The Function of the Histamine H4 Receptor in Inflammatory and Inflammation-Associated Diseases of the Gut
Abstract
:1. Introduction
2. H4R Basics
3. Histamine in the Intestine
4. Histamine Receptors in the Intestine
5. Histamine-Responsive Cells
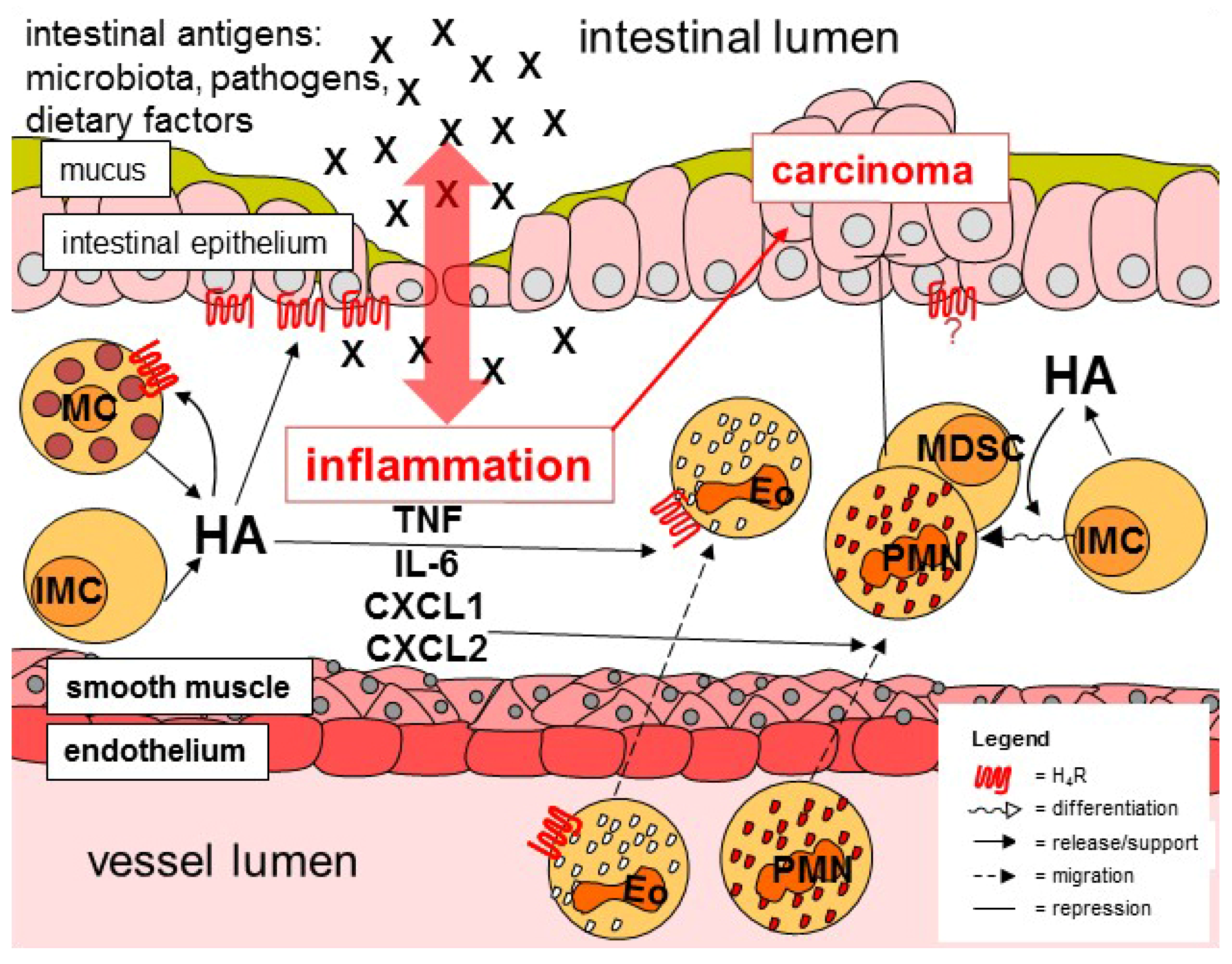
6. Histamine and Carcinogenesis
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Schayer, R.W. The metabolism of histamine in various species. Br. J. Pharmacol. Chemother. 1956, 11, 472–473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schayer, R.W.; Karjala, S.A. Ring N methylation; a major route of histamine metabolism. J. Biol. Chem. 1956, 221, 307–313. [Google Scholar] [CrossRef]
- Best, C.H.; McHenry, E.W. The inactivation of histamine. J. Physiol. 1930, 7, 349–372. [Google Scholar] [CrossRef]
- Riley, J.F.; West, G.B. The presence of histamine in tissue mast cells. J. Physiol. 1953, 120, 528–537. [Google Scholar] [CrossRef] [PubMed]
- Graham, H.T.; Lowry, O.H.; Wahl, N.; Priebat, M.K. Mast cells as sources of tissue histamine. J. Exp. Med. 1955, 102, 307–318. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hakanson, R.; Owman, C. Concomitant histochemical demonstration of histamine and catecholamines in enterochromaffin-like cells of gastric mucosa. Life Sci. 1967, 6, 759–766. [Google Scholar] [CrossRef]
- Kwiatkowski, H. Histamine in nervous tissue. J. Physiol. 1943, 102, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Fawcott, D.W. Cytological and pharmacological observations on the release of histamine by mast cells. J. Exp. Med. 1954, 100, 217–224. [Google Scholar] [CrossRef] [Green Version]
- Aborg, C.H.; Novotny, J.; Uvnäs, B. Ionic binding of histamine in mast cell granules. Acta Physiol. Scand. 1967, 69, 276–283. [Google Scholar] [CrossRef] [PubMed]
- Dvorak, A.M.; McLeod, R.S.; Ondcrdonk, A.; Monahan-Earley, R.A.; Cullen, J.B.; Antonioli, D.A.; Morgari, E.; Blair, J.E.; Estrella, P.; Cisneros, R.L.; et al. Ultrastructural evidence for piecemeal and anaphylactic degranulation of human gut mucosal mast cells in vivo. Int. Arch. Allergy Immunol. 1992, 99, 74–83. [Google Scholar] [CrossRef]
- Sahid, M.N.A.; Kiyoi, T. Mast cell activation markers for in vitro study. J. Immunoass. Immunochem. 2020, 41, 7798–7816. [Google Scholar] [CrossRef]
- Jutel, M.; Akdis, M.; Akdis, C.A. Histamine, histamine receptors and their role in immune pathology. Clin. Exp. Allergy 2009, 39, 1786–18000. [Google Scholar] [CrossRef]
- Deng, X.; Wu, X.; Yu, Z.; Arai, I.; Sasano, T.; Sugawara, S.; Endo, Y. Inductions of histidine decarboxylase in mouse tissues following systemic antigen challenge: Contributions made by mast cells, non-mast cells and IL-1. Int. Arch. Allergy Immunol. 2007, 144, 69–78. [Google Scholar] [CrossRef]
- Wu, X.; Yoshida, A.; Sasano, T.; Iwakura, Y.; Endo, Y. Histamine production via mast cell-independent induction of histidine decarboxylase in response to lipopolysaccharide and interleukin-1. Int. Immunopharmacol. 2004, 4, 516–520. [Google Scholar] [CrossRef] [PubMed]
- Hirasawa, N. Expression of histidine decarboxylase and its roles in inflammation. Int. J. Mol. Sci. 2019, 20, 376. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moriguchi, T.; Takai, J. Histamine and histidine decarboxylase: Immunomodulatory functions and regulatory mechanisms. Genes to Cells 2020, 25, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Calvielli Castelo Branco, A.C.; Seiti, F.; Yoshikawa, Y.; Pietrobon, A.J.; Sato, M.N. Role of histamine in modulating the immune response and inflammation. Mediat. Inflamm. 2018, 2018, e9524075. [Google Scholar]
- Seifert, R.; Strasser, A.; Schneider, E.H.; Neumann, D.; Dove, S.; Buschauer, A. Molecular and cellular analysis of human histamine receptor subtypes. Trends Pharmacol. Sci. 2013, 34, 33–58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tiligada, E.; Ennis, M. Histamine pharmacology: From Sir Henry Dale to the 21st century. Br. J. Pharmacol. 2020, 177, 469–489. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Panula, P. Histamine, histamine H3 receptor, and alcohol use disorder. Br. J. Pharmacol. 2020, 177, 634–641. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schirmer, B.; Lindemann, L.; Bittkau, K.S.; Isaev, R.; Bösche, D.; Juchem, M.; Seifert, R.; Neumann, D. Mouse colonic epithelial cells functionally express the histamine H4 receptor. J. Pharmacol. Exp. Ther. 2020, 373, 167–174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mehta, P.; Miszta, P.; Rzodkiewicz, P.; Michalak, O.; Krzeczyński, P.; Filipek, S. Enigmatic histamine receptor H4 for potential treatment of multiple inflammatory, autoimmune, and related diseases. Life 2020, 10, 50. [Google Scholar] [CrossRef]
- Wechsler, J.B.; Szabo, A.; Hsu, C.L.; Krier-Burris, R.A.; Schroeder, H.A.; Wang, M.Y.; Carter, R.G.; Velez, T.E.; Aguiniga, L.M.; Brown, J.B.; et al. Histamine drives severity of innate inflammation via histamine 4 receptor in murine experimental colitis. Mucosal Immunol. 2018, 11, 861–870. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sander, L.E.; Lorentz, A.; Sellge, G.; Coëffier, M.; Neipp, M.; Veres, T.; Frieling, T.; Meier, P.N.; Manns, M.P.; Bischoff, S.C. Selective expression of histamine receptors H1R, H2R, and H4R, but not H3R, in the human intestinal tract. Gut 2006, 55, 498–504. [Google Scholar] [CrossRef] [PubMed]
- Breunig, E.; Michel, K.; Zeller, F.; Seidl, S.; Weyhern, C.W.; Schemann, M. Histamine excites neurones in the human submucous plexus through activation of H1, H2, H3 and H4 receptors. J. Physiol. 2007, 583, 731–742. [Google Scholar] [CrossRef]
- Zhu, Y.; Michalovich, D.; Wu, H.; Tan, K.B.; Dytko, G.M.; Mannan, I.J.; Boyce, R.; Alston, J.; Tierney, L.A.; Li, X.; et al. Cloning, expression, and pharmacological characterization of a novel human histamine receptor. Mol. Pharmacol. 2001, 59, 434–441. [Google Scholar] [CrossRef] [PubMed]
- Gutzmer, R.; Mommert, S.; Gschwandtner, M.; Zwingmann, K.; Stark, H.; Werfel, T. The histamine H4 receptor is functionally expressed on TH2 cells. J. Allergy Clin. Immunol. 2009, 123, 619–625. [Google Scholar] [CrossRef] [PubMed]
- Schaper-Gerhardt, K.; Wohlert, M.; Mommert, S.; Kietzmann, M.; Werfel, T.; Gutzmer, R. Stimulation of histamine H4 receptors increases the production of IL-9 in Th9 polarized cells. Br. J. Pharmacol. 2020, 177, 614–622. [Google Scholar] [CrossRef]
- Mommert, S.; Gschwandtner, M.; Koether, B.; Gutzmer, R.; Werfel, T. Human memory Th17 cells express a functional histamine H4 receptor. Am. J. Pathol. 2012, 180, 177–185. [Google Scholar] [CrossRef] [PubMed]
- Cowden, J.M.; Yu, F.; Banie, H.; Farahani, M.; Ling, P.; Nguyen, S.; Riley, J.P.; Zhang, M.; Zhu, J.; Dunford, P.J.; et al. The histamine H4 receptor mediates inflammation and Th17 responses in preclinical models of arthritis. Ann. Rheum. Dis. 2014, 73, 600–608. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, S.H.; Hur, M.S.; Kim, M.J.; Kim, B.M.; Kim, K.W.; Kim, H.R.; Choe, Y.B.; Ahn, K.J.; Lee, Y.W. Preliminary study of histamine H4 receptor expressed on human CD4+ T cells and its immunomodulatory potency in the IL-17 pathway of psoriasis. J. Dermatol. Sci. 2017, 88, 29–35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morgan, R.K.; McAllister, B.; Cross, L.; Green, D.S.; Kornfeld, H.; Center, D.M.; Cruikshank, W.W. Histamine 4 receptor activation induces recruitment of FoxP3+ T cells and inhibits allergic asthma in a murine model. J. Immunol. 2007, 178, 8081–8089. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- del Rio, R.; Noubade, R.; Saligrama, N.; Wall, E.H.; Krementsov, D.N.; Poynter, M.E.; Zachary, J.F.; Thurmond, R.L.; Teuscher, C. Histamine H4 receptor optimizes T regulatory cell frequency and facilitates anti-inflammatory responses within the central nervous system. J. Immunol. 2012, 188, 541–547. [Google Scholar] [CrossRef] [Green Version]
- Truta-Feles, K.; Lagadari, M.; Lehmann, K.; Berod, L.; Cubillos, S.; Piehler, S.; Herouy, Y.; Barz, D.; Kamradt, T.; Maghazachi, A.; et al. Histamine modulates γδ-T lymphocyte migration and cytotoxicity, via Gi and Gs protein-coupled signalling pathways. Br. J. Pharmacol. 2010, 161, 1291–1300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gantner, F.; Sakai, K.; Tusche, M.W.; Cruikshank, W.W.; Center, D.M.; Bacon, K.B. Histamine H4 and H2 receptors control histamine-induced interleukin-16 release from human CD8+ T cells. J. Pharmacol. Exp. Ther. 2002, 303, 300–307. [Google Scholar] [CrossRef] [PubMed]
- Glatzer, F.; Mommert, S.; Köther, B.; Gschwandtner, M.; Stark, H.; Werfel, T.; Gutzmer, R. Histamine downregulates the Th1-associated chemokine IP-10 in monocytes and myeloid dendritic cells. Int. Arch. Allergy Immunol. 2014, 163, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Gutzmer, R.; Diestel, C.; Mommert, S.; Köther, B.; Stark, H.; Wittmann, M.; Werfel, T. Histamine H4 receptor stimulation suppresses IL-12p70 production and mediates chemotaxis in human monocyte-derived dendritic cells. J. Immunol. 2005, 174, 5224–5232. [Google Scholar] [CrossRef]
- Ling, P.; Ngo, K.; Nguyen, S.; Thurmond, R.L.; Edwards, J.P.; Karlsson, L.; Fung-Leung, W.P. Histamine H4 receptor mediates eosinophil chemotaxis with cell shape change and adhesion molecule upregulation. Br. J. Pharmacol. 2004, 142, 161–171. [Google Scholar] [CrossRef] [Green Version]
- Dunford, P.J.; O’Donnell, N.; Riley, J.P.; Williams, K.N.; Karlsson, L.; Thurmond, R.L. The histamine H4 receptor mediates allergic airway inflammation by regulating the activation of CD4+ T cells. J. Immunol. 2006, 176, 7062–7070. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hartwig, C.; Munder, A.; Glage, S.; Wedekind, D.; Schenk, H.; Seifert, R.; Neumann, D. The histamine H4 receptor (H4R) regulates eosinophilic inflammation in ovalbumin-induced experimental allergic asthma in mice. Eur. J. Immunol. 2015, 45, 1129–1140. [Google Scholar] [CrossRef]
- Lundberg, K.; Broos, S.; Greiff, L.; Borrebaeck, C.A.K.; Lindstedt, M. Histamine H4 receptor antagonism inhibits allergen-specific T-cell responses mediated by human dendritic cells. Eur. J. Pharmacol. 2011, 651, 197–204. [Google Scholar] [CrossRef]
- Gschwandtner, M.; Mommert, S.; Köther, B.; Werfel, T.; Gutzmer, R. The histamine H4 receptor is highly expressed on plasmacytoid dendritic cells in psoriasis and histamine regulates their cytokine production and migration. J. Investig. Dermatol. 2011, 131, 1668–1676. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Damaj, B.B.; Becerra, C.B.; Esber, H.J.; Wen, Y.; Maghazachi, A.A. Functional expression of H4 histamine receptor in human natural killer cells, monocytes, and dendritic cells. J. Immunol. 2007, 179, 7907–7915. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dijkstra, D.; Stark, H.; Chazot, P.L.; Shenton, F.C.; Leurs, R.; Werfel, T.; Gutzmer, R. Human inflammatory dendritic epidermal cells express a functional histamine H4 receptor. J. Investig. Dermatol. 2008, 128, 1696–1703. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bäumer, W.; Wendorff, S.; Gutzmer, R.; Werfel, T.; Dijkstra, D.; Chazot, P.; Stark, H.; Kietzmann, M. Histamine H4 receptors modulate dendritic cell migration through skin—Immunomodulatory role of histamine. Allergy 2008, 63, 1387–1394. [Google Scholar] [CrossRef] [PubMed]
- Simon, T.; Jelinek, I.; Apponyi, G.; László, V.; Rajnavölgyi, É.; Falus, A. Expression and function of histamine H4 receptor in mouse splenic dendritic cells. Inflamm. Res. 2010, 59 (Suppl. S2), S201–S203. [Google Scholar] [CrossRef]
- Gschwandtner, M.; Schäkel, K.; Werfel, T.; Gutzmer, R. Histamine H4 receptor activation on human slan-dendritic cells down-regulates their pro-inflammatory capacity. Immunology 2011, 132, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Leite-de-Moraes, M.C.; Diem, S.; Michel, M.-L.; Ohtsu, H.; Thurmond, R.L.; Schneider, E.; Dy, M. Cutting Edge: Histamine receptor H4 activation positively regulates in vivo IL-4 and IFN-γ production by invariant NKT cells. J. Immunol. 2009, 182, 1233–1236. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hofstra, C.L.; Desai, P.J.; Thurmond, R.L.; Fung-Leung, W.P. Histamine H4 receptor mediates chemotaxis and calcium mobilization of mast cells. J. Pharmacol. Exp. Ther. 2003, 305, 1212–1221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jemima, E.A.; Prema, A.; Thangam, E.B. Functional characterization of histamine H4 receptor on human mast cells. Mol. Immunol. 2014, 62, 19–28. [Google Scholar] [CrossRef]
- Kay, L.J.; Suvarna, S.K.; Peachell, P.T. Histamine H4 receptor mediates chemotaxis of human lung mast cells. Eur. J. Pharmacol. 2018, 837, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Honda, T.; Nishio, Y.; Sakai, H.; Asagiri, M.; Yoshimura, K.; Inui, M.; Kuramasu, A. Calcium/calmodulin-dependent regulation of Rac GTPases and Akt in histamine-induced chemotaxis of mast cells. Cell. Signal. 2021, 83, 1099733. [Google Scholar] [CrossRef] [PubMed]
- Lippert, U.; Artuc, M.; Grützkau, A.; Babina, M.; Guhl, S.; Haase, I.; Blaschke, V.; Zachmann, K.; Knosalla, M.; Middel, P.; et al. Human skin mast cells express H2 and H4, but not H3 receptors. J. Investig. Dermatol. 2004, 123, 116–123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Godot, V.; Arock, M.; Garcia, G.; Capel, F.; Flys, C.; Dy, M.; Emilie, D.; Humbert, M. H4 histamine receptor mediates optimal migration of mast cell precursors to CXCL12. J. Allergy Clin. Immunol. 2007, 120, 827–834. [Google Scholar] [CrossRef] [PubMed]
- Desai, P.; Thurmond, R.L. Histamine H4 receptor activation enhances LPS-induced IL-6 production in mast cells via ERK and PI3K activation. Eur. J. Immunol. 2011, 41, 1764–1773. [Google Scholar] [CrossRef] [PubMed]
- Mirzahosseini, A.; Dalmadi, B.; Csutora, P. Histamine receptor H4 regulates mast cell degranulation and IgE induced FcεRI upregulation in murine bone marrow-derived mast cells. Cell. Immunol. 2013, 283, 38–44. [Google Scholar] [CrossRef]
- Aldi, S.; Takano, K.I.; Tomita, K.; Koda, K.; Chan, N.Y.K.; Marino, A.; Salazar-Rodriguez, M.; Thurmond, R.L.; Levi, R. Histamine H4-receptors inhibit mast cell renin release in ischemia/reperfusion via protein kinase Cε-dependent aldehyde dehydrogenase type-2 activation. J. Pharmacol. Exp. Ther. 2014, 349, 508–517. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ebenezer, A.J.; Arunachalam, P.; Elden, B.T. H4R activation utilizes distinct signaling pathways for the production of RANTES and IL-13 in human mast cells. J. Recept. Signal Transduct. 2017, 37, 133–140. [Google Scholar] [CrossRef]
- Kuramasu, A.; Wakabayashi, M.; Inui, M.; Yanai, K. Distinct roles of small GTPases RAC1 and RAC2 in histamine H4 receptor–mediated chemotaxis of mast cells. J. Pharmacol. Exp. Ther. 2018, 367, 9–19. [Google Scholar] [CrossRef]
- Oda, T.; Morikawa, N.; Saito, Y.; Masuho, Y.; Matsumoto, S. Molecular cloning and characterization of a novel type of histamine receptor preferentially expressed in leukocytes. J. Biol. Chem. 2000, 275, 36781–36786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dib, K.; Perecko, T.; Jenei, V.; McFarlane, C.; Comer, D.; Brown, V.; Katebe, M.; Scheithauer, T.; Thurmond, R.L.; Chazot, P.L.; et al. The histamine H4 receptor is a potent inhibitor of adhesion-dependent degranulation in human neutrophils. J. Leukoc. Biol. 2014, 96, 411–418. [Google Scholar] [CrossRef] [Green Version]
- Dijkstra, D.; Leurs, R.; Chazot, P.; Shenton, F.C.; Stark, H.; Werfel, T.; Gutzmer, R. Histamine downregulates monocyte CCL2 production through the histamine H4 receptor. J. Allergy Clin. Immunol. 2007, 120, 300–307. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Wang, J.; Ye, X.Y.; Cheng, J.; Huang, C.Z.; Li, L.Y.; Li, T.Y.; Li, C.W. Histamine H4 receptor regulates IL-6 and INF-γ secretion in native monocytes from healthy subjects and patients with allergic rhinitis. Clin. Transl. Allergy 2019, 9, 49. [Google Scholar] [CrossRef] [Green Version]
- Czerner, C.P.; Klos, A.; Seifert, R.; Neumann, D. Histamine induces chemotaxis and phagocytosis in murine bone marrow-derived macrophages and RAW 264.7 macrophage-like cells via histamine H4-receptor. Inflamm. Res. 2014, 63, 239–247. [Google Scholar] [CrossRef]
- Mommert, S.; Ratz, L.; Stark, H.; Gutzmer, R.; Werfel, T. The histamine H4 receptor modulates the differentiation process of human monocyte-derived M1 macrophages and the release of CCL4/MIP-1β from fully differentiated M1 macrophages. Inflamm. Res. 2018, 67, 503–513. [Google Scholar] [CrossRef]
- Mommert, S.; Hüer, M.; Schaper-Gerhardt, K.; Gutzmer, R.; Werfel, T. Histamine up-regulates oncostatin M expression in human M1 macrophages. Br. J. Pharmacol. 2020, 177, 600–613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mommert, S.; Aslan, D.; Ratz, L.; Stark, H.; Gutzmer, R.; Werfel, T. The anaphylatoxin C3a receptor expression on human M2 macrophages is down-regulated by stimulating the histamine H4 receptor and the IL-4 receptor. J. Innate Immun. 2018, 10, 349–362. [Google Scholar] [CrossRef] [PubMed]
- Schneider, E.; Rolli-Derkinderen, M.; Arock, M.; Dy, M. Trends in histamine research: New functions during immune responses and hematopoiesis. Trends Immunol. 2002, 23, 255–263. [Google Scholar] [CrossRef]
- Mommert, S.; Kleiner, S.; Gehring, M.; Eiz-Vesper, B.; Stark, H.; Gutzmer, R.; Werfel, T.; Raap, U. Human basophil chemotaxis and activation are regulated via the histamine H4 receptor. Allergy 2016, 71, 1264–1273. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Ma, X.; Jiang, X.; Wilson, S.J.; Hofstra, C.L.; Blevitt, J.; Pyati, J.; Li, X.; Chai, W.; Carruthers, N.; et al. Cloning and pharmacological characterization of a fourth histamine receptor (H4) expressed in bone marrow. Mol. Pharmacol. 2001, 59, 420–426. [Google Scholar] [CrossRef] [PubMed]
- O’Reilly, M.; Alpert, R.; Jenkinson, S.; Gladue, R.P.; Foo, S.; Trim, S.; Peter, B.; Trevethick, M.; Fidock, M. Identification of a histamine H4 receptor on human eosinophils-role in eosinophil chemotaxis. J. Recept. Signal Transduct. Res. 2002, 22, 431–448. [Google Scholar] [CrossRef]
- Buckland, K.F.; Williams, T.J.; Conroy, D.M. Histamine induces cytoskeletal changes in human eosinophils via the H4 receptor. Br. J. Pharmacol. 2003, 140, 1117–1127. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reher, T.M.; Neumann, D.; Buschauer, A.; Seifert, R. Incomplete activation of human eosinophils via the histamine H4-receptor: Evidence for ligand-specific receptor conformations. Biochem. Pharmacol. 2012, 84, 192–203. [Google Scholar] [CrossRef] [PubMed]
- Schaper-Gerhardt, K.; Köther, B.; Wolff, L.; Kabatas, A.; Gehring, M.; Nikolouli, E.; Mommert, S.; Werfel, T.; Gutzmer, R. The H4R is highly expressed on eosinophils from AD patients and IL-4 upregulates expression and function via the JAK/STAT pathway. Allergy 2021, 76, 1261–1264. [Google Scholar] [CrossRef]
- Grosicki, M.; Adami, M.; Micheloni, C.; Głuch-Lutwin, M.; Siwek, A.; Latacz, G.; Łażewska, D.; Więcek, M.; Reiner-Link, D.; Stark, H.; et al. Eosinophils adhesion assay as a tool for phenotypic drug screening—The pharmacology of 1,3,5–triazine and 1H-indole like derivatives against the human histamine H4 receptor. Eur. J. Pharmacol. 2021, 890, 173611. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, T.; Kato, Y.; Hieshima, K.; Nagakubo, D.; Kunori, Y.; Fujisawa, T.; Yoshie, O. Liver-expressed chemokine/CC chemokine ligand 16 attracts eosinophils by interacting with histamine H4 receptor. J. Immunol. 2004, 173, 2078–2083. [Google Scholar] [CrossRef] [PubMed]
- Barnard, R.; Barnard, A.; Salmon, G.; Liu, W.; Sreckovic, S. Histamine-induced actin polymerization in human eosinophils: An imaging approach for histamine H4 receptor. Cytometry A 2008, 73, 299–304. [Google Scholar] [CrossRef] [PubMed]
- Mommert, S.; Dittrich-Breiholz, O.; Stark, H.; Gutzmer, R.; Werfel, T. The histamine H4 receptor regulates chemokine production in human natural killer cells. Int. Arch. Allergy Immunol. 2015, 166, 225–230. [Google Scholar] [CrossRef]
- Ehling, S.; Roßbach, K.; Dunston, S.M.; Stark, H.; Bäumer, W. Allergic inflammation is augmented via histamine H4 receptor activation: The role of natural killer cells in vitro and in vivo. J. Dermatol. Sci. 2016, 83, 106–115. [Google Scholar] [CrossRef] [PubMed]
- Raible, D.G.; Lenahan, T.; Fayvilevich, Y.; Kosinski, R.; Schulman, E.S. Pharmacologic characterization of a novel histamine receptor on human eosinophils. Am. J. Respir. Crit. Care Med. 1994, 149, 1506–1511. [Google Scholar] [CrossRef]
- Morse, K.L.; Behan, J.; Laz, T.M.; West, R.E.; Greenfeder, S.A.; Anthes, J.C.; Umland, S.; Wan, Y.; Hipkin, R.W.; Gonsiorek, W.; et al. Cloning and characterization of a novel human histamine receptor. J. Pharmacol. Exp. Ther. 2001, 296, 1058–1066. [Google Scholar] [PubMed]
- Nakamura, T.; Itadani, H.; Hidaka, Y.; Ohta, M.; Tanaka, K. Molecular cloning and characterization of a new human histamine receptor, HH4R. Biochem. Biophys. Res. Commun. 2000, 279, 615–620. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.; Shapiro, D.A.; George, S.R.; Setola, V.; Lee, D.K.; Cheng, R.; Rauser, L.; Lee, S.P.; Lynch, K.R.; Roth, B.L.; et al. Discovery of a novel member of the histamine receptor family. Mol. Pharmacol. 2001, 59, 427–433. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Verweij, E.W.E.; Al Araaj, B.; Prabhata, W.R.; Prihandoko, R.; Nijmeijer, S.; Tobin, A.B.; Leurs, R.; Vischer, H.F. Differential role of serines and threonines in intracellular loop 3 and C-terminal tail of the histamine H4 receptor in β-arrestin and G protein-coupled receptor kinase interaction, internalization, and signaling. ACS Pharmacol. Transl. Sci. 2020, 3, 321–333. [Google Scholar] [CrossRef] [Green Version]
- Beermann, S.; Bernhardt, G.; Seifert, R.; Buschauer, A.; Neumann, D. Histamine H1- and H4-receptor signaling cooperatively regulate MAPK activation. Biochem. Pharmacol. 2015, 98, 432–439. [Google Scholar] [CrossRef]
- Beermann, S.; Vauth, M.; Hein, R.; Seifert, R.; Neumann, D. Distinct Signalling Pathways of Murine Histamine H1- and H4-receptors expressed at comparable levels in HEK293 cells. PLoS ONE 2014, 9, e107481. [Google Scholar] [CrossRef] [Green Version]
- Schneider, E.H.; Seifert, R. Pharmacological characterization of human histamine receptors and histamine receptor mutants in the Sf9 cell expression system. In Histamine and Histamine Receptors in Health and Disease; Handbook of Experimental Pharmacology; Hattori, Y., Seifert, R., Eds.; Springer: Cham, Switzerland, 2017; Volume 241, pp. 63–118. [Google Scholar]
- Rovedatti, L.; Kudo, T.; Biancheri, P.; Sarra, M.; Knowles, C.; Rampton, D.; Corazza, G.; Monteleone, G.; Di Sabatino, A.; Macdonald, T. Differential regulation of interleukin 17 and interferon gamma production in inflammatory bowel disease. Gut 2009, 58, 1629–1636. [Google Scholar] [CrossRef] [PubMed]
- Schirmer, B.; Rezniczek, T.; Seifert, R.; Neumann, D. Proinflammatory role of the histamine H4 receptor in dextrane sodium sulfate-induced acute colitis. Biochem. Pharmacol. 2015, 98, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Wunschel, E.J.; Schirmer, B.; Seifert, R.; Neumann, D. Lack of histamine H4-receptor expression aggravates TNBS-induced acute colitis symptoms in mice. Front. Pharmacol. 2017, 8, 642. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thurmond, R.L.; Desai, P.J.; Dunford, P.J.; Fung-Leung, W.-P.P.; Hofstra, C.L.; Jiang, W.; Nguyen, S.; Riley, J.P.; Sun, S.; Williams, K.N.; et al. A potent and selective histamine H4 receptor antagonist with anti-inflammatory properties. J. Pharmacol. Exp. Ther. 2004, 309, 404–413. [Google Scholar] [CrossRef] [Green Version]
- Rosethorne, E.M.; Charlton, S.J. Agonist-biased signaling at the histamine H4 receptor: JNJ7777120 recruits β-arrestin without activating G proteins. Mol. Pharmacol. 2011, 79, 749–757. [Google Scholar] [CrossRef] [Green Version]
- Seifert, R.; Schneider, E.H.; Dove, S.; Brunskole, I.; Neumann, D.; Strasser, A.; Buschauer, A. Paradoxical stimulatory effects of the “standard” histamine H4-receptor antagonist JNJ7777120: The H4 receptor joins the club of 7 transmembrane domain receptors exhibiting functional selectivity. Mol. Pharmacol. 2011, 79, 631–638. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wifling, D.; Löffel, K.; Nordemann, U.; Strasser, A.; Bernhardt, G.; Dove, S.; Seifert, R.; Buschauer, A. Molecular determinants for the high constitutive activity of the human histamine H4 receptor: Functional studies on orthologues and mutants. Br. J. Pharmacol. 2015, 172, 785–798. [Google Scholar] [CrossRef] [Green Version]
- Cowden, J.M.; Riley, J.P.; Ma, J.Y.; Thurmond, R.L.; Dunford, P.J. Histamine H4 receptor antagonism diminishes existing airway inflammation and dysfunction via modulation of Th2 cytokines. Respir. Res 2010, 11, 86. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kollmeier, A.; Francke, K.; Chen, B.; Dunford, P.J.; Greenspan, A.J.; Xia, Y.; Xu, X.L.; Zhou, B.; Thurmond, R. The H4 receptor antagonist, JNJ 39758979, is effective in reducing histamine-induced pruritus in a randomized clinical study in healthy subjects. J. Pharmacol. Exp. Ther. 2014, 350, 181–187. [Google Scholar] [CrossRef] [PubMed]
- Reher, T.M.; Brunskole, I.; Neumann, D.; Seifert, R. Evidence for ligand-specific conformations of the histamine H2 receptor in human eosinophils and neutrophils. Biochem. Pharmacol. 2012, 84, 1174–1185. [Google Scholar] [CrossRef] [PubMed]
- Xie, G.; Wang, F.; Peng, X.; Liang, Y.; Yang, H.; Li, L. Modulation of mast cell toll-like receptor 3 expression and cytokines release by histamine. Cell. Physiol. Biochem. 2018, 46, 2401–2411. [Google Scholar] [CrossRef] [PubMed]
- Podolsky, D.K. Inflammatory bowel disease. N. Engl. J. Med. 2002, 247, 417–429. [Google Scholar] [CrossRef] [PubMed]
- Baumgart, D.C.; Sandborn, W.J. Inflammatory bowel disease: Clinical aspects and established and evolving therapies. Lancet 2007, 369, 1641–1657. [Google Scholar] [CrossRef]
- Baumgart, D.C.; Carding, S.R. Inflammatory bowel disease: Cause and immunobiology. Lancet 2007, 369, 1627–1640. [Google Scholar] [CrossRef]
- Xavier, R.J.; Podolsky, D.K. Unravelling the pathogenesis of inflammatory bowel disease. Nature 2007, 448, 427–434. [Google Scholar] [CrossRef]
- Cobrin, G.M.; Abreu, M.T. Defects in mucosal immunity leading to Crohn’s disease. Immunol. Rev. 2005, 206, 277–295. [Google Scholar] [CrossRef] [PubMed]
- Elson, C.O.; Weaver, C.T. Experimental mouse models of inflammatory bowel disease: New insights into pathogenic mechanisms. In Inflammatory Bowel Disease: From Bench to Bedside; Springer: Berlin/Heidelberg, Germany, 2006; ISBN 0387258078. [Google Scholar]
- Pizarro, T.T.; Cominelli, F. Cytokine therapy for Crohn’s disease: Advances in translational research. Annu. Rev. Med. 2007, 58, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Eaden, J.A.; Abrams, K.R.; Mayberry, J.F. The risk of colorectal cancer in ulcerative colitis: A meta-analysis. Gut 2001, 48, 526–535. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rahier, J.F. Prevention and management of infectious complications in IBD. Dig. Dis. 2012, 30, 408–414. [Google Scholar] [CrossRef]
- Baenkler, H.W.; Lux, G.; Günthner, R.; Kohlhäufl, M.; Matek, W. Biopsy histamine in ulcerative colitis and Crohn’s disease. Hepatogastroenterology 1987, 34, 289–290. [Google Scholar]
- Raithel, M.; Matek, M.; Baenkler, H.W.; Jorde, W.; Hahn, E.G. Mucosal histamine content and histamine secretion in Crohn’s disease, ulcerative colitis and allergic enteropathy. Int. Arch. Allergy Immunol. 1995, 108, 127–133. [Google Scholar] [CrossRef]
- Bene, L.; Sápi, Z.; Bajtai, A.; Buzás, E.; Szentmihályi, A.; Arató, A.; Tulassay, Z.; Falus, A. Partial protection against dextran sodium sulphate induced colitis in histamine-deficient, histidine decarboxylase knockout mice. J. Pediatr. Gastroenterol. Nutr. 2004, 39, 171–176. [Google Scholar] [CrossRef] [PubMed]
- Keubler, L.M.; Buettner, M.; Häger, C.; Bleich, A. A multihit model: Colitis lessons from the interleukin-10-deficient mouse. Inflamm. Bowel Dis. 2015, 21, 1967–1975. [Google Scholar] [CrossRef]
- Buechler, G.; Wos-Oxley, M.L.; Smoczek, A.; Zschemisch, N.-H.; Neumann, D.; Pieper, D.H.; Hedrich, H.J.; Bleich, A. Strain-specific colitis susceptibility in IL10-deficient mice depends on complex gut microbiota-host interactions. Inflamm. Bowel Dis. 2012, 18, 943–954. [Google Scholar] [CrossRef]
- Barcik, W.; Pugin, B.; Westermann, P.; Perez, N.R.; Ferstl, R.; Wawrzyniak, M.; Smolinska, S.; Jutel, M.; Hessel, E.M.; Michalovich, D.; et al. Histamine-secreting microbes are increased in the gut of adult asthma patients. J. Allergy Clin. Immunol. 2016, 138, 1491–1494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beermann, S.; Seifert, R.; Neumann, D. Commercially available antibodies against human and murine histamine H4 receptor lack specificity. Naunyn Schmiedebergs Arch. Pharmacol. 2012, 385, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Gutzmer, R.; Werfel, T.; Bäumer, W.; Kietzmann, M.; Chazot, P.L.; Leurs, R. Well characterized antihistamine 4 receptor antibodies contribute to current knowledge of the expression and biology of the human and murine histamine 4 receptor. Naunyn Schmiedebergs Arch. Pharmacol. 2012, 385, 853–860. [Google Scholar] [CrossRef]
- Michel, M.C.; Wieland, T.; Tsujimoto, G. How reliable are G-protein-coupled receptor antibodies? Naunyn Schmiedebergs Arch. Pharmacol. 2009, 379, 385–388. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hashemipetroudi, S.H.; Nematzadeh, G.; Ahmadian, G.; Yamchi, A.; Kuhlmann, M. Assessment of DNA contamination in RNA samples based on ribosomal DNA. J. Vis. Exp. 2018, 22, 55451. [Google Scholar] [CrossRef] [Green Version]
- Maier, T.; Güell, M.; Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef] [Green Version]
- Strasser, A.; Wittmann, H.J.; Buschauer, A.; Schneider, E.H.; Seifert, R. Species-dependent activities of G-protein-coupled receptor ligands: Lessons from histamine receptor orthologs. Trends Pharmacol. Sci. 2013, 34, 13–32. [Google Scholar] [CrossRef]
- Sullivant, A.; Mackin, A.; Pharr, T.; Cooley, J.; Wills, R.; Archer, T. Identification of histamine receptors in the canine gastrointestinal tract. Vet. Immunol. Immunopathol. 2016, 182, 29–36. [Google Scholar] [CrossRef]
- Kim, H.; Dwyer, L.; Song, J.H.; Martin-Cano, F.E.; Bahney, J.; Peri, L.; Britton, F.C.; Sanders, K.M.; Koh, S.D. Identification of histamine receptors and effects of histamine on murine and simian colonic excitability. Neurogastroenterol. Motil. 2011, 23, 949–e409. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smolinska, S.; Groeger, D.; Perez, N.R.; Schiavi, E.; Ferstl, R.; Frei, R.; Konieczna, P.; Akdis, C.A.; Jutel, M.; OʼMahony, L. Histamine receptor 2 is required to suppress innate immune responses to bacterial ligands in patients with inflammatory bowel disease. Inflamm. Bowel Dis. 2016, 22, 1575–1586. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deiteren, A.; De Man, J.G.; Pelckmans, P.A.; De Winter, B.Y. Histamine H₄ receptors in the gastrointestinal tract. Br. J. Pharmacol. 2015, 172, 1165–1178. [Google Scholar] [CrossRef] [Green Version]
- Varga, C.; Horvath, K.; Berko, A.; Thurmond, R.L.; Dunford, P.J.; Whittle, B.J. Inhibitory effects of histamine H4 receptor antagonists on experimental colitis in the rat. Eur. J. Pharmacol. 2005, 522, 130–138. [Google Scholar] [CrossRef]
- Cianchi, F.; Cortesini, C.; Schiavone, N.; Perna, F.; Magnelli, L.; Fanti, E.; Bani, D.; Messerini, L.; Fabbroni, V.; Perigli, G.; et al. The role of cyclooxygenase-2 in mediating the effects of histamine on cell proliferation and vascular endothelial growth factor production in colorectal cancer. Clin. Cancer Res. 2005, 11, 6807–6815. [Google Scholar] [CrossRef] [Green Version]
- Boer, K.; Helinger, E.; Helinger, A.; Pocza, P.; Pos, Z.; Demeter, P.; Baranyai, Z.; Dede, K.; Darvas, Z.; Falus, A. Decreased expression of histamine H1 and H4 receptors suggests disturbance of local regulation in human colorectal tumours by histamine. Eur. J. Cell Biol. 2008, 87, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Lamas, D.J.M.; Carabajal, E.; Prestifillipo, J.P.; Rossi, L.; Elverdin, J.C.; Merani, S.; Bergoc, R.M.; Rivera, E.S.; Medina, V.A. Protection of radiation-unduced damage to the hematopoietic system, small intestine and salivary glands in rats by JNJ7777120 compound, histamine H4 ligand. PLoS ONE 2013, 8, e69106. [Google Scholar]
- Ji, Y.; Sakata, Y.; Li, X.; Zhang, C.; Yang, Q.; Xu, M.; Wollin, A.; Langhans, W.; Tso, P. Lymphatic diamine oxidase secretion stimulated by fat absorption is linked with histamine release. Am. J. Physiol. Gastrointest. Liver Physiol. 2013, 304, G732–G740. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deiteren, A.; De Man, J.G.; Ruyssers, N.E.; Moreels, T.G.; Pelckmans, P.A.; De Winter, B.Y. Histamine H4 and H1 receptors contribute to postinflammatory visceral hypersensitivity. Gut 2014, 63, 1873–1882. [Google Scholar] [CrossRef]
- Wang, M.; Han, J.; Domenico, J.; Shin, Y.S.; Jia, Y.; Gelfand, E.W. Combined blockade of the histamine H1 and H4 receptor suppresses peanut-induced intestinal anaphylaxis by regulating dendritic cell function. Allergy 2016, 71, 1561–1574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schirmer, B.; Rother, T.; Bruesch, I.; Bleich, A.; Werlein, C.; Jonigk, D.; Seifert, R.; Neumann, D. Genetic deficiency of the histamine H4-receptor reduces experimental colorectal carcinogenesis in mice. Cancers 2020, 12, 912. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thurmond, R.L.; Gelfand, E.W.; Dunford, P.J. The role of histamine H1 and H4 receptors in allergic inflammation: The search for new antihistamines. Nat. Rev. Drug Discov. 2008, 7, 41–53. [Google Scholar] [CrossRef]
- Rijnierse, A.; Nijkamp, F.P.; Kraneveld, A.D. Mast cells and nerves tickle in the tummy: Implications for inflammatory bowel disease and irritable bowel syndrome. Pharmacol. Ther. 2007, 116, 207–235. [Google Scholar] [CrossRef]
- van Hoboken, E.A.; Thijssen, A.Y.; Verhaaren, R.; van der Veek, P.P.; Prins, F.A.; Verspaget, H.W.; Masclee, A.A. Symptoms in patients with ulcerative colitis in remission are associated with visceral hypersensitivity and mast cell activity. Scand. J. Gastroenterol. 2011, 46, 981–987. [Google Scholar] [CrossRef] [PubMed]
- Karhausen, J.; Qing, M.; Gibson, A.; Moeser, A.J.; Griefingholt, H.; Hale, L.P.; Abraham, S.N.; MacKensen, G.B. Intestinal mast cells mediate gut injury and systemic inflammation in a rat model of deep hypothermic circulatory arrest. Crit. Care Med. 2013, 41, e200–e210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eissmann, M.F.; Dijkstra, C.; Jarnicki, A.; Phesse, T.; Brunnberg, J.; Poh, A.R.; Etemadi, N.; Tsantikos, E.; Thiem, S.; Huntington, N.D.; et al. IL-33-mediated mast cell activation promotes gastric cancer through macrophage mobilization. Nat. Commun. 2019, 10, 2735. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chin, K.W.; Barrett, K.E. Mast cells are not essential to inflammation in murine model of colitis. Dig. Dis. Sci. 1994, 39, 513–525. [Google Scholar] [CrossRef] [PubMed]
- Minocha, A.; Thomas, C.; Omar, R. Lack of crucial role of mast cells in pathogenesis of experimental colitis in mice. Dig. Dis. Sci. 1995, 40, 1757–1762. [Google Scholar] [CrossRef] [PubMed]
- Araki, Y.; Andoh, A.; Fujiyama, Y.; Bamba, T. Development of dextran sulphate sodium-induced experimental colitis is suppressed in genetically mast cell-deficient Ws/Ws rats. Clin. Exp. Immunol. 2000, 119, 264–269. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, S.; Tanimoto, A.; Arima, N.; Xu, H.; Murata, Y.; Hamada, T.; Makishima, K.; Sasaguri, Y. Effects of histamine and interleukin-4 synthesized in arterial intima on phagocytosis by monocytes/macrophages in relation to atherosclerosis. FEBS Lett. 2001, 505, 217–222. [Google Scholar] [CrossRef] [Green Version]
- Zwadlo-Klarwasser, G.; Vogts, M.; Hamann, W.; Belke, K.; Baron, J.; Schmutzler, W. Generation and subcellular distribution of histamine in human blood monocytes and monocyte subsets. Inflamm. Res. 1998, 74, 434–439. [Google Scholar] [CrossRef]
- Yang, X.D.; Ai, W.; Asfaha, S.; Bhagat, G.; Friedman, R.A.; Jin, G.; Park, H.; Shykind, B.; Diacovo, T.G.; Falus, A.; et al. Histamine deficiency promotes inflammation-associated carcinogenesis through reduced myeloid maturation and accumulation of CD11b+Ly6G+ immature myeloid cells. Nat. Med. 2011, 17, 87–95. [Google Scholar] [CrossRef] [Green Version]
- Amaral, M.M.; Davio, C.; Ceballos, A.; Salamone, G.; Cañones, C.; Geffner, J.; Vermeulen, M. Histamine improves antigen uptake and cross-presentation by dendritic cells. J. Immunol. 2007, 179, 3425–3433. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bischoff, S.C.; Gebhardt, T. Role of mast cells and eosinophils in neuroimmune interactions regulating mucosal inflammation in inflammatory bowel disease. Adv. Exp. Med. Biol. 2006, 579, 177–208. [Google Scholar] [PubMed]
- Masterson, J.C.; McNamee, E.N.; Fillon, S.A.; Hosford, L.; Harris, R.; Fernando, S.D.; Jedlicka, P.; Iwamoto, R.; Jacobsen, E.; Protheroe, C.; et al. Eosinophil-mediated signalling attenuates inflammatory responses in experimental colitis. Gut 2015, 64, 1236–1247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loktionov, A. Eosinophils in the gastrointestinal tract and their role in the pathogenesis of major colorectal disorders. World J. Gastroenterol. 2019, 24, 3503–3526. [Google Scholar] [CrossRef] [PubMed]
- Grosicki, M.; Wójcik, T.; Chlopicki, S.; Kieć-Kononowicz, K. In vitro study of histamine and histamine receptor ligands influence on the adhesion of purified human eosinophils to endothelium. Eur. J. Pharmacol. 2016, 777, 49–59. [Google Scholar] [CrossRef]
- Schirmer, B.; Bringmann, L.; Seifert, R.; Neumann, D. In vivo evidence for partial activation of eosinophils via the histamine H4 receptor: Adoptive transfer experiments using eosinophils from H4R−/− and H4R+/+ mice. Front. Immunol. 2018, 9, 2119. [Google Scholar] [CrossRef] [PubMed]
- Gay, J.; Kokkotou, E.; O’Brien, M.; Pothoulakis, C.; Karalis, K.P. Interleukin-6 Genetic ablation protects from trinitrobenzene sulfonic acid-induced colitis in mice. Neuroimmunomodulation 2006, 13, 114–121. [Google Scholar] [CrossRef] [PubMed]
- Huang, E.; Liu, R.; Lu, Z.; Liu, J.; Liu, X.; Zhang, D.; Chu, Y. NKT cells mediate the recruitment of neutrophils by stimulating epithelial chemokine secretion during colitis. Biochem. Biophys. Res. Commun. 2016, 474, 252–258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Werner, K.; Neumann, D.; Buschauer, A.; Seifert, R. No evidence for histamine H4 receptor in human monocytes. J. Pharmacol. Exp. Ther. 2014, 351, 519–526. [Google Scholar] [CrossRef] [Green Version]
- Takeshita, K.; Sakai, K.; Bacon, K.B.; Gantner, F. Critical role of histamine H4 receptor in leukotriene B4 production and mast cell-dependent neutrophil recruitment induced by zymosan in vivo. J. Pharmacol. Exp. Ther. 2003, 307, 1072–1078. [Google Scholar] [CrossRef] [Green Version]
- Peterson, L.W.; Artis, D. Intestinal epithelial cells: Regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 2014, 14, 141–153. [Google Scholar] [CrossRef]
- Odenwald, M.A.; Turner, J.R. The intestinal epithelial barrier: A therapeutic target? Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 9–21. [Google Scholar] [CrossRef]
- Landy, J.; Ronde, E.; English, N.; Clark, S.K.; Hart, A.L.; Knight, S.C.; Al-Hassi, H.O. Tight junctions in inflammatory bowel diseases and inflammatory bowel disease associated colorectal cancer. World J. Gastroenterol. 2016, 22, 3117–3126. [Google Scholar] [CrossRef] [PubMed]
- Buckley, A.; Turner, J.R. Cell biology of tight junction barrier regulation and mucosal disease. Cold Spring Harb. Perspect. Biol. 2018, 10, a029314. [Google Scholar] [CrossRef] [PubMed]
- Otegbeye, E.E.; Mitchem, J.B.; Park, H.; Chaudhuri, A.A.; Kim, H.; Mutch, M.G.; Ciorba, M.A. Immunity, immunotherapy, and rectal cancer: A clinical and translational science review. Transl. Res. 2021, 231, 124–138. [Google Scholar] [CrossRef]
- Kimiz-Gebologlu, I.; Gulce-Iz, S.; Biray-Avci, C. Monoclonal antibodies in cancer immunotherapy. Mol. Biol. Rep. 2018, 45, 2935–2940. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, S.; Imaeda, Y.; Umemoto, S.; Kobayashi, K.; Suzuki, H.; Okannoto, T. Cimetidine increases survival of colorectal cancer patients with high levels of sialyl Lewis-X and sialyl Lewis-A epitope expression on tumour cells. Br. J. Cancer 2002, 86, 161–167. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ali, A.H.; Hale, L.; Yalamanchili, B.; Ahmed, M.; Ahmed, M.; Zhou, R.; Wright, S.E. The effect of perioperative cimetidine administration on time to colorectal cancer recurrence. Am. J. Ther. 2018, 25, e405–e411. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.P.H.; Perry, C.M. Spotlight on histamine dihydrochloride in acute myeloid leukaemia. Drugs Aging 2011, 28, 325–329. [Google Scholar] [CrossRef]
- Medina, V.A.; Rivera, E.S. Histamine as a potential adjuvant to immuno and radiotherapy for cancer treatment: Discovering new functions for the oldest biogenic amine. Curr. Immunol. Rev. 2010, 6, 357–370. [Google Scholar] [CrossRef]
- Massari, N.A.; Nicoud, M.B.; Medina, V.A. Histamine receptors and cancer pharmacology: An update. Br. J. Pharmacol. 2020, 177, 516–538. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Medina, V.A.; Coruzzi, G.; Lamas, D.J.M.; Massari, N.; Adami, M.; Levi-Schaffer, F.; Ben-Zimra, M.; Schwelberger, H.; Rivera, E.S. Histamine in cancer. In Histamine H4 Receptor: A Novel Drug Target in Immunoregulation and Inflammation; De Gruyter Open Poland: Warsaw, Poland, 2013; ISBN 9788376560564. [Google Scholar]
- Vacante, M.; Ciuni, R.; Basile, F.; Biondi, A. Gut microbiota and colorectal cancer development: A closer look to the adenoma-carcinoma sequence. Biomedicines 2020, 8, 489. [Google Scholar] [CrossRef] [PubMed]
- Janney, A.; Powrie, F.; Mann, E.H. Host–microbiota maladaptation in colorectal cancer. Nature 2020, 585, 509–517. [Google Scholar] [CrossRef] [PubMed]
- Mathers, J.C.; Movahedi, M.; Macrae, F.; Mecklin, J.-P.; Moeslein, G.; Olschwang, S.; Eccles, D.; Evans, G.; Maher, E.R.; Bertario, L.; et al. Long-term effect of resistant starch on cancer risk in carriers of hereditary colorectal cancer: An analysis from the CAPP2 randomised controlled trial. Lancet Oncol. 2012, 13, 1242–1249. [Google Scholar] [CrossRef]
- Chan, A.T.; Lippman, S.M. Aspirin and colorectal cancer prevention in Lynch syndrome. Lancet 2011, 378, 2051–2052. [Google Scholar] [CrossRef]
- Masini, E.; Fabbroni, V.; Giannini, L.; Vannacci, A.; Messerini, L.; Perna, F.; Cortesini, C.; Cianchi, F. Histamine and histidine decarboxylase up-regulation in colorectal cancer: Correlation with tumor stage. Inflamm. Res. 2005, 54, S80–S81. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Ganesh, B.P.; Shi, Z.; Shah, R.R.; Fultz, R.; Major, A.; Venable, S.; Lugo, M.; Hoch, K.; Chen, X.; et al. Gut microbe–mediated suppression of inflammation-associated colon carcinogenesis by luminal histamine production. Am. J. Pathol. 2017, 187, 232–236. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Churchill, M.J.; Nagar, K.K.; Tailor, Y.H.; Chu, T.; Rush, B.S.; Jiang, Z.; Wang, E.B.C.; Renz, B.W.; Wang, H.; et al. IL-17 producing mast cells promote the expansion of myeloid-derived suppressor cells in a mouse allergy model of colorectal cancer. Oncotarget 2015, 6, 32966–32979. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, G.H.; Ding, J.Q.; Zhang, X.; Xu, W.M.; Lin, X.Q.; Huang, M.J.; Feng, J.; Wang, P.; Cai, W.K. Activation of histamine H4 receptor suppresses the proliferation and invasion of esophageal squamous cell carcinoma via both metabolism and non-metabolism signaling pathways. J. Mol. Med. 2018, 96, 951–964. [Google Scholar] [CrossRef]
- Tanaka, T.; Ishikawa, H. Mast cells and inflammation-associated colorectal carcinogenesis. Semin. Immunopathol. 2013, 35, 245–254. [Google Scholar] [CrossRef]
- Malfettone, A.; Silvestris, N.; Saponaro, C.; Ranieri, G.; Russo, A.; Caruso, S.; Popescu, O.; Simone, G.; Paradiso, A.; Mangia, A. High density of tryptase-positive mast cells in human colorectal cancer: A poor prognostic factor related to protease-activated receptor 2 expression. J. Cell. Mol. Med. 2013, 17, 1025–1037. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tanaka, T.; Kochi, T.; Shirakami, Y.; Mori, T.; Kurata, A.; Watanabe, N.; Moriwaki, H.; Shimizu, M. Cimetidine and clobenpropit attenuate inflammation-associated colorectal carcinogenesis in male ICR Mice. Cancers 2016, 8, 25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, C.; Xiong, Y.; Li, J.; Yang, Y.; Liu, L.; Wang, W.; Wang, L.; Li, M.; Fang, Z. Deletion and down-regulation of HRH4 gene in gastric carcinomas: A potential correlation with tumor progression. PLoS ONE 2012, 7, e31207. [Google Scholar] [CrossRef] [PubMed]
| Cell Type | Mouse | Human | Reference |
|---|---|---|---|
| CD4+ T cell | x | [26] | |
| Th2 | x | [27] | |
| Th9 | x | [28] | |
| Th17 | x | x | [29,30,31] |
| Treg | x | x | [32,33] |
| γδ T cell | x | [34] | |
| CD8+ T cell | x | [35] | |
| DC | x | x | [36,37,38,39,40,41,42,43,44,45,46,47] |
| NKT | x | [48] | |
| Mast cell | x | x | [26,49,50,51,52,53,54,55,56,57,58,59] |
| Neutrophil | x | x | [23,26,60,61] |
| Monocyte | x | [26,36,60,62,63] | |
| MΦ | x | [64] | |
| M1 | x | [65,66] | |
| M2 | x | [67] | |
| Basophil | x | [49,68,69] | |
| Eosinophil | x | x | [26,38,49,60,70,71,72,73,74,75,76,77] |
| NK cell | x | [43,78,79] |
| Main Findings | Species | Ref. |
|---|---|---|
| H2R and H4R are pro-proliferative and pro-angiogenic in HT29, Caco-2, and HCT116 colon cancer cell lines | human | [125] |
| H4R possesses pro-inflammatory role in TNBS-induced colitis in rats | rat | [124] |
| H1R, H2R, H4R are expressed in the human gastrointestinal tract; expression is altered in patients with gastrointestinal diseases | human | [24] |
| H1R, H2R, H3R, H4R activation excite human enteric neurons | human | [25] |
| H1R, H2R, and H4R are expressed in colon carcinoma and in adjacent normal mucosa; H1R and H4R expression is reduced in carcinoma compared to normal colon | human | [126] |
| H1R, H2R, and H4R are expressed in simian colon smooth muscle cells | mouse, monkey | [121] |
| H4R activity contributes to radiation-induced cytotoxic and genotoxic damages in small intestine | rat | [127] |
| H4R stimulates the release of DAO, contributing to histamine deamination during fat absorption | rat | [128] |
| H4R and H1R contribute to post-inflammatory visceral hypersensitivity | rat | [129] |
| H4R possesses a pro-inflammatory role in DSS-induced colitis in mice | mouse | [89] |
| H1R and H4R regulate the DC-CD4+ T-cell axis in peanut-induced intestinal allergic responses | mouse | [130] |
| H4R possesses an anti-inflammatory role in TNBS-induced colitis in mice | mouse | [90] |
| H4R possesses a pro-inflammatory role in experimental colitis in mice | mouse | [23] |
| H4R is functionally expressed on colon epithelial cells, affecting epithelial barrier integrity | mouse | [22] |
| H4R is involved in carcinogenesis of chemically-induced colitis-associated colorectal carcinoma | mouse | [131] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schirmer, B.; Neumann, D. The Function of the Histamine H4 Receptor in Inflammatory and Inflammation-Associated Diseases of the Gut. Int. J. Mol. Sci. 2021, 22, 6116. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22116116
Schirmer B, Neumann D. The Function of the Histamine H4 Receptor in Inflammatory and Inflammation-Associated Diseases of the Gut. International Journal of Molecular Sciences. 2021; 22(11):6116. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22116116
Chicago/Turabian StyleSchirmer, Bastian, and Detlef Neumann. 2021. "The Function of the Histamine H4 Receptor in Inflammatory and Inflammation-Associated Diseases of the Gut" International Journal of Molecular Sciences 22, no. 11: 6116. https://0-doi-org.brum.beds.ac.uk/10.3390/ijms22116116






