Milk in Shaanxi Province, China" Antibiotics 8, no. 3: 141. https://0-doi-org.brum.beds.ac.uk/10.3390/antibiotics8030141





