Microbiota of the Tongue and Systemic Connections: The Examination of the Tongue as an Integrated Approach in Oral Medicine
Abstract
:1. Introduction
2. Materials and Methods
2.1. General Characteristics
2.2. Inclusion and Exclusion Criteria
- Scientific articles published without time limit;
- International articles in English;
- Experimental studies with no age limits;
- In vivo experimental studies and in vitro experimental studies;
- Expert review;
- Literature reviews;
- Narrative reviews;
- Historical reviews and milestone papers;
- Scientific articles that dealt more generally with the microbiota of the oral cavity;
- Short communications;
- Opinion papers;
- Book chapter/congress proceedings.
3. Results
3.1. Summary of the Search Output
3.2. Characteristics of the Included Studies
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sun, B.; Zhou, D.; Tu, J.; Lu, Z. Evaluation of the Bacterial Diversity in the Human Tongue Coating Based on Genus-Specific Primers for 16S rRNA Sequencing. BioMed Res. Int. 2017, 2017, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Anastasi, J.K.; Currie, L.M.; Kim, G.H. Understanding diagnostic reasoning in TCM practice: Tongue diagnosis. Altern. Ther. Heal. Med. 2009, 15, 18–28. [Google Scholar]
- Ye, J.; Cai, X.; Yang, J.; Sun, X.; Hu, C.; Xia, J.; Shen, J.; Su, K.; Yan, H.; Xu, Y.; et al. Bacillus as a potential diagnostic marker for yellow tongue coating. Sci. Rep. 2016, 6, 32496. [Google Scholar] [CrossRef] [Green Version]
- Asakawa, M.; Takeshita, T.; Furuta, M.; Kageyama, S.; Takeuchi, K.; Hata, J.; Ninomiya, T.; Yamashita, Y. Tongue Microbiota and Oral Health Status in Community-Dwelling Elderly Adults. mSphere 2018, 3, e00332-18. [Google Scholar] [CrossRef] [Green Version]
- Welch, J.L.M.; Utter, D.; Rossetti, B.J.; Welch, D.B.M.; Eren, A.M.; Borisy, G.G. Dynamics of tongue microbial communities with single-nucleotide resolution using oligotyping. Front. Microbiol. 2014, 5, 568. [Google Scholar] [CrossRef] [Green Version]
- Cui, J.; Cui, H.; Yang, M.; Du, S.; Li, J.; Li, Y.; Liu, L.; Zhang, X.; Li, S. Tongue coating microbiome as a potential biomarker for gastritis including precancerous cascade. Protein Cell 2019, 10, 496–509. [Google Scholar] [CrossRef] [Green Version]
- Kageyama, S.; Asakawa, M.; Takeshita, T.; Ihara, Y.; Kanno, S.; Hara, T.; Takahashi, I.; Yamashita, Y. Transition of Bacterial Diversity and Composition in Tongue Microbiota during the First Two Years of Life. mSphere 2019, 4, e00187-19. [Google Scholar] [CrossRef] [Green Version]
- Jiang, B.; Liang, X.; Chen, Y.; Ma, T.; Liu, L.; Li, J.; Jiang, R.; Chen, T.; Zhang, X.; Li, S. Integrating next-generation sequencing and traditional tongue diagnosis to determine tongue coating microbiome. Sci. Rep. 2012, 2, srep00936. [Google Scholar] [CrossRef] [Green Version]
- Hsu, C.-H.; Yu, M.-C.; Lee, C.-H.; Lee, T.-C.; Yang, S.-Y. High Eosinophil Cationic Protein Level in Asthmatic Patients with “Heat” Zheng. Am. J. Chin. Med. 2003, 31, 277–283. [Google Scholar] [CrossRef]
- Zhao, Y.; Mao, Y.-F.; Tang, Y.-S.; Ni, M.-Z.; Liu, Q.-H.; Wang, Y.; Feng, Q.; Peng, J.-H.; Hu, Y.-Y. Altered oral microbiota in chronic hepatitis B patients with different tongue coatings. World J. Gastroenterol. 2018, 24, 3448–3461. [Google Scholar] [CrossRef]
- Farrell, J.J.; Zhang, L.; Zhou, H.; Chia, D.; Elashoff, D.; Akin, D.; Paster, B.J.; Joshipura, K.; Wong, D.T.W. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut 2011, 61, 582–588. [Google Scholar] [CrossRef]
- Mitsuhashi, K.; Nosho, K.; Sukawa, Y.; Matsunaga, Y.; Ito, M.; Kurihara, H.; Kanno, S.; Igarashi, H.; Naito, T.; Adachi, Y.; et al. Association ofFusobacteriumspecies in pancreatic cancer tissues with molecular features and prognosis. Oncotarget 2015, 6, 7209–7220. [Google Scholar] [CrossRef] [Green Version]
- Ballini, A.; Di Palma, G.; Isacco, C.G.; Boccellino, M.; Di Domenico, M.; Santacroce, L.; Nguyễn, K.C.; Scacco, S.; Calvani, M.; Boddi, A.; et al. Oral Microbiota and Immune System Crosstalk: A Translational Research. Biology 2020, 9, 131. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, J.H.; Parra, B.; Gutiérrez, S.; Arce, R.M.; Jaramillo, A.; Ariza, Y.; Contreras, A. Biomarkers of cardiovascular disease are increased in untreated chronic periodontitis: A case control study. Aust. Dent. J. 2014, 59, 29–36. [Google Scholar] [CrossRef]
- Fava, F.; Danese, S. Intestinal microbiota in inflammatory bowel disease: Friend of foe? World J. Gastroenterol. 2011, 17, 557–566. [Google Scholar] [CrossRef]
- Ahn, J.; Chen, C.Y.; Hayes, R.B. Oral microbiome and oral and gastrointestinal cancer risk. Cancer Causes Control. 2012, 23, 399–404. [Google Scholar] [CrossRef] [PubMed]
- Besnard, P.; Christensen, J.E.; Brignot, H.; Bernard, A.; Passilly-Degrace, P.; Nicklaus, S.; De Barros, J.-P.P.; Collet, X.; Lelouvier, B.; Servant, F.; et al. Author Correction: Obese Subjects With Specific Gustatory Papillae Microbiota and Salivary Cues Display an Impairment to Sense Lipids. Sci. Rep. 2018, 8, 9773. [Google Scholar] [CrossRef]
- Chen, L.Y.; Lu, G.Z. Analysis of Tongue Images in 114 Patients with Gastric Cancer. Zhong Yi Za Zhi 2011, 52, 1935–1938. [Google Scholar]
- Miyazaki, H.; Sakao, S.; Katoh, Y.; Takehara, T. Correlation Between Volatile Sulphur Compounds and Certain Oral Health Measurements in the General Population. J. Periodontol. 1995, 66, 679–684. [Google Scholar] [CrossRef]
- Xu, J.; Xiang, C.; Zhang, C.; Xu, B.; Wu, J.; Wang, R.; Yang, Y.; Shi, L.; Zhang, J.; Zhan, Z. Microbial biomarkers of common tongue coatings in patients with gastric cancer. Microb. Pathog. 2019, 127, 97–105. [Google Scholar] [CrossRef]
- Huang, S.; Yang, F.; Zeng, X.; Chen, J.; Li, R.; Wen, T.; Li, C.; Wei, W.; Liu, J.; Chen, L.; et al. Preliminary characterization of the oral microbiota of Chinese adults with and without gingivitis. BMC Oral Health 2011, 11, 33. [Google Scholar] [CrossRef] [Green Version]
- Kistler, J.O.; Booth, V.; Bradshaw, D.J.; Wade, W.G. Bacterial Community Development in Experimental Gingivitis. PLoS ONE 2013, 8, e71227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, Y.; Gong, D.; Shi, C.; Shao, F.; Shi, J.; Fei, J. Dysbiosis of oral buccal mucosa microbiota in patients with oral lichen planus. Oral Dis. 2017, 23, 674–682. [Google Scholar] [CrossRef]
- Cao, Y.; Qiao, M.; Tian, Z.; Yu, Y.; Xu, B.; Lao, W.; Ma, X.; Li, W. Comparative Analyses of Subgingival Microbiome in Chronic Periodontitis Patients with and Without IgA Nephropathy by High Throughput 16S rRNA Sequencing. Cell. Physiol. Biochem. 2018, 47, 774–783. [Google Scholar] [CrossRef]
- Healy, C.; Moran, G.P. The microbiome and oral cancer: More questions than answers. Oral Oncol. 2019, 89, 30–33. [Google Scholar] [CrossRef]
- Mager, D.L.; Haffajee, A.D.; Devlin, P.M.; Norris, C.M.; Posner, M.R.; Goodson, J.M. The salivary microbiota as a diagnostic indicator of oral cancer: A descriptive, non-randomized study of cancer-free and oral squamous cell carcinoma subjects. J. Transl. Med. 2005, 3, 27. [Google Scholar] [CrossRef] [Green Version]
- Allali, I.; Boukhatem, N.; Bouguenouch, L.; Hardi, H.; Boudouaya, H.A.; Cadenas, M.B.; Ouldim, K.; Amzazi, S.; Azcarate-Peril, M.A.; Ghazal, H. Gut microbiome of Moroccan colorectal cancer patients. Med Microbiol. Immunol. 2018, 207, 211–225. [Google Scholar] [CrossRef]
- Downes, J.; Sutcliffe, I.C.; Booth, V.; Wade, W.G. Prevotella maculosa sp. nov., isolated from the human oral cavity. Int. J. Syst. Evol. Microbiol. 2007, 57, 2936–2939. [Google Scholar] [CrossRef] [Green Version]
- Tsigarida, A.; Dabdoub, S.; Nagaraja, H.; Kumar, P. The Influence of Smoking on the Peri-Implant Microbiome. J. Dent. Res. 2015, 94, 1202–1217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ehrenfest, D.M.D.; Del Corso, M.; Inchingolo, F.; Charrier, J.-B. Selecting a relevant in vitro cell model for testing and comparing the effects of a Choukroun’s platelet-rich fibrin (PRF) membrane and a platelet-rich plasma (PRP) gel: Tricks and traps. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. Endodontol. 2010, 110, 409–411. [Google Scholar] [CrossRef]
- Aas, J.A.; Griffen, A.L.; Dardis, S.R.; Lee, A.M.; Olsen, I.; Dewhirst, F.E.; Leys, E.J.; Paster, B.J. Bacteria of Dental Caries in Primary and Permanent Teeth in Children and Young Adults. J. Clin. Microbiol. 2008, 46, 1407–1417. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, N.; Nyvad, B. The Role of Bacteria in the Caries Process: Ecological perspectives. J. Dent. Res. 2010, 90, 294–303. [Google Scholar] [CrossRef] [PubMed]
- Goodson, J.; Groppo, D.; Halem, S.; Carpino, E. Is Obesity an Oral Bacterial Disease? J. Dent. Res. 2009, 88, 519–523. [Google Scholar] [CrossRef]
- Takeshita, T.; Kageyama, S.; Furuta, M.; Tsuboi, H.; Takeuchi, K.; Shibata, Y.; Shimazaki, Y.; Akifusa, S.; Ninomiya, T.; Kiyohara, Y.; et al. Bacterial diversity in saliva and oral health-related conditions: The Hisayama Study. Sci. Rep. 2016, 6, 22164. [Google Scholar] [CrossRef] [Green Version]
- Carlsson, J.; Grahnén, H.; Jonsson, G.; Wikner, S. Early Establishment of Streptococcus salivarius in the Mouths of Infants. J. Dent. Res. 1970, 49, 415–418. [Google Scholar] [CrossRef] [PubMed]
- Segata, N.; Haake, S.K.; Mannon, P.; Lemon, K.P.; Waldron, L.; Gevers, D.; Huttenhower, C.; Izard, J. Composition of the adult digestive tract bacterial microbiome based on seven mouth surfaces, tonsils, throat and stool samples. Genome Biol. 2012, 13, R42. [Google Scholar] [CrossRef] [Green Version]
- Chu, D.M.; Ma, J.; Prince, A.L.; Antony, K.M.; Seferovic, M.D.; Aagaard, K.M. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 2017, 23, 314–326. [Google Scholar] [CrossRef] [Green Version]
- Mason, M.R.; Chambers, S.; Dabdoub, S.; Thikkurissy, S.; Kumar, P.S. Characterizing oral microbial communities across dentition states and colonization niches. Microbiome 2018, 6, 1–10. [Google Scholar] [CrossRef]
- Seerangaiyan, K.; Jüch, F.; Winkel, E.G. Tongue coating: Its characteristics and role in intra-oral halitosis and general health—A review. J. Breath Res. 2017, 12, 034001. [Google Scholar] [CrossRef]
- Loesche, W.J. Microbiology and treatment of halitosis. Curr. Infect. Dis. Rep. 2003, 5, 220–226. [Google Scholar] [CrossRef]
- NISC Comparative Sequencing Program; Oh, J.; Byrd, A.L.; Deming, C.; Conlan, S.; Kong, H.H.; Segre, J.A. Biogeography and individuality shape function in the human skin metagenome. Nature 2014, 514, 59–64. [Google Scholar] [CrossRef] [Green Version]
- Ballini, A.; Gnoni, A.; De Vito, D.; DiPalma, G.; Cantore, S.; Isacco, C.G.; Saini, R.; Santacroce, L.; Topi, S.; Scarano, A.; et al. Effect of probiotics on the occurrence of nutrition absorption capacities in healthy children: A randomized double-blinded placebo-controlled pilot study. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 8645–8657. [Google Scholar]
- Inchingolo, F.; Martelli, F.S.; Isacco, C.G.; Borsani, E.; Cantore, S.; Corcioli, F.; Boddi, A.; Nguyễn, K.C.; De Vito, D.; Aityan, S.K.; et al. Chronic Periodontitis and Immunity, Towards the Implementation of a Personalized Medicine: A Translational Research on Gene Single Nucleotide Polymorphisms (SNPs) Linked to Chronic Oral Dysbiosis in 96 Caucasian Patients. Biomedicines 2020, 8, 115. [Google Scholar] [CrossRef]
- Cantore, S.; Ballini, A.; De Vito, D.; Abbinante, A.; Altini, V.; DiPalma, G.; Inchingolo, F.; Saini, R. Clinical results of improvement in periodontal condition by administration of oral probiotics. J. Biol. Regul. Homeost. Agents 2018, 32, 1329–1334. [Google Scholar]
- Inchingolo, F.; Santacroce, L.; Ballini, A.; Topi, S.; DiPalma, G.; Haxhirexha, K.; Bottalico, L.; Charitos, I.A. Oral Cancer: A Historical Review. Int. J. Environ. Res. Public Health 2020, 17, 3168. [Google Scholar] [CrossRef]
- Topi, S.; Santacroce, L.; Bottalico, L.; Ballini, A.; Inchingolo, A.D.; Dipalma, G.; Charitos, I.A.; Inchingolo, F. Gastric Cancer in History: A Perspective Interdisciplinary Study. Cancers 2020, 12, 264. [Google Scholar] [CrossRef] [Green Version]
- Boccellino, M.; Di Stasio, D.; Di Palma, G.; Cantore, S.; Ambrosio, P.; Coppola, M.; Quagliuolo, L.; Scarano, A.; Malcangi, G.; Borsani, E.; et al. Steroids and growth factors in oral squamous cell carcinoma: Useful source of dental-derived stem cells to develop a steroidogenic model in new clinical strategies. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 8730–8740. [Google Scholar]
- Yang, I.; Nell, S.; Suerbaum, S. Survival in hostile territory: The microbiota of the stomach. FEMS Microbiol. Rev. 2013, 37, 736–761. [Google Scholar] [CrossRef]
- Gibbons, R.J.; Houte, J.V. Bacterial Adherence in Oral Microbial Ecology. Annu. Rev. Microbiol. 1975, 29, 19–42. [Google Scholar] [CrossRef]
- Cabre, M.; Serra-Prat, M.; Palomera, E.; Almirall, J.; Pallares, R.; Clavé, P. Prevalence and prognostic implications of dysphagia in elderly patients with pneumonia. Age Ageing 2010, 39, 39–45. [Google Scholar] [CrossRef] [Green Version]
- Abe, S.; Ishihara, K.; Adachi, M.; Okuda, K. Tongue-coating as risk indicator for aspiration pneumonia in edentate elderly. Arch. Gerontol. Geriatr. 2008, 47, 267–275. [Google Scholar] [CrossRef] [Green Version]
- Shimazaki, Y.; Tomioka, M.; Saito, T.; Nabeshima, F.; Ikematsu, H.; Koyano, K.; Yamashita, Y. Influence of oral health on febrile status in long-term hospitalized elderly patients. Arch. Gerontol. Geriatr. 2009, 48, 411–414. [Google Scholar] [CrossRef]
- Marik, P.E.; Kaplan, D. Aspiration Pneumonia and Dysphagia in the Elderly. Chest 2003, 124, 328–336. [Google Scholar] [CrossRef] [Green Version]
- Sjögren, P.; Nilsson, E.; Forsell, M.; Johansson, O.; Hoogstraate, J.; Sjögren, P. A Systematic Review of the Preventive Effect of Oral Hygiene on Pneumonia and Respiratory Tract Infection in Elderly People in Hospitals and Nursing Homes: Effect Estimates and Methodological Quality of Randomized Controlled Trials: Oral hygiene and pneumonia in elderly. J. Am. Geriatr. Soc. 2008, 56, 2124–2130. [Google Scholar] [CrossRef]
- Zhou, Y.; Gao, H.; Mihindukulasuriya, A.K.; La Rosa, P.S.; Wylie, K.M.; Vishnivetskaya, T.; Podar, M.; Warner, B.; Tarr, I.P.; Nelson, E.D.; et al. Biogeography of the ecosystems of the healthy human body. Genome Biol. 2013, 14, R1. [Google Scholar] [CrossRef] [Green Version]
- Humphrey, L.L.; Fu, R.; Buckley, D.I.; Freeman, M.; Helfand, M. Periodontal Disease and Coronary Heart Disease Incidence: A Systematic Review and Meta-analysis. J. Gen. Intern. Med. 2008, 23, 2079–2086. [Google Scholar] [CrossRef] [Green Version]
- Ide, R.; Hoshuyama, T.; Wilson, D.; Takahashi, K.; Higashi, T. Periodontal Disease and Incident Diabetes: A Seven-Year Study. J. Dent. Res. 2011, 90, 41–46. [Google Scholar] [CrossRef]
- Yoneda, M.; Naka, S.; Nakano, K.; Wada, K.; Endo, H.; Mawatari, H.; Imajo, K.; Nomura, R.; Hokamura, K.; Ono, M.; et al. Involvement of a periodontal pathogen, Porphyromonas gingivalis on the pathogenesis of non-alcoholic fatty liver disease. BMC Gastroenterol. 2012, 12, 16. [Google Scholar] [CrossRef] [Green Version]
- Iwauchi, M.; Horigome, A.; Ishikawa, K.; Mikuni, A.; Nakano, M.; Xiao, J.; Odamaki, T.; Hironaka, S. Relationship between oral and gut microbiota in elderly people. Immun. Inflamm. Dis. 2019, 7, 229–236. [Google Scholar] [CrossRef] [Green Version]
- Dawes, C. Salivary flow patterns and the health of hard and soft oral tissues. J. Am. Dent. Assoc. 2008, 139, 18S–24S. [Google Scholar] [CrossRef]
- Schwabe, R.F.; Jobin, C. The microbiome and cancer. Nat. Rev. Cancer 2013, 13, 800–812. [Google Scholar] [CrossRef] [Green Version]
- Zhao, Y.; Gao, X.; Guo, J.; Yu, D.; Xiao, Y.; Wang, H.; Li, Y. Helicobacter pyloriinfection alters gastric and tongue coating microbial communities. Helicobacter 2019, 24, e12567. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Han, S.; Chen, Y.; Ji, Z. Variations of Tongue Coating Microbiota in Patients with Gastric Cancer. BioMed Res. Int. 2015, 2015, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Mashima, I.; Theodorea, C.F.; Thaweboon, B.; Thaweboon, S.; Nakazawa, F. Identification of Veillonella Species in the Tongue Biofilm by Using a Novel One-Step Polymerase Chain Reaction Method. PLoS ONE 2016, 11, e0157516. [Google Scholar] [CrossRef]
- Said, H.S.; Suda, W.; Nakagome, S.; Chinen, H.; Oshima, K.; Kim, S.; Kimura, R.; Iraha, A.; Ishida, H.; Fujita, J.; et al. Dysbiosis of Salivary Microbiota in Inflammatory Bowel Disease and Its Association With Oral Immunological Biomarkers. DNA Res. 2013, 21, 15–25. [Google Scholar] [CrossRef] [Green Version]
- Isacco, C.G.; Ballini, A.; De Vito, D.; Nguyen, K.C.D.; Cantore, S.; Bottalico, L.; Quagliuolo, L.; Boccellino, M.; Di Domenico, M.; Santacroce, L. Rebalance the Oral Microbiota as Efficacy Tool in Endocrine, Metabolic, and Immune Disorders. Endocr. Metab. Immune Dis. Drug Targets 2021, 21, 777–784. [Google Scholar] [CrossRef]
- Hajishengallis, G.; Darveau, R.P.; Curtis, M.A. The keystone-pathogen hypothesis. Nat. Rev. Microbiol. 2012, 10, 717–725. [Google Scholar] [CrossRef]
- Ceccarelli, F.; Orrù, G.; Pilloni, A.; Bartosiewicz, I.; Perricone, C.; Martino, E.; Lucchetti, R.; Fais, S.; Vomero, M.; Olivieri, M.; et al. Porphyromonas gingivalisin the tongue biofilm is associated with clinical outcome in rheumatoid arthritis patients. Clin. Exp. Immunol. 2018, 194, 244–252. [Google Scholar] [CrossRef] [Green Version]
- Han, S.; Yang, X.; Qi, Q.; Pan, Y.; Chen, Y.; Shen, J.; Liao, H.; Ji, Z. Potential screening and early diagnosis method for cancer: Tongue diagnosis. Int. J. Oncol. 2016, 48, 2257–2264. [Google Scholar] [CrossRef] [Green Version]
- Tribble, G.D.; Angelov, N.; Weltman, R.; Wang, B.-Y.; Eswaran, S.V.; Gay, I.C.; Parthasarathy, K.; Dao, D.-H.V.; Richardson, K.N.; Ismail, N.M.; et al. Frequency of Tongue Cleaning Impacts the Human Tongue Microbiome Composition and Enterosalivary Circulation of Nitrate. Front. Cell. Infect. Microbiol. 2019, 9, 39. [Google Scholar] [CrossRef] [Green Version]
- Han, S.; Chen, Y.; Hu, J.; Ji, Z. Tongue images and tongue coating microbiome in patients with colorectal cancer. Microb. Pathog. 2014, 77, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Ren, Z.; Li, A.; Zhang, H.; Jiang, J.; Xu, S.; Luo, Q.; Zhou, K.; Sun, X.; Zheng, S.; et al. Deep sequencing reveals microbiota dysbiosis of tongue coat in patients with liver carcinoma. Sci. Rep. 2016, 6, 33142. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, H.; Parthasarathy, D.; Torregrossa, A.C.; Mian, A.; Bryan, N.S. Analytical Techniques for Assaying Nitric Oxide Bioactivity. J. Vis. Exp. 2012, 3722. [Google Scholar] [CrossRef] [Green Version]
- Ma, L.; Hu, H.L.; Feng, X.; Wang, S. Nitrate and Nitrite in Health and Disease. Aging Dis. 2018, 9, 938–945. [Google Scholar] [CrossRef] [Green Version]
- Epstein, F.H.; Moncada, S.; Higgs, A. The L-Arginine-Nitric Oxide Pathway. N. Engl. J. Med. 1993, 329, 2002–2012. [Google Scholar] [CrossRef]
- Torregrossa, A.C.; Aranke, M.; Bryan, N.S. Nitric oxide and geriatrics: Implications in diagnostics and treatment of the elderly. J. Geriatr. Cardiol. 2011, 8, 230–242. [Google Scholar] [CrossRef] [Green Version]
- Graudal, N.A.; Hubeck-Graudal, T.; Jurgens, G. Effects of low sodium diet versus high sodium diet on blood pressure, renin, aldosterone, catecholamines, cholesterol, and triglyceride. Cochrane Database Syst. Rev. 2017, 4, CD004022. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Cai, X.; Cao, P. Problems and prospects of current studies on the microecology of tongue coating. Chin. Med. 2014, 9, 9. [Google Scholar] [CrossRef] [Green Version]
- Santacroce, L.; Inchingolo, F.; Topi, S.; Del Prete, R.; Di Cosola, M.; Charitos, I.A.; Montagnani, M. Potential beneficial role of probiotics on the outcome of COVID-19 patients: An evolving perspective. Diabetes Metab. Syndr. Clin. Res. Rev. 2021, 15, 295–301. [Google Scholar] [CrossRef]
- Santacroce, L.; Charitos, I.A.; Ballini, A.; Inchingolo, F.; Luperto, P.; De Nitto, E.; Topi, S. The Human Respiratory System and its Microbiome at a Glimpse. Biology 2020, 9, 318. [Google Scholar] [CrossRef]
- Contaldo, M.; Fusco, A.; Stiuso, P.; Lama, S.; Gravina, A.G.; Itro, A.; Federico, A.; Itro, A.; Dipalma, G.; Inchingolo, F.; et al. Oral Microbiota and Salivary Levels of Oral Pathogens in Gastro-Intestinal Diseases: Current Knowledge and Exploratory Study. Microorganisms 2021, 9, 1064. [Google Scholar] [CrossRef] [PubMed]
- Santacroce, L.; Sardaro, N.; Topi, S.; Pettini, F.; Bottalico, L.; Cantore, S.; Cascella, G.; Del Prete, R.; Di Palma, G.; Inchingolo, F. The pivotal role of oral microbiota in health and disease. J. Biol. Regul. Homeost. Agents 2020, 34, 733–737. [Google Scholar] [PubMed]
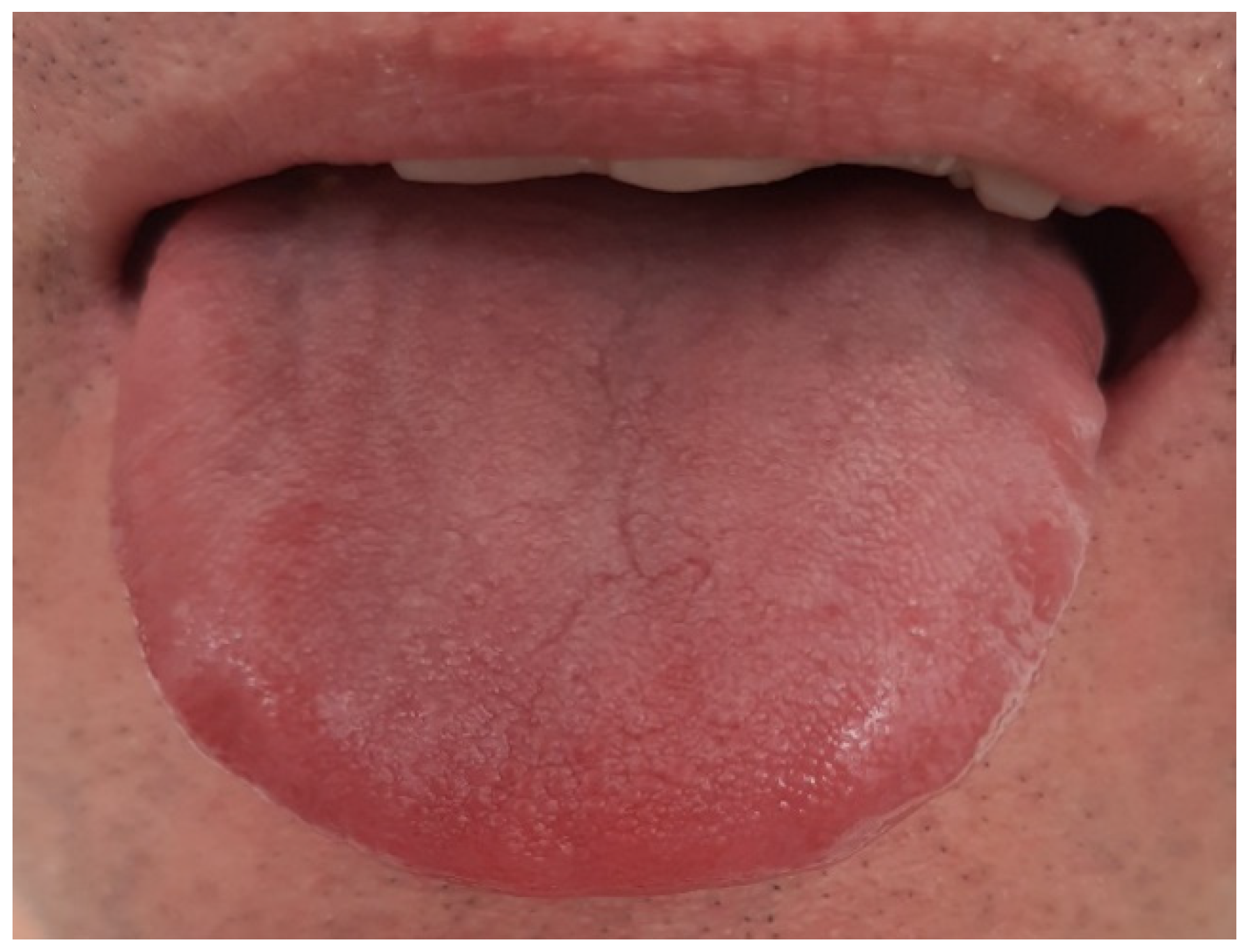
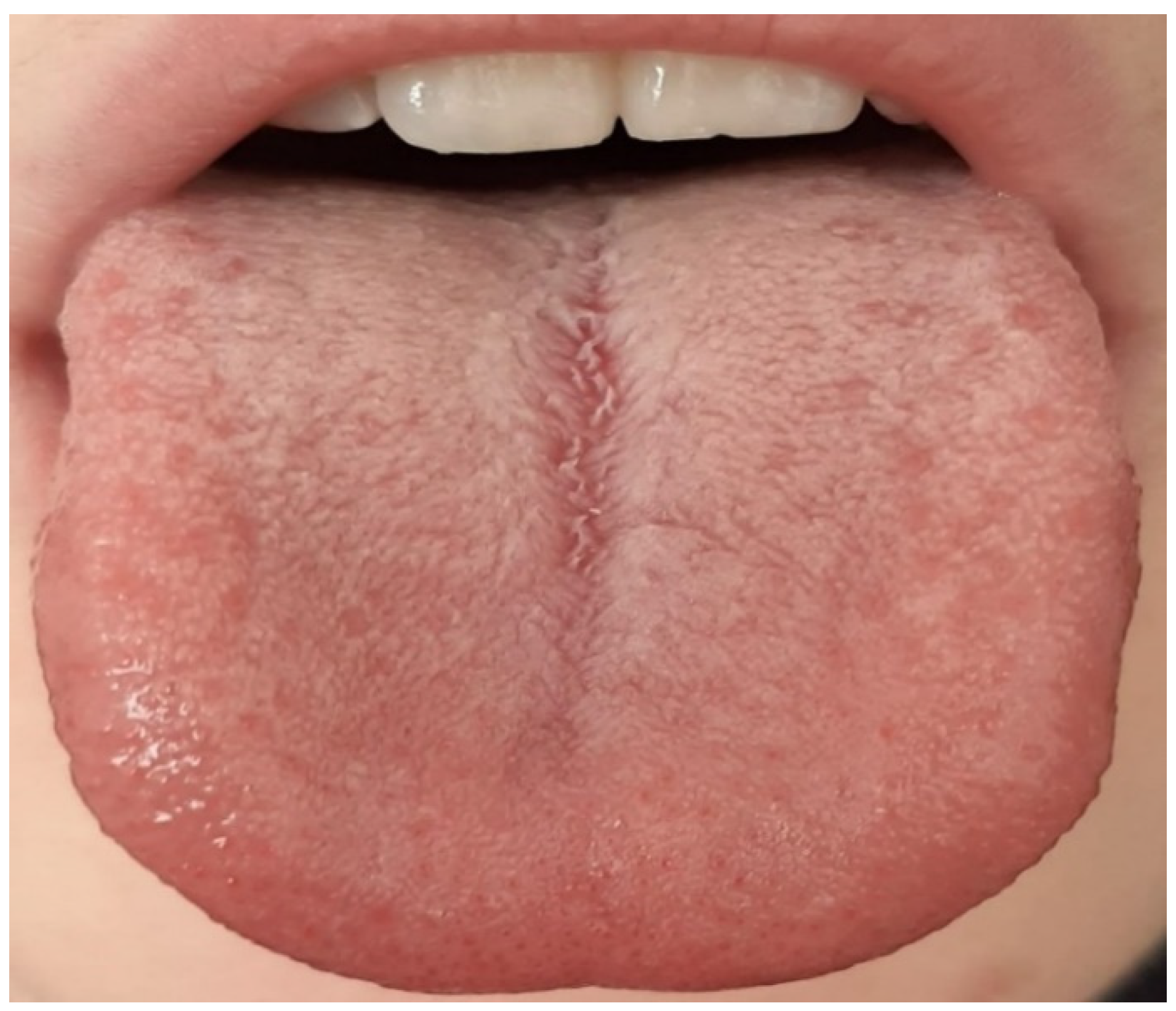
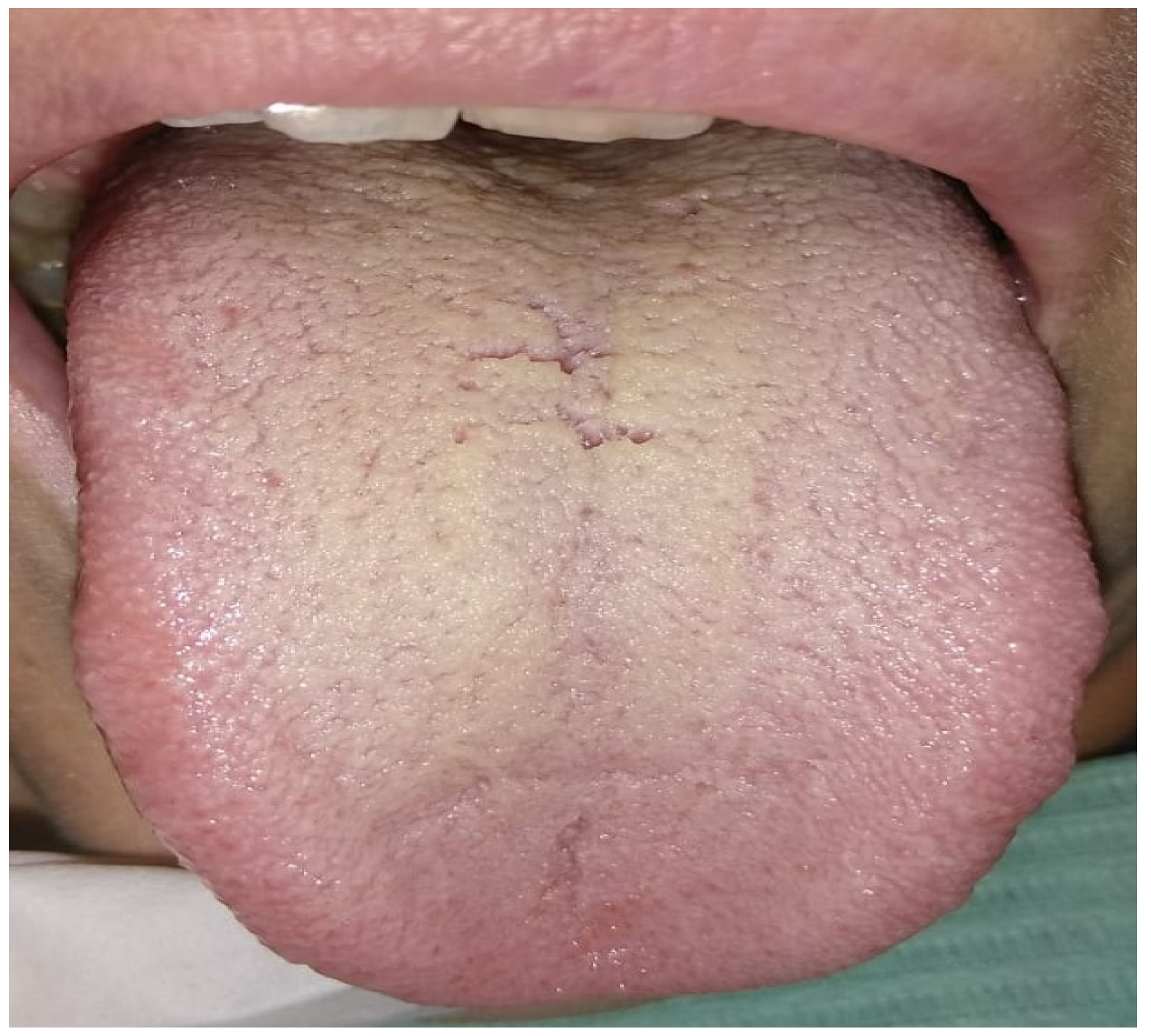

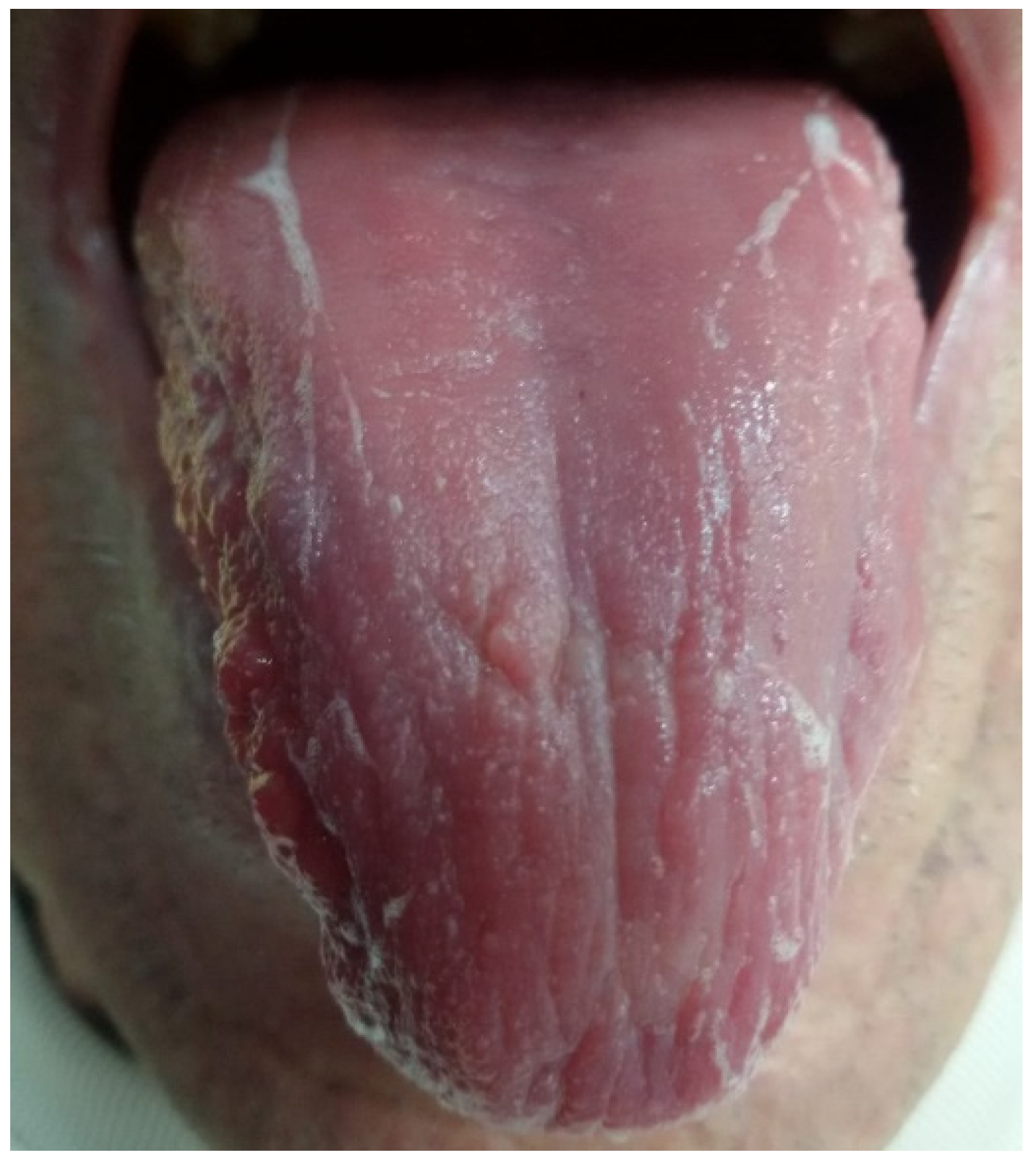


| Description |
|---|
| Thin white, thick white, and fat white tongue coating |
| Tongue coating thin yellow, thick yellow, and fat yellow |
| Thin black and gray tongue coating and thick gray black |
| Value | Description |
|---|---|
| 0 | No visible patina on the back of the tongue |
| 1 | Patina present only on the posterior III of the tongue |
| 2 | Patina that completely covers the dorsal surface of the tongue but does not mask the underlying mucosa |
| 3 | Very thick patina covering the entire dorsal surface of the tongue |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Casu, C.; Mosaico, G.; Natoli, V.; Scarano, A.; Lorusso, F.; Inchingolo, F. Microbiota of the Tongue and Systemic Connections: The Examination of the Tongue as an Integrated Approach in Oral Medicine. Hygiene 2021, 1, 56-68. https://0-doi-org.brum.beds.ac.uk/10.3390/hygiene1020006
Casu C, Mosaico G, Natoli V, Scarano A, Lorusso F, Inchingolo F. Microbiota of the Tongue and Systemic Connections: The Examination of the Tongue as an Integrated Approach in Oral Medicine. Hygiene. 2021; 1(2):56-68. https://0-doi-org.brum.beds.ac.uk/10.3390/hygiene1020006
Chicago/Turabian StyleCasu, Cinzia, Giovanna Mosaico, Valentino Natoli, Antonio Scarano, Felice Lorusso, and Francesco Inchingolo. 2021. "Microbiota of the Tongue and Systemic Connections: The Examination of the Tongue as an Integrated Approach in Oral Medicine" Hygiene 1, no. 2: 56-68. https://0-doi-org.brum.beds.ac.uk/10.3390/hygiene1020006






