Assessing the Effect of Real Spatial Resolution of In Situ UAV Multispectral Images on Seedling Rapeseed Growth Monitoring
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Area
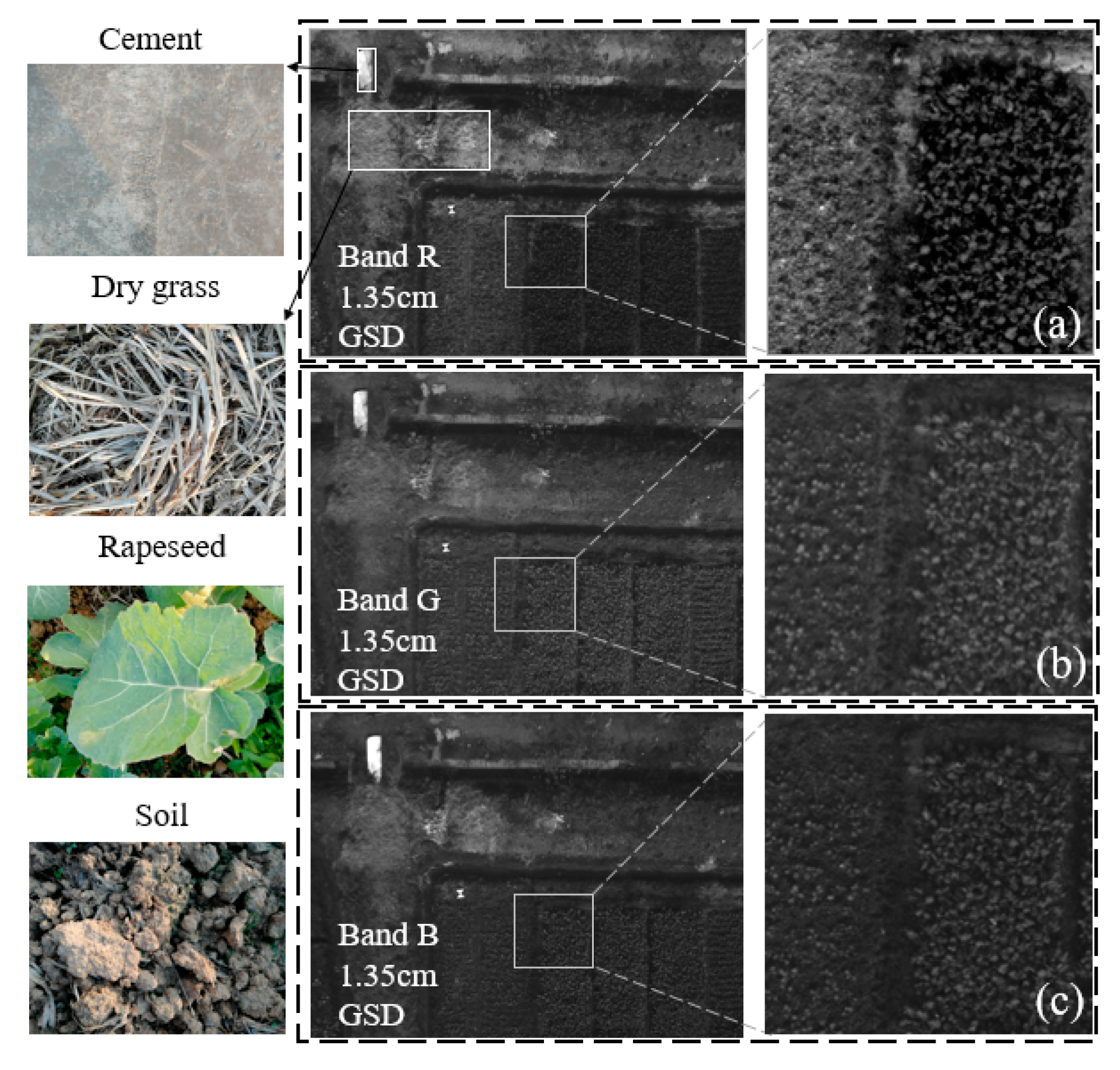
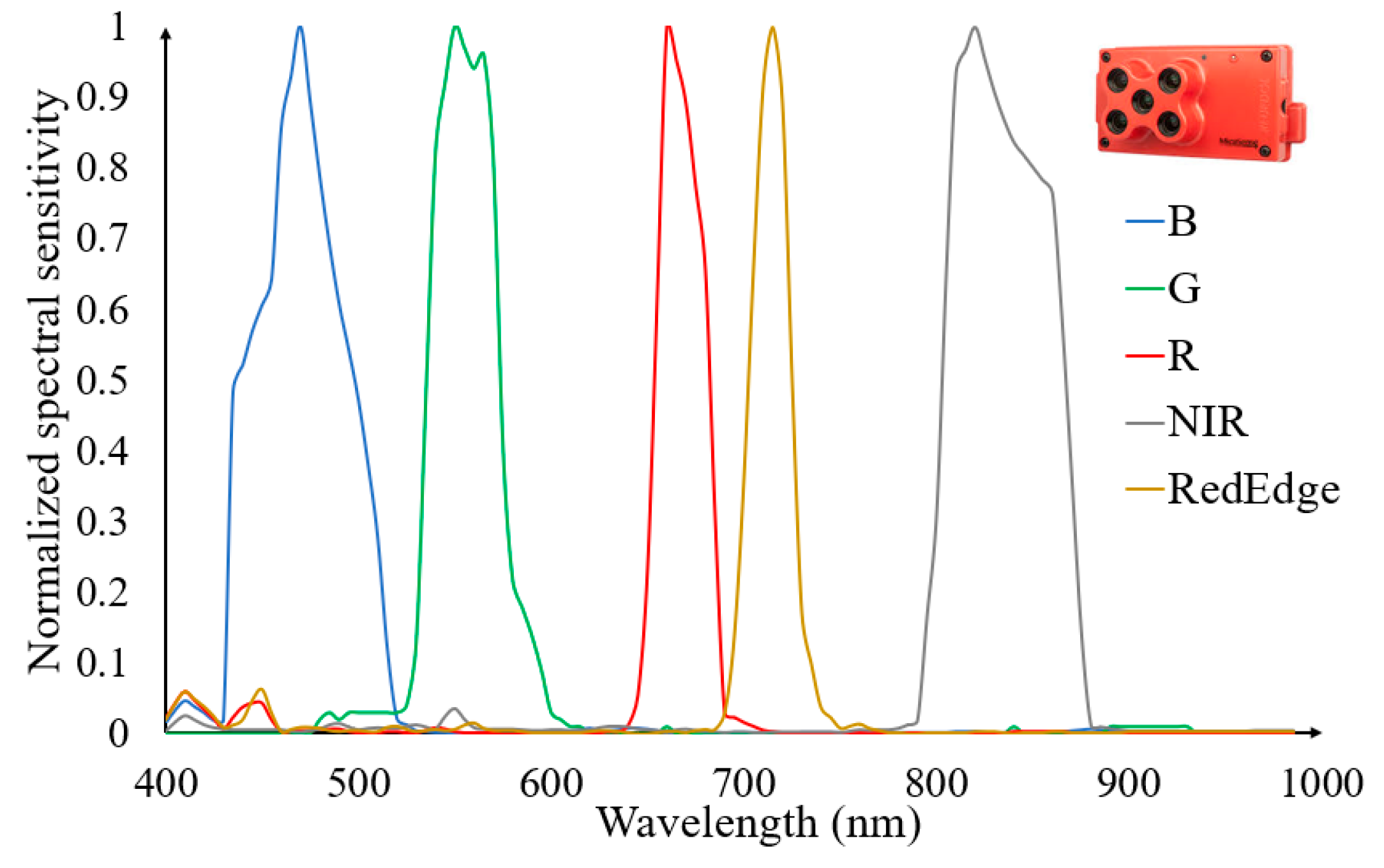
2.2. UAV Image Acquisition
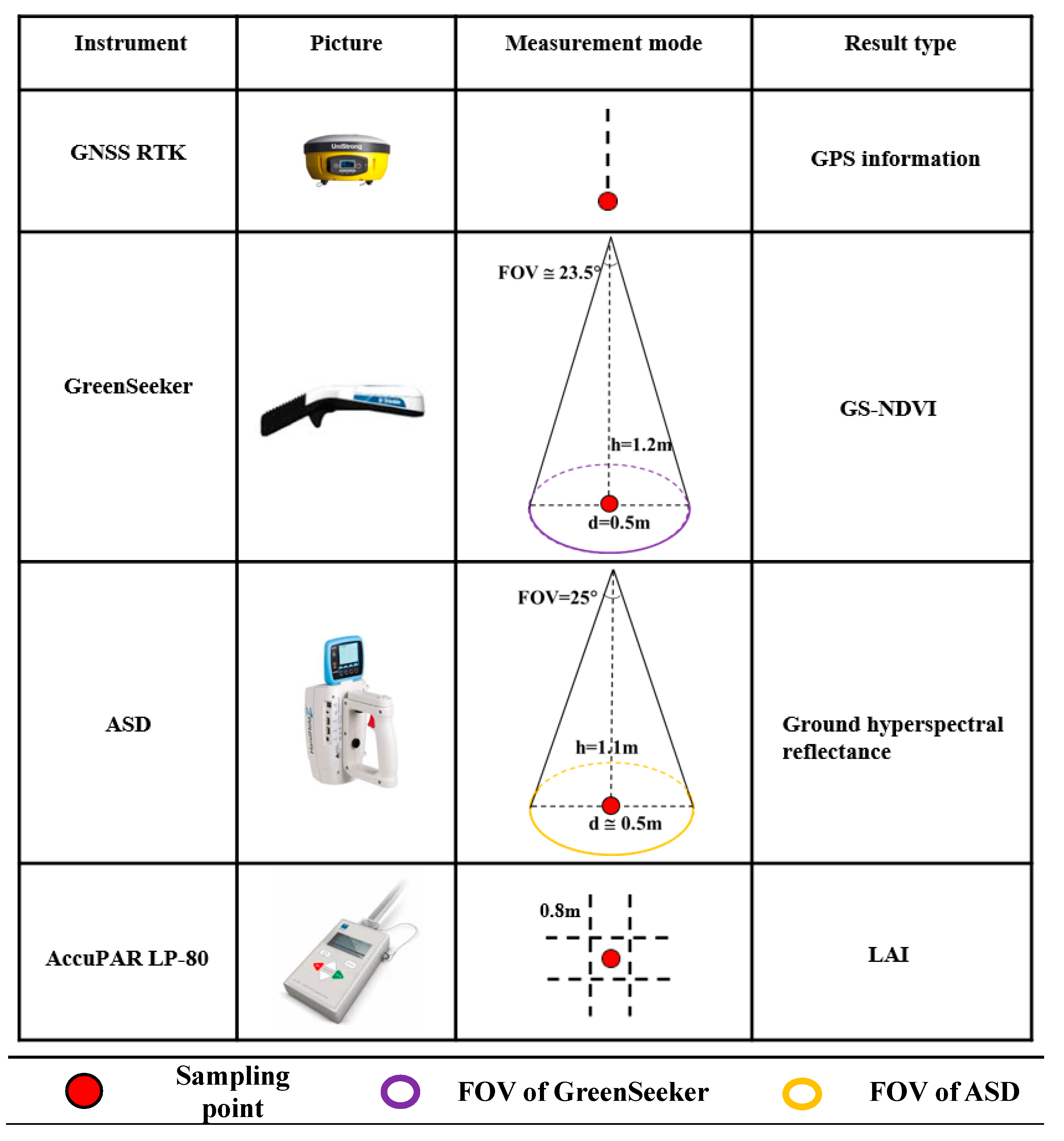
2.3. Field Data Acquisition
2.4. Data Pre-Processing
2.5. Regression Models of GS-NDVI and LAI
2.5.1. Estimation of GS-NDVI and LAI by UAV-VIs
2.5.2. Estimation of GS-NDVI and LAI by ASD-VIs
2.5.3. Estimation of LAI by UAV-VIs*PHDSM
3. Results
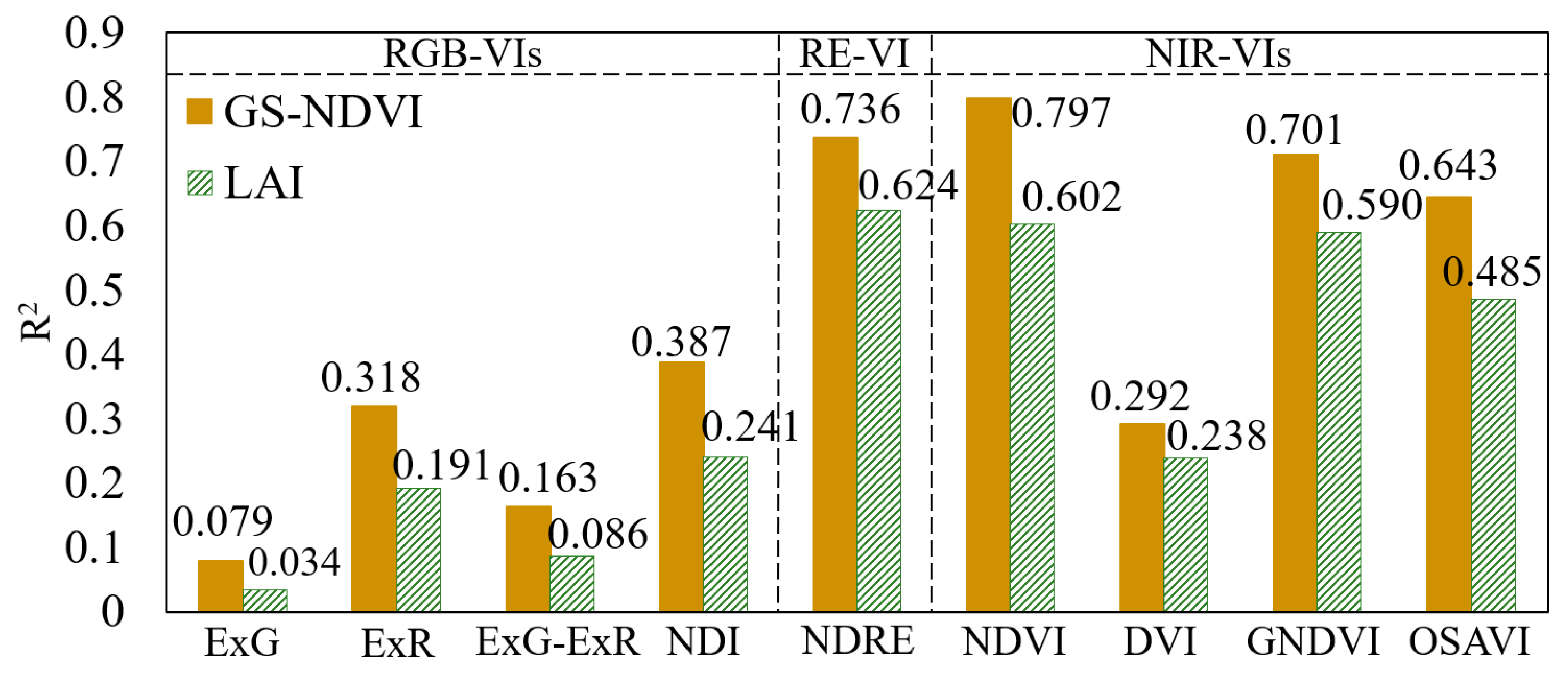
3.1. Performance of Different UAV-VIs for LAI and GS-NDVI Estimation
3.2. Performance of Different ASD-VIs for LAI and GS-NDVI Estimation
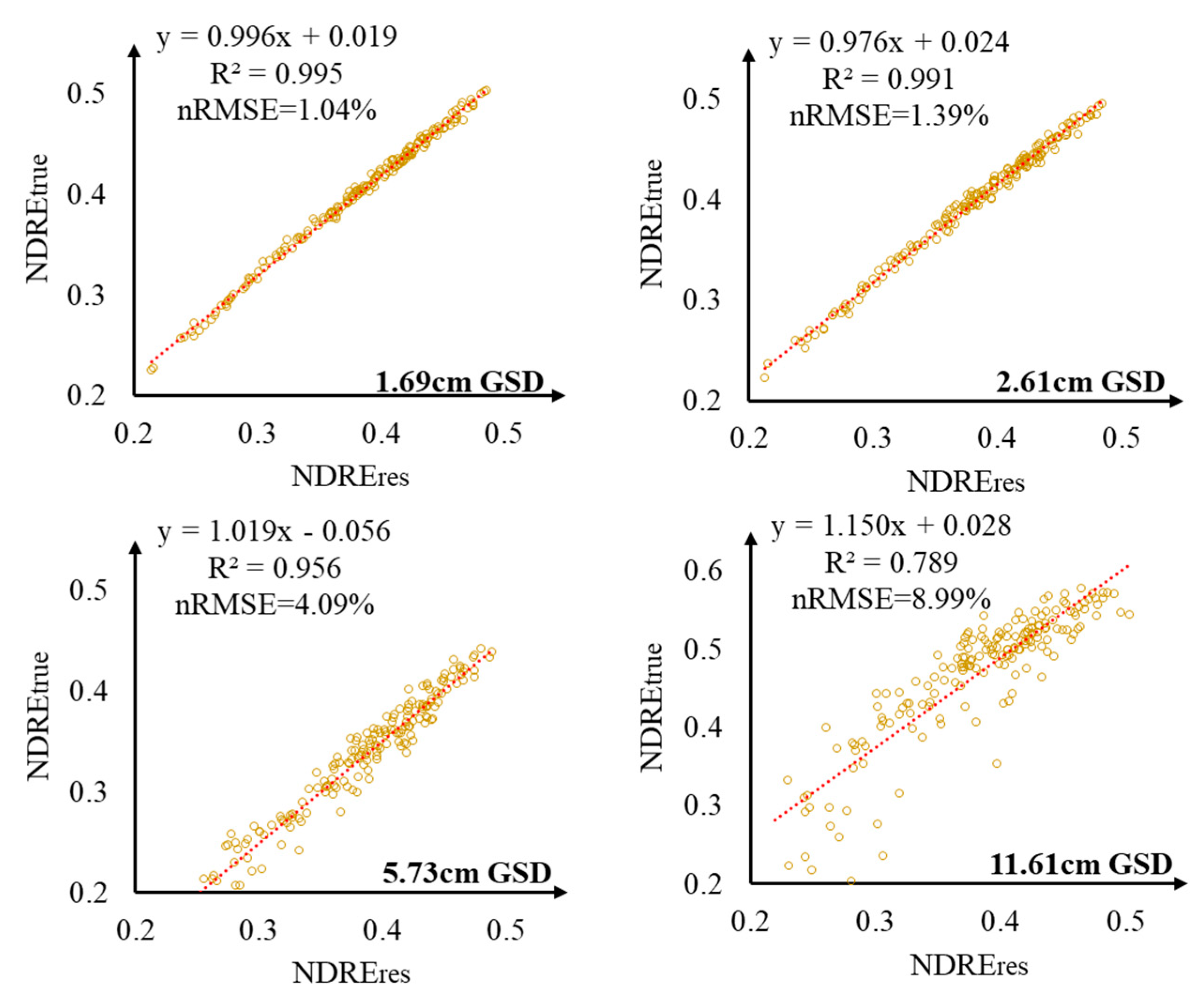
3.3. Performance of Optimal VIs under Different GSDs for LAI and GS-NDVI Estimation
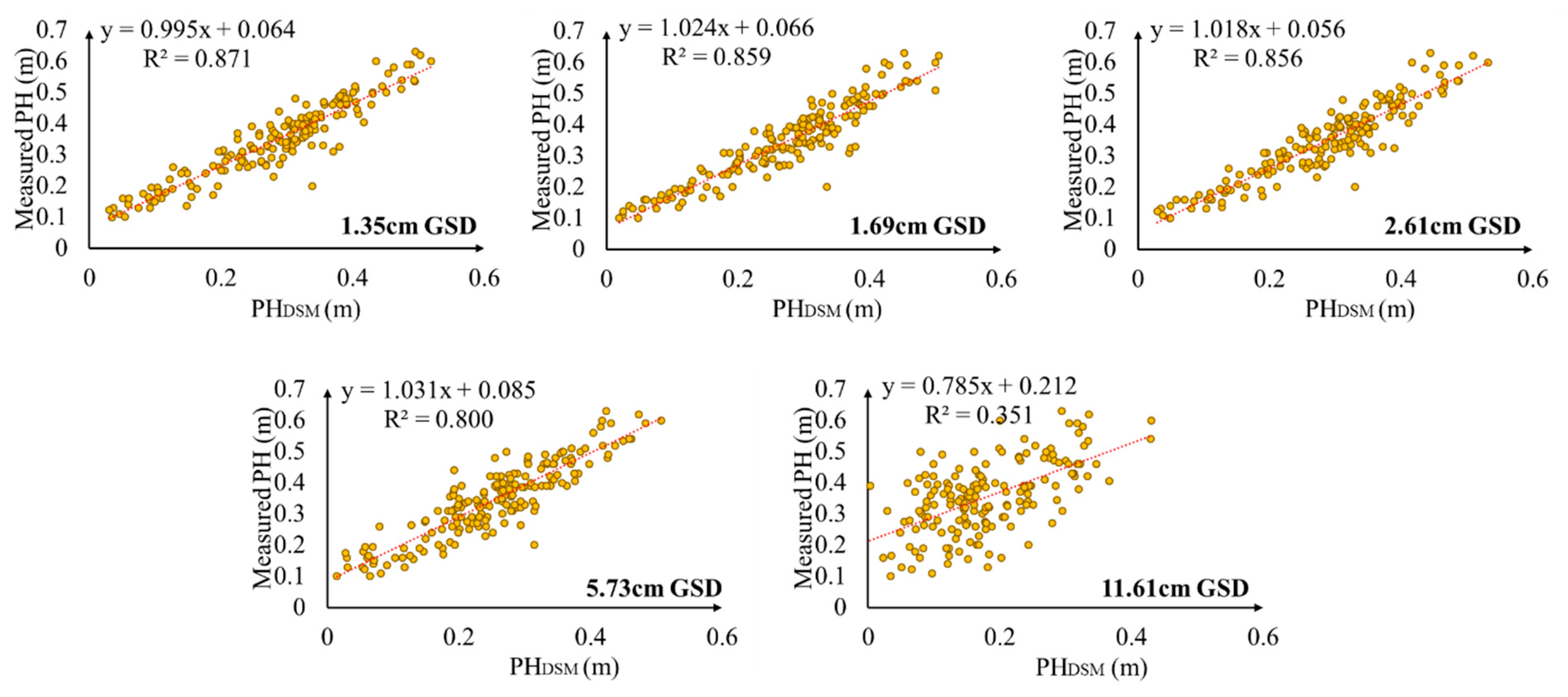
3.4. Performance of PHDSM under Different GSDs for PH Estimation
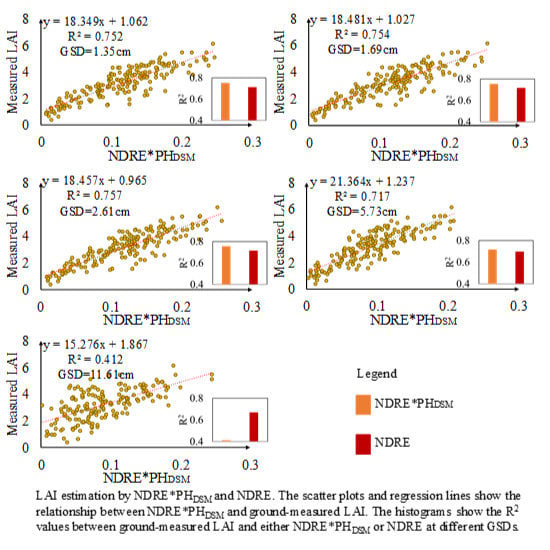
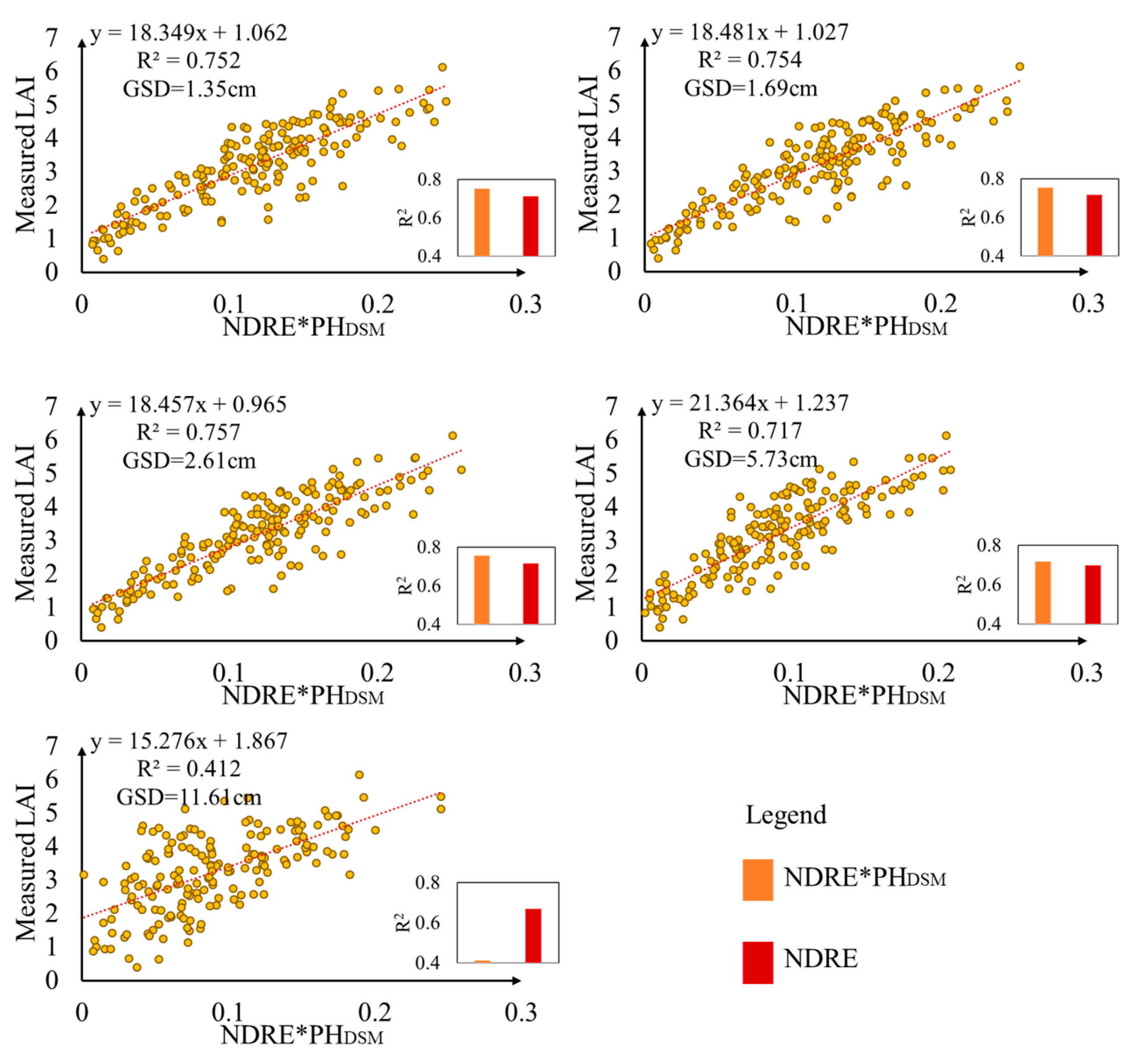
3.5. Performance of VIs*PHDSM under Different GSDs for LAI Estimation
4. Discussion
4.1. Effect of VI Type on GS-NDVI and LAI Estimation
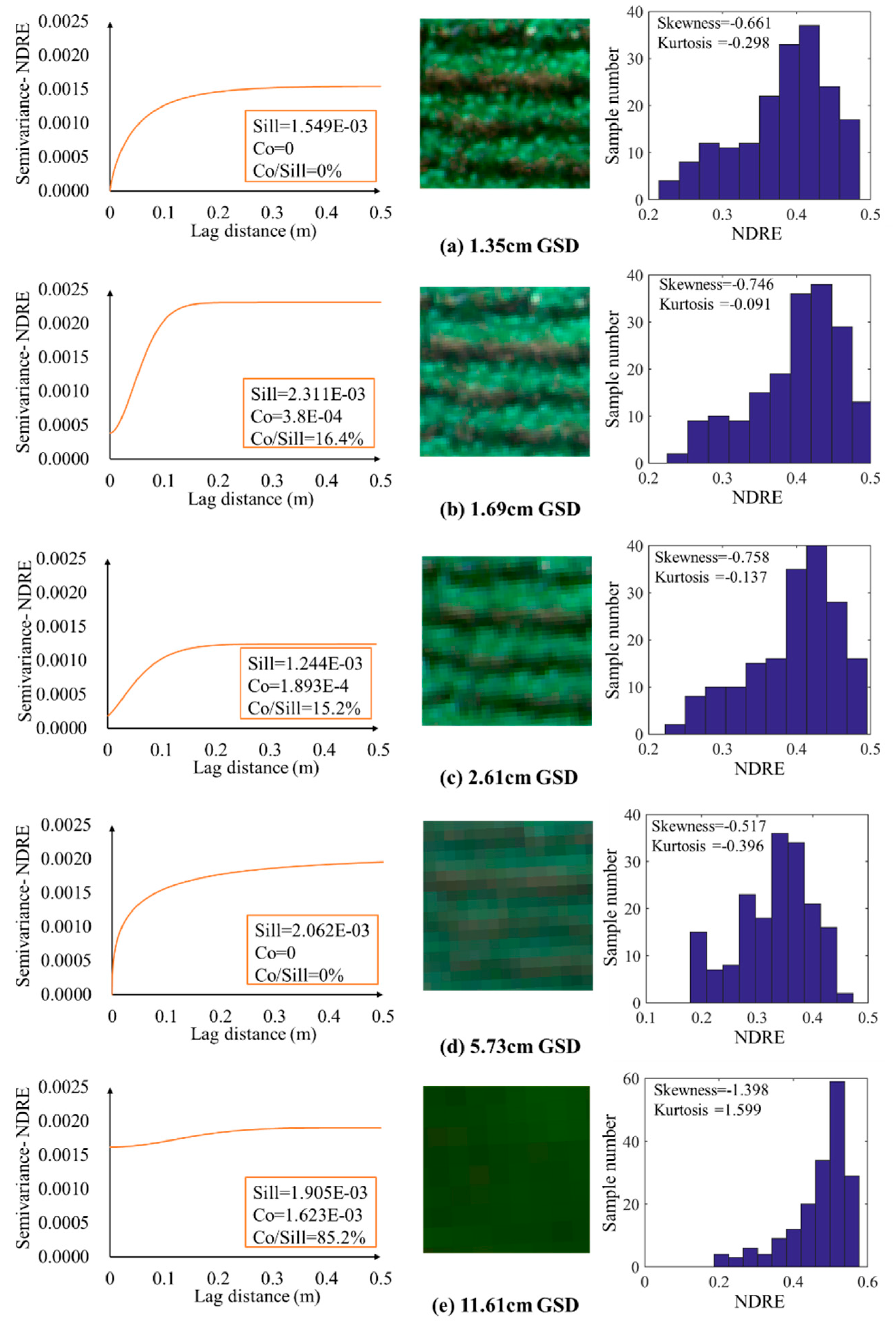
4.2. Effect of VIs under Different GSDs on GS-NDVI and LAI Estimation
4.3. Effect of PHDSM under Different GSDs on LAI Estimation
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Namany, S.; Al-Ansari, T.; Govindan, R. Optimisation of the energy, water, and food nexus for food security scenarios. Comput. Chem. Eng. 2019, 129, 106513. [Google Scholar] [CrossRef]
- Zhang, C.; Kovacs, J.M. The application of small unmanned aerial systems for precision agriculture: A review. Precis. Agric. 2012, 13, 693–712. [Google Scholar] [CrossRef]
- Jonckheere, I.; Fleck, S.; Nackaerts, K.; Muys, B.; Coppin, P.; Weiss, M.; Baret, F. Review of methods for in situ leaf area index determination. Agric. For. Meteorol. 2004, 121, 19–35. [Google Scholar] [CrossRef]
- Nelson, B.W.; Mesquita, R.; Pereira, J.L.; De Souza, S.G.A.; Batista, G.T.; Couto, L.B. Allometric regressions for improved estimate of secondary forest biomass in the central Amazon. For. Ecol. Manag. 1999, 117, 149–167. [Google Scholar] [CrossRef]
- Uddling, J.; Gelang-Alfredsson, J.; Piikki, K.; Pleijel, H. Evaluating the relationship between leaf chlorophyll concentration and SPAD-502 chlorophyll meter readings. Photosynth. Res. 2007, 91, 37–46. [Google Scholar] [CrossRef]
- Pradhan, S.; Sehgal, V.K.; Bandyopadhyay, K.K.; Sahoo, R.N.; Panigrahi, P.; Parihar, C.M.; Jat, S.L. Comparison of vegetation indices from two ground based sensors. J. Indian Soc. Remote Sens. 2018, 46, 321–326. [Google Scholar] [CrossRef]
- Zheng, H.; Cheng, T.; Yao, X.; Deng, X.; Tian, Y.; Cao, W.; Zhu, Y. Detection of rice phenology through time series analysis of ground-based spectral index data. Field Crops Res. 2016, 198, 131–139. [Google Scholar] [CrossRef]
- Verhulst, N.; Govaerts, B.; Nelissen, V.; Sayre, K.D.; Crossa, J.; Raes, D.; Deckers, J. The effect of tillage, crop rotation and residue management on maize and wheat growth and development evaluated with an optical sensor. Field Crops Res. 2011, 120, 58–67. [Google Scholar] [CrossRef]
- Marshall, M.; Thenkabail, P. Developing in situ non-destructive estimates of crop biomass to address issues of scale in remote sensing. Remote Sens. 2015, 7, 808–835. [Google Scholar] [CrossRef] [Green Version]
- Jay, S.; Baret, F.; Dutartre, D.; Malatesta, G.; Héno, S.; Comar, A.; Weiss, M.; Maupas, F. Exploiting the centimeter resolution of UAV multispectral imagery to improve remote-sensing estimates of canopy structure and biochemistry in sugar beet crops. Remote Sens. Environ. 2018, 231, 110898. [Google Scholar] [CrossRef]
- Kanning, M.; Kühling, I.; Trautz, D.; Jarmer, T. High-resolution UAV-based hyperspectral imagery for LAI and chlorophyll estimations from wheat for yield prediction. Remote Sens. 2018, 10, 2000. [Google Scholar] [CrossRef] [Green Version]
- Peng, Y.; Li, Y.; Dai, C.; Fang, S.; Gong, Y.; Wu, X.; Zhu, R.; Liu, K. Remote prediction of yield based on LAI estimation in oilseed rape under different planting methods and nitrogen fertilizer applications. Agric. For. Meteorol. 2019, 271, 116–125. [Google Scholar] [CrossRef]
- Li, S.; Yuan, F.; Ata-UI-Karim, S.T.; Zheng, H.; Cheng, T.; Liu, X.; Tian, Y.; Zhu, Y.; Cao, W.; Cao, Q. Combining color indices and textures of UAV-based digital imagery for rice LAI estimation. Remote Sens. 2019, 11, 1763. [Google Scholar] [CrossRef] [Green Version]
- Aasen, H.; Burkart, A.; Bolten, A.; Bareth, G. Generating 3D hyperspectral information with lightweight UAV snapshot cameras for vegetation monitoring: From camera calibration to quality assurance. ISPRS J. Photogramm. Remote Sens. 2015, 108, 245–259. [Google Scholar] [CrossRef]
- Verger, A.; Vigneau, N.; Chéron, C.; Gilliot, J.-M.; Comar, A.; Baret, F. Green area index from an unmanned aerial system over wheat and rapeseed crops. Remote Sens. Environ. 2014, 152, 654–664. [Google Scholar] [CrossRef]
- Hunt, E.R.; Cavigelli, M.; Daughtry, C.S.T.; Mcmurtrey, J.E.; Walthall, C.L. Evaluation of digital photography from model aircraft for remote sensing of crop biomass and nitrogen status. Precis. Agric. 2005, 6, 359–378. [Google Scholar] [CrossRef]
- Deng, L.; Mao, Z.; Li, X.; Hu, Z.; Duan, F.; Yan, Y. UAV-based multispectral remote sensing for precision agriculture: A comparison between different cameras. ISPRS J. Photogramm. Remote Sens. 2018, 146, 124–136. [Google Scholar] [CrossRef]
- Yao, X.; Wang, N.; Liu, Y.; Cheng, T.; Tian, Y.; Chen, Q.; Zhu, Y. Estimation of wheat LAI at middle to high levels using unmanned aerial vehicle narrowband multispectral imagery. Remote Sens. 2017, 9, 1304. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Li, L.; Gao, W.; Zhang, Y.; Liu, Y.; Wang, S.; Lu, J. Diagnosis of nitrogen status in winter oilseed rape (Brassica Napus L.) using in-situ hyperspectral data and unmanned aerial vehicle (UAV) multispectral images. Comput. Electron. Agric. 2018, 151, 185–195. [Google Scholar] [CrossRef]
- Zhou, K.; Cheng, T.; Zhu, Y.; Cao, W.; Ustin, S.L.; Zheng, H.; Yao, X.; Tian, Y. Assessing the impact of spatial resolution on the estimation of leaf nitrogen concentration over the full season of paddy rice using near-surface imaging spectroscopy Data. Front. Plant Sci. 2018, 9, 964. [Google Scholar] [CrossRef] [Green Version]
- Jay, S.; Gorretta, N.; Morel, J.; Maupas, F.; Bendoula, R.; Rabatel, G.; Dutartre, D.; Comar, A.; Baret, F. Estimating leaf chlorophyll content in sugar beet canopies using millimeter- to centimeter-scale reflectance imagery. Remote Sens. Environ. 2017, 198, 173–186. [Google Scholar] [CrossRef]
- Bendig, J.; Yu, K.; Aasen, H.; Bolten, A.; Bennertz, S.; Broscheit, J.; Gnyp, M.L.; Bareth, G. Combining UAV-based plant height from crop surface models, visible, and near infrared vegetation indices for biomass monitoring in barley. Int. J. Appl. Earth Obs. Geoinformation 2015, 39, 79–87. [Google Scholar] [CrossRef]
- Yue, J.; Yang, G.; Li, C.; Li, Z.; Wang, Y.; Feng, H.; Xu, B. Estimation of winter wheat above-ground biomass using unmanned aerial vehicle-based snapshot hyperspectral sensor and crop height improved models. Remote Sens. 2017, 9, 708. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Niu, Z.; Chen, H.; Li, D.; Wu, M.; Zhao, W. Remote estimation of canopy height and aboveground gbiomass of maize using high-resolution stereo images from a low-cost unmanned aerial vehicle system. Ecol. Indic. 2016, 67, 637–648. [Google Scholar] [CrossRef]
- Croft, H.; Chen, J.M.; Zhang, Y.; Simic, A. Modelling leaf chlorophyll content in broadleaf and needle leaf canopies from ground, CASI, Landsat TM 5 and MERIS reflectance data. Remote Sens. Environ. 2013, 133, 128–140. [Google Scholar] [CrossRef]
- Jayathunga, S.; Owari, T.; Tsuyuki, S. Digital aerial photogrammetry for uneven-aged forest management: Assessing the potential to reconstruct canopy structure and estimate living biomass. Remote Sens. 2019, 11, 338. [Google Scholar] [CrossRef] [Green Version]
- Ming, D.; Yang, J.; Li, L.; Song, Z. Modified ALV for selecting the optimal spatial resolution and its scale effect on image classification accuracy. Math. Comput. Model. 2011, 54, 1061–1068. [Google Scholar] [CrossRef]
- Lu, D. The potential and challenge of remote sensing-based biomass estimation. Int. J. Remote Sens. 2006, 27, 1297–1328. [Google Scholar] [CrossRef]
- Singh, I.; Srivastava, A.K.; Chandna, P.; Gupta, R.K. Crop Sensors for Efficient Nitrogen Management in Sugarcane: Potential and Constraints. Sugar Tech. 2006, 8, 299–302. [Google Scholar] [CrossRef]
- Teal, R.K.; Tubana, B.; Girma, K.; Freeman, K.W.; Arnall, D.B.; Walsh, O.; Raun, W.R. In-season prediction of corn grain yield potential using normalized difference vegetation index. Agron. J. 2006, 98, 1488–1494. [Google Scholar] [CrossRef] [Green Version]
- Yao, Y.; Miao, Y.; Huang, S.; Gao, L.; Ma, X.; Zhao, G.; Jiang, R.; Chen, X.; Zhang, F.; Yu, K. Active canopy sensor-based precision N management strategy for rice. Agron. Sustain. Dev. 2012, 32, 925–933. [Google Scholar] [CrossRef] [Green Version]
- Levy, P.E.; Jarvis, P.G. Direct and indirect measurements of LAI in millet and fallow Vegetation in HAPEX-Sahel. Agric. For. Meteorol. 1999, 97, 199–212. [Google Scholar] [CrossRef]
- Jiang, J.; Liu, D.; Gu, J.; Süsstrunk, S. What is the space of spectral sensitivity functions for digital color cameras? In Proceedings of the 2013 IEEE Workshop on Applications of Computer Vision, Tampa, USA, 15–17 January 2013; pp. 168–179. [Google Scholar]
- Zhou, X.; Zheng, H.B.; Xu, X.Q.; He, J.Y.; Ge, X.K.; Yao, X.; Cheng, T.; Zhu, Y.; Cao, W.X.; Tian, Y.C. Predicting grain yield in rice using multi-temporal vegetation indices from UAV-based multispectral and digital Imagery. ISPRS J. Photogramm. Remote Sens. 2017, 130, 246–255. [Google Scholar] [CrossRef]
- Coburn, C.A.; Smith, A.M.; Logie, G.S.; Kennedy, P. Radiometric and spectral comparison of inexpensive camera systems used for remote sensing. Int. J. Remote Sens. 2018, 39, 4869–4890. [Google Scholar] [CrossRef]
- Rouse, J., Jr.; Haas, R.H.; Schell, J.A.; Deering, D.W. Monitoring vegetation systems in the Great Plains with ERTS. J. Water Resource Protection 1974, 351, 309–317. [Google Scholar]
- Gitelson, A.A.; Kaufman, Y.J.; Merzlyak, M.N. Use of a green channel in remote sensing of global vegetation from EOS-MODIS. Remote Sens. Environ. 1996, 58, 289–298. [Google Scholar] [CrossRef]
- Richardson, A.J.; Everitt, J.H. Using spectral vegetation indices to estimate rangeland productivity. Geocarto Int. 1992, 7, 63–69. [Google Scholar] [CrossRef]
- Rondeaux, G.; Steven, M.; Baret, F. Optimization of soil-adjusted vegetation indices. Remote Sens. Environ. 1996, 55, 95–107. [Google Scholar] [CrossRef]
- Woebbecke, D.M.; Meyer, G.E.; Von Bargen, K.; Mortensen, D.A. Color indices for weed identification under various soil, residue, and lighting conditions. Trans. ASAE 1995, 38, 259–269. [Google Scholar] [CrossRef]
- Meyer, G.E.; Hindman, T.W.; Laksmi, K. Machine vision detection parameters for plant species identification. In Precision Agriculture and Biological Quality; Society of Photo Optical: San Francisco, CA, USA, 1999; Volume 3543, pp. 327–336. [Google Scholar]
- Meyer, G.E.; Neto, J.C. Verification of color vegetation indices for automated crop imaging applications. Comput. Electron. Agric. 2008, 63, 282–293. [Google Scholar] [CrossRef]
- Bannari, A.; Morin, D.; Bonn, F.; Huete, A.R. A review of vegetation indices. Remote Sens. Rev. 1995, 13, 95–120. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Merzlyak, M.N. Spectral reflectance changes associated with autumn senescence of Aesculushippocatanum I and Hacer plantanoides I. leaves. Spectral features and relation to chlorophyll estimation. J. Plant Physiol. 1994, 143, 286–292. [Google Scholar] [CrossRef]
- Hollberg, J.; Schellberg, J. Distinguishing intensity levels of grassland fertilization using vegetation indices. Remote Sens. 2017, 9, 81. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.W.; Yun, H.; Jeong, S.J.; Kwon, Y.S.; Kim, S.G.; Lee, W.; Kim, H.J. Modeling and testing of growth status for Chinese cabbage and white radish with UAV-based RGB imagery. Remote Sens. 2018, 10, 563. [Google Scholar] [CrossRef] [Green Version]
- Zhao, B.; Zhang, J.; Yang, C.; Zhou, G.; Ding, Y.; Shi, Y.; Zhang, D.; Xie, J.; Liao, Q. Rapeseed seedling stand counting and seeding performance evaluation at two early growth stages based on unmanned aerial vehicle imagery. Front. Plant Sci. 2018, 9, 1362. [Google Scholar] [CrossRef]
- Rizeei, H.M.; Pradhan, B. Urban mapping accuracy enhancement in high-rise built-up areas deployed by 3D-orthorectification correction from WorldView-3 and LiDAR imageries. Remote Sens. 2019, 11, 692. [Google Scholar] [CrossRef] [Green Version]
- Xiaoyan, L.; Shuwen, Z.; Zongming, W.; Huilin, Z. Spatial variability and pattern analysis of soil properties in Dehui city, Jilin province. J. Geogr. Sci. 2004, 14, 503–511. [Google Scholar] [CrossRef]
- Hasinoff, S.W.; Sharlet, D.; Geiss, R.; Adams, A.; Barron, J.T.; Kainz, F.; Chen, J.; Levoy, M. Burst photography for high dynamic range and low-light imaging on mobile cameras. ACM Trans. Graph. 2016, 35, 192. [Google Scholar] [CrossRef]
- Monno, Y.; Teranaka, H.; Yoshizaki, K.; Tanaka, M.; Okutomi, M. Single-sensor RGB-NIR imaging: High-quality system design and prototype implementation. IEEE Sens. J. 2019, 19, 497–507. [Google Scholar] [CrossRef]
- Nijland, W.; de Jong, R.; de Jong, S.M.; Wulder, M.A.; Bater, C.W.; Coops, N.C. Monitoring plant condition and phenology using infrared sensitive consumer grade digital cameras. Agric. For. Meteorol. 2014, 184, 98–106. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Wang, C.; Yang, C.; Jiang, Z.; Zhou, G.; Wang, B.; Zhang, D.; You, L.; Xie, J. Evaluation of a UAV-mounted consumer grade camera with different spectral modifications and two handheld spectral sensors for rapeseed growth monitoring: Performance and influencing factors. Precis. Agric. 2020, 1–29. [Google Scholar] [CrossRef]
- Dong, T.; Liu, J.; Shang, J.; Qian, B.; Ma, B.; Kovacs, J.M.; Walters, D.; Jiao, X.; Geng, X.; Shi, Y. Assessment of red-edge vegetation indices for crop leaf area index estimation. Remote Sens. Environ. 2019, 222, 133–143. [Google Scholar] [CrossRef]
- Dash, J.; Pearse, G.; Watt, M. UAV multispectral imagery can complement satellite data for monitoring forest health. Remote Sens. 2018, 10, 1216. [Google Scholar] [CrossRef] [Green Version]
- Brell, M.; Segl, K.; Guanter, L.; Bookhagen, B. 3D hyperspectral point cloud generation: Fusing airborne laser scanning and hyperspectral imaging sensors for improved object-based information extraction. ISPRS J. Photogramm. Remote Sens. 2019, 149, 200–214. [Google Scholar] [CrossRef]
- Zou, J.; Lan, J. A multiscale hierarchical model for sparse hyperspectral unmixing. Remote Sens. 2019, 11, 500. [Google Scholar] [CrossRef] [Green Version]
- Downey, R.K. Agricultural and genetic potentials of cruciferous oilseed crops. J. Am. Oil Chem. Soc. 1971, 48, 718–722. [Google Scholar] [CrossRef]






| Flight Number | Altitude (m) | Image Acquisition Efficiency (min) | Number of Images | Image Processing Efficiency (min) | Image GSD (cm) |
|---|---|---|---|---|---|
| 1 | 22 | 7.45 | 2400 | 100 | 1.35 |
| 2 | 29 | 4.02 | 1300 | 59 | 1.69 |
| 3 | 44 | 2.02 | 760 | 35 | 2.61 |
| 4 | 88 | 1.45 | 550 | 30 | 5.73 |
| 5 | 176 | 0.7 | 470 | 25 | 11.61 |
| Ground Measured Data | Sample Number | Min | Max | Mean | Std | CV (%) |
|---|---|---|---|---|---|---|
| NDVI measured by GreenSeeker (GS-NDVI) | 180 | 0.53 | 0.78 | 0.72 | 0.06 | 8 |
| Leaf area index (LAI) | 180 | 0.41 | 6.15 | 3.17 | 1.20 | 38 |
| Plant height (PH) (m) | 180 | 0.10 | 0.63 | 0.35 | 0.12 | 34 |
| VI | Formula 1 | NIR-VI | RE-VI | RGB-VI |
|---|---|---|---|---|
| Normalized difference vegetation Index | NDVI = (NIR – R) / (NIR + R) [36] | √ | ||
| Green Normalized Difference Vegetation Index | GNDVI = (NIR – G) / (NIR + G) [37] | √ | ||
| Difference Vegetation Index | DVI = NIR − R [38] | √ | ||
| Optimized Soil Adjusted Vegetation Index | OSAVI = (1 + 0.16) × (NIR-R) / (NIR + R + 0.16) [39] | √ | ||
| Excess Green index | ExG = 2G* – R* – B* [40] | √ | ||
| Excess Red index | ExR=1.4R* – G* [41] | √ | ||
| ExG – ExR | ExG – ExR [42] | √ | ||
| Normalized Difference Index | NDI = (G-R)/(G + R) [43] | √ | ||
| Red-edge Normalized Difference Vegetation Index | NDRE = (NIR-RE) / (NIR + RE) [44] | √ |
| VI Type | VI Name | R2 | ||||
|---|---|---|---|---|---|---|
| GSD = 1.35 cm GS-NDVI/LAI | GSD = 1.69 cm GS-NDVI/LAI | GSD = 2.61 cm GS-NDVI/LAI | GSD = 5.73 cm GS-NDVI/LAI | GSD = 11.61 cm GS-NDVI/LAI | ||
| RGB-VIs | EXG | 0.055/0.011 | 0.061/0.014 | 0.061/0.013 | 0.002/0.007 | 0.063/0.015 |
| EXR | 0.292/0.166 | 0.317/0.186 | 0.313/0.179 | 0.237/0.101 | 0.308/0.209 | |
| EXG-EXR | 0.135/0.055 | 0.148/0.063 | 0.148/0.061 | 0.051/0.005 | 0.155/0.076 | |
| NDI | 0.344/0.208 | 0.384/0.240 | 0.362/0.218 | 0.292/0.137 | 0.280/0.184 | |
| RE-VI | NDRE | 0.790/0.712 | 0.808/0.717 | 0.812/0.716 | 0.760/0.698 | 0.706/0.670 |
| NIR-VIs | NDVI | 0.817/0.664 | 0.826/0.666 | 0.821/0.651 | 0.804/0.661 | 0.664/0.609 |
| DVI | 0.728/0.558 | 0.712/0.554 | 0.713/0.501 | 0.740/0.587 | 0.655/0.605 | |
| GNDVI | 0.787/0.714 | 0.794/0.704 | 0.806/0.700 | 0.763/0.699 | 0.630/0.627 | |
| OSAVI | 0.782/0.621 | 0.774/0.621 | 0.773/0.573 | 0.777/0.634 | 0.667/0.617 | |
| RGB-VIs | RE-VI | NIR-VIs | ||||
| VI Type | VI Name | RMSE | ||||
|---|---|---|---|---|---|---|
| GSD = 1.35 cm GS-NDVI/LAI | GSD = 1.69 cm GS-NDVI/LAI | GSD = 2.61 cm GS-NDVI/LAI | GSD = 5.73 cm GS-NDVI/LAI | GSD = 11.61 cm GS-NDVI/LAI | ||
| RGB-VIs | EXG | 0.058/1.235 | 0.058/1.235 | 0.058/1.235 | 0.059/1.216 | 0.058/1.236 |
| EXR | 0.050/1.208 | 0.049/1.205 | 0.049/1.205 | 0.052/1.219 | 0.049/1.191 | |
| EXG-EXR | 0.055/1.232 | 0.055/1.230 | 0.055/1.230 | 0.058/1.234 | 0.055/1.229 | |
| NDI | 0.048/1.193 | 0.047/1.184 | 0.047/1.190 | 0.050/1.207 | 0.050/1.200 | |
| RE-VI | NDRE | 0.027/0.701 | 0.026/0.695 | 0.025/0.697 | 0.029/0.732 | 0.032/0.741 |
| NIR-VIs | NDVI | 0.025/0.742 | 0.024/0.744 | 0.025/0.769 | 0.026/0.767 | 0.034/0.777 |
| DVI | 0.032/0.897 | 0.033/0.906 | 0.033/0.967 | 0.032/0.888 | 0.035/0.838 | |
| GNDVI | 0.027/0.696 | 0.027/0.704 | 0.026/0.714 | 0.029/0.731 | 0.036/0.772 | |
| OSAVI | 0.028/0.817 | 0.028/0.821 | 0.028/0.890 | 0.028/0.821 | 0.034/0.782 | |
| RGB-VIs | RE-VI | NIR-VIs | ||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Wang, C.; Yang, C.; Xie, T.; Jiang, Z.; Hu, T.; Luo, Z.; Zhou, G.; Xie, J. Assessing the Effect of Real Spatial Resolution of In Situ UAV Multispectral Images on Seedling Rapeseed Growth Monitoring. Remote Sens. 2020, 12, 1207. https://0-doi-org.brum.beds.ac.uk/10.3390/rs12071207
Zhang J, Wang C, Yang C, Xie T, Jiang Z, Hu T, Luo Z, Zhou G, Xie J. Assessing the Effect of Real Spatial Resolution of In Situ UAV Multispectral Images on Seedling Rapeseed Growth Monitoring. Remote Sensing. 2020; 12(7):1207. https://0-doi-org.brum.beds.ac.uk/10.3390/rs12071207
Chicago/Turabian StyleZhang, Jian, Chufeng Wang, Chenghai Yang, Tianjin Xie, Zhao Jiang, Tao Hu, Zhibang Luo, Guangsheng Zhou, and Jing Xie. 2020. "Assessing the Effect of Real Spatial Resolution of In Situ UAV Multispectral Images on Seedling Rapeseed Growth Monitoring" Remote Sensing 12, no. 7: 1207. https://0-doi-org.brum.beds.ac.uk/10.3390/rs12071207