Synonymous Dinucleotide Usage: A Codon-Aware Metric for Quantifying Dinucleotide Representation in Viruses
Abstract
:1. Introduction
2. Materials and Methods
2.1. Synonymous Dinucleotide Usage (SDU)
- An SDU of 1 indicates that the representation of the dinucleotide of interest in the given frame position is equal to that expected under the null hypothesis of equal synonymous codon usage;
- an SDU of 0 indicates that the dinucleotide of interest is completely absent in the given frame position across the sequence;
- an SDU greater than 1 indicates that the dinucleotide of interest is over-represented in the given frame position, compared to the representation expected under the null hypothesis;
- an SDU between 0 and 1 indicates that the dinucleotide of interest is under-represented in the given frame position, compared to the representation expected under the null hypothesis.
2.2. Relative Synonymous Dinucleotide Usage (RSDU)
- An RSDU of 1 indicates that only the dinucleotide of interest is being used in the sequence, all the other synonymous dinucleotides being absent for the given position;
- an RSDU of 0, similar to the SDU, indicates that the dinucleotide of interest is completely absent in the given frame position.
2.3. Measuring the Null Distribution
2.4. DinuQ Python Package
2.5. Applying the SDU on Viral Sequences
3. Results
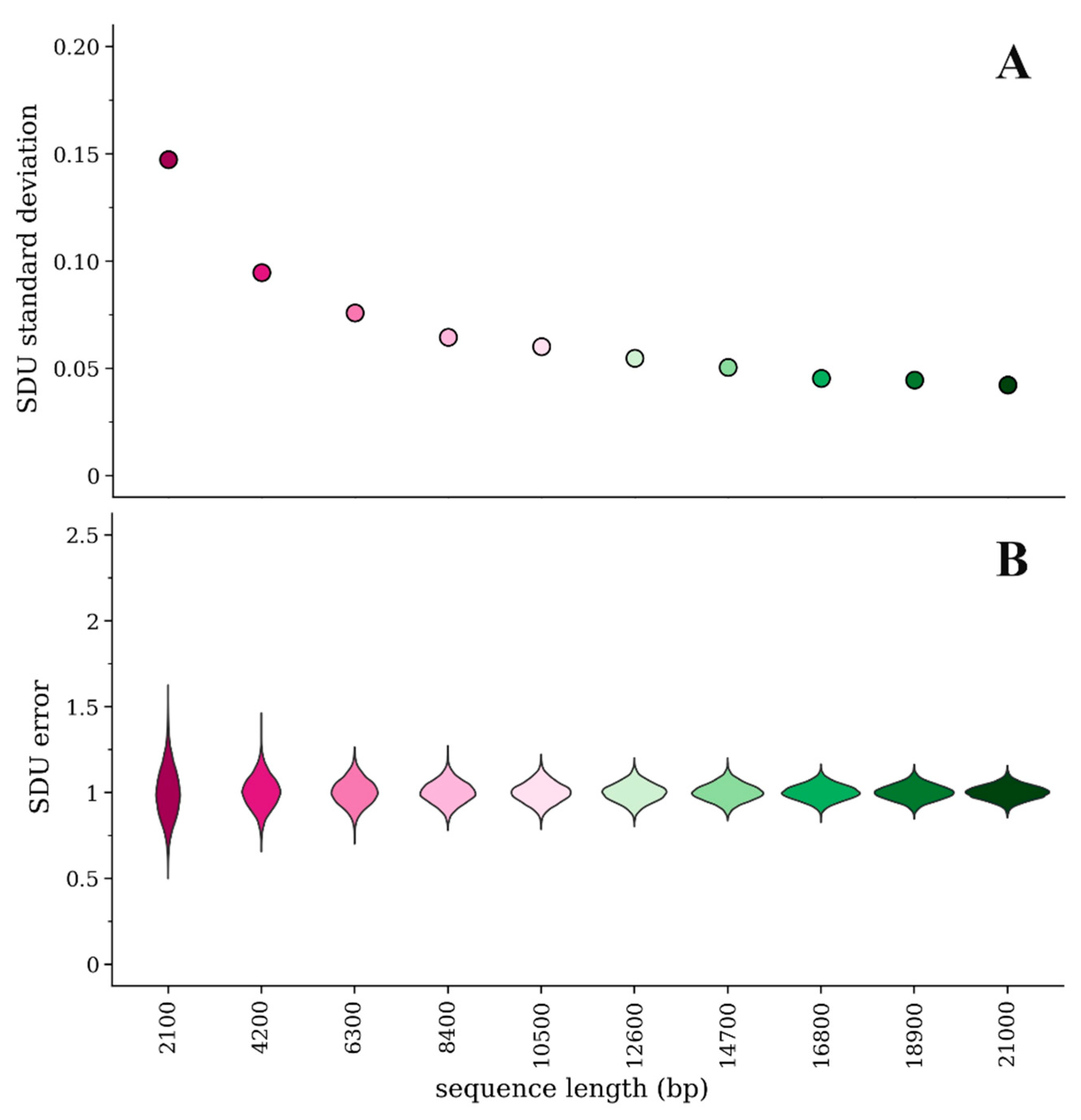
3.1. Effect of Sequence Length on SDU Error
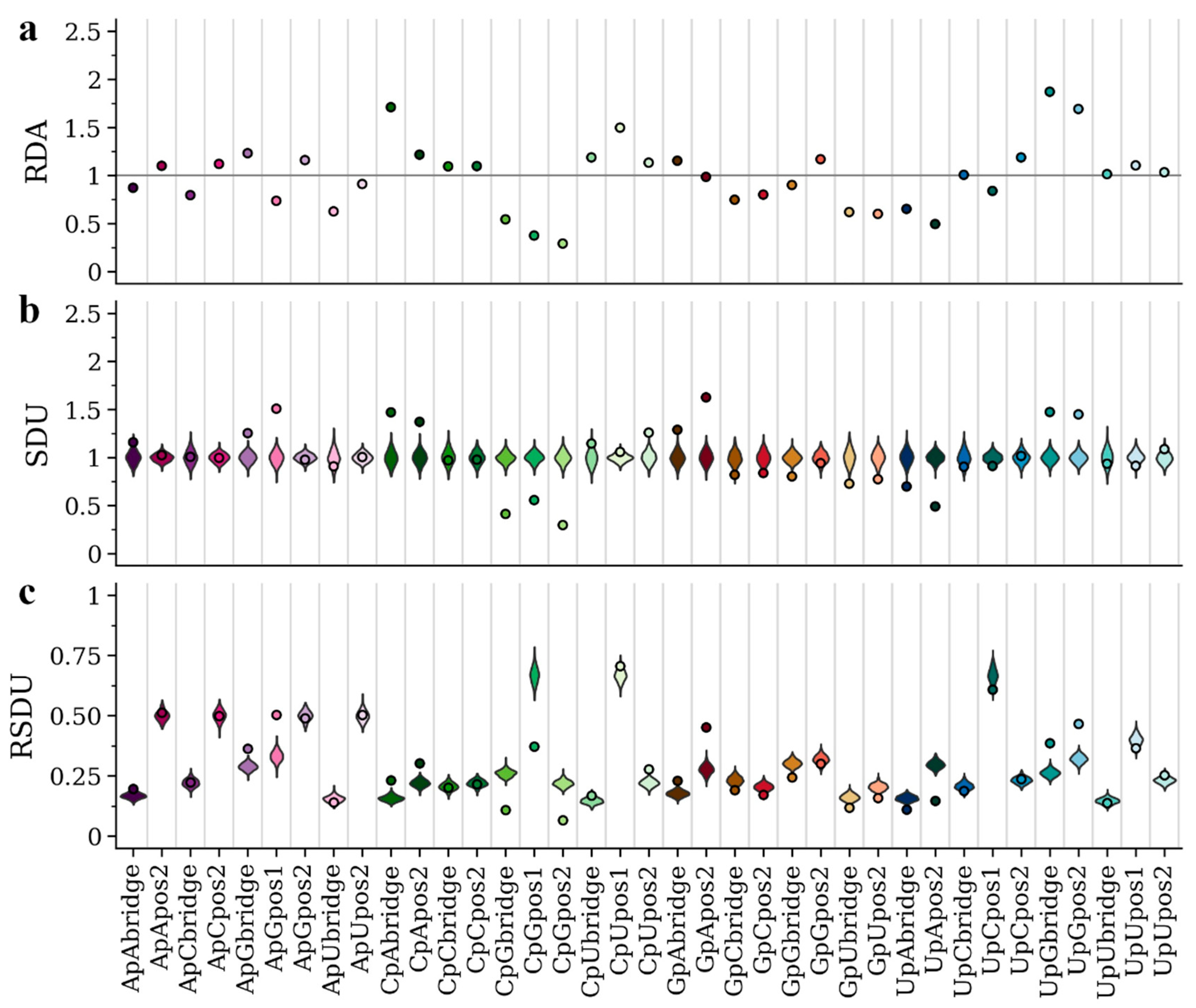
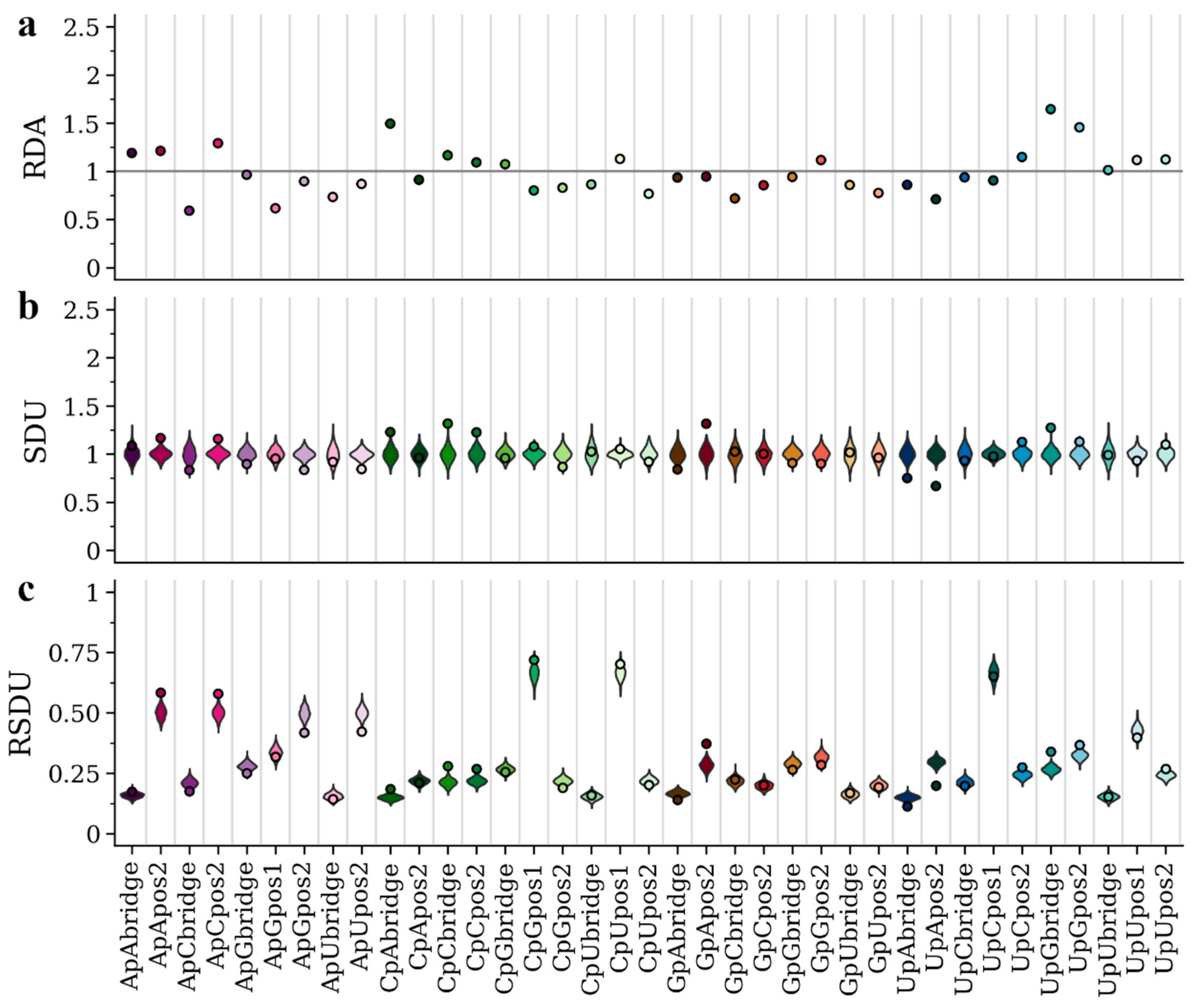
3.2. Metric Comparisons Using an Insect- and a Vertebrate-Specific Flavivirus
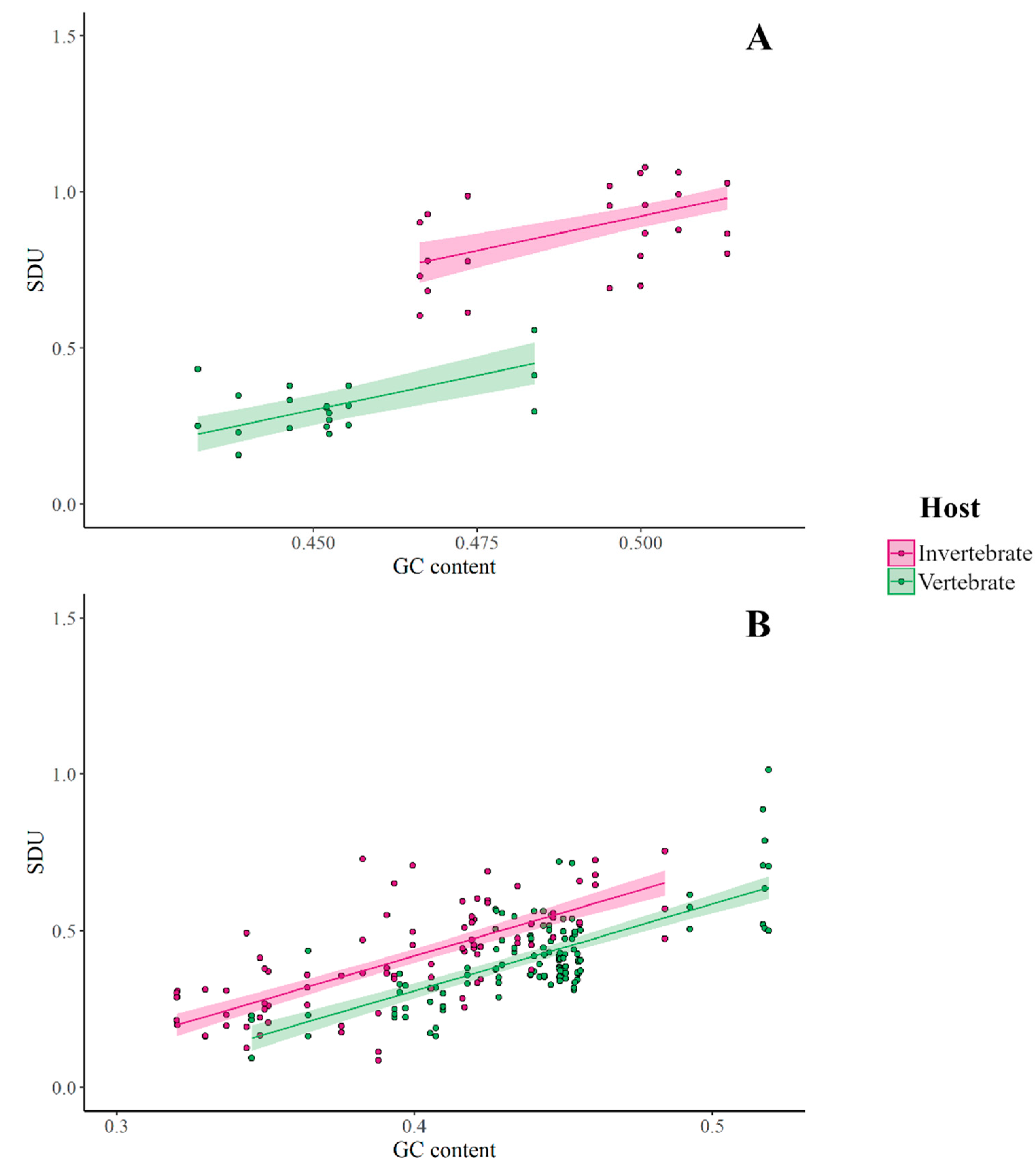
3.3. SDU Shows Consistent CpG Differences between Insect- and Vertebrate-Specific Viruses
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Computer Code and Software
References
- Beutler, E.; Gelbart, T.; Han, J.H.; Koziol, J.A.; Beutler, B. Evolution of the genome and the genetic code: Selection at the dinucleotide level by methylation and polyribonucleotide cleavage. Proc. Natl. Acad. Sci. USA 1989, 86, 192–196. [Google Scholar] [CrossRef] [Green Version]
- Karlin, S.; Doerfler, W.; Cardon, L.R. Why is CpG suppressed in the genomes of virtually all small eukaryotic viruses but not in those of large eukaryotic viruses? J. Virol. 1994, 68, 2889–2897. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, X.; Virk, N.; Chen, W.; Ji, S.; Ji, S.; Sun, Y.; Wu, X. CpG Usage in RNA Viruses: Data and Hypotheses. PLoS ONE 2013, 8, e74109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bird, A. DNA methylation and the frequency of CpG in animal DNA. Nucleic Acids Res. 1980, 8, 1499–1504. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.N.; Krawczak, M. Cytosine methylation and the fate of CpG dinucleotides in vertebrate genomes. Qual. Life Res. 1989, 83, 181–188. [Google Scholar] [CrossRef]
- Shaw, G.; Kamen, R. A conserved AU sequence from the 3’ untranslated region of GM-CSF mRNA mediates selective mRNA degradation. Cell 1986, 46, 659–667. [Google Scholar] [CrossRef]
- Duan, J.; Antezana, M. Mammalian Mutation Pressure, Synonymous Codon Choice, and mRNA Degradation. J. Mol. Evol. 2003, 57, 694–701. [Google Scholar] [CrossRef]
- Simmonds, P.; Xia, W.; Baillie, J.K.; McKinnon, K. Modelling mutational and selection pressures on dinucleotides in eukaryotic phyla –selection against CpG and UpA in cytoplasmically expressed RNA and in RNA viruses. BMC Genom. 2013, 14, 610. [Google Scholar] [CrossRef] [Green Version]
- Atkinson, N.J.; Witteveldt, J.; Evans, D.J.; Simmonds, P. The influence of CpG and UpA dinucleotide frequencies on RNA virus replication and characterization of the innate cellular pathways underlying virus attenuation and enhanced replication. Nucleic Acids Res. 2014, 42, 4527–4545. [Google Scholar] [CrossRef]
- Tulloch, F.; Atkinson, N.J.; Evans, D.J.; Ryan, M.D.; Simmonds, P. RNA virus attenuation by codon pair deoptimisation is an artefact of increases in CpG/UpA dinucleotide frequencies. eLife 2014, 3, 04531. [Google Scholar] [CrossRef] [Green Version]
- Witteveldt, J.; Martin-Gans, M.; Simmonds, P. Enhancement of the Replication of Hepatitis C Virus Replicons of Genotypes 1 to 4 by Manipulation of CpG and UpA Dinucleotide Frequencies and Use of Cell Lines Expressing SECL14L2 for Antiviral Resistance Testing. Antimicrob. Agents Chemother. 2016, 60, 2981–2992. [Google Scholar] [CrossRef] [Green Version]
- Gaunt, E.; Wise, H.M.; Zhang, H.; Ni Lee, L.; Atkinson, N.J.; Nicol, M.Q.; Highton, A.J.; Klenerman, P.; Beard, P.; Dutia, B.M.; et al. Elevation of CpG frequencies in influenza A genome attenuates pathogenicity but enhances host response to infection. eLife 2016, 5. [Google Scholar] [CrossRef]
- Klitting, R.; Riziki, T.; Moureau, G.; Piorkowski, G.; A Gould, E.; De Lamballerie, X. Exploratory re-encoding of yellow fever virus genome: New insights for the design of live-attenuated viruses. Virus Evol. 2018, 4, vey021. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takata, M.A.; Carneiro, D.G.; Zang, T.M.; Soll, S.J.; York, A.; Blanco-Melo, D.; Bieniasz, P.D. CG dinucleotide suppression enables antiviral defence targeting non-self RNA. Nature 2017, 550, 124–127. [Google Scholar] [CrossRef] [PubMed]
- Odon, V.; Fros, J.J.; Goonawardane, N.; Dietrich, I.; Ibrahim, A.; Alshaikhahmed, K.; Nguyen, D.; Simmonds, P. The role of ZAP and OAS3/RNAseL pathways in the attenuation of an RNA virus with elevated frequencies of CpG and UpA dinucleotides. Nucleic Acids Res. 2019, 47, 8061–8083. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.-T.; Chiweshe, S.; McCormick, D.; Raper, A.R.; Wickenhagen, A.; DeFillipis, V.; Gaunt, E.; Simmonds, P.; Wilson, S.J.; Grey, F.; et al. Human cytomegalovirus evades ZAP detection by suppressing CpG dinucleotides in the major immediate early genes. bioRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Shackelton, L.A.; Parrish, C.R.; Holmes, E.C. Evolutionary Basis of Codon Usage and Nucleotide Composition Bias in Vertebrate DNA Viruses. J. Mol. Evol. 2006, 62, 551–563. [Google Scholar] [CrossRef]
- Fros, J.; Dietrich, I.; Alshaikhahmed, K.; Passchier, T.; Evans, D.J.; Simmonds, P. CpG and UpA dinucleotides in both coding and non-coding regions of echovirus 7 inhibit replication initiation post-entry. eLife 2017, 6. [Google Scholar] [CrossRef] [Green Version]
- Sharp, P.M.; Averof, M.; Lloyd, A.T.; Matassi, G.; Peden, J.F. DNA sequence evolution: The sounds of silence. Philos. Trans. R. Soc. B Boil. Sci. 1995, 349, 241–247. [Google Scholar] [CrossRef]
- Sharp, P.M.; Tuohy, T.M.; Mosurski, K.R. Codon usage in yeast: Cluster analysis clearly differentiates highly and lowly expressed genes. Nucleic Acids Res. 1986, 14, 5125–5143. [Google Scholar] [CrossRef]
- Greenbaum, B.D.; Levine, A.J.; Bhanot, G.; Rabadan, R. Patterns of Evolution and Host Gene Mimicry in Influenza and Other RNA Viruses. PLOS Pathog. 2008, 4, e1000079. [Google Scholar] [CrossRef] [PubMed]
- Greenbaum, B.D.; Cocco, S.; Levine, A.; Monasson, R. Quantitative theory of entropic forces acting on constrained nucleotide sequences applied to viruses. Proc. Natl. Acad. Sci. USA 2014, 111, 5054–5059. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kariin, S.; Burge, C. Dinucleotide relative abundance extremes: A genomic signature. Trends Genet. 1995, 11, 283–290. [Google Scholar] [CrossRef]
- Cock, P.J.; Antao, T.; Chang, J.T.; Chapman, B.; Cox, C.J.; Dalke, A.; Friedberg, I.; Hamelryck, T.; Kauff, F.; Wilczynski, B.; et al. Biopython: Freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 2009, 25, 1422–1423. [Google Scholar] [CrossRef]
- Simón, D.; Fajardo, Á.; Soñora, M.; Delfraro, A.; Musto, H. Host influence in the genomic composition of flaviviruses: A multivariate approach. Biochem. Biophys. Res. Commun. 2017, 492, 572–578. [Google Scholar] [CrossRef]
- Blitvich, B.; Firth, A.E. Insect-Specific Flaviviruses: A Systematic Review of Their Discovery, Host Range, Mode of Transmission, Superinfection Exclusion Potential and Genomic Organization. Viruses 2015, 7, 1927–1959. [Google Scholar] [CrossRef] [Green Version]
- Billoir, F.; De Micco, P.; Tolou, H.; De Chesse, R.; De Lamballerie, X.; Gould, E.A. Phylogeny of the genus Flavivirus using complete coding sequences of arthropod-borne viruses and viruses with no known vector. J. Gen. Virol. 2000, 81, 781–790. [Google Scholar] [CrossRef]
- International Committee on Taxonomy of Viruses (ICTV). Available online: https://talk.ictvonline.org/taxonomy/vmr/ (accessed on 27 February 2020).
- Ibrahim, A.; Fros, J.; Bertran, A.; Sechan, F.; Odon, V.; Torrance, L.; Kormelink, R.; Simmonds, P. A functional investigation of the suppression of CpG and UpA dinucleotide frequencies in plant RNA virus genomes. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef]
| Symbol | Description |
|---|---|
| i | Amino acid or amino acid pair |
| j | Dinucleotide |
| h | Dinucleotide frame position |
| ni | Number of occurrences of amino acid or amino acid pair i in the sequence |
| k | Set of different amino acids or amino acid pairs present in the sequence |
| oi,j,h | Synonymous proportion of dinucleotide j in frame position h for amino acid or amino acid pair i observed in the sequence |
| ei,j,h | Synonymous proportion of dinucleotide j in frame position h for amino acid or amino acid pair i expected under equal synonymous codon usage |
| N | Total number of amino acids or amino acid pairs present in the sequence |
| APOIV | AEFV | APOIV | AEFV | APOIV | AEFV | APOIV | AEFV | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| UUU | 1.11 | 1.08 | UCU | 1.01 | 0.75 | UAU | 0.98 | 1.00 | UGU | 0.95 | 1.04 |
| UUC | 0.89 | 0.92 | UCC | 0.73 | 1.13 | UAC | 1.02 | 1.00 | UGC | 1.05 | 0.96 |
| UUA | 0.38 | 0.56 | UCA | 1.57 | 0.96 | UAA | STOP | STOP | UGA | STOP | STOP |
| UUG | 1.39 | 1.23 | UCG | 0.34 | 1.06 | UAG | STOP | STOP | UGG | 1.00 | 1.00 |
| CUU | 1.08 | 0.83 | CCU | 1.19 | 0.89 | CAU | 1.25 | 1.05 | CGU | 0.48 | 1.19 |
| CUC | 1.00 | 1.42 | CCC | 0.86 | 1.00 | CAC | 0.75 | 0.95 | CGC | 0.39 | 1.19 |
| CUA | 0.67 | 0.79 | CCA | 1.69 | 1.32 | CAA | 0.91 | 1.23 | CGA | 0.68 | 1.05 |
| CUG | 1.48 | 1.17 | CCG | 0.25 | 0.79 | CAG | 1.09 | 0.77 | CGG | 0.68 | 0.88 |
| AUU | 1.10 | 1.22 | ACU | 1.13 | 0.98 | AAU | 0.91 | 0.84 | AGU | 1.04 | 0.96 |
| AUC | 1.15 | 1.05 | ACC | 1.27 | 1.07 | AAC | 1.09 | 1.16 | AGC | 1.32 | 1.13 |
| AUA | 0.75 | 0.73 | ACA | 1.29 | 1.00 | AAA | 1.01 | 1.23 | AGA | 2.19 | 1.00 |
| AUG | 1.00 | 1.00 | ACG | 0.32 | 0.95 | AAG | 0.99 | 0.77 | AGG | 1.58 | 0.69 |
| GUU | 1.08 | 1.29 | GCU | 1.65 | 1.07 | GAU | 0.97 | 0.67 | GGU | 0.73 | 0.74 |
| GUC | 1.00 | 0.98 | GCC | 1.03 | 1.62 | GAC | 1.03 | 1.33 | GGC | 0.73 | 0.73 |
| GUA | 0.29 | 0.62 | GCA | 1.05 | 0.68 | GAA | 1.09 | 1.07 | GGA | 1.75 | 1.57 |
| GUG | 1.63 | 1.11 | GCG | 0.27 | 0.63 | GAG | 0.91 | 0.93 | GGG | 0.80 | 0.96 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lytras, S.; Hughes, J. Synonymous Dinucleotide Usage: A Codon-Aware Metric for Quantifying Dinucleotide Representation in Viruses. Viruses 2020, 12, 462. https://0-doi-org.brum.beds.ac.uk/10.3390/v12040462
Lytras S, Hughes J. Synonymous Dinucleotide Usage: A Codon-Aware Metric for Quantifying Dinucleotide Representation in Viruses. Viruses. 2020; 12(4):462. https://0-doi-org.brum.beds.ac.uk/10.3390/v12040462
Chicago/Turabian StyleLytras, Spyros, and Joseph Hughes. 2020. "Synonymous Dinucleotide Usage: A Codon-Aware Metric for Quantifying Dinucleotide Representation in Viruses" Viruses 12, no. 4: 462. https://0-doi-org.brum.beds.ac.uk/10.3390/v12040462





